Abstract
Increasing anthropogenic pressure on coral reefs is creating an urgent need to understand how and where corals can proliferate both now and under future scenarios. Resolving environmental limits of corals has progressed through the accurate identification of corals’ ‘realised ecological niche’. Here we expand the ecological niche concept to account for corals’ ‘biogeochemical niche’ (BN), defined as the chemical space in which a coral is adapted to survive, and which is identifiable by a unique quantity and proportion of elements (termed “elementome”). BN theory has been commonly applied to other taxa, successfully predicting species distributions and stress responses by their elementomes. Here, we apply the BN theory to corals for the first time, by using dry combustion and inductively coupled plasma–mass spectrometry (ICP-MS) to determine five key macronutrients and thirteen trace elements of four diverse scleractinian coral species from the Great Barrier Reef (GBR): Acropora hyacinthus; Echinopora lamellosa; Pocillopora cf. meandrina; and Pocillopora cf. verrucosa. The elementomes were investigated in both host and Symbiodiniaceae, and the latter had the highest elemental concentrations (except molybdenum). Each coral species associated with distinct members of the genus Cladocopium (determined by ITS2 analysis) with photo-physiological data suggesting specialisation of Cladocopium functional biology. Distinct endosymbiont community structure and functioning between corals with different elementomes confirms that BN theory holds as metabolic compatibility alters across host–symbiont associations. Additional work is needed to understand the plasticity of coral elementomes, and in turn BN, over space and time to aid predictions on coral distribution and survival with environmental change.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Coral reefs are experiencing rapid environmental change driving an urgent need to understand where and how corals can flourish, both now and under future scenarios. The capacity of corals to thrive in reef systems is defined as the realised ecological niche or as the ecological space occupied by any given species where it is physiologically capable and competitively superior over other species (Hutchinson 1957; Soberón and Arroyo-Peña 2017). Whilst resolving the ecological niche for corals has been the focus of reef studies for decades (e.g. Done 1982; Georgian et al. 2014), it remains largely unknown how this translates to their corresponding biogeochemical niche (BN; sensu Peñuelas et al. 2019). The BN is the multidimensional chemical space in which an organism operates, formed by its environmental variables and ecological and functional strategies (Peñuelas et al. 2019). The BN is characterised by a unique stoichiometry of elements, often referred to as the “elementome” (proportion and abundance of all, or functionally essential, elements; Peñuelas et al. 2019) that influences a species' morphological and functional adaptations to the abiotic and biotic conditions that define their ecological niche.
Nutritional strategies (e.g. heterotrophy versus autotrophy; Houlbrèque and Ferrier‐Pagès 2009) and elemental acquisition rates are likely pivotal in diverse ecological niche exploitation by corals (Suggett et al. 2017). Such factors have been demonstrated to explain functional differences between coral species such as, growth rates of coral host and their dinoflagellate algal symbionts of the family Symbiodiniaceae (Dubinsky and Jokiel 1994; Houlbrèque and Ferrier‐Pagès 2009; Wong et al. 2021; Camp et al. 2022), ability to maintain and restore energy reserves (Houlbrèque and Ferrier‐Pagès 2009), regulation and stability of the coral host-Symbiodiniaceae symbiosis (Dubinsky and Jokiel 1994; Morris et al. 2019), and tolerance and resilience capacities to thermal stress (Ezzat et al. 2016; Gibbin et al. 2018; Morris et al. 2019). However, whether distinct BNs can be identified for corals, as evidenced by unique elementomes, remains unexplored. This is surprising given the transformative information that has been obtained for other ecosystems by exploring species’ elementomes in relation to fitness success across different environments. For example, plant physiological performance across different environments has been predicted by their elementome (Ågren and Weih 2020), with certain elements commonly acquired during stress (e.g. potassium (K+) under reactive oxygen species (ROS) accumulation; Hasanuzzaman et al. 2018). Exploration of animal elementomes has found, carbon (C) and calcium (Ca) relative to phosphorus (P) concentrations to be greatest within the shells of faster growing species of freshwater mussels (Hopper et al. 2021), and arthropod trophic groups (i.e. herbivores, detritivores and predators) have been demonstrated to have unique elementomes, determined by function and morphology (Bartrons et al. 2018). Resolving species’ elementomes and in turn their BN can help to predict the impacts of environmental change on species function and distribution and allow elemental stress diagnostics to be identified (Wang et al. 2013; Urbina et al. 2015; Peñuelas et al. 2019).
Investigating the elementome of corals (host and algal symbiont) requires describing multiple macronutrients and trace elements simultaneously. Both carbon:nitrogen (C:N) (e.g. Szmant and Gassman 1990; Tanaka et al. 2006) and nitrogen:phosphorous (N:P) (e.g. Ezzat et al. 2016; Blanckaert et al. 2020) ratios have been widely studied for corals, revealing the susceptibility of species to N and/or P limitation (Rädecker et al. 2015; Rosset et al. 2017), thereby providing evidence for distinct elemental profiles at least based on these few key macronutrients. C:N:P ratios of both the coral host and algal symbiont on the Great Barrier Reef (GBR) have also revealed that coral taxonomy is a significant driver of host-algal symbiont stoichiometric variations at the species level (Blanckaert et al. 2020). However, the elemental signature of other macronutrients (beyond C:N:P) and trace elements has received far less attention and remains almost entirely unknown for coral. Early work revealed differential uptake of key trace elements by coral (skeleton and generally tissue and symbiont combined; Amiel et al. 1973; Hanna and Muir 1990) and examined toxicological levels of physiological stress (e.g. Entsch et al. 1983). More recent work has built upon this basis identifying differences in trace metal content and uptake in symbiont species (Reich et al. 2020). However, inclusion of multiple macronutrients and trace elements is ultimately needed to accurately resolve the elementome of a species and predict their BN (Hofmann et al. 2021).
Prevailing BN theory proposes three different rules: (i) Greater taxonomic distance and evolutionary time between species increases differences between elementomes; (ii) Coexisting species, when at equilibrium, display distinct elementomes, as a strategy to minimise competitive pressure; (iii) Various strategies (e.g. elemental acquisition and storage) are used to adopt certain trade-offs which increase competitive success and result in species differences in homeostasis and plasticity across stable or fluctuating environments (Peñuelas et al. 2019). These three rules underlie the species-specific nature of the elementome and BN and is evident across plants (Sardans et al. 2015; Urbina et al. 2017), phytoplankton (Bonachela et al. 2016; Meunier et al. 2017), animals (Bartrons et al. 2018; González et al. 2018) and microbes (Lennon et al. 2012; Pereira and Berry 2017).
In this study, we tested applicability of this BN theory to corals. Four biologically diverse and important coral species (Acropora hyacinthus, Echinopora lamellosa, Pocillopora cf. meandrina and Pocillopora cf. verrucosa) were obtained from the northern GBR (Opal Reef) and analysed to resolve the elementome (five key macronutrients and thirteen trace elements) for both the coral host (tissue and mucus) and algal symbiont. Analysis of host and algal symbiont elementomes was important for this BN context as it provides evidence on the relative trading and restriction of essential nutrients between the symbiont and host (Blanckaert et al. 2020), thus providing insight into how their stoichiometries diverge. A combination of different techniques including symbiont metabarcoding, Fast Repetition Rate fluorescence (FRRf), photographic analysis, total C/N analysis and inductively coupled plasma-mass spectrometry (ICP-MS) was employed to explore the elementome and BN theory for these corals and their associated Symbiodiniaceae. In doing so, our approach captures a ‘snapshot’ of the elementome, enabling contrast of the elemental profiles for corals with overlapping ecological niche space.
Methods
Study site and sample collection
Sampling was conducted at Opal Reef (16° 12′37.62″ S, 145° 52ʹ52.752″ E, “Mojo” site 16° 12ʹ23.4″ S, 145°53ʹ53.786″ E) (GBR) in February 2020. All activities were conducted under permit G18/40023.1 (issued to EFC, DJS). Three scleractinian coral species were selected (A. hyacinthus, E. lamellosa, Pocillopora sp. based on the three criteria outlined in Peñuelas et al. 2019) to anticipate distinct elementomes and thus BNs. Specifically, they are taxonomically distinct species, coexisting, and have diverse life histories and physiologies (summarised in Supp. Table 1). After collection, visual taxonomic assessment along with returned ITS2 type profiles (see below) were used to separate Pocillopora sp. into P. meandrina and P. verrucosa, which are subsequently referred to as Pocillopora cf. meadrina and Pocillopora cf. verrucosa. A minimum of 20 samples (each ≤ 5 cm and from separate colonies) were collected for each coral genus (see Supp. Table 2) by SCUBA from 2 to 5 m depth. For each sample, a fragment was removed from the edge of a parent colony and placed in a numbered zip-lock bag containing native seawater. Once on the research vessel, the zip-lock bags were opened and maintained on deck in an aquarium with aerated native seawater prior to analysis. From each sample, a 2 mm fragment was collected for Fast Repetition Rate fluorescence (FRRf; Soliense Inc., USA). The remaining material was immediately flash frozen in liquid N2 (preservation method commonly utilised; e.g. Ferrier‐Pagès et al. 2018a; Wall et al. 2020; Ferrier-Pagès et al. 2021), to preserve the spatial distribution of elements, and stored in a dry shipper. The International Atomic Energy Agency trace metal clean procedures (IAEA 2003) were followed: all plasticwares were soaked for 20 min in 2% Micro-90 concentrated cleaning solution (Sigma-Aldrich, Castle Hill, NSW, Australia), acid-washed and soaked in 10% hydrochloric acid for 24 h and rinsed with Milli-Q water (as per Reich et al. 2020).
Fast Repetition Rate fluorescence (FRRf)
Samples were analysed using FRRf to describe photophysiological performance of the algal endosymbiont communities, using a Light Induced Fluorescence Transient FRRf (LIFT-FRRf; Soliense Inc., USA) as per Suggett et al. 2022 (see also Suggett et al. 2015). Briefly, LIFT-FRRf continuously delivered single turnover induction and relaxation sequences (excitation 445 + 505 nm) consisting of 100 flashlets of 1.6 μs at 2.5 μs and 127 flashlets of 1.6 μs (at exponentially increasing intervals over 30 ms), respectively. Irradiance of increasing intensity (0, 10, 25, 50, 100, 150, 250, 500, 750, 1000 and 1250 µmol photons m−2 s−1) was delivered by blue LEDs (445 nm) to assay the light-response dynamics. Acquisitions were recorded as the average of 10 induction sequences and then fit to a custom-designed application software package (Z. Kolber, Pers. Comm.) to retrieve physiological parameters; specifically, the minimum (F0, F´) and maximum (Fm, Fm′) fluorescence yields under light and darkness, respectively, maximum photochemical efficiency ([Fm − F0]/Fm). From these data we then determined the photosynthetic performance strategy as the photochemical (1-C, operating efficiency = [Fm′ − F′]/[Fm′ − F0′]) versus non-photochemical (1-Q = ([Fm′ − F0′]/Fm′)/[[Fm − F0]/Fm]) energy dissipation (Suggett et al. 2015). Each coral fragment was initially low light acclimated for at least 20 min in aerated tanks, after which 2 mm biopsies of intact corallite and tissue (Suggett et al. 2022) were removed and placed within the LIFT-FRRf optical chamber.
Preparation of samples for ITS2 and elemental analysis
Methods were conducted on a bench with class 100 laminar flow, following the clean laboratories and clean rooms for analysis of radionuclides and trace elements protocol (IAEA 2003). Each coral fragment was split into two for elemental and DNA analysis and initially air picked in 10 mL of buffer (0.1 N sterile TRIS buffer for elemental, phosphate-buffered saline (PBS) for DNA analysis (3 tablets/100 mL)) to remove the coral tissue from the skeleton. TRIS buffer was utilised for preparation of coral host and algal symbiont fractions for elemental analysis to prevent element migration between fractions by keeping the stability of the ionic content of the samples, therefore reducing possible movement of ions from one fraction to another via osmosis. The resulting slurry was poured into 15 mL centrifuge tubes, and wash steps to separate the host (tissue) and symbiont (algal pellet) fractions were performed.
For elemental analysis, samples were centrifuged for 5 min at 27 °C and 3500 RPM in a Rotanta 460R centrifuge (Hettich, Germany). The liquid host tissue fractions (supernatant) were collected in 15 mL centrifuge tubes. Five millilitres of sterile TRIS buffer was added to the centrifuge tubes containing the algal pellet and resuspended before repeat centrifugation. This step was further repeated to ensure the host tissue fraction was removed from the algal pellet fraction. All samples were stored at − 80 °C after processing. For ITS2 analysis, samples were centrifuged for 5 min at 4 °C and 3500 RPM and the liquid host tissue fractions were discarded. The remaining algal pellets were stored at − 80 °C.
Symbiodiniaceae DNA analysis
DNA was extracted from the algal pellet samples (see previous section) utilising the DNeasy PowerPlant Kit (Qiagen, Germany) following manufacturer’s instructions with modifications as per Camp et al. (2019) (see Supplementary Methods). The PCR amplification of the internal transcribed spacer 2 (ITS2) region was done using the primers ITSintfor2 and ITS2-reverse (Illumina sequencing adapters underlined) and following the PCR conditions of Arif et al. (2014). For individual reactions 1 µL of DNA was added to 12.5 µL Qiagen Mix, 0.4 µM each forward and reverse primer, and volume adjusted to 25 µL with DNase-free water. To visualise successful amplification, 3 µL of each PCR product was run on a 1% agarose gel for 40 min at 80 V. Additional PCR optimisation was required for some samples (Supplementary Methods). Samples were sequenced by the Australian Genome Research Facility using 250 bp paired end sequencing on a MiSeq instrument (Illumina) and then analysed utilising the SymPortal analytical framework (Hume et al. 2019).
Elemental analysis
In preparation for coral elemental analysis, frozen samples (see above preparation of samples for elemental analysis) were placed on an Alpha 2–4 LDplus freeze dryer (CHRIST, Germany) and removed once fully dried. Dry combustion with a LECO TruMac Carbon Nitrogen Analyser (LECO Castle Hill, Australia) was utilised to determine total C and N. Mean (± SE) host tissue and algal symbiont mass utilised for total C and N analysis were 42.2 ± 1.8 and 33.9 ± 2.9 mg, respectively. Manufacturer methods for soil and plant material when utilising a furnace temperature of 1200 °C were followed. A calibration standard (LECO) was used, and sample blanks and standards were utilised prior to sampling, and every 15–20 samples through a sampling run to check for instrument drift.
For proof of concept, the identification and quantification of additional macronutrients (sulphur (S) and Ca) and a suite of trace elements (cadmium (Cd), cobalt (Co), copper (Cu), iron (Fe), lead (Pb), manganese (Mn), molybdenum (Mo), nickel (Ni), selenium (Se), strontium (Sr), tin (Sn), vanadium (V) and zinc (Zn)) present in the samples were performed by acid digestion followed by ICP-MS analysis. Although Cd, Sn and Pb are non-essential elements, they were included in the analysis for being important indicators of pollution as they may bioaccumulate (Debelius et al. 2009; Bajguz 2011). Broadening the scope of elemental quantification would be of use in future studies to gain greater insight into the coral elementome. An average of 30 mg and 15 mg of host tissue and algal symbiont fraction, respectively, was used for ICP-MS analysis. Host tissue samples were digested with 200 µL of each ultra-pure HNO3 (67–69% w/w, Seastar Chemicals, BC, Canada) and H2O2 (30–32% w/w, Seastar Chemicals), whilst 100 µL of both HNO3 and H2O2 was added to each algal symbiont sample. Varying volumes were used for greater biomass tissue samples which required more acid to complete the digestion. Samples were left to digest overnight. After digestion, 4 mL and 2 mL of Milli-Q water (18.2 MΩ cm) obtained from a Sartorius 611 arium® pro water generation system (Sartorius Lab Instruments GmbH and Co. KG, Goettingen, Germany) were added to the tissue and algal symbiont samples, respectively. To avoid adsorption effects, only sample containers and filters made of polypropylene were used. For external calibration, high-purity ICP-MS calibration standards were purchased from Choice Analytical (Thornleigh, NSW, Australia) and diluted in aqueous solution of 3.4% HNO3 and 1.5% H2O2. Digested samples were analysed by flow injection analysis using an Agilent 1200 Series HPLC system coupled on-line with an Agilent 8900 series ICP-MS/MS instrument (Santa Clara, CA, USA). The injection volume was 20 μL and blanks were run periodically to ensure the absence of carry over and cross-contamination.
Data analysis
All following statistical analyses were run in RStudio Team version 4.0.2 (RStudio, PBC. Boston, MA). For all statistical tests, differences were considered significant when P-values < 0.05. Significant differences in the photochemical (1-C) and non-photochemical (1-Q) energy dissipation between the four coral species were identified by using one-way ANOVA and post hoc Tukey HSD on the last (1-C) and (1-Q) actinic light step values. Shapiro–Wilk normality test and Levene’s test of homogeneity of variance were used to check data passed ANOVA assumptions. All elemental analyses were conducted on element-to-phosphorous (E:P) ratios following Redfield ratios (Redfield 1934) as P was analysed simultaneously in ICP-MS (as per; Redfield 1934; Finkel et al. 2006; Quigg et al. 2011; Reich et al. 2020). Macronutrient to P ratios are given in mol Element (E):mol P, whilst trace element to P ratios are given in mmol E:mol P. Whilst E:P is the common reporting format for stoichiometry studies, we also analysed the absolute element concentration to ensure normalisation to P was appropriate. Absolute P was slightly elevated in the host and symbiont of P. cf. meandrina and P. cf. verrucosa (P < 0.05 for interactions of P. cf. meandrina and P. cf. verrucosa with the two other species; Supp. Table 3); however, it did not alter the observed analytical trends (Supp. Figure 1, 2; Supp. Tables 3–6). All absolute data are provided in Supplementary Materials. Outliers were removed from absolute concentration (n = 2) and E:P (n = 1) data based on: (i) If after log(x + 1) transformation outliers were identified using Mahalanobis distance (as per Reich et al. 2021) in RStudio using the rstatix package (grouped coral species separately for both host and symbiont components); (ii) If replicates had multiple (≥ 2) outliers across elements. Replicates (n = 6) where C and N samples were lost, due to low biomass, were omitted from all visual and statistical analyses.
Variations in E:P values within host tissue and algal symbiont components, across and within the four coral species, were investigated by utilising the car package to perform Kruskal–Wallis and post hoc Dunn tests (with Benjamini–Hochberg adjustment), as data did not meet the assumptions of parametric statistics. ANOVA with a post hoc Tukey HSD was used if assumptions were met. Elementomes of the four species were assessed using principal component analysis (PCA). Packages ggfortify and ggplot2 were used to perform PCA on E:P values, which were log(x + 1) transformed to homogenise variance (as per Finkel et al. 2006). As the data did not meet the assumptions of parametric tests, Kruskal–Wallis and post hoc Dunn tests (with Benjamini–Hochberg adjustment) were used to compare the extracted ordination axes (PC1 and PC2) across species. If data met assumptions, ANOVA with a post hoc Tukey HSD was used. PCA on E:P values of the algal symbiont component was additionally conducted with ITS2 type profiles for coral species with more than one ITS2 type profile (Supp. Figure 3), to determine whether ITS2 type profiles influence the algal symbiont elementome. Finally, the elementome profile of each coral (host and symbiont) was visually represented by radar plots created in RStudio using the fmsb package. Differences between host and symbiont were assessed via fold change analysis of host to symbiont E:P. Mean E:P values and host to symbiont ratios were log10 transformed to homogenise variance. Mo:P was excluded from elementome profiles and host to symbiont ratios, due to the inability to log10 transform Mo:P mean values of zero, of the algal symbiont component of P. cf. meandrina and P. cf. verrucosa (see Supp. Figure 4 for elementome profiles utilising log(x + 1) transformation on E:P means to include Mo:P).
Results
Taxonomically and functionally diverse coral-symbiont associations
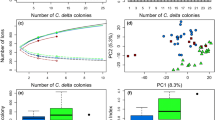
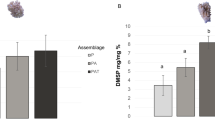
Corals often display symbiont specificity, which results in varying host–symbiont combinations. ITS2 type profiles of Symbiodiniaceae differed between coral species (Fig. 1). Symbiodiniaceae of the genus Cladocopium were prevalent: C3-C3.10-C3b-C29, C40-C3-C115 and C1d/C1/C42.2/C3-C1b-C3cg-C45c-C115k-C1au-C41p were the dominant profiles for A. hyacinthus, E. lamellosa and P. cf. verrucosa, respectively (Fig. 1). Of all species, P. cf. meandrina exhibited the greatest variance of ITS2 type profile between replicates, associating with three distinct ITS2 type profiles (Fig. 1), also of the genus Cladocopium. Different dominant ITS2 type profiles between coral species were consistent with alternate strategies of photosynthetic performance based on preferential photochemical (1-C) versus non-photochemical (1-Q) energy dissipation (Fig. 2). Whilst all four species dissipated a greater proportion of absorbed excitation energy via (1-C) than (1-Q), the extent of (1-C) varied between species (Tukey HSD, P < 0.05 for all interactions; Supp. Table 7a), except for between P. cf. meandrina and P. cf. verrucosa (Tukey HSD, P > 0.05; Supp. Table 7a). Energy dissipated via (1-Q) did not differ between E. lamellosa and both Pocillopora spp. (Tukey HSD, P > 0.05; Supp, Table 5b), whereas A. hyacinthus directed significantly more relative excitation pressure across (1-Q) compared to the other three species (Tukey HSD, P < 0.001 for all interactions with A. hyacinthus; Supp. Table 7b). Despite E. lamellosa associating with several ITS2 type profiles, distinct clustering of replicate elementomes by ITS2 type profiles was not found (Supp. Figure 3). Clustering by ITS2 type profiles was found for A. hyacinthus and P. cf. meandrina; however, further investigation is required, due to low replication (Supp. Figure 3).
Recovered ITS2 sequences and predicted ITS2 type profiles for Acropora hyacinthus, Echinopora lamellosa, Pocillopora cf. meandrina and Pocillopora cf. verrucosa. Each column represents a sample. Plotted above the horizontal black line is the relative abundance of recovered ITS2 sequences, whilst relative abundance of predicted ITS2 type profiles is below. Samples with no data recovered for either ITS2 sequence or ITS2 type profile were removed
Identification of coral host and Symbiodiniaceae species via their elementomes
Elementome (E:P; 5 macronutrients and 13 trace elements) analysis of either the coral host or algal symbiont revealed species-specific elemental signatures, despite some overlap between species (Fig. 3). Just under 50% of the host elementome variance between coral species was accounted for by the first and second principal components collectively (PC1 and PC2, respectively; Fig. 3a). PC1 accounted for 29.05% of the total elemental variance of the host component, with Sr:P (12.9%), Mn:P (12.3%), Cd:P (11.6%) and Ca:P (11.1%) contributing the largest loadings to this vector (Fig. 3a). A further 18.26% of the total elemental variance of the host component was accounted for by PC2, with Ni:P (21.7%), Co:P (17.7%), Mo:P (12.2%) and C:P (7.7%) contributing the largest loadings to this vector (Fig. 3a). No separation was identified between P. cf. meandrina and P. cf. verrucosa; however, both Pocillopora spp. separated from the other species by PC1 (P < 0.05 for interactions of P. cf. meandrina and P. cf. verrucosa with other species; Supp. Table 8a), whilst A. hyacinthus was separated from the other species by PC2 (P < 0.05 for all interactions with A. hyacinthus; Supp. Table 8a).
Principal component analysis (PCA) of the five macronutrients and thirteen trace elements normalised to P, for a host tissue (n = 15 (Ah), 20 (El), 6 (Pm), 12 (Pv)) and b algal symbiont (n = 15 (Ah), 18 (El), 6 (Pm), 10 (Pv)), comparing Acropora hyacinthus (Ah), Echinopora lamellosa (El), Pocillopora cf. meandrina (Pm) and Pocillopora cf. verrucosa (Pv)
For the algal symbiont fraction of each coral species, PC1 and PC2 accounted for over 50% of variance, with 32.49% and 22.55% by PC1 and PC2, respectively (Fig. 3b). Specifically, Co:P (13.1%), Sn:P (11.0%), Cd:P (10.9%) and N:P (10.4%) contributed the largest loadings to PC1, whilst Sr:P (15.8%), Ca:P (13.1%), Zn:P (10.5%) and Mo:P (7.9%) contributed the largest loadings to PC2 (Fig. 3b). Dunn test identified PC1 to separate A. hyacinthus algal symbionts from algal symbionts of both Pocillopora spp. (P < 0.05 for interactions of A. hyacinthus with P. cf. meandrina and P. cf. verrucosa; Supp. Table 8b), whilst the algal symbionts of E. lamellosa were separated from those of A. hyacinthus and P. cf. verrucosa by PC2 (P < 0.05 for said interactions; Supp. Table 8b).
Macronutrient content variation between coral species and host and symbiont fractions
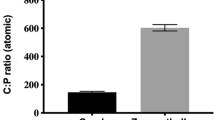
Macronutrient content varied across coral species, as well as between coral host and symbiont fractions within coral species (P < 0.05; Supp. Tables 9–12). C:N:P was highest for the algal symbiont fractions (511:65:1, 621:41:1, 271:33:1 and 279:36:1, for A. hyacinthus, E. lamellosa, P. cf. meandrina and P. cf. verrucosa, respectively), and all were significantly higher than the reported Redfield ratios of 106:16:1. Host fraction C:N:P values of 302:50:1, 186:41:1, 138:28:1 and 164:32:1, for A. hyacinthus, E. lamellosa, P. cf. meandrina and P. cf. verrucosa, respectively, were also higher than the Redfield ratios. E:P ratios revealed that all macronutrients (except for E. lamellosa and Pocillopora spp. N:P) were elevated in the algal symbiont compared to the host tissue (P < 0.05; Supp. Tables 10, 12).
Species differences in macronutrient content of host fractions were predominately driven by the significantly higher C:P and Ca:P of A. hyacinthus (302.1 ± 34.5 and 4.8 ± 0.4 mol p−1, respectively; Figs. 4, 5; P < 0.05, Supp. Tables 9a, 11a). Greater C:N of A. hyacinthus (6.3 ± 0.3 mol p−1) also caused for species differences with E. lamellosa (4.7 ± 0.2 mol p−1; Fig. 4; P < 0.05, Supp. Table 9a), and higher N:P of A. hyacinthus (49.7 ± 6.5 mol p−1) found differences with P. cf. meandrina and P. cf. verrucosa (27.8 ± 3.7 and 32.4 ± 2.3 mol p−1, respectively; Fig. 4; P < 0.05, Supp. Table 9a). E. lamellosa (5.6 ± 0.5 mol p−1) and P. cf. verrucosa (2.5 ± 0.1 mol p−1) were found to differ due to higher S:P of E. lamellosa (Fig. 5; P < 0.05, Supp. Table 11a).
Mean (± SE) elemental ratios of C:N, C:P and N:P, calculated from mol E:mol P of macronutrients, carbon (C), nitrogen (N) and phosphorus (P), present within the algal symbiont and host tissue components of coral species (Ah = Acropora hyacinthus, El = Echinopora lamellosa, Pm = Pocillopora cf. meandrina, Pv = Pocillopora cf. verrucosa) collected from Mojo, Opal Reef. Significant differences between the same components across species are indicated by different letters. Asterisks indicate significant difference between components within species. Sample sizes are as follows: algal symbiont (n = 15 (Ah), 18 (El), 6 (Pm), 10 (Pv)) and host tissue (n = 15 (Ah), 20 (El), 6 (Pm), 12 (Pv))
Mean (± SE) mol E:mol P of macronutrients, sulphur and calcium, and mmol E:mol P of trace elements, present within the algal symbiont (n = 15 (Ah), 18 (El), 6 (Pm), 10 (Pv)) and host tissue (n = 15 (Ah), 20 (El), 6 (Pm), 12 (Pv)) components of coral species (Ah = Acropora hyacinthus, El = Echinopora lamellosa, Pm = Pocillopora cf. meandrina, Pv = Pocillopora cf. verrucosa) collected from Mojo, Opal Reef. Significant differences between the same components across species are indicated by different letters. Asterisks indicate significant difference between components within species
Species differences in macronutrient content of algal symbiont fractions were predominately driven from greater C:N of E. lamellosa (16.2 ± 1.7 mol p−1) and higher N:P of A. hyacinthus (65.2 ± 5.1 mol p−1; Fig. 4; P < 0.05, Supp. Table 9b). Separation of C:P content between species was also found, caused by greater C:P of A. hyacinthus (510.9 ± 26.7 mol p−1) compared to P. cf. meandrina (271.4 ± 62.7 mol p−1) and P. cf. verrucosa (278.9 ± 9.7 mol p−1; Fig. 4; P < 0.05, Supp. Table 9b). Additionally, the algal symbiont Ca:P of E. lamellosa was found to be greater than that of P. cf. verrucosa (57.5 ± 11.9 and 10.3 ± 2.8 mol p−1, respectively; Fig. 5; P < 0.05, Supp. Table 11b). No significant differences in S:P ratios were observed between the algal symbiont fractions of species (P > 0.05, Supp. Table 11b; Fig. 5).
Trace element variation between coral species and host and symbiont fractions
Variation of trace element content between the four coral species was found for Cd:P, Cu:P, Fe:P, Mn:P, Pb:P, Sn:P, Sr:P and V:P, for the host (P < 0.05, Supp. Table 11a), and Cd:P, Mo:P, Ni:P, V:P and Zn:P, for the symbiont fractions (P < 0.05, Supp. Table 11b; Fig. 5). Trace elements with the highest content levels across all species were Sr:P and Fe:P (mean range across species 8.8–66.1 and 14.3–36.5 mmol P−1, respectively, for host; 1163.3–9402.5 and 258.5–489.2 mmol P−1, respectively, for symbiont; Fig. 5). Majority of trace element concentrations were higher for the algal symbiont than host fractions for all species (P < 0.05, Supp. Table 12; Fig. 5), with the exception of Mo:P (mean range across species of 0.1–0.4 mmol P−1 and 0–0.1 mmol P−1 for host and symbiont, respectively; Supp. Table 12; Fig. 5). In addition, E:P content of host and symbiont fractions did not significantly differ for Ni:P of E. lamellosa and Pocillopora spp. (P > 0.05, Supp. Table 12b,c,d; Fig. 5), Se:P of A. hyacinthus and P. cf. meandrina (P > 0.05, Supp. Table 12a,c; Fig. 5) and V:P of P. cf. meandrina (P > 0.05, Supp. Table 12c; Fig. 5). Overall, trace elements E:P varied between coral species and their fractions, with trace elements E:P being predominantly greatest in the algal symbiont fraction (Fig. 5).
Elementome profiles for coral host and symbiont fractions
Integration of the macronutrient and trace element contents yielded elementome profiles of each species (as host and symbiont fraction) for our sampling period (Table 1), as visualised with radar plots (Fig. 6a–h). Differences in E:P content between host and symbiont fractions are also shown between species (Fig. 6i–l). Overall, the visualisations highlight that trace elements influence the main differences between the elementome profiles of both host and symbiont species (Fig. 6). Comparison of the four coral species shows P. cf. meandrina and P. cf. verrucosa sharing the most commonalities in their elementome profiles (Fig. 6a–h). Amongst the four coral species, similarities in trends of E:P content between host and symbiont fractions can be observed; however, each species possess unique host–symbiont E:P log fold change signatures (Fig. 6i–l). The log-fold change of elements (E:P) between the host and symbiont fractions may be utilised to pinpoint which elements may be at greater risk of becoming limited to the host and how this varies with coral species. For example, the coral hosts of A. hyacinthus and the Pocillopora spp. show greater risk of S-limitation and lower risk of Fe-limitation within the host–Symbiodiniaceae relationship compared to E. lamellosa (Fig. 6i–l). The four coral species and their fractions demonstrated unique elementome profiles at a point in time from a single site, with more similarities between the taxonomically closer P. cf. meandrina and P. cf. verrucosa (Fig. 6; Supp. Table 1). Overall, host and symbiont fractions display a particular elementome profile, where the same fractions across species share more E:P similarities than comparison of host and symbiont within a species (Fig. 6a–h).
Radar plots of the elementome profiles of the a–d host and e–h algal symbiont components of Acropora hyacinthus, Echinopora lamellosa, Pocillopora cf. meandrina and Pocillopora cf. verrucosa, respectively, and their i–l host to symbiont log-fold change across five macronutrients and twelve trace elements. a–h Mean E:P and i–l fold change of mean host E:P to symbiont E:P, were log10 transformed. Mo:P was removed as values of zero could not be log10 transformed. Macronutrients and trace elements expressed as mol E:mol P and mmol E:mol P, respectively
Discussion
Understanding how corals persist over space and time remains a key goal for reef studies, yet the extent to which co-existing coral species diverge through their elemental profiles remains unexplored. Here, we define the elementome—and in turn biogeochemical niche (BN) separation—of host tissue and algal symbiont fractions of four co-occurring scleractinian coral species (A. hyacinthus, E. lamellosa, P. cf. meandrina and P. cf. verrucosa) at a single time point. We identified that species-specific differences are retained in both fractions; however, the two fractions are defined by different elementomes. Such an outcome is consistent with stoichiometries from other organisms where elementome divergence occurs via taxonomic distinctness, coexistence of species and differences in life history traits (e.g. European forest tree species, Sardans et al. 2015; dwarf shrub and grass species, Urbina et al. 2015; trophic groups from aquatic to terrestrial ecosystems, Bartrons et al. 2018; aquatic macroinvertebrates, González et al. 2018).
Unique biogeochemical niches (BNs) are identifiable
It is evident that species-specific elementomes for both host and symbiont adhere to BN theory and reflect unique BNs (sensu Peñuelas et al. 2019). A. hyacinthus, E. lamellosa, P. cf. meandrina and P. cf. verrucosa co-exist yet show distinct elementomes (Table 1), supporting this hypothesis. The elementome profiles of P. cf. meandrina and P. cf. verrucosa were most similar, likely due to both species belonging to the same genus (Peñuelas et al. 2019). Additionally, the greater similarities of the elementome profiles of the host tissue fractions of Pocillopora spp. and A. hyacinthus, compared to E. lamellosa, are likely explained by the closer taxonomic distance and shared evolutionary origins (Fig. 6; Supp. Table 1; Peñuelas et al. 2019). The distinct endosymbiont community structure (A. hyacinthus: C3-C3.10-C3b-C29, E. lamellosa: C40-C3-C115, P. cf. meandrina: ITS2 profiles with co-dominant C42.2 and C1 sequences (likely corresponding to Cladocopium latusorum sensu Turnham et al. (2021)), and P. cf. verrucosa: C1d/C1/C42.2/C3-C1b-C3cg-C45c-C115k-C1au-C41p, likely corresponding to Cladocopium pacificum sensu Turnham et al. (2021); Fig. 1) and photophysiological traits between species, also supports that the BN theory holds for host–symbiont associations, where both taxa’s BN must be metabolically compatible, and therefore in sync, to support symbiosis. In both cases, the photophysiological data suggest that our four corals represent different ‘ecotypes’ of endosymbiont community (see Suggett et al. 2015, 2022), where E. lamellosa appears relatively low light acclimated (greater reliance on (1-C) over (1-Q); Suggett et al. 2015; Nitschke et al. 2018) and A. hyacinthus relatively high light acclimated (increased reliance on (1-Q) over (1-C)). Such photoacclimation strategies may come with other life-history trade-offs (e.g. Supp. Table 1), including preferential reliance on heterotrophy to meet resource needs in low light acclimated coral (Lesser et al. 2010). However, differing physical characteristics of the host, such as skeletal and tissue structure, provide the symbionts with unique optical environments (Enríquez et al. 2017) causing variance of photosynthetic performance between coral species. Therefore, it is recommended that future studies determine the level of influence that Symbiodiniaceae (e.g. taxonomy, cell size, nutrient assimilation rates) and host (e.g. feeding and reproduction strategies, morphology, nutrient assimilation rates) traits have on elemental differences. Whilst we cannot resolve metabolic mechanisms with which elementomes—and hence BN—varies for the investigated species, the species—and fraction—specific signatures themselves raise important considerations for elemental phenotype plasticity, causing varying nutrient availability, acquisition and utilisation, as well as mechanisms involved in nutrient cycling within the host–symbiont relationship.
C:N:P ratios of the unique host and algal symbiont
Most elemental stoichiometry studies traditionally focus on three macronutrients, C, N and P, given their crucial role in growth, which are explored here in more detail for four coral species and their algal symbionts. Coral host and algal symbiont fractions of all four coral species had higher C:N:P ratios than 106:16:1 which is largely derived from phytoplankton (Redfield 1934; Ptacnik et al. 2010). These higher C:N:P ratios are attributed to the corals’ symbiotic relationship which allows for efficient nutrient cycling and uptake (Rahav et al. 1989; Dubinsky and Jokiel 1994). The algal symbiont consistently had elevated C:N:P ratios compared to the host tissue component (A. hyacinthus: 511:65:1 vs. 302:50:1; E. lamellosa: 621:41:1 vs. 186:41:1; P. cf. meandrina: 271:33:1 vs. 138:28:1; P. cf. verrucosa: 279:36:1 vs. 164:32:1) as also observed by Blanckaert et al. (2020) who reported C:N:P ratios of 603:69:1 (algal symbiont) and 146:23:1 (host tissue) for coral species within the families Merulinidae, Pocilloporidae, Poritidae and Acroporidae combined (Blanckaert et al. 2020). The C:N ratio reported by Redfield was 6.6, and this has generally been found to be tightly conserved in phytoplankton relative to ratios of C:P and N:P (Geider and La Roche 2002). However, previous work on coral has reported variable C:N, ranging from 6 to > 15 (Falkowski et al. 1993; Blanckaert et al. 2020). In our study, C:N of the host fraction varied from 4.68 ± 0.21 to 6.28 ± 0.35 for E. lamellosa and A. hyacinthus, respectively, whilst the symbiont fraction varied from 7.75 ± 0.30 to 16.23 ± 1.65 for P. cf. verrucosa and E. lamellosa, respectively. Our C:N values support recent work conducted on the GBR that show variable C:N across coral species and potential commonalities within family. For example, Blanckaert et al. (2020) found that isolated zooxanthellae from corals belonging to the family Merulinidae had the highest C:N ratios compared to other families (including Acroporidae and Pocilloporidae), a finding we corroborate here.
The N:P and C:P ratio in the marine environment can indicate whether N or P availability is limiting photosynthetic primary production (Godinot et al. 2011; Rosset et al. 2017), as they are essential nutrients for algal symbiont physiological functioning (Wiedenmann et al. 2013; D'Angelo and Wiedenmann 2014). High N:P and C:P ratios are consistent with P limitation, whilst low N:P and C:P ratios are consistent with N limitation (Godinot et al. 2011). We observed consistently high C:P and high or equal N:P for the algal symbiont compared to host fractions suggesting P limitation for the symbionts, with extent variable between taxa. This finding again supports conclusions drawn by Blanckaert et al. (2020), where P limitation was found for the symbionts, with varying degree of limitation among coral taxa.
Coral species have unique host and algal E:P
Expansion beyond C:N:P to consider the elementome (E:P; 5 macronutrients and 13 trace elements) revealed species-specific elemental signatures, despite overlap in elementome profiles, with greater elementome characteristics shared between taxonomically closer species. Our findings demonstrate that both macronutrients and trace elements were significant contributors in resolving host and algal elementomes. Also, three elements considered pollutants (Pb, Sn, Cd) were present in all coral species and within both the host and symbiont fractions. Such pollutants have been measured in corals before, such as in Goniastrea aspera tissues at both ‘polluted’ and ‘non-polluted’ sites on the GBR (Esslemont 2000). Our study found lower Pb, Sn and Cd concentrations in the host tissue of all coral species compared to the symbiont (Supp. Figure 2). This supports prior work indicating that algal symbionts have significant and unique roles in accumulating and regulating metals in corals (Jafarabadi et al. 2018). As the coral host regulates algal symbiont access to elements that are bioavailable within the external environment or obtained by host heterotrophic feeding, greater concentrations of metals in the symbionts suggest their ability to more rapidly separate and absorb metals compared to their host, thus restricting metal levels in host tissues (Reichelt-Brushett and McOrist 2003; Jafarabadi et al. 2018). Pb, Sn and Cd bioaccumulate and hence their presence at outer reef GBR sites warrants further targeted investigation that is beyond the scope of this study. It also demonstrates that efforts to understand the BN of corals need to consider the role non-essential elements (i.e. elements that do not significantly contribute to meeting nutritional demands) play in shaping corals’ biogeochemical space.
When E:P ratios of the coral host were considered, majority differed between coral species, except for Co:P, Mo:P, Ni:P, Se:P and Zn:P. The variation in E:P ratios may reflect different symbiotic partnerships that influence resource acquisition (Suggett et al. 2017; Ros et al. 2020), as well as the unique traits of coral species that impact stoichiometric requirements (Peñuelas et al. 2019). For example, the high C:N of E. lamellosa indicates elevated nutritional storage (Szmant and Gassman 1990) relative to the three other coral species. In contrast to the host fraction, E:P similarities were more prominent between algal symbiont fractions, where S:P, Co:P, Cu:P, Fe:P, Mn:P, Pb:P, Se:P, Sn:P and Sr:P ratios displayed no differences amongst species. The elevated Sr:P observed across all symbionts could not be explained and requires further study. Past studies show that strontium uptake into the coral skeleton is influenced by the algal symbionts residing in the host tissue (Cohen et al., 2002) and symbiont activity, such as photosynthesis, is responsible for up to 65% of variability in skeletal Sr:Ca (Cohen et al., 2002). In this study, calcium and strontium were significantly greatest within the algal symbiont components for each of the four coral species compared to the host tissue, which may be suggestive of the potential role of Symbiodiniaceae in the Sr:Ca ratio of coral skeletons and ultimately coral growth. Still, there is lacking information on the role of calcium and strontium within the host tissue and algal symbiont components. Minimal-to-no skeletal particulate was observed in the sample fractions and as such elemental content is not anticipated to be a bi-product of the methodology. However, this should be investigated in future studies. When grown in culture, Symbiodiniaceae species have shown significant differences in the assimilation of several trace elements (see Reich et al. 2020, 2021; Camp et al. 2022), which likely reflects the different acquisition and storage capacities of Symbiodiniaceae when ex hospite compared to in hospite. Overlap in E:P across Cladocopium assemblages is a likely driver of the host specificity we observed. Further, symbiont elementome similarities suggest occurrence of competitive exclusion that could contribute to the phenomenon that multiple profiles of one Symbiodiniaceae genera are rarely found within a single coral (Fig. 1; Hume et al. 2020). The similarity of Symbiodiniaceae E:P found here requires further targeted study, but likely reflects the ubiquitous importance of certain elements that are typically depleted in reef waters. For example, Fe has been underscored as an essential trace metal for photosynthetic organisms (Martin et al. 1991; Raven et al. 1999; Merchant 2006), as it is a cofactor for photosynthesis. As Fe is required in large amounts for Symbiodiniaceae growth, large Fe reserves are maintained by Symbiodiniaceae (Reich et al. 2020), aligning with the high Fe:P levels measured across species, compared to host tissue, in this study.
Whilst similarities in E:P content amongst species may indicate elements most essential to organism functioning, differences may highlight species-specific elemental phenotype plasticity. In this study, V:P was one of the elemental ratios with differences between coral host elementomes, showing variances across species and fraction, potentially indicative of the ability of coral-associated diazotrophs (bacteria and archaea) to express V-nitrogenases when present under Mo-deficient conditions (Rehder 2000). Although V is not as efficient as Mo as an enzymatic cofactor (Zhao et al. 2006), it can be used in antioxidative haloperoxidase enzymes (Winter and Moore 2009). Such observed differences in coral host and algal symbiont V:P are suggestive of the BN rule that variances in strategies, such as elemental uptake and utilisation, increase competitive success (Peñuelas et al. 2019).
Our findings also demonstrate potential intraspecies variation via expansive dispersion across PCA plots, particularly for E. lamellosa. Previous studies examining phenotypic plasticity have revealed that coral species that demonstrate greater heterotrophic plasticity under stress events (e.g. bleaching; Grottoli et al. 2006) or greater metabolic plasticity in response to food supply (Roder et al. 2010, 2011), display greater resilience to environmental change compared to species possessing limited heterotrophic acclimation potential (e.g. light; Ziegler et al. 2014). This may indicate that, under disturbance, E. lamellosa may be a more successful competitor compared to species with limited elemental phenotype plasticity.
Future perspectives to advance understanding of coral elemental phenotypic plasticity
Considering the BN rules proposed by Peñuelas et al. (2019), our findings of E:P variation amongst species of the coral-Symbiodiniaceae symbiosis, are suggestive of differences in nutritional strategies and plasticity within their given environment. However, methodological advancement currently lacks the ability to measure the elemental signature of bacteria and other microorganisms associated with the host and symbiont fractions, which are known to impact nutrient acquisition (Rädecker et al. 2015; Ainsworth et al. 2017; Morris et al. 2019). As work advances to understand the coral elementome and their BN, it will be important to determine the contribution of associated microorganisms (Ainsworth et al. 2017) to resolve elemental stoichiometries. This is particularly true for elements such as Mo that is an essential biological trace element, for animals, plants and nitrogen-fixing bacteria (Glass et al. 2012; Andersson et al. 2020).
As environmental change puts increasing pressure on the health and survival of coral reefs globally, it is essential to investigate the plasticity of the elementome and BN of corals. Here we establish that coral elementomes are identifiable by looking at a single point in time. However, further work is needed to resolve the plasticity of corals’ BN to understand how changing bioavailability of essential elements will impact their realised ecological niche. For example, the limitation of Mo with ocean acidification will limit N2 fixation, which supports the coral–symbiont relationship, as well as the coral holobiont via provision of limiting nitrogen (Glass et al. 2012). Additionally, changes in bioavailability of Fe, integral to biochemical functioning and acclimation ability of Symbiodiniaceae (Ferrier-Pagès et al. 2018a, b; Reich et al. 2020, 2021) under ocean acidification (Shi et al. 2010) and warmer waters (Liu and Millero 2002), make understanding coral–symbiont elemental requirements essential. Growing evidence demonstrating the interplay between trace elements for biological function (e.g. marine phytoplankton requirement of Cu for Fe uptake and co-limitation of Fe with Mn and Co; Maldonado et al. 2006; Bertrand et al. 2015; Wu et al. 2019) emphasises the need to understand the role all trace elements play to determine how these roles, and therefore biological function, may be impacted under changing ocean chemistry (Sunda and Huntsman 2000; Hoffmann et al. 2012).
Further work is needed to establish how broadly applicable BN theory is across reef locations and coral taxa and whether additional inclusion of elements can further resolve unique species-specific elementomes. Targeted studies to uncouple the taxonomic and functional contributions of both the host and symbiont fractions are also required, to better predict how changing environment will impact the niche space coral can operate. Characterisation of the coral elementome and BN under environmental gradients and stress will also further knowledge on the elemental phenotype plasticity and the key mechanisms and elements that increase coral fitness and competitive success. Such investigation may enable utilisation of the elementome as a novel measure of stress tolerance and resilience in the future.
References
Ågren GI, Weih M (2020) Multi-dimensional plant element stoichiometry—looking beyond carbon, nitrogen, and phosphorus. Front Plant Sci 11:23. https://doi.org/10.3389/fpls.2020.00023
Ainsworth T, Fordyce A, Camp E (2017) The other microeukaryotes of the coral reef microbiome. Trends Microbiol 25:980–991
Amiel AJ, Friedman GM, Miller DS (1973) Distribution and nature of incorporation of trace elements in modern aragonitic corals. Sedimentology 20:47–64
Andersson ER, Stewart JA, Work TM, Woodley CM, Schock TB, Day RD (2020) Morphological, elemental, and boron isotopic insights into pathophysiology of diseased coral growth anomalies. Sci Rep 10:8252
Arif C, Daniels C, Bayer T, Banguera-Hinestroza E, Barbrook A, Howe CJ, LaJeunesse TC, Voolstra CR (2014) Assessing Symbiodinium diversity in scleractinian corals via next-generation sequencing-based genotyping of the ITS2 rDNA region. Mol Ecol 23:4418–4433
Bajguz A (2011) Suppression of Chlorella vulgaris growth by cadmium, lead, and copper stress and its restoration by endogenous brassinolide. Arch Environ Contam Toxicol 60:406–416
Bartrons M, Sardans J, Hoekman D, Penuelas J (2018) Trophic transfer from aquatic to terrestrial ecosystems: a test of the biogeochemical niche hypothesis. Ecosphere 9:e02338
Bertrand EM, McCrow JP, Moustafa A, Zheng H, McQuaid JB, Delmont TO, Post AF, Sipler RE, Spackeen JL, Xu K (2015) Phytoplankton–bacterial interactions mediate micronutrient colimitation at the coastal Antarctic sea ice edge. Proc Natl Acad Sci 112:9938–9943
Blanckaert ACA, Reef R, Pandolfi JM, Lovelock CE (2020) Variation in the elemental stoichiometry of the coral–zooxanthellae symbiosis. Coral Reefs 39:1071–1079
Bonachela JA, Klausmeier CA, Edwards KF, Litchman E, Levin SA (2016) The role of phytoplankton diversity in the emergent oceanic stoichiometry. J Plankton Res 38:1021–1035
Camp EF, Edmondson J, Doheny A, Rumney J, Grima AJ, Huete A, Suggett DJ (2019) Mangrove lagoons of the Great Barrier Reef support coral populations persisting under extreme environmental conditions. Mar Ecol Prog Ser 625:1–14
Camp EF, Nitschke MR, Clases D, Gonzalez de Vega R, Reich HG, Goyen S, Suggett DJ (2022) Micronutrient content drives elementome variability amongst the Symbiodiniaceae. BMC Plant Biol 22(1):1-4
Cohen AL, Owens KE, Layne GD, Shimizu N (2002) The effect of algal symbionts on the accuracy of Sr/Ca paleotemperatures from coral. Science 296(5566):331–333
D’Angelo C, Wiedenmann J (2014) Impacts of nutrient enrichment on coral reefs: new perspectives and implications for coastal management and reef survival. Curr Opin Environ Sustain 7:82–93
Debelius B, Forja JM, DelValls Á, Lubián LM (2009) Toxicity and bioaccumulation of copper and lead in five marine microalgae. Ecotoxicol Environ Saf 72:1503–1513
Done T (1982) Patterns in the distribution of coral communities across the central Great Barrier Reef. Coral Reefs 1:95–107
Dubinsky Z, Jokiel PL (1994) Ratio of energy and nutrient fluxes regulates symbiosis between zooxanthellae and corals. Pac Sci 48:313–324
Enríquez S, Méndez E, Hoegh-Guldberg O, Iglesias-Prieto R (2017) Key functional role of the optical properties of coral skeletons in coral ecology and evolution. Proc R Soc b Biol Sci 284:20161667. https://doi.org/10.1098/rspb.2016.1667
Entsch B, Sim RG, Hatcher BG (1983) Indications from photosynthetic components that iron is a limiting nutrient in primary producers on coral reefs. Mar Biol 73:17–30
Esslemont G (2000) Heavy metals in seawater, marine sediments and corals from the Townsville section, Great Barrier Reef Marine Park, Queensland. Mar Chem 71:215–231
Ezzat L, Maguer JF, Grover R, Ferrier-Pages C (2016) Limited phosphorus availability is the Achilles heel of tropical reef corals in a warming ocean. Sci Rep 6:1–11
Falkowski PG, Dubinsky Z, Muscatine L, McCloskey L (1993) Population control in symbiotic corals: ammonium ions and organic materials maintain the density of zooxanthellae. Bioscience 43:606–611
Ferrier-Pagès C, Sauzeat L, Balter V (2018a) Coral bleaching is linked to the capacity of the animal host to supply essential metals to the symbionts. Glob Change Biol 24:3145–3157
Ferrier-Pagès C, Sauzéat L, Balter V (2018b) Coral bleaching is linked to the capacity of the animal host to supply essential metals to the symbionts. Glob Change Biol 24:3145–3157
Ferrier-Pagès C, Martinez S, Grover R, Cybulski J, Shemesh E, Tchernov D (2021) Tracing the trophic plasticity of the coral-dinoflagellate symbiosis using amino acid compound-specific stable isotope analysis. Microorganisms 9:182
Finkel Z, Quigg A, Raven J, Reinfelder J, Schofield O, Falkowski P (2006) Irradiance and the elemental stoichiometry of marine phytoplankton. Limnol Oceanogr 51:2690–2701
Geider R, La Roche J (2002) Redfield revisited: variability of C:N:P in marine microalgae and its biochemical basis. Eur J Phycol 37:1–17
Georgian SE, Shedd W, Cordes EE (2014) High-resolution ecological niche modelling of the cold-water coral Lophelia pertusa in the Gulf of Mexico. Mar Ecol Prog Ser 506:145–161
Gibbin EM, Krueger T, Putnam HM, Barott KL, Bodin J, Gates RD, Meibom A (2018) Short-term thermal acclimation modifies the metabolic condition of the coral holobiont. Front Mar Sci 5:10. https://doi.org/10.3389/fmars.2018.00010
Glass J, Axler R, Chandra S, Goldman C (2012) Molybdenum limitation of microbial nitrogen assimilation in aquatic ecosystems and pure cultures. Front Microbiol 3:331
Godinot C, Houlbrèque F, Grover R, Ferrier-Pagès C (2011) Coral uptake of inorganic phosphorus and nitrogen negatively affected by simultaneous changes in temperature and pH. PLoS ONE 6:e25024
González AL, Cereghino R, Dezerald O, Farjalla VF, Leroy C, Richardson BA, Richardson MJ, Romero GQ, Srivastava DS (2018) Ecological mechanisms and phylogeny shape invertebrate stoichiometry: a test using detritus-based communities across Central and South America. Funct Ecol 32:2448–2463
Grottoli AG, Rodrigues LJ, Palardy JE (2006) Heterotrophic plasticity and resilience in bleached corals. Nature 440:1186–1189
Hanna R, Muir GL (1990) Red Sea corals as biomonitors of trace metal pollution. Environ Monit Assess 14:211–222
Hasanuzzaman M, Bhuyan M, Nahar K, Hossain MS, Al Mahmud J, Hossen MS, Masud AAC, Moumita FM (2018) Potassium: a vital regulator of plant responses and tolerance to abiotic stresses. Agronomy-Basel 8:31. https://doi.org/10.3390/agronomy8030031
Hoffmann L, Breitbarth E, Boyd P, Hunter K (2012) Influence of ocean warming and acidification on trace metal biogeochemistry. Mar Ecol Prog Ser 470:191–205
Hofmann P, Clark A, Hoffmann P, Chatzinotas A, Harpole W, Dunker S (2021) Beyond nitrogen: phosphorus—estimating the minimum niche dimensionality for resource competition between phytoplankton. Ecol Lett 24:761–771
Hopper GW, Dickinson GK, Atkinson CL (2021) Associations among elements in freshwater mussel shells (Unionidae) and their relation to morphology and life history. Freshw Biol. https://doi.org/10.1111/fwb.13807
Houlbrèque F, Ferrier-Pagès C (2009) Heterotrophy in tropical scleractinian corals. Biol Rev 84:1–17
Hume BCC, Smith EG, Ziegler M, Warrington HJM, Burt JA, LaJeunesse TC, Wiedenmann J, Voolstra CR (2019) SymPortal: a novel analytical framework and platform for coral algal symbiont next-generation sequencing ITS2 profiling. Mol Ecol Resour 19:1063–1080
Hume B, Mejia Restrepo A, Voolstra CR, Berumen ML (2020) Fine-scale delineation of Symbiodiniaceae genotypes on a previously bleached central Red Sea reef system demonstrates a prevalence of coral host-specific associations. Coral Reefs 39(3):583–601
Hutchinson G (1957) Concluding Remarks. Cold Spring Harb Symp Quant Biol 22:415–427
IAEA (2003) Clean laboratories and clean rooms for analysis of radionuclides and trace elements. Austria
Jafarabadi AR, Bakhtiari AR, Maisano M, Pereira P, Cappello T (2018) First record of bioaccumulation and bioconcentration of metals in Scleractinian corals and their algal symbionts from Kharg and Lark coral reefs (Persian Gulf, Iran). Sci Total Environ 640–641:1500–1511
Lennon JT, Aanderud ZT, Lehmkuhl BK, Schoolmaster DR Jr (2012) Mapping the niche space of soil microorganisms using taxonomy and traits. Ecology 93:1867–1879
Lesser MP, Slattery M, Stat M, Ojimi M, Gates RD, Grottoli A (2010) Photoacclimatization by the coral Montastraea cavernosa in the mesophotic zone: light, food, and genetics. Ecology 91:990–1003
Liu X, Millero F (2002) The Solubility of iron in seawater. Mar Chem—MAR CHEM 77:43–54
Maldonado M, Allen A, Chong J, Lin K, Leus D, Karpenko N, Harris S (2006) Copper-dependent iron transport in coastal and oceanic diatoms. Limnol Oceanogr 51:1729–1743
Martin JH, Gordon M, Fitzwater SE (1991) The case for iron. Limnol Oceanogr 36:1793–1802
Merchant S (2006) Trace metal utilization in chloroplasts. The structure and function of plastids. pp 199–218
Meunier CL, Boersma M, El-Sabaawi R, Halvorson HM, Herstoff EM, Van de Waal DB, Vogt RJ, Litchman E (2017) From elements to function: toward unifying ecological stoichiometry and trait-based ecology. Front Environ Sci 5:18. https://doi.org/10.3389/fenvs.2017.00018
Morris LA, Voolstra CR, Quigley KM, Bourne DG, Bay LK (2019) Nutrient availability and metabolism affect the stability of coral-Symbiodiniaceae symbioses. Trends Microbiol 27:678–689
Nitschke MR, Gardner SG, Goyen S, Fujise L, Camp EF, Ralph PJ, Suggett DJ (2018) Utility of photochemical traits as diagnostics of thermal tolerance amongst Great Barrier Reef Corals. Front Mar Sci 5:45. https://doi.org/10.3389/fmars.2018.00045
Peñuelas J, Fernández-Martínez M, Ciais P, Jou D, Piao S, Obersteiner M, Vicca S, Janssens IA, Sardans J (2019) The bioelements, the elementome, and the biogeochemical niche. Ecology 100:e02652
Pereira FC, Berry D (2017) Microbial nutrient niches in the gut. Environ Microbiol 19:1366–1378
Ptacnik R, Andersen T, Tamminen T (2010) Performance of the redfield ratio and a family of nutrient limitation indicators as thresholds for phytoplankton N vs P Limitation. Ecosystems 13:1201–1214
Quigg A, Irwin AJ, Finkel ZV (2011) Evolutionary inheritance of elemental stoichiometry in phytoplankton. Proc R Soc B: Biol Sci 278:526–534
Rädecker N, Pogoreutz C, Voolstra CR, Wiedenmann J, Wild C (2015) Nitrogen cycling in corals: the key to understanding holobiont functioning? Trends Microbiol 23:490–497
Rahav O, Dubinsky Z, Achituv Y, Falkowski P (1989) Ammonium metabolism in the zooxanthellate coral, Stylophora pistillata. Proc R Soc Lond B Biol Sci 236:325–337
Raven JA, Evans MCW, Korb RE (1999) The role of trace metals in photosynthetic electron transport in O2-evolving organisms. Photosynth Res 60:111–150
Redfield AC (1934) On the proportions of organic derivatives in sea water and their relation to the composition of plankton. James Johnstone Memorial, pp 176–192
Rehder D (2000) Vanadium nitrogenase. J Inorg Biochem 80:133–136
Reich HG, Rodriguez IB, LaJeunesse TC, Ho T-Y (2020) Endosymbiotic dinoflagellates pump iron: differences in iron and other trace metal needs among the Symbiodiniaceae. Coral Reefs 39:915–927
Reich HG, Tu WC, Rodriguez IB, Chou Y, Keister EF, Kemp DW, LaJeunesse TC, Ho TY (2021) Iron availability modulates the response of endosymbiotic dinoflagellates to heat stress. J Phycol 57:3–13
Reichelt-Brushett AJ, McOrist G (2003) Trace metals in the living and nonliving components of scleractinian corals. Mar Pollut Bull 46:1573–1582
Roder C, Fillinger L, Jantzen C, Schmidt GM, Khokiattiwong S, Richter C (2010) Trophic response of corals to large amplitude internal waves. Mar Ecol Prog Ser 412:113–128
Roder C, Jantzen C, Schmidt GM, Kattner G, Phongsuwan N, Richter C (2011) Metabolic plasticity of the corals Porites lutea and Diploastrea heliopora exposed to large amplitude internal waves. Coral Reefs 30:57–69
Ros M, Camp EF, Hughes DJ, Crosswell JR, Warner ME, Leggat WP, Suggett DJ (2020) Unlocking the black-box of inorganic carbon-uptake and utilization strategies among coral endosymbionts (Symbiodiniaceae). Limnol Oceanogr 65(8):1747–1763
Rosset S, Wiedenmann J, Reed AJ, D’Angelo C (2017) Phosphate deficiency promotes coral bleaching and is reflected by the ultrastructure of symbiotic dinoflagellates. Mar Pollut Bull 118:180–187
Sardans J, Janssens IA, Alonso R, Veresoglou SD, Rillig MC, Sanders TGM, Carnicer J, Filella I, Farre-Armengol G, Penuelas J (2015) Foliar elemental composition of European forest tree species associated with evolutionary traits and present environmental and competitive conditions. Glob Ecol Biogeogr 24:240–255
Shi D, Xu Y, Hopkinson BM, Morel FM (2010) Effect of ocean acidification on iron availability to marine phytoplankton. Science 327:676–679
Soberón J, Arroyo-Peña B (2017) Are fundamental niches larger than the realized. Testing a 50-year-old prediction by Hutchinson. PLoS ONE 12:e0175138
Suggett DJ, Goyen S, Evenhuis C, Szabó M, Pettay DT, Warner ME, Ralph PJ (2015) Functional diversity of photobiological traits within the genus Symbiodinium appears to be governed by the interaction of cell size with cladal designation. New Phytol 208:370–381
Suggett DJ, Warner ME, Leggat W (2017) Symbiotic dinoflagellate functional diversity mediates coral survival under ecological crisis. Trends Ecol Evol 32:735–745
Suggett D, Nitschke M, Hughes D, Bartels N, Camp E, Dilernia N, Edmondson J, Fitzgerald S, Grima A, Sage A, Warner M (2022) Toward bio-optical phenotyping of reef-forming corals using light-induced fluorescence transient-Fast Repetition Rate fluorometry. Limnol Oceanogr: Methods. https://doi.org/10.1002/lom3.10479
Sunda WG, Huntsman SA (2000) Effect of Zn, Mn, and Fe on Cd accumulation in phytoplankton: implications for oceanic Cd cycling. Limnol Oceanogr 45:1501–1516
Szmant A, Gassman N (1990) The effects of prolonged “bleaching” on the tissue biomass and reproduction of the reef coral Montastrea annularis. Coral Reefs 8:217–224
Tanaka Y, Miyajima T, Koike I, Hayashibara T, Ogawa H (2006) Translocation and conservation of organic nitrogen within the coral-zooxanthella symbiotic system of Acropora pulchra, as demonstrated by dual isotope-labeling techniques. J Exp Mar Biol Ecol 336:110–119
Turnham KE, Wham DC, Sampayo E, LaJeunesse TC (2021) Mutualistic microalgae co-diversify with reef corals that acquire symbionts during egg development. ISME J 15:3271–3285
Urbina I, Sardans J, Beierkuhnlein C, Jentsch A, Backhaus S, Grant K, Kreyling J, Penuelas J (2015) Shifts in the elemental composition of plants during a very severe drought. Environ Exp Bot 111:63–73
Urbina I, Sardans J, Grau O, Beierkuhnlein C, Jentsch A, Kreyling J, Peñuelas J (2017) Plant community composition affects the species biogeochemical niche. Ecosphere 8:e01801
Wall CB, Kaluhiokalani M, Popp BN, Donahue MJ, Gates RD (2020) Divergent symbiont communities determine the physiology and nutrition of a reef coral across a light-availability gradient. ISME J 14:945–958
Wang M, Zheng QS, Shen QR, Guo SW (2013) The critical role of potassium in plant stress response. Int J Mol Sci 14:7370–7390
Wiedenmann J, D’Angelo C, Smith EG, Hunt AN, Legiret FE, Postle AD, Achterberg EP (2013) Nutrient enrichment can increase the susceptibility of reef corals to bleaching. Nat Clim Chang 3:160–164
Winter JM, Moore BS (2009) Exploring the chemistry and biology of vanadium-dependent haloperoxidases. J Biol Chem 284:18577–18581
Wong JC, Enriquez S, Baker DM (2021) Towards a trait-based understanding of Symbiodiniaceae nutrient acquisition strategies. Coral Reefs 40:625–639
Wu M, McCain JSP, Rowland E, Middag R, Sandgren M, Allen AE, Bertrand EM (2019) Manganese and iron deficiency in Southern Ocean Phaeocystis antarctica populations revealed through taxon-specific protein indicators. Nat Commun 10:1–10
Zhao Y, Bian S-M, Zhou H-N, Huang J-F (2006) Diversity of Nitrogenase systems in Diazotrophs. J Integr Plant Biol 48:745–755
Ziegler M, Roder CM, Büchel C, Voolstra CR (2014) Limits to physiological plasticity of the coral Pocillopora verrucosa from the central Red Sea. Coral Reefs 33:1115–1129
Acknowledgements
Thank you to John Edmondson and the crew of Wavelength Reef Cruises and Climate Change Cluster (C3) members at University of Technology Sydney (UTS), Paige Strudwick, Isabel Nunez Lendo and Lorna Howlett, for their help during fieldwork. Field-based sampling was conducted under Great Barrier Reef Marine Park Authority (GBMPRA) permits for Opal Reef (G18/40023.1 to EFC, DJS). We would also like to thank Technical Staff at UTS, particularly Sue Fenech and Helen Price for their support and guidance with sample processing. Additional thanks to Abby Aquilina for her assistance during sample processing and to Dr Hannah Reich for her feedback on the manuscript. Thank you to the anonymous reviewer and to Prof. Todd LaJeunesse who provided helpful and insightful reviews of the manuscript.
Funding
Open Access funding enabled and organized by CAUL and its Member Institutions. This research was funded by the University of Technology Sydney Chancellor’s Postdoctoral Research Fellowship and ARC Discovery Early Career Research Award (DE190100142) to EFC.
Author information
Authors and Affiliations
Contributions
EFC conceived the study with AJG, DJS and EFC designing the experiment. DJS, EFC and MRN collected the coral samples. LIFT-FRRf and carbon and nitrogen analysis were conducted by AJG, whilst all other elemental analyses were conducted by DC and RGV. LIFT-FRRf data processing was conducted by AJG, EFC, MRN and DJS. AJG conducted Symbiodiniaceae DNA extraction and PCR, with SG assisting PCR optimisation. Statistical analysis was conducted by AJG. Figures were produced by AJG. MRN and SG assisted with data analysis. AJG and EFC led writing of the manuscript, with SG providing significant edits. All authors contributed to the final version with their respective expertise.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Topic Editor Lauren T. Toth
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visithttp://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Grima, A.J., Clases, D., Gonzalez de Vega, R. et al. Species-specific elementomes for scleractinian coral hosts and their associated Symbiodiniaceae. Coral Reefs 41, 1115–1130 (2022). https://doi.org/10.1007/s00338-022-02259-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00338-022-02259-2