Abstract
Maternal signals shape embryonic development, and in turn post-natal phenotypes. RNA deposition is one such method of maternal signalling and circadian rhythms are one trait thought to be maternally inherited, through this mechanism. These maternal circadian gene transcripts aid development of a functioning circadian system. There is increasing evidence that maternal signals can be modified, depending on prevailing environmental conditions to optimise offspring fitness. However, currently, it is unknown if maternal circadian gene transcripts, and consequently early embryonic gene transcription, are altered by maternal developmental conditions. Here, using avian mothers who experienced either pre-natal corticosterone exposure, and/or post-natal stress as juveniles we were able to determine the effects of the timing of stress on downstream circadian RNA deposition in offspring. We demonstrated that maternal developmental history does indeed affect transfer of offspring circadian genes, but the timing of stress was important. Avian mothers who experienced stress during the first 2 weeks of post-natal life increased maternally deposited transcript levels of two core circadian clock genes, BMAL1 and PER2. These differences in transcript levels were transient and disappeared at the point of embryonic genome transcription. Pre-natal maternal stress alone was found to elicit delayed changes in circadian gene expression. After activation of the embryonic genome, both BMAL1 and PER2 expression were significantly decreased. If both pre-natal and post-natal stress occurred, then initial maternal transcript levels of BMAL1 were significantly increased. Taken together, these results suggest that developmental stress differentially produces persistent transgenerational effects on offspring circadian genes.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Oviparous embryonic development is initially controlled exclusively by maternal gene products and proteins deposited within the egg (Abrams and Mullins 2009; Langley et al. 2014). Maternal RNA is then progressively degraded in sequential steps, whilst the zygote begins to take control of transcription (Tadros and Lipshitz 2009; Langley et al. 2014). There is increasing evidence suggesting that maternal RNA can provide active information about the maternal environment. Two recent studies using fish have shown that the critical environmental factors can also be communicated by altered maternal RNA composition in eggs. Round Goby’s (Neogobius melanostomus) communicate the temperature of water experienced by the mother through maternally deposited RNA. Specifically, metabolic genes such as C1QTNF3 were found to be the main group of maternally deposited genes which displayed at least a twofold change by maternally experienced water temperature, hypothesised to optimise aspects of offspring metabolism for specific environments (Adrian-Kalchhauser et al. 2018). A further example of the environment altering maternal RNA deposition comes from Cichlid fish (from the Cichlidae family). Maternally deposited RNA of growth-related genes (for 15 haplochromine Cichlid species) were found to significantly increase or decrease depending on trophic specialisation and environment. For example, species residing in the youngest lake were found to deposit significantly lower amounts of glucocorticoid receptor transcripts within eggs. The results were hypothesised to then influence embryonic production of growth-related genes and later adaptive responses (Ahi et al. 2018). Taken together these studies indicate that environmental conditions may shape the levels of maternal signalling via RNA deposition in a wide range of physiological systems; however, to date, there has been little exploration of this phenomenon across species or physiological traits.
One system in oviparous organisms which may be inherited is the circadian system (Delaunay et al. 2000). Circadian behaviour is critical for allowing congruence between internal and external cycling environments, facilitating optimal resource usage and energy expenditure (Mazzoccoli et al. 2012; Patke et al. 2020). Underlying circadian behaviour are molecular oscillations in clock genes and proteins that interact with each other in transcriptional–translational feedback loops. At the simplified core of these loops are Bmal (Brain and Muscle Arnt-like proteins) and Per family genes (Period). Bmal-related genes activate transcription of Per family genes, which then in turn repress Bmal transcription. This autoinhibitory feedback loop results in the circadian system featuring cyclic expression throughout a 24-h period. The rhythmic expression and repression of these circadian clock genes modulates downstream expression of genes associated with behavioural and physiological changes. This ‘clock system’ is ubiquitous throughout an organism and across species (Reppert and Weaver 2003; Ko and Takahashi 2006).
Circadian system ontology is thought to originate from maternally deposited RNA, specifically Bmal and Per family transcripts (Delaunay et al. 2000; Shang and Zhdanova 2007). This appears to be a conserved developmental mechanism. Maternal transcripts of circadian genes have been identified in oocytes and early embryonic stages prior to activation of the zygotic genome in mammals (Mus musculus, Oryctolagus cuniculus), fish (Perca fluviatilis and Danio rerio), birds (Gallus gallus) and amphibians (Xenopus laevis) (Delaunay et al. 2000; Curran et al. 2008; Dekens and Whitmore 2008; Amano et al. 2009, 2010; Gonçalves et al. 2012; Nishikawa et al. 2013; Almeida et al. 2019). It has been suggested that maternally deposited circadian genes may be involved in phase synchronisation of zygotic clock gene transcription. For example, accumulation of maternal Per3 was suggested to aid initiation of the phase of the free running clock in zebrafish embryos (Delaunay et al. 2000).
The circadian system is sensitive to alteration by external and internal cues, allowing for adaptability. One such factor that the system is particularly sensitive to is stress (Razzoli et al. 2014; Tahara et al. 2015; Koch et al. 2017). Preliminary studies suggest pre-natal stress interacts with elements of the post-natal circadian system, such as the endogenous corticosterone rhythm (Koehl et al. 1997, 1999; Kiryanova et al. 2017). Following maternal stress, offspring have been found to feature hyperactivity, and diminished shifting abilities of circadian rhythms (Kiryanova et al. 2017; Morley-Fletcher et al. 2019). Attenuated expression of circadian clock genes and phase shifted circadian rhythms have also been identified (Yun et al. 2020). This suggests that circadian clock genes and consequently circadian behaviour is susceptible to alteration following exposure to both pre-natal and post-natal developmental stress.
As the circadian system appears to be inherited through maternal RNA, there is a strong theoretical basis for the idea that stressful maternal environments may alter circadian RNA deposition and potentially disrupt zygotic transcription of clock genes. However, this has yet to be tested.
In this study we investigated the effects of maternal developmental history on the deposition of maternal RNA and initiation of embryonic circadian gene transcription. We used a well-established comparative model of circadian neurobiology, the Japanese quail (Coturnix japonica) and investigated the expression of two core circadian clock genes, Bmal1 (a main positive clock element) and Per2 (a main negative clock element) from the point of maternal RNA deposition to early embryonic genome activation.
Materials and methods
All procedures and housing of animals complied with the local ethics committee at the University of St. Andrews and in accordance with the Animals (Scientific Procedures) Act 1986 ASPA regulations under PIL IE1CF3B75 held by JHC and PPL 70/8159 and PAF9F705D held by KAS.
Generation of maternal developmental environments
Pre-natal maternal manipulations
53 fertile Japanese quail eggs (supplier Moonridge farm, Exeter, UK) were incubated (Ova-Easy 190A, Brinsea Products Ltd, UK) in complete darkness whilst on a rotating platform at 37.4 °C with 60% humidity. Fertility was confirmed at day 5 of incubation via an egg candling torch. Eggs were injected at the apex under sterile conditions, as per Zimmer et al. (2013). Injections occurred at day 5 of incubation (Fig. 1). The experimental treatment (N = 29) consisted of 10 µl of 850 ng/ml corticosterone (CORT) prepared in sterile peanut oil (Sigma Aldrich, Poole, UK). The total dose administered was 8.5 ng, as per Zimmer et al. 2013. Administration of 8.5 ng increases the CORT levels 1.8 SD above control CORT yolk levels (Zimmer et al. 2013).
Schematic for experiment. Eggs were injected with 10 µl of 850 ng/ml CORT (orange) or 10 µL peanut oil (blue) at embryonic (E) day 5. Chicks were then subjected to an unpredictable food removal paradigm, or allowed ad libitum food access at post-natal day 19, for 14 days. The feeder and surrounding area were blocked by a panel to prevent access to food for 25% of daylight hours. Eggs collection commenced at post-natal day (PN) 120
Control eggs (N = 24) were injected with 10 µL sterile peanut oil. Needle punctures were sealed (Germolene New Skin, UK), eggs were labelled with their treatment and returned to incubator within 30 min of being removed. At day 13 of incubation, eggs were removed from the Ova-Easy incubator and placed in a treatment specific hatcher (Janoel24 incubator, 80% humidity) and egg turning was ceased. Hatchers were covered with black plastic to ensure the eggs were not exposed to light during the final days of incubation.
Quail housing
All chicks hatched within 48 h. Hatched chicks (Control n = 18, hatch rate = 75%; CORT n = 18, hatch rate = 62%) remained in the hatcher until feathers were dry, a period of approximately 24 h. After this they were transferred to 1m2 floor area pens. CORT and control chicks were moved at the same time into four pre-natal treatment specific pens (two pre-natal CORT and two pre-natal control with a 1 m2 floor area). Chicks were provided with ad libitum food (ground up chick crumb, Dodson & Horrell Ltd, UK) water and an electric contact brooder (Comfort chicks and hatchers, UK). Chicks were kept on a 12:12 light cycle at all times.
At 3 days of age, chicks were moved into larger treatment specific pens (120 cm × 180 cm). One pen from each pre-natal condition was allocated to one of two post-natal treatments; food removal or control (Fig. 1). This gave four groups with 9 chicks in each (pre-natal control/post-natal control n = 9, pre-natal control/post-natal stress n = 9, pre-natal CORT injected/post-natal control n = 9, pre-natal CORT injected/post-natal stress n = 9). In the larger pens, two electric contact brooders were provided for each pen, alongside ad libitum food (chick crumb, Dodson & Horrell Ltd, UK) and water.
Post-natal maternal manipulations
At 19 days of age, chicks in the post-natal stress conditions (pre-natal control n = 9, pre-natal CORT n = 9) were subjected to an unpredictable random food removal paradigm. For 25% of daylight hours food was removed unpredictably for 14 consecutive days, as per Zimmer et al. (2013). Feeders were kept in the same location and a panel was placed in front of the food to prevent access. It was ensured any scattered grain was also blocked by the panel (Fig. 1).
The experiment aimed to investigate maternal transfer of RNA. Therefore, only females were kept in each treatment once sex could be determined (pre-natal control/post-natal control: males n = 5; females n = 4. Pre-natal control/post-natal stress: males n = 6; females n = 3. Pre-natal CORT injected/post-natal control: males n = 5; females n = 4. Pre-natal CORT injected/post-natal stress: males n = 4; females n = 5.) In addition 4 control males (who had received control peanut oil injections and no post-natal food manipulation) were retained to produce fertile eggs for the generation of embryos to study maternal transfer. Of the four remaining control males, one male was placed in each treatment. Males were rotated between conditions approximately every 30 days throughout the experiment to encourage breeding interests.
Generation of embryos to study maternal transfer
Egg collection began at 120 days, when eggs were laid regularly for all conditions. Eggs older than 1 week were not incubated. Eggs were incubated every 6 days; the oldest eggs were 6 days, whilst the youngest were 1 day. Eggs had to be incubated in this manner to allow for sufficient sample size. Not all hens produced equally fertile eggs; therefore, collection over multiple days enabled fertile replicates from each quail to be obtained. Eggs were incubated (Ova-Easy 190A, Brinsea Products Ltd, UK) in complete darkness at 37.4 °C with 60% humidity. Eggs were not rotated whilst incubated to help stabilise the embryo for dissections.
Embryo dissections
The Hamburger and Hamilton (HH) staging system for chickens was used for dissections (Hamburger and Hamilton 1951; Bellairs and Osmond 2005; Hill 2021). Quail and chicken follow the same development time until HH19 (approximately 72 h) Ainsworth et al. (2010). Eggs collected from the quail were incubated for 2 h, 18 h, 23 h and 55 h, corresponding to HH stages 1, 4, 6 and 15, respectively. Maternally deposited transcripts are present at HH1. HH4 marks the onset of gastrulation, with increasing embryonic RNA (and decreasing maternal transcripts) by HH6. HH15 consists of only embryonic RNA (Gonçalves et al. 2012). Single timepoints at each developmental stage were used as clock genes do not oscillate until late embryonic development (Okabayashi et al. 2003; Csernus et al. 2007; Zeman et al. 2009).
Hamburger and Hamilton (HH) stages 1,4 and 6 dissections
Eggs were dissected using a stereoscopic microscope (SMZ745T, Nikon) under red light only. All dissections were carried out within the same hour. To control for order effects, dissection of control and CORT embryos were carried out alternately. Eggs were held at 5 °C prior to dissection. Blackout curtain lining (Black Blackout Thermal 3 Pass Curtain Lining Fabric, Amazon) was sealed around the microscope to ensure no light entered the dissecting area. Thin red Acetate (A4 Red Acetate 200 Micron × 5 Sheets—UK Card Crafts) was sealed around the lamp (Photonic PL2000). Red light was used as it has been demonstrated that light sensitive circadian clock genes, such as PER2, are least effected by red light exposure compared to other wavelengths (Di Rosa et al. 2015). A ~ 2 cm window was cut into the egg and approximately 4 ml of albumen was removed. If the embryo was not found by inspection of the yolk, dye solution (3:10 Indian ink: PBS (Lee et al. 2005) was injected under the chorion to reveal the location of the embryo to ensure precise dissection. The embryo was then dissected out of the chorion and placed immediately into RNAlater (AM7020, Invitrogen). For early developmental timepoints dissecting proto-nervous system tissue was not possible via light microscope due to accuracy. In addition, embryos prior to HH15 were not solid enough to remain intact for dissections. Therefore, the whole embryo was used. Whole embryos have been used successfully for clock gene expression in early chicken development (Gonçalves et al. 2012).
Hamburger and Hamilton stage 15 dissections
For HH15 dissections, Indian ink:PBS dye solution was injected into the egg, and the whole embryo was removed from the yolk as above. The anterior region of the embryo was dissected out to reveal the anterior portion of the embryo (Fig. 2). The anterior region of the embryo (Fig. 2F) was then removed using pulled glass capillaries (Standard Glass Capillaries—4 in, OD 1.5 mm, Filament, World Precision Instruments and Narishige PC-100). Dissected tissue was immediately placed in RNAlater.
HH15 embryo dissection. A Egg injected with dye solution to reveal embryo. B Embryo dissected out and placed in dish with PBS. C Excess tissue removed. Embryo removed from Amnion. D Dorsal view of dissected embryo (posterior section discarded). E Dorsal view of final dissected region. F Side view of final dissected region. Section consists of; (1) prosencephalon, (2) optic vesicle, (3) metencephalon and (4) mesencephalon. Tissue ends just below metencephalon. Embryo was exposed to white light specifically for images
RNA extraction and qPCR
Total RNA was extracted via Absolutely RNA miniprep kit (Agilent, UK) as per the manufacturer’s instructions. RNA concentrations for all samples were analysed on a QuBit 2.0 flourometer using RNA HS Assay Kit (Thermofisher, UK). A subset of 30 samples were analysed for integrity using a 2100 BioAnalyzer system, RNA 6000 nanokit (Agilent, UK). The mean RIN was 3.9 and ranged from 1 to 10.
2 ng of cDNA was then synthesised (nanoscript2 reverse transcription kits, primer design, UK). qPCR analysis was then performed using the synthesised cDNA. Specific primers were designed and validated (PrimerDesign Ltd, UK), amplifying single products only for Bmal1 and Per2, Bmal1-Foward: GTACGTTTCTCGACATGCAATAGA Reverse: AGGTGTCCTATATCATCTTGATGGAA, Per2-Forward: CTGGCAAACCTGAAAGTGTTGTA Reverse: CTCCACTTGGACCATCTTCTATCA.
βactin (ACTβ) was used as the reference gene due to being previously identified as the most stable via gnorm (Zimmer and Spencer 2014). All qPCR runs were conducted in duplicate, with a 20 ul total reaction volume (10 uL PrecisionPLUS qPCR Master Mix, 1 uL primer mix (300 ng/20 uL reaction), 8 uL RNAse free H2O and 1uL (0.45 nG) DNA. The following amplification settings were used as per manufactures instructions (Presicion plus mastermix, primerdesign); an initial 2 min at 95 °C, and then 40 cycles of 10 s at 95 °C, 60 s at 60 °C, followed by a melt curve analysis. All standard curve efficiencies were above 96%, and r2 > 0.7. ΔCT was calculated as \(\mathrm{\Delta CT}={2}^{-( \overline{x}\mathrm{ Gene of interest }- \overline{x}\mathrm{ housekeeping })}\)
Statistical analysis
All statistics were conducted in RStudio version 1.4.17. Residuals did not follow a normal distribution, so transcript expression data sets were log transformed to fulfilled parametric model assumptions. Stability of the housekeeping gene was assessed. ACTβ was found to be differentially expressed between age groups: F(3,127) = 11.72, P < 0.0001. Expression was stable across treatments (Prenatal stress: F(1,127) = 0.510, P = 0.48, Postnatal stress: F(1,127) = 0.23, P = 0.63). Due to unstable expression of ACTβ across developmental time, age specific models were conducted. To compare relative transcript expression, age specific ANOVAs were conducted for each gene. Pre-natal and post-natal treatments were included as main effects. Interactions were also included between pre-natal and post-natal treatments (gene ~ pre-natal + post-natal + pre-natal*post-natal). The room quail were housed in was included as a blocking factor in the ANOVAs (Mangiafico 2016). If significant effects were found, Tukey multiple comparisons of means with a 95% confidence was run as a post hoc test, where appropriate.
Results
Maternal RNA transcript deposition at HH1 is affected by developmental history
Bmal1 and Per2 were found in all samples at HH1, 2 h after incubation. At HH1, maternal developmental history significantly affected deposition of both Bmal1 and Per2 in the developing embryo (Fig. 3). For Bmal1, a significant interaction between maternal pre-natal and post-natal stress was identified (F(1,26) = 14.33, P = 0.00082). Post hoc analysis revealed a significantly higher level of BMAL1 in embryos whose mothers had experienced both pre-natal CORT and post-natal stress, compared to all other conditions (pre-natal CORT/post-natal stress—pre-natal control/post-natal control (P = 0.025), pre-natal CORT/post-natal stress—pre-natal control/post-natal stress (P = 0.0047), and pre-natal CORT/post-natal stress—pre-natal CORT/post-natal control (P = 0.00015), supplementary Table 1). Pre-natal maternal stress was not found to effect BMAL1 expression (F(1,26) = 1.18, P = 0.29). Post-natal maternal stress, however, was found to significantly increase Bmal1 expression within embryos from post-natally stressed mothers, compared to controls (F(1,26) = 11.63, P = 0.0021). This marked difference is likely due to the strong increase of Bmal1 seen in pre-natal CORT/post-natal stress embryos.
Effects of maternal developmental history on Bmal1 gene deposition at HH1 using whole embryos. Normalized, untransformed Delta CT values are shown. a Pre-natal CORT exposure only. b Post-natal stress only. c Interactions between pre-natal CORT exposure and post-natal stress. Boxplots depict median relative expression levels and the 25th and 75th percentiles. Whiskers are 1.5× the interquartile range, data points outside this range are marked as outliers (circles). ***Indicate significant difference (P < 0.001), **Indicate significant differences (P < 0.01) and *Indicates significant differences (P < 0.05)
For Per2 expression, no significant interactions between pre-natal and post-natal stress were identified (F(1,28) = 2.36, P = 0.14). Similar to Bmal1, Per2 expression at HH1 was not significantly affected by pre-natal maternal stress (F(1,28) = 0.3, P = 0.59). A significant increase in relative Per2 expression was identified in embryos whose mothers had experienced post-natal stress (F(1,28) = 10.15, P = 0.0035) (Fig. 4).
Effects of maternal developmental history on Per2 gene deposition at HH1 using whole embryos. Normalized, untransformed Delta CT values are shown. a Pre-natal CORT exposure only. b Post-natal stress only. c Interactions between pre-natal CORT exposure and post-natal stress. Boxplots depict median relative expression levels and the 25th and 75th percentiles. Whiskers are 1.5× the interquartile range, data points outside this range are marked as outliers (circles). ***Indicate significant difference (P < 0.001), **Indicate significant differences (P < 0.01) and *Indicates significant differences (P < 0.05)
Circadian clock gene RNA at embryonic stage HH4 is affected by maternal developmental history
At HH4, no significant interaction was identified between pre- and post-natal stress and Bmal1 expression (F(1,31) = 0.156, P = 0.7). Bmal1 expression was significantly affected by the developmental timing a stress was received (Fig. 5) (pre-natal: F(1,31) = 8.26, P = 0.0073, post-natal: F(1,31) = 9.03, P = 0.0052). Embryos from mothers that had experienced pre-natal stress alone were found to have a significant increase in Bmal1 expression, whereas a decreased Bmal1 expression was found in embryos whose mothers experienced post-natal stress.
Effects of maternal developmental history on Bmal1 relative expression at HH4 using whole embryos. Normalized, untransformed Delta CT values are shown. a Pre-natal stress only. b Post-natal stress only. c Interactions between pre-natal CORT exposure and post-natal stress. Boxplots depict median relative expression levels and the 25th and 75th percentiles. Whiskers are 1.5× the interquartile range, data points outside this range are marked as outliers (circles). ***Indicate significant difference (P < 0.001), **Indicate significant differences (P < 0.01) and *Indicates significant differences (P < 0.05)
For Per2 expression, a significant interaction between pre-natal and post-natal maternal stress was identified (F(1,31) = 5.71, P = 0.023, Fig. 6). Post hoc analysis revealed Per2 was also significantly higher in embryos from mothers who has experienced both pre-natal CORT and post-natal stress compared to those who had experienced post-natal stress alone (P = 0.0017) (Supplementary Table 1). Similar to Bmal1, Per2 expression was significantly increased in whole embryos from pre-natal stress in mothers (F(1,31) = 10.77, P = 0.0026). Post-natal stress was not found to cause any significant differences in Per2 expression (F(1,31) = 0.23, P = 0.64).
Effects of maternal developmental history on Per2 relative expression at HH4 using whole embryos. Normalized, untransformed Delta CT values are shown. a Pre-natal CORT only. b Post-natal stress only. c Pre-natal and post-natal interactions. Boxplots depict median relative expression levels and the 25th and 75th percentiles. Whiskers are 1.5× the interquartile range, data points outside this range are marked as outliers. ***Indicate significant difference (P < 0.001), **Indicate significant differences (P < 0.01) and *Indicates significant differences (P < 0.05)
Maternal developmental history does not alter circadian clock gene levels at HH6
At HH6, no significant interactions between pre-natal and post-natal stress were identified (Bmal1: F(1,23) = 0.59, P = 0.52; Fig. 7, Per2: F(1,25) = 2.07, P = 0.16; Fig. 8). No differences in Bmal1 or Per2 expression were identified at HH6 following pre-natal maternal stress (Bmal1: F(1,23) = 0.53, P = 0.48, Per2: F(1,25) = 0.2, P = 0.66) or post-natal maternal stress (Bmal1: F(1,23) = 1.35, P = 0.26, Per2: F(1,25) = 1.49, P = 0.23).
Effects of maternal developmental history on Bmal1 gene deposition at HH6 using whole embryos. Normalized, untransformed Delta CT values are shown. a Pre-natal CORT exposure only. b Post-natal stress only. c Interactions between pre-natal CORT exposure and post-natal stress. Boxplots depict median relative expression levels and the 25th and 75th percentiles. Whiskers are 1.5X the interquartile range, data points outside this range are marked as outliers (circles). ***Indicate significant difference (P < 0.001), **Indicate significant differences (P < 0.01) and *Indicates significant differences (P < 0.05)
Effects of maternal developmental history on Per2 gene deposition at HH6 using whole embryos. Normalized, untransformed Delta CT values are shown. a Pre-natal CORT exposure only. b Post-natal stress only. c Interactions between pre-natal CORT exposure and post-natal stress. Boxplots depict median relative expression levels and the 25th and 75th percentiles. Whiskers are 1.5X the interquartile range, data points outside this range are marked as outliers (circles). ***Indicate significant difference (P < 0.001), **Indicate significant differences (P < 0.01) and *Indicates significant differences (P < 0.05)
Maternal developmental history alters embryonic circadian gene transcription at HH15
By HH15, levels of Bmal1 within the dorsal region of the embryo were not found to be significantly affected by interactions between maternal pre- and post-natal stress (F(1,30) = 1.89, P = 0.18, Fig. 9). Bmal1 levels were, however, found to be significantly decreased if the mother had been exposed to pre-natal CORT (F(1,30) = 7.13, P = 0.012). No significant effect of post-natal stress alone was identified (F(1,30) = 0.2, P = 0.66). Per2 featured the same patterns of expression; no significant interaction between maternal pre-natal and post-natal stress identified (F(1,29) = 0.13, P = 0.72, Fig. 10). A significant decrease was present following pre-natal maternal CORT exposure (F(1,29) = 5.86, P = 0.022), and no significant differences in Per2 expression were identified for post-natal stress (F(1,29) = 0.37, P = 0.29).
Effects of maternal developmental history on Bmal1 gene deposition at HH15 using whole embryos. Normalized, untransformed Delta CT values are shown. a Pre-natal CORT exposure only. b Post-natal stress only. c Interactions between pre-natal CORT exposure and post-natal stress. Boxplots depict median relative expression levels and the 25th and 75th percentiles. Whiskers are 1.5X the interquartile range, data points outside this range are marked as outliers (circles). ***Indicate significant difference (P < 0.001), **Indicate significant differences (P < 0.01) and *Indicates significant differences (P < 0.05)
Effects of maternal developmental history on Per2 gene deposition at HH15 using whole embryos. Normalized, untransformed Delta CT values are shown. a Pre-natal CORT exposure only. b Post-natal stress only. c Interactions between pre-natal CORT exposure and post-natal stress. Boxplots depict median relative expression levels and the 25th and 75th percentiles. Whiskers are 1.5X the interquartile range, data points outside this range are marked as outliers (circles). ***Indicate significant difference (P < 0.001), **Indicate significant differences (P < 0.01) and *Indicates significant differences (P < 0.05)
Discussion
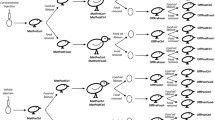
We present the first evidence that environmental conditions experienced by the mother can drive later RNA deposition and ontogenesis of circadian clock genes, and in turn early embryonic transcription of circadian clock genes (Fig. 11). Pre-natal stress appears to feature longer term modifications to the embryonic genome, whilst post-natal stress elicits a change in maternal gene deposition which appears more transient in nature. Such pre-natal communication of environmental information between parents and offspring offers a fitness advantage, by ‘preparing’ the offspring for a specific post-natal environment (Dzialowski and Sotherland 2004; Horner and Wolfner 2008; Hargitai et al. 2009; Abrams and Mullins 2009; Sharda et al. 2021). This may then give rise to adaptive potential by phenotypic alterations which are appropriate for the maternally communicated environment (Mousseau and Fox 1998; Räsänen and Kruuk 2007; Shama et al. 2014; English et al. 2015; Kuijper and Hoyle 2015; Van Dooren et al. 2016; Lubzens et al. 2017). This is further supported by both pre-natal and post-natal stress elicit cumulative increases in initial Bmal1 expression, and Per2 expression during initial embryonic transcription (HH1 and HH4). Communicating environments through non-genomic means is a relatively new avenue of research; therefore, the full extent of non-genomic maternal communication in shaping adaptive potential is a largely unknown (Kuijper and Hoyle 2015; Moore et al. 2019). The downstream effects of the altered circadian expression identified in this study remain unknown and further studies investigating post-natal circadian hormone activity and behaviour should be conducted to confirm if these mRNA findings translate to a meaningful biological response.
Graphical depiction of results. Mothers exposed to pre-natal stress only were not found to deposit different levels of Bmal1 and Per2 transcripts in ovo (HH1). Embryos of mothers exposed to pre-natal stress only were found to have significantly increased levels of both Bmal1 and Per2 compared to controls at HH4. No differences were identified at HH6. At HH15 embryos featured significantly decreased Bmal1 and Per2. Mothers who had experienced post-natal stress in early life were found to deposit in ovo (HH1) higher levels of Bmal1 and Per2 transcripts than controls. At HH4 Bmal1 levels were significantly decreased when compared to controls. No further differences were identified at HH6 and HH15. Mothers who experienced both pre-natal and post-natal stress were found to initially deposit larger quantities of Bmal1 when compared to all other conditions. At HH4, increased levels of Per2 were identified when compared to mothers who had experienced post-natal stress only. No further differences were found at HH6 and HH15
Post-natal maternal stress elicits changes in Circadian clock gene RNA deposition
Post-natal maternal stress was found to elicit transient changes in circadian RNA levels, with all differences disappearing by HH6. Goncalves et al., have previously confirmed circadian RNA at these early developmental timepoints are maternally deposited (Gonçalves et al. 2012). The altered levels of both Bmal1 and Per2 we identified provides strong evidence that post-natal maternal stress altered levels of maternally deposited circadian clock gene RNA in ovo. For initial maternal transcripts, at HH1, both Bmal1 and Per2 were found to be significantly increased in embryos from mothers who had experienced post-natal stress exposure, but not from those who had experiences pre-natal stress. Due to the immature state of the circadian system at this point in development the autoinhibitory expression of both Bmal1 and Per2 is not yet fully functional (Okabayashi et al. 2003). It is, therefore, not surprising that both the positive and negative regulator are increased in expression.
The increased expression of both Bmal1 and Per2 from mothers experiencing post-natal stress only, suggests that stress features different effects on maternal transmission depending on the developmental time (pre- or post-natal) it was experienced. A significant interaction between pre-natal and post-natal stress was present for Bmal1 expression. Bmal1 was significantly increased in pre-natal CORT/post-natal stress compared to all other conditions.
At HH4, Bma1 expression was again altered following post-natal maternal stress. A significant decrease in Bmal1 expression was identified. Per2 expression was unaffected. Pre-natal and post-natal maternal stress significantly interacted at HH4; both pre-natal and post-natal stress resulted in a higher expression of Per2 compared to post-natal stress alone. It is thought the majority of embryonic circadian clock genes become active at HH6 (Gonçalves et al. 2012). At this point, maternal developmental history was not found to alter Bmal1 or Per2 expression.
Pre-natal maternal stress causes changes to embryonic circadian transcript levels
Pre-natal maternal stress appears to have a delayed effect on circadian clock gene expression. This is likely due to changes in the embryonic genome (which gradually activates) rather than alteration to initial maternal transcript deposits. Pre-natal maternal stress effects became apparent at HH4. Both Bmal1 and Per2 were found to be significantly increased following maternal exposure to pre-natal stress. Maternal to zygotic genome activation occurs in gradual steps; therefore, it is not unlikely this may indicate changes in the embryonic genome. By HH15 the embryo is presumed to be transcribing only the embryonic genome. Pre-natal stress caused a significant decrease in both Bmal1 and Per2 expression. Again given the immature state of the autoinhibitory feedback loop of these genes it is not surprising the same effects were seen in both.
Pre-natal and post-natal stress elicit cumulative increases in circadian clock gene RNA
Exposure to both pre- and post-natal stress appears to elicit a cumulative increase in maternal Bmal1 deposition. Per2 did not display such increases, indicating the maternal Bmal1 deposition is most susceptible to environmental stress. The increased levels of Bmal1 may then alter the phase of the embryonic circadian clock. Synchronization of embryonic clocks have been found to be altered by different levels of maternally deposited Per3 in zebrafish (Delaunay et al. 2000). The cumulative effect of stress may be a method of communicating reliably stressful maternal environments to offspring. Such cumulative stress effects were also seen in Per2 at HH4; maternal pre-natal and post-natal stress elicited significantly higher Per2 levels compared to post-natal stress alone.
Taken together the results suggest that the developmental period a stress is received alters the quantities of maternal circadian RNA deposited in eggs. Post-natal stress elicits a significant increase in circadian gene deposition. These differences disappear as the embryo begins to transcribe the embryonic genome. Pre-natal stress alone, however, features little impact until the end of maternal RNA transcription at HH4, where a significant increase is present in both genes. Pre-natal maternal stress appears to elicit long term effects on embryonic circadian gene transcription; significantly reducing the amount synthesised at HH15. This may indicate maternally experienced embryonic stress causes a more permanent change in future maternal communication than post-natal stress. This may be due to epigenetic modifications to the embryonic genome (Aiken and Ozanne 2014) which are not communicated through varying transcript deposition levels, and do not appear until activation of the embryonic genome. This would contrast to post-natal developmental stress which is communicated by varying levels of maternal RNA.
Quail who received both pre-natal and post-natal stress deposited significantly higher amounts of transcripts into the embryo. This suggests that the type of stress received influences maternal deposition of mRNA; consistent stress (from pre-natal CORT/post-natal stress) is better communicated to the embryo, compared to inconsistent stress (from pre-natal control/post-natal stress or pre-natal control/post-natal stress).
Recent studies have found in ovo development to be more flexible than previously thought. Signals during incubation, such as acoustics, clutch size and incubation patterns can alter post-natal phenotypes (Gorman and Nager 2004; Reid et al. 2008; Noguera 2019; Mariette 2020). Such external factors likely effect the cumulative stress effects seen in this experiment and may potentially reinforce altered levels of circadian clock genes, allowing effects seen to persist to embryonic transcription.
For the most part, however, maternal signals are communicated before oviposition in the form of RNA, proteins and hormones (Ahi et al. 2018; Costantini et al. 2014; Monaghan 2007; Valcu et al. 2019). It is thought for optimal survival of offspring, the ‘one off’ signals put into the egg need to reliably ‘predict’ the future post-natal environment. Maternal signals can rapidly alter offspring phenotypes (Räsänen and Kruuk. 2007). This is thought to be problematic when in an unpredictable environment. A caveat of this rapid alteration is offspring may be altered to optimize fitness in one extreme post-natal environment and may hatch into the opposite, reducing fitness (Monaghan 2007). Unlike placental development there is no continuous biochemical communication from the mother (Groothuis et al. 2019, 2020). Oviparous maternal signals need to reliably predict the future post-natal environment without direct maternal feedback.
To date there is little research on the transgenerational effects of stress and the circadian system. A recent study in mice showed that offspring from mothers exposed to chronic stress during pregnancy had advanced circadian phase behaviour, increased PER1 Suprachiasmatic nuclei (SCN) expression and finally significant variation in activity onset (Yun et al. 2020). Other findings in mammals confirm that stress exposure during pregnancy alters offspring circadian behaviour. Alterations included fragmented sleep and phase advance in both activity and corticosterone rhythm (Koehl et al. 1997; Dugovic et al. 1999; Kiryanova et al. 2017; Morley-Fletcher et al. 2019). Currently there is no research into maternal developmental history and the effects on offspring circadian gene expression in oviparous organisms. Our results suggest that maternal circadian gene transfer is altered if the maternal life history consists of both a pre-natal exposure to CORT, and a post-natal stressor.
If the environment of an embryo predisposed to altered circadian clock genes is poor this may signal for altered regulation. Altered circadian clock gene expression is generally regarded as detrimental to long-term health (Rijo-Ferreira and Takahashi 2019). Altered levels of PER2 are associated with altered circadian activity, such as disrupted sleep rhythms and phase delays (Balsalobre et al. 2000; Chen et al. 2009; Cheon et al. 2013). This may, however, be beneficial to survival in a poor environment. A disrupted rhythm may increase fitness when there are no predictable sources of necessities, for example, food and temperature. As an example, if the organism follows predictable diurnal rhythms, but food is unpredictable, this may decrease chances of obtaining food (by sleeping when sporadic food becomes present). In addition, if the poor environment correlates with increased predators, it may be beneficial to rest for short periods interspersed with moving to a new location, compared to constant sleep for 8 h. This gives a short-term advantage to survival and is, therefore, not selected against as the animal will still reproduce. The disrupted rhythms may act as a trade-off between reaching sexual maturity and reproducing with a shortened lifespan. This ‘grow now pay later’ trade-off is known to occur from other maternal signals (Metcalfe and Monaghan 2001; Monaghan 2007).
Data availability
Research data and R analysis code is available at DOI: https://doi.org/10.17630/792c7a01-54d3-48dd-8588-b29bbee39608.
References
Abrams EW, Mullins MC (2009) Early zebrafish development: it’s in the maternal genes. Curr Opin Genet Dev 19:396–403. https://doi.org/10.1016/J.GDE.2009.06.002
Adrian-Kalchhauser I, Walser J-C, Schwaiger M, Burkhardt-Holm P (2018) RNA sequencing of early round goby embryos reveals that maternal experiences can shape the maternal RNA contribution in a wild vertebrate. BMC Evol Biol 181(18):1–14. https://doi.org/10.1186/S12862-018-1132-2
Ahi EP, Singh P, Lecaudey LA, Gessl W, Sturmbauer C (2018) Maternal mRNA input of growth and stress-response-related genes in cichlids in relation to egg size and trophic specialization. EvoDevo 91(9):1–17. https://doi.org/10.1186/S13227-018-0112-3
Aiken CE, Ozanne SE (2014) Transgenerational developmental programming. Hum Reprod Update 20:63–75. https://doi.org/10.1093/HUMUPD/DMT043
Ainsworth SJ, Stanley RL, Evans DJR (2010) Developmental stages of the Japanese quail. J Anat 216:3–15. https://doi.org/10.1111/j.1469-7580.2009.01173.x
Amano T, Matsushita A, Hatanaka Y, Watanabe T, Oishi K, Ishida N, Anzai M, Mitani T, Kato H, Kishigami S, Saeki K, Hosoi Y, Iritani A, Matsumoto K (2009) Expression and functional analyses of circadian genes in mouse oocytes and preimplantation embryos: Cry1 is involved in the meiotic process independently of circadian clock regulation. Biol Reprod 80:473–483. https://doi.org/10.1095/BIOLREPROD.108.069542
Amano T, Tokunaga K, Kakegawa R, Yanagisawa A, Takemoto A, Tatemizo A, Watanabe T, Hatanaka Y, Matsushita A, Kishi M, Anzai M, Kato H, Mitani T, Kishigami S, Saeki K, Hosoi Y, Iritani A, Matsumoto K (2010) Expression analysis of circadian genes in oocytes and preimplantation embryos of cattle and rabbits. Anim Reprod Sci 121:225–235. https://doi.org/10.1016/J.ANIREPROSCI.2010.05.020
Balsalobre A, Brown SA, Marcacci L, Tronche F, Kellendonk C, Reichardt HM, Schütz G, Schibler U (2000) Resetting of circadian time in peripheral tissues by glucocorticoid signaling. Science 289:2344–2347. https://doi.org/10.1126/SCIENCE.289.5488.2344
Bellairs R, Osmond M (2005) Atlas of Chick Development: Second Edition. Atlas Chick Dev Third Ed 1–660. https://doi.org/10.1016/C2010-0-65149-2
Chen R, Schirmer A, Lee Y, Lee H, Kumar V, Yoo S-H, Takahashi JS, Lee C (2009) Rhythmic PER abundance defines a critical nodal point for negative feedback within the circadian clock mechanism. Mol Cell 36:417–430. https://doi.org/10.1016/J.MOLCEL.2009.10.012
Cheon S, Park N, Cho S, Kim K (2013) Glucocorticoid-mediated Period2 induction delays the phase of circadian rhythm. Nucleic Acids Res 41:6161–6174. https://doi.org/10.1093/nar/gkt307
Costantini D, Monaghan P, Metcalfe NB (2014) Prior hormetic priming is costly under environmental mismatch. Biol Lett. https://doi.org/10.1098/RSBL.2013.1010
Csernus VJ, Nagy AD, Faluhelyi N (2007) Development of the rhythmic melatonin secretion in the embryonic chicken pineal gland. Gen Comp Endocrinol 152:148–153. https://doi.org/10.1016/J.YGCEN.2007.01.008
Curran KL, LaRue S, Bronson B, Solis J, Trow A, Sarver N, Zhu H (2008) Circadian genes are expressed during early development in xenopus laevis. PLoS ONE 3:e2749. https://doi.org/10.1371/JOURNAL.PONE.0002749
de Almeida TR, Alix M, Le CA, Klopp C, Montfort J, Toomey L, Ledoré Y, Bobe J, Chardard D, Schaerlinger B, Fontaine P (2019) Domestication may affect the maternal mRNA profile in unfertilized eggs, potentially impacting the embryonic development of Eurasian perch (Perca fluviatilis). PLoS ONE 14:e0226878. https://doi.org/10.1371/JOURNAL.PONE.0226878
Dekens MPS, Whitmore D (2008) Autonomous onset of the circadian clock in the zebrafish embryo. EMBO J 27:2757–2765. https://doi.org/10.1038/emboj.2008.183
Delaunay F, Thisse C, Marchand O, Laudet V, Thisse B (2000) An inherited functional circadian clock in zebrafish embryos. Science 289:297–300. https://doi.org/10.1126/science.278.5343.1632
Di Rosa V, Frigato E, López-Olmeda JF, Sánchez-Vázquez FJ, Bertolucci C (2015) The light wavelength affects the ontogeny of clock gene expression and activity rhythms in Zebrafish Larvae. PLoS ONE 10:e0132235. https://doi.org/10.1371/JOURNAL.PONE.0132235
Dugovic C, Maccari S, Weibel L, Turek FW, Van Reeth O, Van Reeth O (1999) High corticosterone levels in prenatally stressed rats predict persistent paradoxical sleep alterations. J Neurosci 19:8656–8664. https://doi.org/10.1523/JNEUROSCI.19-19-08656.1999
Dzialowski EM, Sotherland PR (2004) Maternal effects of egg size on emu Dromaius novaehollandiae egg composition and hatchling phenotype. J Exp Biol 207:597–606. https://doi.org/10.1242/JEB.00792
English S, Pen I, Shea N, Uller T (2015) The information value of non-genetic inheritance in plants and animals. PLoS ONE 10:e0116996. https://doi.org/10.1371/JOURNAL.PONE.0116996
Gonçalves L, Vinhas M, Pereira R, Pais De Azevedo T, Bajanca F, Palmeirim I (2012) Circadian clock genes Bmal1 and clock during early chick development. Dev Dyn 241:1365–1373. https://doi.org/10.1002/dvdy.23821
Gorman H, Nager R (2004) Prenatal developmental conditions have long-term effects on offspring fecundity. Proceedings Biol Sci 271:1923–1928. https://doi.org/10.1098/RSPB.2004.2799
Groothuis TGG, Hsu B-Y, Kumar N, Tschirren B (2019) Revisiting mechanisms and functions of prenatal hormone-mediated maternal effects using avian species as a model. Philos Trans R Soc B. https://doi.org/10.1098/RSTB.2018.0115
Groothuis TG, Kumar N, Hsu BY (2020) Explaining discrepancies in the study of maternal effects: the role of context and embryo. Curr Opin Behav Sci 36:185–192. https://doi.org/10.1016/J.COBEHA.2020.10.006
Hamburger V, Hamilton HL (1951) A series of normal stages in the development of the chick embryo. J Morphol 88(1):49–92
Hargitai R, Arnold KE, Herényi M, Prechl J, Török J (2009) Egg composition in relation to social environment and maternal physiological condition in the collared flycatcher. Behav Ecol Sociobiol 63:869–882. https://doi.org/10.1007/S00265-009-0727-4/FIGURES/5
Hill M (2021) Hamburger hamilton stages. In: embryology. https://embryology.med.unsw.edu.au/embryology/index.php/Hamburger_Hamilton_Stages. Accessed Jan 2021
Horner VL, Wolfner MF (2008) Transitioning from egg to embryo: triggers and mechanisms of egg activation. Dev Dyn 237:527–544. https://doi.org/10.1002/DVDY.21454
Kiryanova V, Smith VM, Dyck RH, Antle MC (2017) Circadian behavior of adult mice exposed to stress and fluoxetine during development. Psychopharmacology 234:793–804. https://doi.org/10.1007/s00213-016-4515-3
Ko CH, Takahashi JS (2006) Molecular components of the mammalian circadian clock. Hum Mol Genet 15:R271–R277. https://doi.org/10.1093/hmg/ddl207
Koch CE, Leinweber B, Drengberg BC, Blaum C, Oster H (2017) Interaction between circadian rhythms and stress. Neurobiol Stress 6:57–67. https://doi.org/10.1016/J.YNSTR.2016.09.001
Koehl M, Barbazanges A, Le Moal M, Maccari S (1997) Prenatal stress induces a phase advance of circadian corticosterone rhythm in adult rats which is prevented by postnatal stress. Brain Res 759:317–320. https://doi.org/10.1016/S0006-8993(97)00394-6
Koehl M, Darnaudery M, Dulluc J, Van Reeth O, Le MM, Maccari S, Darnaudéry M, Dulluc J, Van RO, Le MM, Maccari S, Darnaudery M, Dulluc J, Van Reeth O, Le MM, Maccari S (1999) Prenatal stress alters circadian activity of hypothalamo–pituitary–adrenal axis and hippocampal corticosteroid receptors in adult rats of both gender. J Neurobiol 40:302–315. https://doi.org/10.1002/(SICI)1097-4695(19990905)40:3%3c302::AID-NEU3%3e3.0.CO;2-7
Kuijper B, Hoyle RB (2015) When to rely on maternal effects and when on phenotypic plasticity? Evolution 69:950–968. https://doi.org/10.1111/EVO.12635
Langley AR, Smith JC, Stemple DL, Harvey SA (2014) New insights into the maternal to zygotic transition. Development 141:3834–3841. https://doi.org/10.1242/dev.102368
Lee VM, Bronner-Fraser M, Baker CVH (2005) Restricted response of mesencephalic neural crest to sympathetic differentiation signals in the trunk. Dev Biol 278:175–192. https://doi.org/10.1016/j.ydbio.2004.10.024
Lubzens E, Bobe J, Young G, Sullivan CV (2017) Maternal investment in fish oocytes and eggs: the molecular cargo and its contributions to fertility and early development. Aquaculture 472:107–143. https://doi.org/10.1016/J.AQUACULTURE.2016.10.029
Mangiafico SS (2016) Summary and Analysis of Extension Program Evaluation in R, version 1
Mariette MM (2020) Acoustic developmental programming: implications for adaptive plasticity and the evolution of sensitive periods. Curr Opin Behav Sci 36:129–134. https://doi.org/10.1016/J.COBEHA.2020.09.010
Mazzoccoli G, Pazienza V, Vinciguerra M (2012) Clock genes and clock-controlled genes in the regulation of metabolic rhythms. Chronobiol Int 29:227–251. https://doi.org/10.3109/07420528.2012.658127
Metcalfe NB, Monaghan P (2001) Compensation for a bad start: grow now, pay later? Trends Ecol Evol 16:254–260. https://doi.org/10.1016/S0169-5347(01)02124-3
Monaghan P (2007) Early growth conditions, phenotypic development and environmental change. Philos Trans R Soc B Biol Sci 363:1635–1645. https://doi.org/10.1098/rstb.2007.0011
Moore MP, Whiteman HH, Martin RA (2019) A mother’s legacy: the strength of maternal effects in animal populations. Ecol Lett 22:1620–1628. https://doi.org/10.1111/ELE.13351
Morley-Fletcher S, Mairesse J, Van Camp G, Reynaert M-L, Gatta E, Marrocco J, Bouwalerh H, Nicoletti F, Maccari S (2019) Perinatal stress programs sex differences in the behavioral and molecular chronobiological profile of rats maintained under a 12-h light-dark cycle. Front Mol Neurosci 12:89. https://doi.org/10.3389/fnmol.2019.00089
Mousseau TA, Fox CW (1998) The adaptive significance of maternal effects. Trends Ecol Evol 13:403–407. https://doi.org/10.1016/S0169-5347(98)01472-4
Nishikawa S, Hatanaka Y, Tokoro M, Shin S-W, Shimzu N, Nishihara T, Kato R, Takemoto A, Amano T, Anzai M, Kishigami S, Hosoi Y, Matsumoto K (2013) Functional Analysis of nocturnin, a circadian deadenylase, at maternal-to-zygotic transition in mice. J Reprod Dev 59:2013–3001. https://doi.org/10.1262/JRD.2013-001
Noguera JC, Velando A (2019) Bird embryos perceive vibratory cues of predation risk from clutch mates. Nat Ecol Evol 38(3):1225–1232. https://doi.org/10.1038/s41559-019-0929-
Okabayashi N, Yasuo S, Watanabe M, Namikawa T, Ebihara S, Yoshimura T (2003) Ontogeny of circadian clock gene expression in the pineal and the suprachiasmatic nucleus of chick embryo. Brain Res 990:231–234. https://doi.org/10.1016/S0006-8993(03)03531-5
Patke A, Young MW, Axelrod S (2020) Molecular mechanisms and physiological importance of circadian rhythms. Nat Rev Mol Cell Biol 21(2):67–84. https://doi.org/10.1038/s41580-019-0179-2
Räsänen K, Kruuk LEB (2007) Maternal effects and evolution at ecological time-scales. Funct Ecol 21:408–421. https://doi.org/10.1111/j.1365-2435.2007.01246.x
Razzoli M, Karsten C, Yoder JM, Bartolomucci A, Engeland WC (2014) Chronic subordination stress phase advances adrenal and anterior pituitary clock gene rhythms. Am J Physiol Regul Integr Comp Physiol 307(2):R198–R205. https://doi.org/10.1152/ajpregu.00101.2014
Reid JM, Monaghan P, Ruxton GD (2008) The consequences of clutch size for incubation conditions and hatching success in starlings. Funct Ecol 14:560–565. https://doi.org/10.1046/j.1365-2435.2000.t01-1-00446.x
Reppert SM, Weaver DR (2003) Molecular analysis of mammalian circadian rhythms. Annu Rev Physiol 63:647–676. https://doi.org/10.1146/ANNUREV.PHYSIOL.63.1.647
Rijo-Ferreira F, Takahashi JS (2019) Genomics of circadian rhythms in health and disease. Genome Med 111(11):1–16. https://doi.org/10.1186/S13073-019-0704-0
Shama LNS, Strobel A, Mark FC, Wegner KM (2014) Transgenerational plasticity in marine sticklebacks: maternal effects mediate impacts of a warming ocean. Funct Ecol 28:1482–1493. https://doi.org/10.1111/1365-2435.12280
Shang EH, Zhdanova IV (2007) The circadian system is a target and modulator of prenatal cocaine effects. PLoS ONE 2:e587. https://doi.org/10.1371/JOURNAL.PONE.0000587
Sharda S, Zuest T, Erb M, Taborsky B (2021) Predator-induced maternal effects determine adaptive antipredator behaviors via egg composition. Proc Natl Acad Sci USA. https://doi.org/10.1073/PNAS.2017063118/-/DCSUPPLEMENTAL
Tadros W, Lipshitz HD (2009) The maternal-to-zygotic transition: a play in two acts. Development 136:3033–3042. https://doi.org/10.1242/DEV.033183
Tahara Y, Shiraishi T, Kikuchi Y, Haraguchi A, Kuriki D, Sasaki H, Motohashi H, Sakai T, Shibata S (2015) Entrainment of the mouse circadian clock by sub-acute physical and psychological stress. Sci Rep 5:11417. https://doi.org/10.1038/srep11417
Valcu C-M, Scheltema RA, Schweiggert RM, Valcu M, Teltscher K, Walther DM, Carle R, Kempenaers B (2019) Life history shapes variation in egg composition in the blue tit Cyanistes caeruleus. Commun Biol 21(2):1–14. https://doi.org/10.1038/s42003-018-0247-8
Van Dooren TJM, Hoyle RB, Plaistow SJ (2016) Maternal effects. Encycl Evol Biol. https://doi.org/10.1016/B978-0-12-800049-6.00051-2
Yun S, Lee EJ, Choe HK, Son GH, Kim K, Chung S (2020) Programming effects of maternal stress on the circadian system of adult offspring. Exp Mol Med 523(52):473–484. https://doi.org/10.1038/s12276-020-0398-9
Zeman M, Szántóová K, Herichová I (2009) Ontogeny of circadian oscillations in the heart and liver in chicken. Comp Biochem Physiol Part A Mol Integr Physiol 154:78–83
Zimmer C, Spencer KA (2014) Modifications of glucocorticoid receptors mRNA expression in the hypothalamic-pituitary-adrenal axis in response to early-life stress in female Japanese quail. J Neuroendocrinol 26:853–860. https://doi.org/10.1111/jne.12228
Zimmer C, Boogert NJ, Spencer KA (2013) Developmental programming: cumulative effects of increased pre-hatching corticosterone levels and post-hatching unpredictable food availability on physiology and behaviour in adulthood. Horm Behav 64:494–500. https://doi.org/10.1016/J.YHBEH.2013.07.002
Acknowledgements
We thank the St Mary’s Unit animal care staff for providing bird husbandry. We also thank Dr. Martin Zwaart for dissection training and use of laboratory equipment.
Funding
This work was supported by the UKRI Biotechnology and Biological Sciences Research Council (BBSRC) Eastbio Doctoral Training Programme grant number BB/M010996/1.
Author information
Authors and Affiliations
Contributions
JHC wrote the main manuscript text, conducted experimental work and analysis. All authors were involved in study design, manuscript creation. KAS and TJS provided input on statistical analysis.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare there are no financial or non-financial interests that are directly or indirectly related to the work submitted for publication.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Handling Editor: Charlotte Helfrich-Förster.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Harvey-Carroll, J., Stevenson, T.J. & Spencer, K.A. Maternal developmental history alters transfer of circadian clock genes to offspring in Japanese quail (Coturnix japonica). J Comp Physiol A 210, 399–413 (2024). https://doi.org/10.1007/s00359-023-01666-2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00359-023-01666-2