Abstract
Widespread coral bleaching occurred in the central Red Sea in 2010 and 2015. During both events, a cross-shelf and depth gradient of bleaching severity was identified within the Thuwal reef system, central Red Sea, Saudi Arabia. While bleaching and survival of coral taxa were monitored, neither in situ reef temperatures nor coral-associated algal communities (family Symbiodiniaceae) were characterized. Here, we determined coral host-associated Symbiodiniaceae communities and monitored temperatures along the same cross-shelf and depth gradient on six reefs in 2017 to better understand the role of these factors in the observed bleaching patterns and to generate a baseline for further studies. We characterized > 600 coral–algal associations across winter and summer in six genera of scleractinian coral (Pocillopora, Stylophora, Seriatopora, Galaxea, Gardineroseris, and Porites) and one fire coral (Family Milleporidae) using ITS2 next-generation sequencing in conjunction with the SymPortal analytical framework. We show that previous bleaching patterns correlate poorly with the largely coral host-specific structure of the 2017 Symbiodiniaceae community and are in better agreement with absolute and intraday sea water temperature variations monitored on the reefs. We demonstrate a greater distinctiveness of Symbiodiniaceae communities at the more severely bleached inshore reefs compared to those reefs further offshore. However, the potential Symbiodiniaceae community changes at these reefs prior to our sampling prevent us from evaluating this distinctiveness as determinative of the differences in bleaching severities. Based on our analyses, we discuss how fine-scale delineation of algal genotypes, including host-specific putative genotypes of Durusdinium trenchii that represent alluring targets for further taxonomic identification, corroborate a niche-adapted rather than generalist character of many coral–Symbiodiniaceae associations. In conclusion, as studies such as this one continue to build the global catalogue of coral–Symbiodiniaceae associations, we may be afforded a better oversight of how specialized coral–algal associations really are and how restricted their modification may be, both of which are critical considerations in predicting the adaptive potential of corals and the reef ecosystems they build.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Within reef-building corals, there exists a considerable variation in resistance to stressors (Rowan et al. 1997; Palumbi et al. 2014). The degree of resistance afforded to any given coral is likely determined by the animal itself in combination with the significant number of other symbiotic organisms that consider it host (Rohwer et al. 2002; Bang et al. 2018). Of particular note, it would appear that resistance of the coral holobiont (the consideration of the animal host and its associates as a single unit) is functionally linked to the genotypes of intracellular dinoflagellate algae (family Symbiodiniaceae) present in the association (Sampayo et al. 2008; LaJeunesse et al. 2014; D’Angelo et al. 2015; Silverstein et al. 2015; LaJeunesse et al. 2018). As such, the study of these associations is necessary to understand the fate of corals in an increasingly inhospitable ocean.
In particular, different genotypes of these algal symbionts may afford a greater resistance and resilience of the coral host to coral bleaching, the process in which the coral–algal symbiosis breaks down and the coral host loses a considerable proportion of its algal symbiont partners (Tchernov et al. 2004; Sampayo et al. 2008; Takahashi et al. 2009; Hume et al. 2016; Diaz-Almeyda et al. 2017; Morris et al. 2019). Given the importance of this association, considerable attention has been paid to identifying Symbiodiniaceae genotypes and associating these genotypes to particular phenotypes (Thornhill et al. 2014; Suggett et al. 2015; Smith et al. 2017).
Within the context of molecular identification, the second internal transcribed spacer (ITS2) region of the nrDNA (and to a much lesser extent the ITS1 region Hume et al. 2018a) is the most commonly used genetic marker to assess Symbiodiniaceae diversity (e.g., LaJeunesse 2001, 2002; Arif et al. 2014; Swain et al. 2016; Cunning et al. 2017; Varasteh et al. 2017; Teschima et al. 2019). This can be attributed to the marker’s ability to amplify across the entire Symbiodiniaceae taxonomic range using a single set of primers while still being able to resolve at and below the species level (LaJeunesse 2001; Smith et al. 2017; Hume et al. 2018b, 2019). The most recent analytical approaches, such as SymPortal (Hume et al. 2019), make use of the considerable intragenomic sequence diversity found within every known Symbiodiniaceae genome (Thornhill et al. 2007; Arif et al. 2014) to differentiate between and characterize putative genotypes that represent subspecies taxonomic resolutions. There is currently no other single marker available that can match this level of practicability.
In this study, we apply the SymPortal analytical framework to Symbiodiniaceae ITS2 NGS amplicon sequencing data collected from the Thuwal reef system in the central Red Sea. Reef systems such as this one, which exist in extreme thermal environments, are often the target of investigations to better understand how coral reef ecosystems will react to increasing sea surface temperatures. These corals are already living in environments that are as extreme or more extreme than the environments predicted for the majority of coral harboring waters in the years to come (Hume et al. 2013).
This particular reef system has seen considerable bleaching as part of two wider bleaching events in the central Red Sea that were putatively triggered by uncharacteristically warm SSTs in 2010 and 2015 (Furby et al. 2013; Monroe et al. 2018). Bleaching in both years was more severe on inshore reefs and at shallower depths than at reefs further from shore and at greater depths. This cross-shelf and depth gradient pattern was thought to be related to differences in water temperatures, but this hypothesis was not explicitly tested. Additionally, while the degree to which specific coral taxa underwent bleaching was characterized, no information about the Symbiodiniaceae community before, during, or after the bleaching event was collected.
To better understand to what extent Symbiodiniaceae community structure and spatial differences in temperature may have played a role in the observed bleaching patterns, and to generate a baseline for further studies in the region, here we characterize the host-associated Symbiodiniaceae community and monitor temperatures along a cross-shelf and depth gradient in both summer and winter in six genera of scleractinian coral (Pocillopora, Stylophora, Seriatopora, Galaxea, Gardineroseris, and Porites) and one fire coral (Family Milleporidae).
Materials and methods
Study area
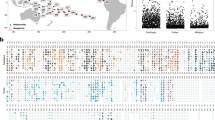
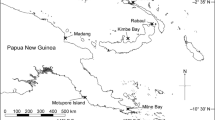
Six reefs near Thuwal, Saudi Arabia, in the central Red Sea were sampled to assess temperature and Symbiodiniaceae community variation along a depth and cross-shelf gradient (Fig. 1). Two offshore reefs, Abu Madafi (22° 05.209′ N 38° 46.700′ E) and Shi’b Nazar (22° 19.352′ N 38°51.257′ E), were located on the edge of the continental shelf, approximately 15–20 km from shore; two mid-shelf reefs, Qita al-Kirsh (22° 25.843′ N 38° 59.564′ E) and Al Fahal (22° 18.374′ N 38° 57.632′ E), were located approximately 8–9 km from shore; and two inshore reefs, Tahla (22° 17.420′ N 39°03.271′ E) and Fsar (22° 13.957 N 39° 01.816′ E), were 2–4 km away from shore. In the offshore reefs and Qita al Kirsh, loggers were deployed, and coral samples were taken at three different depths, 1–5 m, 10–15 m, and 25–30 m, while in the other reefs only the first two depths were possible due to the reef topography (Supplementary Fig. 1). All sampling and logger deployment was done on the exposed side of the reefs.
Location of the reefs monitored and sampled as part of this study. Reefs are classified as two inshore reefs (Fsar and Tahla), two mid-shelf reefs (Al Fahal and Qita al-Kirsh), and two offshore reefs (Shi’b Nazar and Abu Madafi). Latitude and longitude are given in decimal degrees. Selected reefs from the UNEP World Conservation Monitoring Centre-curated Global Distribution of Coral Reefs data set (Spalding et al. 2001; IMaRS-USF 2005; IMaRS-USF, IRD 2005; UNEP-WCMC, WorldFish Centre, WRI, TNC 2018) are highlighted in green
Sample collection
Samples for six scleractinian coral species, Pocillopora verrucosa, Stylophora pistillata, Seriatopora hystrix, Gardineroseris planulata, Galaxea fascicularis, and Porites spp., and one hydrozoan with calcareous skeleton, Millepora dichotoma, were collected during the first week of March and the last week of July 2017, at all reefs and at all depths available. Collection of all species at all sites, depths, and both seasons was not possible due to heterogeneity in species distributions (Supplementary Fig. 1). Coral species identification was done visually in the field. At each sampling location (reef, season, depth), colonies were sampled at least 5 m apart with hammer and chisel and a maximum of five colonies were collected. Samples were transported in buckets filled with seawater to the laboratory, where they were fixed in ethanol 96% the same day and stored until DNA extraction.
Temperature measurement
For each reef and available depth, one temperature logger was deployed during the first week of March 2017 to log the temperature every hour. For 1 and 15 m depths, HOBO pendant Temp/Alarm 64 K loggers were deployed, while for the 30 m depth, HOBO Water Temp Pro v2 loggers (Onset Computer Corporation, Bourne, MA) were used, since the HOBO pendant loggers were not suitable for this depth. The loggers were collected the last week of July 2017, after almost six months of recording temperature. Some of the temperature loggers were not recovered. Those were the 1 m and 15 m depths at Fsar; and the 1 m and 30 m depths at Abu Madafi. For clarity of visualization, each of the logged hourly temperature datasets was plotted as daily averages and average intraday temperature ranges were considered separately.
Information relating to degree heating weeks (DHWs) was accessed from NOAA’s Coral Reef Watch product via their National Environmental Satellite, Data, and Information Service (NOAA Corl Reef Watch 2019). Daily Global 5 km Satellite Sea Surface Temperature (a.k.a. ‘CoralTemp’) data from the Coral Reef Watch product was also downloaded for the same period of time that the temperature loggers were deployed. To assess how well these remotely sensed SST measurements represented in situ temperatures, the two were directly compared.
Statistical comparison of temperature profiles was done using a regression analysis approach with days since first measurement as a continuous independent variable (time) and temperature as a dependent variable (temp). Full details of the tests conducted can be found in the Supplementary Methods. In brief, we assessed whether Tahla had a hotter profile than the other sites either through a higher rate of temperature increase (significant difference in slope coefficients) or an overall higher profile (nonsignificant slope coefficient but significant effect of site). We did pairwise site comparisons separately for each of the two depths at which profile data were available for Tahla. An ANOVA was run on the linear model (temp ~ time*site) before conducting Tukey corrected pairwise tests. If, and only if, for any paired combination (i.e., Tahla and another site) there was a nonsignificant difference in slope coefficient, an ANCOVA was conducted using the same model to verify no interaction effect between site and time, before assessing significance of the site factor.
Using a similar approach, we additionally tested whether shallower profiles were warmer than deeper profiles, whether shallower sites had higher intraday temperature ranges (the independent variable was the intraday range), and whether the remotely sensed profiles were lower than their associated locally documented profiles. All regression-based analyses were conducted in R (R Core Team 2019). Raw temperature data and processed data used for hypothesis testing are figure creation that can be found in Supplementary Files 1–14.
DNA extraction and PCR
Symbiodiniaceae DNA was extracted from coral fragments using the Qiagen DNeasy Plant Mini Kit (Qiagen, Hilden, Germany) following the manufacturer’s protocol with minor modifications. Coral fragments were crushed using mortar and pestle and transferred to 1.5-mL Eppendorf tubes containing 200 μl of sterile glass beads (BioSpec, Bartlesville, OK) and 400 μl of Buffer AP1. Then, 4 μl of RNase A was added to each sample and they were vortexed for 1 min. The DNA extractions were continued according to manufacturer’s instructions, and the final product was analyzed using the NanoDrop 2000 Spectrophotometer (ThermoFisher Scientific, Wilmington, DE) to check the DNA concentrations and quality.
The PCR amplification of the ITS2 gene marker was performed using the primers ITSintfor2 (LaJeunesse and Trench, 2000) and ITS2-reverse (Coleman et al., 1994). For each sample, three PCRs were performed using 11 μl of Qiagen Multiplex PCR Master Mix 2X (Qiagen, Hilden, Germany), 9 μl of RNase-free water, 2 μl of each primer at 10 mM concentration, and 1 μl of DNA extract. The PCR conditions were 15 min at 95 °C, followed by 27 cycles of 94 °C for 30 s, 51 °C for 30 s, 72 °C for 30 s, and a final extension step of 10 min at 72 °C. PCR products were checked in the QIAxcel, with the DNA Screening Kit (2400) using a 3 kb marker, and the AM320 method (Qiagen, Hilden, Germany) to visualize successful amplification. For each sample, 20 μl of each PCR product was pooled together for a final volume of 60 μl used for the library preparation.
Library preparation and MiSeq ITS2 sequencing
Pooled samples were cleaned with Agencourt AMPure XP magnetic bead system (Beckman Coulter, Brea, CA). Nextera XT indexing and sequencing adapters were added via PCR (8 cycles, total PCR cycles for all samples = 35) following the manufacturer’s instructions. Samples were normalized using the SequalPrep Normalization Plate (96) Kit (Invitrogen, Frederick, Maryland, USA), and then all samples for each library were pooled together and the library was concentrated to a final volume of 100 μl using CentriVap Complete Vacuum Concentrator (Labconco, Kansas City, MO). Libraries were quantified on the Agilent Bioanalyzer 2100 (Agilent Technologies, Santa Clara, CA) and qPCR (ThermoFisher Scientific, Wilmington, DE) to check the concentration and average fragment size. The libraries were diluted to 7 pM with 15% phiX for sequencing on the Illumina MiSeq 2 × 300 bp paired-end version 3 according to the manufacturer’s specifications. Raw sequencing data are available under NCBI BioProject ID PRJNA565621 (https://www.ncbi.nlm.nih.gov/sra/PRJNA565621).
Sequencing data processing
The sequences from the Illumina MiSeq were submitted to SymPortal for quality control and analysis (SymPortal.org; Hume et al. 2019). The SymPortal analysis is conducted within a framework provided by a database that contains sequencing data for all previously run analyses. The data held within this database are used to improve each analysis’ ability to identify Symbiodiniaceae genotype-representative ITS2-type profiles (shortened to ‘profiles’ here when used repetitively). These profiles are characterized by unique combinations of intragenomic sequence variants referred to as defining intragenomic [sequence] variants (DIVs). Both sequence abundances and identities are considered in the characterization of the profiles. For more details, refer to Hume et al. (2019). For a specific example of how several closely related ITS2-type profiles are predicted and delineated in this analysis, readers may refer to Fig. 2. The framework, and outputs from the framework, were used as detailed below. The quality control pipelines and profile identification algorithms as well as the latest releases of source code can be accessed via the Zenodo platform (Hume 2019) or on GitHub.
Resolution of closely related Symbiodinium ITS2-type profiles by identifying co-occurring sets of ITS2 intragenomic sequence variants. a Six groups of samples. Each group contains multiple samples that contained the same most abundant ITS2-type profile (annotated in the bottom left of each group). For ease of annotation and discussion, each group has been arbitrarily labeled as Group1–6. Within each group, each column of stacked bars represents one sample. The bar heights within each column represent the relative proportion of a given sequence in the sample. For clarity, only Symbiodinium sequences are shown and normalized to a total proportion of 1. That is, samples may contain sequences from other Symbiodiniaceae genera. For each group, the sequences are plotted in the order of their relative abundance across the entire set of samples (i.e., all groups combined). For all of the samples in this figure, only one Symbiodinium ITS2-type profile was predicted. However, ITS2-type profiles from other genera may also have been predicted. For each group, the coral host species is given above the ITS2-type profile name. For all but Group 1, all samples within each group were from the same coral host species. For Group1, two samples were from coral species that were not Pocillopora verrucosa. These are denoted by downward facing black arrows annotated as either ‘P’ or ‘S,’ for the sample from Porites spp. or Seriatopora hystrix, respectively. b Schematic to visualize and emphasize how the presence and absence of less abundant ITS2 intragenomic sequence variants may be used to delineate closely related ITS2-type profiles. Here, exactly the same group layout is followed as in (a); however, bars in columns denote the presence or absence of a given sequence in a sample regardless of abundance. If a bar is present, the sequence was present, else a white bar has been plotted (non-visible). For each group, the sequences have been plotted in order of their relative abundance within each group. For both a and b, all sequences that were found in all samples within a group are annotated in the legend. Some rare sequences are not annotated. The ITS2-type profiles explored in this figure are annotated in Fig. 6. The comparison of Groups 1, 4, 5, and 6 represents the delineation of closely related genotypes found within different coral host species, whereas the comparisons of Groups 1, 2, and 3 represent the delineation of even more closely related genotypes found within the same coral host species
Symbiodiniaceae community composition analysis
Testing for Symbiodiniaceae community structure
To investigate whether genetic structure in the coral-harbored Symbiodiniaceae communities could be associated with the factors of this study (reef type, reef, species, depth and season), we implemented permutational analysis of variance (PERMANOVA) testing (Anderson 2001). This testing was performed on between-sample genetic distances generated in a genus-separated manner (i.e., one distance matrix per Symbiodiniaceae genus) using a UniFrac-based (Lozupone et al. 2011) methodology (Supplementary Files 15–17). Separate genus matrices were used, rather than a single across genus matrix, so that sensitivity to determine sequence similarity within Symbiodiniaceae genus was not compromised by between genera signal. These distance matrices were automatically output as part of the SymPortal pipeline with a square root transformation applied to increase the weight of lesser abundant sequences. We applied this transformation to afford lower abundance intragenomic sequence variants a resolving power closer to that of higher abundance intragenomic sequence variants while maintaining sequence abundance information. Some of the lesser abundant sequences within samples may represent lower abundance taxa (rather than lower abundance intragenomic sequence variants from the most abundant taxon). As such, the reader should be aware that this transformation could also afford a greater significance to low-level taxa in determining the similarity/dissimilarity between samples. As a reference, between-sample UniFrac-derived matrices were also output without this transformation (Supplementary Files 18–23) and the associated PERMANOVAs and tests for heteroscedasticity were applied. Results from these tests can be found in Supplementary Tables 1 and 2. So as not to overlook the biological pertinence of coral hosts harboring differing proportions of Symbiodiniaceae genera, we additionally computed a between-sample distance matrix derived from the proportion of genera (Symbiodinium, Cladocopium, Durusdinium; formerly Clade A, Clade C, and Clade D, respectively) contained in samples. This matrix (Supplementary File 24) was generated programmatically using the relative ITS2 sequence abundance count tables output from the SymPortal pipeline (Supplementary File 25) applying a square root transformation and using a Bray–Curtis methodology. The genera proportions were also plotted in the form of a ternary plot using the python package python-ternary (https://doi.org/10.5281/zenodo.34938). By working with a combination of the genus-separated distances and the genera proportion matrix, between-sample dissimilarities were essentially partitioned into within- and between-genus variation.
Two additional modifications to the above analyses were also performed in parallel. Firstly, to test what effect using a UniFrac rather than a Bray–Curtis methodology for creating between-sample distances would have, we ran all analyses conducted on the UniFrac-generated matrices on a set of Bray–Curtis-derived matrices as well (Supplementary Files 26–28). Secondly, a set of genera-separated matrices were analyzed that contained only samples that had the genus in question as their most abundant genus. For example, the Symbiodinium matrix contained only samples with a Symbiodinium sequence majority (Supplementary Files 29–31). This analysis was performed to test whether the inclusion of less abundant genus sequence collections in the matrices may be complicating the detection of patterns of association between structure in the matrices and the study’s factors. Full results for both of these tests can be found in the Supplemental Results. In brief, only the UniFrac-derived distance matrices and the matrices without sample exclusions were used in the remainder of the study primarily due to less frequent and smaller significant within-factor differences in group dispersions.
The PERMANOVAs were implemented in R using the adonis method from the vegan package (Oksanen et al. 2019) and run on pairs of distance matrices and their corresponding meta information (Supplementary Files 32–40). Variance was partitioned between five factors: reef type, reef, species, depth, and season, with the R model defined in that order. This order ensured that variance was apportioned to reef type before reef. In this manner, we could partition variation due to reef type and additional variation to reef (independent of the paired reef-type constraints; i.e., Fsar with Tahla, Qita al Kirsh with Al Fahal, and Abu Madafi with Shi’b Naxar). This order also meant that apportion of variance was biased toward reef type and reef, over species and the other factors.
We tested each of our PERMANOVA analyses to evaluate how much of the variance proportion may be due to unbalanced design and within-factor heterogeneity of group dispersions (Anderson and Walsh 2013). Full details of our testing and results can be found in the Supplemental Methods and Results. In brief, we conclude that the apportioned variances of the PERMANOVA analyses are likely robust and represent an accurate partitioning of structure in the Symbiodiniaceae distance matrices to the study’s factors.
To visualize the matrices upon which the PERMANOVA analyses were operating and to validate the results we additionally produced, plots of the principal coordinate analyses (PCoA) coordinates output by SymPortal based on the between-sample distance matrices (UniFrac-derived based on square-root-transformed abundance data; Supplementary Files 41–43, Supplementary Figs. 2–6).
Visualizing associations between Symbiodiniaceae genotypes and study factors
In addition to PERMANOVA testing, we used the predicted ITS2-type profiles output from the SymPortal analysis (e.g., Fig. 2) as proxies for Symbiodiniaceae genotypes (Supplementary File 44). To investigate specificity between these genotypes and the factors of this study, we produced a schematic to visually associate profiles with the categorical levels of the samples in which they were found. We decided to produce two versions of this schematic, one representing a collection of sample-profile instances found at higher abundances (> 0.40) and one representing a collection of associations at lower abundances (0.05 < abundance < 0.40). These thresholds were selected visually from the examined distribution of profile instance abundances across all samples (Fig. 3). The rationale behind separating the sample-profile associations into those at a higher and lower abundance was to minimize potentially misleading associations between low abundance profiles and a given sample’s classification, when that classification may largely be driven by a more abundant profile in the sample. This rationale relies on the assumption that a more abundant Symbiodiniaceae genotype is more likely to have a larger effect on host phenotype than a lesser abundant genotype. Of course, it may be that Symbiodiniaceae genotypes have a disproportional effect (disproportional to their abundance) on host phenotype and lesser abundant profiles are just as likely as abundant ones to demonstrate particular associations to host factors. To test whether more abundant profile instances were more likely to associate with a given host factor than lesser abundant instances, we calculated the average number of different groups a profile was associated with for each factor (instances associating with only one sample were discounted; testing by five separate Mann–Whitney tests to α = 0.01). In producing the lower abundance set of associations, an absolute minimum cutoff of 0.05 was used. This cutoff was put in place due to an unexpectedly large number of profiles being found at relative abundances < 0.05 (Fig. 3). Given the unexpectedly high frequency of instances at this low abundance, we chose to investigate this abundance fraction in a stand-alone manner by creating a third version of the schematic containing only the profile instances found < 0.05, referred to here as the background profile instances. This unexpectedly high number of background profiles may be caused by an increase in the prediction of profiles characterized by a smaller number of defining intragenomic variants (DIVs). To test this, we contrasted the distribution of profiles in relation to the number of DIVs they were characterized by in a collection of profiles found below and in a collection found above, a 0.05 relative abundance threshold. This contrast was visualized by means of two histograms and corresponding kernel density estimations.
Distribution of predicted ITS2-type profile relative (within sample) abundances. The left plot contains all ITS2-type profile instances. The right plot contains only those instances with a relative abundance in the samples they were predicted in above 0.05. Distributions are plotted across 50 bins as gray bars. A kernel density estimation is plotted as a black line. The black arrow indicates a relative abundance of 0.40, the value chosen for the lower boundary of the ‘high abundance’ ITS2-type profile schematic as detailed in the “Materials and methods” section
To additionally assess the ecological relevance of the phylogenetic level at which the profile predictions were being made and to assess whether more closely related profiles were more likely to be associated with similar factor levels, we incorporated between-profile distances into the schematic. To output a custom set of between-ITS2-type-profile distances concerned with only those profiles found within each abundance collection (< 0.05, 0.05-0.40, > 0.40), SymPortal’s between_type_distances_cct_set command was used. These distances were UniFrac-derived based on the square-root-transformed average abundances of the defining intragenomic variant sequences used to characterize the profiles (Supplementary Files 45–47). Dendrograms were made to represent the hierarchical clustering of these distances using the python SciPy library. Briefly, the square-form distance matrix output by SymPortal was converted to a condensed form using scipy.spatial.distance.squareform, hierarchical clustering was performed using scipy.cluster.hierarchy.linkage, and a dendrogram was produced using a modified version (to incorporate a variable leaf thickness) of scipy.cluster.hierarchy.dendrogram.
Calculating a reef-type specificity metric
As a metric for how reef-type-specific the ITS2-type profiles found at a given reef type were (i.e., not found in the other reef types), a reef-type specificity metric was calculated. This metric was the summed number of profile instances that were found at exactly the two reefs of the reef type in question, divided by the total number of profile instances found at either of the reefs in question. This metric was calculated for the high abundance cutoff set of profiles (0.40–1.00), as detailed above.
Computational resources
Scripts used to process the data and make figures can be found at the following GitHub repository: https://github.com/didillysquat/restrepo_2019.
Results
Reef thermal profiles
Averaged across all sites, the seasonal increase from March to July was approximately ~ 7 °C with minimum and maximum daily average temperatures ranging from 25.18 °C (at the Shi’b Nazar 30 m site on the 8th March, 2017) to 32.8 °C (at the Tahla 1 m site on the 21st July, 2017; Fig. 4). For those sites at which loggers were recovered, averaged daily temperature profiles generally showed a temperature decrease at corresponding depths from inshore to offshore. However, of note was inshore site Tahla that had an elevated temperature profile (Fig. 4a). This elevation was to a degree that temperatures at Tahla at 15 m depth were generally higher than those on all other sites at 1 m at the corresponding time. For all pairwise statistical comparisons of Tahla and other sites, Tahla demonstrated either a significantly faster heating rate or a warmer overall profile (Supplemental Table 3).
Average daily temperature profiles by depth (a) and reef (b). Downward-facing black arrows represent the coral sampling points in the first week of March and last week of July. Upward-facing black arrows represent the end of the approximately 2 °C drop in temperature noted across all profiles. c Average intraday range as individual reef/depth combinations and averaged across all sites at sampled depths. d Difference between 1 m in situ temperatures logged as part of this study and the Daily Global 5 km Satellite Sea Surface Temperature (a.k.a. ‘CoralTemp’) for the same day. Difference between the daily average (black line), daily maximum, and daily minimum (maximum and minimum of the gray-shaded area) of the in situ logged temperature and the remotely sensed SST are plotted
In general, the average daily temperature of profiles decreased with depth for all sites. In both of the 30 m profiles, a plateauing of average daily temperature was observed in June/July with final temperatures ending at approximately 29 °C. In contrast, a continued increase in temperature was observed into these months at the shallower depths. In all pairwise statistical comparisons for profiles across depths, shallow temperatures always demonstrated either a significantly faster heating rate or a warmer overall profile (Supplemental Table 4).
Intraday temperature was variable between and within logging locations (Fig. 4c). Intraday variations ranged from < 0.5 °C to > 2.5 °C. The 1 m sites demonstrated a higher average intraday range than the deeper sites that had similar intraday ranges. Pairwise comparisons of the intraday profiles grouped by depth showed a statistical difference in all comparisons (Supplemental Tables 5 and 6).
Locally documented temperature profiles at 1 m were higher than corresponding temperature profiles from the remotely sensed data (Fig. 4d; Supplemental Table 7) for the four sites at which these data were available (Tahla, Qita al Kirsh, Al Fahal, and Shi’b Nazar).
At all locations and depths, a dramatic drop in temperature of approximately 2 °C was observed over approximately a 5-day period in the last week of May.
Symbiodiniaceae community composition
SymPortal analysis: overview
In total, 62 million contiguous amplicon sequences were returned from the 603 successfully sequenced samples giving an average read depth of 102 k sequences per sample (Supplementary File 25). From these sequences, a total of 160 different ITS2-type profiles were predicted that represented 1507 unique sample ITS2-type profile instances (Supplementary File 44). Four of these profiles, each with a single representative instance, were from Clade F and had an average abundance of < 1% in the samples they were found in. These profiles and the Clade F sequences were not considered further in this analysis. Further summary of the SymPortal analysis output can be found in the Supplemental Results.
Symbiodiniaceae genus composition of coral samples
From the 603 samples, 552 of them contained sequences from each of Symbiodinium, Cladocopium, and Durusdinium, 47 contained sequences representing two of the genera, and only 4 samples contained sequences from a single genus. However, per sample, the majority of sequences were usually from a single genus (average abundances of the 1st, 2nd, and 3rd most abundant family in a sample were 0.92, 0.07, and 0.01), with a lesser number of samples containing a substantial mix of two genera and very few containing a close to even mix of three genera (Fig. 5). While a single genus was usually dominant, samples dominated by Cladocopium or Durusdinium had a lower number of sequences from the other genera than samples dominated by Symbiodinium. In addition, when more substantial proportions of a secondary genus were present, Symbiodinium–Durusdinium mixes were noticeably less common than those of Symbiodinium–Cladocopium or Cladocopium–Durusdinium (Fig. 5).
Resolved ITS2-type profiles
In total, 381 Symbiodinium ITS2-type profile instances were identified that represented 44 different profiles. The 44 profiles were unevenly distributed (long tail) across samples with the most abundant profiles (top 50%) representing 91% percent of the instances. For Cladocopium, 800 profile instances were predicted representing 74 different profiles where the most abundant profiles represented 85% of the instances. Finally, 322 Durusdinium profile instances were predicted that represented 38 different profiles where the most abundant profiles represented 87% of the instances. When considering only the profile instances found above a relative abundance of 0.05, Symbiodinium profiles were documented, on average, at higher relative abundances (0.75, 0.47, and 0.52 for Symbiodinium, Cladocopium, and Durusdinium, respectively). Relatedly, the average rank of a Symbiodinium profile in a sample was 1.12 compared to 1.51 and 1.59 for Cladocopium and Durusdinium, respectively.
Specificity of resolved ITS2-type profiles to study factors
After implementing the higher and lower relative abundance cutoffs (0.40–1.00 and 0.05–0.40, respectively) as used to assess the specificity of the lesser and more abundant sample-profile instances to this study’s experimental factors (Fig. 6 and Supplementary Fig. 7), 94 and 72 different ITS2-type profiles remained representing 605 and 202 sample-profile instances, respectively. The higher and lower abundance sample-profile collections produced accounted for 92.80 and 5.54% of the total sequencing reads used to characterize profiles in this study, respectively. When testing whether ITS2-type profile instances from the higher abundance collection were more likely to associate with a smaller number of groups from any given factor than those from the lower abundance collection, nonsignificant test results were returned for reef type, reef, depth, and season (p > 0.01). However, the collection of higher abundance sample-profile instances demonstrated a greater specificity to groups within the species factor (p < 0.001). For this reason, and given the large proportion of the study’s sequencing that the higher abundance collection of profile instances represents, we chose to work with the higher abundance ITS2-type profile schematic (Fig. 6) for the remainder of the study to identify associations between Symbiodiniaceae genotypes and specific host categorical groups and to validate the PERMANOVA results.
Prevalence of ITS2-type profiles (as proxies for Symbiodiniaceae genotypes) found at a within-sample relative abundance of > 0.40. The schematic is presented in a genus-separated manner. One ITS2-type profile is presented per row of the schematic. For each ITS2-type profile, its name is given followed by the number of samples it was found in enclosed in parentheses. The name of each ITS2-type profile is made up of the names of the defining intragenomic [sequence] variants (DIVs) in order of average abundance. Sequence names preceded by a ‘/’ were found as the most abundant sequence in at least one of the samples in which the ITS2-type profile was found; sequences followed by a ‘–’were not. For each ITS2-type profile, the categorical information associated with the samples it was found in is presented to the right. This categorical information is organized by factor (species, reef, reef type, depth, and season; from left to right). The full species names are: Stylophora pistillata, Seriatopora hystrix, Pocillopora verrucosa, Porites spp., Galaxea fascicularis, Gardineroseris planulata, and Millepora dichotoma. The height of bars within each column (levels of the factors) corresponds to the proportion of samples that were of this level of factor per ITS2-type profile. Therefore, the height and number of bars per ITS2-type profile row, and per categorical factor, represent the degree of specificity of a given profile to a given study factor. For example, profile A1/A1k-A1b-A1z, annotated with a black arrow, shows a high host specificity (all samples from M. dichotoma) but low reef, depth, and season specificity (found across all levels of each factor). The relatedness of the ITS2-type profiles is presented to the left of the profile names. This dendrogram is the hierarchical clustering of UniFrac-derived between-profile distances calculated from the average abundance of defining intragenomic variants for each of the profiles. Thickness of leaves correspond to the abundance of the ITS2-type profile. The delineation of six of the Symbiodinium ITS2-type profiles, denoted by asterisks, is further detailed in Fig. 2
Symbiodiniaceae community structure—genus proportion
In the PERMANOVA analysis of the genus proportion-derived sample distances, all single factors were significant except for season (Table 1). Six out of nine of the two-factor interactions were also significant, with only reef/depth, species/season, and depth/season returning nonsignificant results. The species factor was apportioned a much larger variation than any other factor (57%). 19% of variation was apportioned to residuals, while the next largest variances were associated with reef type/species and reef/species interactions (5 and 3%, respectively). This genus-specific, species-driven pattern was also observed in the predicted ITS2-type profiles with host species containing profiles primarily from a single genus or lacking profiles from a given genus. For example, M. dichotoma was dominated by Symbiodinium profiles while only Cladocopium and Durusdinium profiles were returned from G. planulata (Fig. 6).
Symbiodiniaceae community structure—species and reef-driven community structure
Results from the genus-separated, between-sample PERMANOVA testing were largely significant for each factor, and interaction between factors (Table 1; Supplementary Tables 8–11). Variance apportioned to residuals accounted for between 26 and 30% in each of the genera. While species appeared to be the primary driver in Symbiodinium and Cladocopium, reef type and reef played a larger role in Durusdinium.
Within Symbiodinium, genotypes demonstrated a species specificity with a lesser correlation to the other factors. This can be seen in Fig. 6 by the mutually exclusive association of profiles to species groups in contrast to the prevalence of most abundant profiles across multiple groups of the other factors. In the PCoA plots (Supplementary Figs. 2–6), we indeed see a clear clustering by species. However, clustering by reef type and reef are also apparent. Unlike the species clustering though, these clusters take the form of multiple clusters per level (e.g., each of the reef types cluster into multiple groups, whereas each species more commonly cluster into a single predominant group). This would indicate interaction effects in concordance with the reef type/species and reef/species interactive factor that were both apportioned 7–8% of the models variance.
Within Cladocopium, less variance was apportioned to the reef type and reef factors via interactions and rather species as a single factor was apportioned the majority of variance. Similar to Symbiodinium, the species specificity of genotypes that were otherwise distributed across multiple groups of the other factors is in concordance with this finding (Fig. 6). While the patterns of clustering in the PCoA support this relatively large species effect, some evidence for interactive effects is also apparent. For example, for the species group S. pistillata and the reef group Shi’b Nazar, multiple clusters are evident.
Within Durusdinium, the factor species was apportioned less than half of the variance that it was in either Symbiodinium or Cladocopium in the PERMANOVA analysis. By contrast, reef type and reef were apportioned more than double, and the species-reef effect was comparable. Structuring by each of these factors can be seen in the predicted ITS2-type profiles (Fig. 6). Of note, much of this signal is likely driven by the profiles associating with G. planulata at Shi’b Nazar. In the PCoA plots, the considerable overlap of species groups is in agreement with the lesser species factor effect. However, reef and reef-type stand-alone effects and interaction effects with species are difficult to identify by eye.
Across all three of the genera, species structuring was also apparent in the dissimilarity dendrograms (Fig. 6). More closely related ITS2-type profiles often associated with the same species and subclusters contained a greater proportion of profiles associated with a single or several species than would occur through random sorting. These subclusters contained a high proportion of ITS2-type profile from similar species either in the form of single predicted profiles (e.g., A1k/A1-A1b; M. dichotoma) or as multiple closely related profiles (e.g., the A1-A1c-defined profiles associated with P. verrucosa). In some cases, these profiles were evenly distributed across the other categorical factors (e.g., the C15 profiles of the Porites spp. corals), while in other, closely related, species-specific profile groupings appeared to be more specific to a particular reef (e.g., the G. planulata, Durusdinium ITS2-type profiles).
Symbiodiniaceae community structure—Stability across depth and season
Both depth and season as factors, although significant, were apportioned a very small degree of the variation from the PERMANOVA analysis (less than 2% each in every genus), suggesting that they were not substantial drivers of the structure seen in the Symbiodiniaceae community data. The PERMANOVA results are supported by both the PCoA analysis and the distribution of genotypes. For depth, little clustering is seen in the PCoA analysis and the majority of the more abundant and well-defined ITS2-type profiles (characterized by multiple DIVs) are found at multiple depths. Similar to depth, there was little clear clustering in the PCoA for this factor, and almost all abundant ITS2-type profiles were sampled in both seasons.
Characteristics of the background ITS2-type profile collection
The ITS2-type profile instances found below the 0.05 abundance cutoff accounted for 1.66% of the total sequencing reads used to characterize profiles in this study. Of the 160 profiles predicted across the whole SymPortal analysis, 76 profiles were found in common between the background and more abundant sample-profile instances (Fig. 7). However, these 76 profiles represented the majority of background sample-profile instances (1215 out of 1502 instances).
Characterization of background ITS2-type profiles. a ITS2-type profiles found in common, or not, between the background and non-background collection of ITS2-type profiles. Every predicted ITS2-type profile of the data set is plotted as a product of the number of its instances that were documented at an abundance < 0.05 (x-axis; background) or > 0.05 (y-axis; non-background). Darker points represent a larger number of ITS2-type profiles at the given coordinates. (a-inset) ITS2-type profiles found in common, or not, between the background (B) and non-background (NB) collection of ITS2-type profiles. Commonality is shown on the left for ITS2-type profiles and on the right for sample-profile instances. (bi and bii) Distribution of ITS2-type profiles as a product of the number of defining intragenomic variants (DIVs) they are characterized by in the background collection of types (< 0.05; bi) and for all other profiles (relative abundances > 0.05; bii). Histograms of the distributions are plotted and correspond to each left y-axis, while the corresponding kernel density estimations plotted as lines correspond to the right y-axes
A moderate increase in the number of profiles defined by 1 or 2 DIVs was seen in the background collection of profiles (Fig. 7) compared to profiles from the more abundant collections. This was complemented by a slight overall decrease in the relative number of profiles characterized by a higher number of DIVs in the background collection. However, in both profile collections profiles defined by 3, 4, 5, and 6 DIVs were the most common.
Finally, profiles from the background collection were associated with a greater number of groups per factor than those profiles in the more abundant collections (Fig. 6, Supplementary Fig. 8 and Supplementary Table 27). Further general characterizations of the background profiles can be found in the Supplemental Results.
Discussion
In this study, we have documented and characterized the temperature profiles and Symbiodiniaceae communities of six reefs that form part of the Thuwal reef system in the central Red Sea to better understand to what extent Symbiodiniaceae community structure and spatial differences in temperature may have played a role in previously observed bleaching patterns and to generate a baseline for further studies in the region.
Characterization of thermal profiles: correlation to bleaching severities
In 2010 and 2015, an uneven bleaching response was documented both within and across the reefs studied (Furby et al. 2013; Monroe et al. 2018). A more severe bleaching was reported at 5 versus 15 m, and reefs closest to the coast (inshore) had higher bleaching susceptibilities than those further from the shore (midshelf and offshore). In concordance with these findings, we documented a decrease in the daily average and intraday temperatures with depth and distance from shore (Fig. 4). Of particular note was the thermal profile at Tahla (both average daily and intraday) that was considerably higher than any other reef (a finding also documented in Roik et al. 2016). This higher temperature aligns with the particularly severe bleaching noted at this reef in 2010 and 2015.
The 2010 coral bleaching that was documented at the Thuwal reef system was hypothesized to be thermally driven due to the extreme degree heating weeks (DHWs) calculated for the region (between 8 and 11 DHWs, sufficient to categorize the region at Bleaching Alert Level 2, the highest risk level assigned by NOAA’s DHW charts). Yet, during the 2015 bleaching event 1 DHW was not exceeded. Monroe et al. (2018) posited that this discrepancy could be due to fine-scale differences in temperatures experienced by these inshore reefs not captured by the remotely sensed data used for the Coral Reef Watch (CRW) DHW predictions. Indeed, our comparison of the CRW SSTs with the in situ temperatures logged as part of this study (Fig. 4d) demonstrates that reefs in the system generally experience higher temperatures than the remotely sensed data would predict. This discrepancy is the largest at Tahla and could explain how this reef underwent the most severe bleaching of any documented reef without passing 0 DHW (Monroe et al. 2018).
Symbiodiniaceae community characterization: depth
The host-associated Symbiodiniaceae community of the Thuwal reef system appears to show little structuring by depth (apportioned < 2% of variation in all PERMANOVA results, although single two-factor interactions including depth were apportioned up to 4% in some instances). It would therefore seem unlikely that the Symbiodiniaceae community had a considerable influence on the greater bleaching intensity seen at the shallow depths. Indeed, if the Symbiodiniaceae community were to have had a considerable influence, a set of genotypes, specific to this depth, may be expected. However, the genotypes found at 5 m were more often found in common with the 15 m and 30 m depths than in isolation. Exceptions to this were seen in some Porites spp. samples associating with C15/C22b profiles at the shallow depth and several of the G. planulata and G. fascicularis samples associating with Durusdinium genotypes (D1/D4 profiles; putatively D. trenchii). While these exceptions may represent genuine ecological occurrences of specialization/isolation to these shallower depths, this result could be a product of the relatively low number of samples these profiles were found in. That is, a higher sampling may reveal the profiles across multiple depths. Indeed, when looking at closely related yet more abundant profiles, e.g., C15/C15r, C22b/C15-C15r, D1-D4-D6-D2b-D2a-D1d, and D1-D4-D6-D6b-D1d-D1i-D1j-D10 (Fig. 6) we can see that these profiles are found across multiple depths. Should this specialization be genuine, the thermotolerant nature of the D. trenchii- or C15 radiation-containing coral associations may be a causal factor in this structuring (LaJeunesse et al. 2003, 2014).
Symbiodiniaceae community characterization: reef type and reef
With regards to specificity to individual reefs, genotypes that make up the Symbiodiniaceae community of this reef system appear more cosmopolitan in nature than endemic. Although a relatively large number of distinct genotypes were found at only one reef type (Fig. 6), these genotypes were often predicted in only one or two samples. (A genotype found in one sample cannot be found at multiple reefs.) The most abundant and well-defined (multiple DIVs) genotypes from this study are found across all reefs and depths. Indeed, the number of samples a profile was found in was a good predictor of the number of reefs the profile was found at (simple linear regression; p = 0.00, R = 0.71, R2 = 0.49). Examples of these cosmopolitan genotypes include the A1/A1c profiles largely specific to P. verrucosa, the A1/A1k profile specific to M. dichotoma, and the C15/C15y profiles specific to Porites spp. The stark exception to this trend is the most abundant Durusdinium profile (likely a D. trenchii genotype, and with high fidelity to G. planulata) that appears only at Shi’b Nazar. The high host fidelity and the lack of cosmopolitan genotypes found in this genus likely explain the higher variance apportioned to the reef factor in Durusdinium compared to the other genera.
Interestingly, the reef-type specificity metric (indicating the distinctiveness of the Symbiodiniaceae community found at this reef type) was considerably higher for the inshore reef type than the others (0.24, 0.03, and 0.01, for inshore, midshelf, and offshore, respectively). Such distinctiveness is concordant with what would be expected if the Symbiodiniaceae community had a considerable influence on the greater bleaching intensities seen at the inshore sites, although the timing of the study complicates understanding cause versus consequence (see “Inference limitations”).
Symbiodiniaceae community characterization: species
The structure of the Thuwal reef system’s Symbiodiniaceae community appears largely coral host determined. This is evidenced by a combination of the variance apportioned in the PERMANOVA results, the clustering in the PCoA analyses, and the host-specific nature of predicted genotypes. Direct comparison between the Symbiodiniaceae community structure identified here and patterns in coral host bleaching susceptibilities during the 2010 and 2015 bleaching events are restricted due to the lack of overlap in taxa investigated between this study and those documented in the bleaching surveys (three out of the six most common coral family in 2010; three out of the six most common genera in 2015). However, at the inshore reefs in 2010 and at all reefs at 10 m in 2015, patterns in bleaching severity appeared independent of coral host: close to complete bleaching of the six most common families and even bleaching across the six most common genera. Therefore, while an effect of host-specific Symbiodiniaceae genotypes on bleaching susceptibilities cannot be definitively ruled out, it would seem unlikely that the Symbiodiniaceae community determined much of the bleaching and survival rates of these reef systems during the previous bleaching events.
Inference limitations
The time elapsed between our current characterization of coral–algal associations and the prior bleaching events limits our ability to assess to what extent the Symbiodiniaceae community may have played a determinative role in observed bleaching patters. Changes in coral community composition since the time of bleaching would likely have been the greatest precisely in those areas where bleaching was most severe. Identified correlations between the Symbiodiniaceae community structure and this study’s factors can therefore be viewed from a causal or consequential perspective. For example, of the structure identified in this analysis, perhaps the best concordance to bleaching severity was seen with the reef-type factor. However, there is no way of knowing whether the comparative distinctiveness of the Symbiodiniaceae communities seen at the inshore reefs may be a product of a higher bleaching-induced mortality rate seen on these reefs (Furby et al. 2013) or by contrast, whether an existing distinctiveness of community at these inshore reefs could have been a causal agent in the higher observed bleaching. That being said, a combination of the temporal stability documented in this study, the high host specificity of associations, and the relative similarity of the most abundant host taxa in the 2010 and 2015 surveys (Furby et al. 2013; Monroe et al. 2018) suggests that the 2017 Symbiodiniaceae community may still be representative of the community during the bleaching episodes, especially on those reefs and at depths that experienced lower mortalities. Framed within this context, the distribution of well-defined Symbiodiniaceae genotypes across all sites could be interpreted as evidence for the recolonization of the inshore reefs by coral individuals from the deeper and less severely bleached reefs. Alternatively, this homogeneity could evidence that these genotypes were already present across all reefs before the bleaching event, and thus recolonization may also be posited to have occurred via a more endogenous mode (e.g., recolonization by surviving individuals at the same location).
Within-sample genus–genus interactions
Given the wide phenotypic and genotypic diversity harbored by each of the well-sampled genera within the Symbiodiniaceae, the attribution of specific phenotypic characteristics at the genus level is generally unadvised and can be misleading (LaJeunesse et al. 2014; Thornhill et al. 2014; Hume et al. 2015). However, looking at the distributions of the three genera that are the main attention of this study, it would appear that there are some interesting genus–genus interactions occurring within samples. Samples containing considerable proportions of Symbiodinium and Durusdinium were less common than those containing Cladocopium and Symbiodinium, or Cladocopium and Durusdinium (Fig. 6). This result may be interpreted from two contrasting perspectives. On the one hand, genotypes of genera more likely to be found in combination with each other may share phenotypic traits in common and thus be suited to similar environments (e.g., coral hosts). On the other hand, these genotypes may represent contrasting phenotypic traits and thus association to several of these genotypes may be interpreted as the leverage of these different phenotypes by the coral host to acclimate to a wider range of environments or environmental variability. Considering the converse conclusions, for example, that similar phenotypes may represent redundancy, and contrasting phenotypes may represent poor suitability for a given environment, the smaller proportion of Symbiodinium–Durusdinium associations may also be explored within this theoretical framework. However, given that this study concerns only a tiny proportion of the extant coral–Symbiodiniaceae diversity, the validity of these patterns must be reassessed in a more statistically powerful context.
Characteristics of the background ITS2-type profile collection
The question of whether or not rare Symbiodiniaceae genotypes (those found at a very low abundance within their coral host) have the potential to modulate the stress resistance and/or resilience of their host has generated considerable scientific interest (Quigley et al. 2014; Lee et al. 2016; Ziegler et al. 2017). However, difficulties in identifying and characterizing this compartment (see Supplemental Discussion for further detail) mean that claims of its potential importance are largely based on theoretical grounds. However, given the unexpectedly high number of background profiles documented here, and the continued demand to address this rare compartment, it is useful to critically evaluate to what extent the SymPortal rare profiles may represent an accurate identification of genotypes.
SymPortal is primarily intended as a means for identifying dominant genotypes of Symbiodiniaceae (Hume et al. 2019), but given sufficient sequencing depth, there is no reason why it should not work in the same way on less abundant sequence collections. However, as sequencing depth decreases, profile predictions may become less accurate with a bias toward shorter profile assignments due to an inability to detect diagnostic sequences. While SymPortal implements a minimum sequence threshold under which it will not attempt profiles prediction, the slight increase in the relative number of background profile instances characterized by 1 or 2 DIVs compared to instances of the more abundant profiles documented here could indicate that sequencing depth is impairing accurate profile prediction in some cases (Fig. 7). However, the fact that the majority of profile instances are characterized by a similar number of DIVs in both profile collections, and that such a large proportion of the profile instances are found in common between the two collections, may suggest that the majority of background predictions may be as viable as the more abundant instances (Fig. 7). As such, the number of sequences upon which profile predictions are made must be taken into consideration when making any sort of biological inferences from these background collections. For further discussion on considerations when predicting low-level profiles alongside more abundant profiles of the same genus in a single sample, see the Supplemental Discussion.
Further characterization of coral host-specific Symbiodiniaceae genotypes
Our use of the SymPortal pipeline in conjunction with relatively deep NGS amplicon sequencing has enabled fine-scale resolutions of Symbiodiniaceae communities and genotypes in the Thuwal reef system. Resources to which these resolutions may be directly compared are limited (but see Gardner et al. 2019; Terraneo et al. 2019; Camp et al. 2020) given the only recent availability of such analytical approaches (approaches able to appropriately handle Symbiodiniaceae intragenomic ITS2 variability), but some genotypes may be related to previous resolutions if ITS2 intragenomic characteristics have been utilized. For example, the definitions of C. thermophilum or multiple Durusdinium species all include ITS2 intragenomic characterizations (LaJeunesse et al. 2014; Hume et al. 2015). Indeed, from this study, a single instance of a C. thermophilum genotype was found and all the predicted Durusdinium profiles contain both the D1 and D4 sequences characteristic of D. trenchii. However, considering these Durusdinium profiles further, the D6 sequence that has been employed in other Durusdinium species definitions is also widely found. As such, to determine whether the Durusdinium genotypes associated with the G. planulata, and to a lesser extent the G. fasciculari, P. verrucosa, and Porites spp. corals, represent genotypes of D. trenchii or another Durusdinium species, either defined or undefined, will necessitate the use of additional genetic markers (sensu LaJeunesse and Thornhill 2011; Lewis et al. 2019). This particular additional characterization would seem especially alluring given that D. trenchii, which is usually considered to be a host generalist (LaJeunesse et al. 2014), may be made up of many host species-specific genotypes.
In general, the collections of closely related, species-specific genotypes from Symbiodinium and Cladocopium also make interesting cases for further characterization. At present, these genotypes can only loosely be associated with the broad group of Symbiodiniaceae taxa associated with the basal ITS2 sequences A1, C1, and C15 (associated with the species definitions of Symbiodinium microadriaticum, Cladocopium goreauii and the so-called C15-radiation commonly associated with Porites spp. Trench and Blank 1987). Given the likely large number of genetically and phenotypically distinct lineages that share the A1, C1, or C15 sequence in common as their most abundant ITS2 sequence (sensu the C3 and C1 groupings in Thornhill et al. 2014; LaJeunesse 2005), assigning our fine-scale genotypes to these groupings offers little or no insight into the specific phenotypes that our resolutions may represent.
Previous assignments of closely related ITS2-type profiles (i.e., only differentiated by lesser abundant DIVs) have demonstrated agreement with subspecies delineations achieved using highly similar sequences of the hypervariable chloroplastic psbA noncoding region (psbAncr; i.e., only ~ 3 features different between the psbAncr sequences used to delineate; Smith et al. 2017; Hume et al. 2019). As such, ITS2-type profile-based resolutions may be considered comparable in some respects to psbAncr resolutions and able to delineate down to a subspecies level. Using the psbAncr, Thornhill et al. 2014 presented many genetically disparate clades (lineages) that showed a high host specificity within the ITS2 C1 and C3 radiations (although note the discussion on C. goreauii). The taxonomic level to which these lineages relate was not speculated upon. Given the host-specific structuring of the ITS2-type profile hierarchical subclusters seen in this study (Fig. 6), and the comparable nature of psbAncr and ITS2 profile resolutions, it would seem that the single abundant profiles (e.g., A1–A1x–A1r–A1u–A1g, Fig. 6; Group 6, Fig. 2), or the collections of closely related profiles from the same host species, represent ecologically relevant delineations (be they at a species or subspecies level; A1–A1c–A1h–A1q–A1a, A1–A1c–A1h–A1i, A1–A1c–A1h–A1q, A1–A1c–A1k–A1h–A1q, A1–A1c–A1h–A1cv, Fig. 6; Groups 1, 2 and 3, Fig. 2). However, continued genotyping efforts using the ITS2 marker across as a wide range of environments as possible, and in combination with additional population-level genetic characterizations, will be required to further address the currently unresolved questions that relate to these resolutions. For example, at what taxonomic levels do the highly similar genotypes relate? Do they represent phenotypically differentiated genotypes or rather fine-scale resolutions between populations of little phenotypic distinction? Perhaps most exciting, are these resolutions able to detect host specialist genotypes within Symbiodiniaceae species that are often considered host generalist?
As delineations of coral–Symbiodiniaceae associations continue to be conducted at more fine-scale resolutions, high-fidelity associations as seen here continue to be resolved (Smith et al. 2017; Lewis et al. 2019; Terraneo et al. 2019). Such characterizations evidence a considerable pool of phenotypically distinct associations upon which mechanisms of adaptation may act and build upon the considerable evidence pointing toward a prevalence of host specificity within the Symbiodiniaceae (Santos et al. 2004; Smith et al. 2017; Thornhill et al. 2017). Relatively fast mechanisms of acclimation/adaptation posited to be available to the coral host through the modulation of their algal complement (Quigley et al. 2018) may therefore have a restricted viability in those corals that exhibit such specificity (see Pettay et al. 2015).
References
Anderson MJ (2001) A new method for non-parametric multivariate analysis of variance. Austral Ecol 26:32–46
Anderson MJ, Walsh DCI (2013) PERMANOVA, ANOSIM, and the Mantel test in the face of heterogeneous dispersions: What null hypothesis are you testing? Ecol Monogr 83:557–574
Arif C, Daniels C, Bayer T, Banguera-Hinestroza E, Barbrook A, Howe CJ, Lajeunesse TC, Voolstra CR (2014) Assessing Symbiodinium diversity in scleractinian corals via next-generation sequencing-based genotyping of the ITS2 rDNA region. Mol Ecol 23:4418–4433
Bang C, Dagan T, Deines P, Dubilier N, Duschl WJ, Fraune S, Hentschel U, Hirt H, Hulter N, Lachnit T, Picazo D, Pita L, Pogoreutz C, Radecker N, Saad MM, Schmitz RA, Schulenburg H, Voolstra CR, Weiland-Brauer N, Ziegler M, Bosch TCG (2018) Metaorganisms in extreme environments: do microbes play a role in organismal adaptation? Zoology 127:1–19
Camp EF, Suggett DJ, Pogoreutz C, Nitschke MR, Houlbreque F, Hume BCC, Gardner SG, Zampighi M, Rodolfo-Metalpa R, Voolstra CR (2020) Corals exhibit distinct patterns of microbial reorganisation to thrive in an extreme inshore environment. Coral Reefs. https://doi.org/10.1007/s00338-019-01889-3
Cunning R, Gates RD, Edmunds PJ (2017) Using high-throughput sequencing of ITS2 to describe Symbiodinium metacommunities in St. John, US Virgin Islands. Peerj 5
D’Angelo C, Hume BCC, Burt J, Smith EG, Achterberg EP, Wiedenmann J (2015) Local adaptation constrains the distribution potential of heat-tolerant Symbiodinium from the Persian/Arabian Gulf. ISME J 9:2551–2560
Diaz-Almeyda EM, Prada C, Ohdera AH, Moran H, Civitello DJ, Iglesias-Prieto R, Carlo TA, LaJeunesse TC, Medina M (2017) Intraspecific and interspecific variation in thermotolerance and photoacclimation in Symbiodinium dinoflagellates. P R Soc B 284:20171767
Furby KA, Bouwmeester J, Berumen ML (2013) Susceptibility of central Red Sea corals during a major bleaching event. Coral Reefs 32:505–513
Gardner SG, Camp EF, Smith DJ, Kahlke T, Osman EO, Gendron G, Hume BCC, Pogoreutz C, Voolstra CR, Suggett DJ (2019) Coral microbiome diversity reflects mass coral bleaching susceptibility during the 2016 El Nino heat wave. Ecol Evol 9:938–956
Hume B, D’Angelo C, Burt J, Baker AC, Riegl B, Wiedenmann J (2013) Corals from the Persian/Arabian Gulf as models for thermotolerant reef-builders: prevalence of clade C3 Symbiodinium, host fluorescence and ex situ temperature tolerance. Mar Pollut Bull 72:313–322
Hume BCC (2019) SymPortal. Zenodo 10.5281/zenodo.2552177
Hume BCC, D’Angelo C, Burt JA, Wiedenmann J (2018a) Fine-scale biogeographical boundary delineation and sub-population resolution in the Symbiodinium thermophilum coral symbiont group from the Persian/Arabian Gulf and Gulf of Oman. Front Mar Sci 5:138
Hume BCC, D’Angelo C, Smith EG, Stevens JR, Burt J, Wiedenmann J (2015) Symbiodinium thermophilum sp nov., a thermotolerant symbiotic alga prevalent in corals of the world’s hottest sea, the Persian/Arabian Gulf. Sci Rep-Uk 5:1–8
Hume BCC, Voolstra CR, Arif C, D’Angelo C, Burt JA, Eyal G, Loya Y, Wiedenmann J (2016) Ancestral genetic diversity associated with the rapid spread of stress-tolerant coral symbionts in response to Holocene climate change. P Natl Acad Sci USA 113:4416–4421
Hume BCC, Smith EG, Ziegler M, Warrington HJM, Burt JA, LaJeunesse TC, Wiedenmann J, Voolstra CR (2019) SymPortal: a novel analytical framework and platform for coral algal symbiont next-generation sequencing ITS2 profiling. Mol Ecol Res 19:1063–1080
Hume BCC, Ziegler M, Poulain J, Pochon X, Romac S, Boissin E, de Vargas C, Planes S, Wincker P, Voolstra CR (2018b) An improved primer set and amplification protocol with increased specificity and sensitivity targeting the Symbiodinium ITS2 region. Peerj 6:e4816
IMaRS-USF, IRD (Institut de Recherche pour le Developpement) (2005) Millennium Coral Reef Mapping Project. Validated maps. Cambridge (UK): UNEP World Conservation Monitoring Centre
IMaRS-USF (Institute for Marine Remote Sensing-University of South Florida) (2005) Millennium Coral Reef Mapping Project. Unvalidated maps. These maps are unendorsed by IRD, but were further interpreted by UNEP World Conservation Monitoring Centre. Cambridge (UK): UNEP World Conservation Monitoring Centre
LaJeunesse TC (2001) Investigating the biodiversity, ecology, and phylogeny of endosymbiotic dinoflagellates in the genus Symbiodinium using the its region: in search of a “species” level marker. J Phycol 37:866–880
LaJeunesse TC (2002) Diversity and community structure of symbiotic dinoflagellates from Caribbean coral reefs. Mar Biol 141:387–400
LaJeunesse TC (2005) “Species” radiations of symbiotic dinoflagellates in the Atlantic and Indo-Pacific since the miocene-pliocene transition. Mol Biol Evol 22:570–581
LaJeunesse TC, Thornhill DJ (2011) Improved resolution of reef-coral endosymbiont (Symbiodinium) species diversity, ecology, and evolution through psbA non-coding region genotyping. Plos One 6:e29013
LaJeunesse TC, Loh WKW, van Woesik R, Hoegh-Guldberg O, Schmidt GW, Fitt WK (2003) Low symbiont diversity in southern Great Barrier Reef corals, relative to those of the Caribbean. Limnol Oceanogr 48:2046–2054
LaJeunesse TC, Wham DC, Pettay DT, Parkinson JE, Keshavmurthy S, Chen CA (2014) Ecologically differentiated stress-tolerant endosymbionts in the dinoflagellate genus Symbiodinium (Dinophyceae) Clade D are different species. Phycologia 53:305–319
LaJeunesse TC, Parkinson JE, Gabrielson PW, Jin Jeong H, Reimer JD, Voolstra CR, Santos SR (2018) Systematic revision of Symbiodiniaceae highlights the antiquity and diversity of coral endosymbionts. Curr Biol 28:2570–2580
Lee MJ, Jeong HJ, Jang SH, Lee SY, Kang NS, Lee KH, Kim HS, Wham DC, LaJeunesse TC (2016) Most low-abundance “background” Symbiodinium spp. are transitory and have minimal functional significance for symbiotic corals. Microb Ecol 71:771–783
Lewis AM, Chan AN, LaJeunesse TC (2019) New species of closely related endosymbiotic dinoflagellates in the greater Caribbean have niches corresponding to host coral phylogeny. J Eukaryot Microbiol 66:469–482
Lozupone C, Lladser ME, Knights D, Stombaugh J, Knight R (2011) UniFrac: an effective distance metric for microbial community comparison. ISME J 5:169–172
Monroe AA, Ziegler M, Roik A, Rothig T, Hardenstine RS, Emms MA, Jensen T, Voolstra CR, Berumen ML (2018) In situ observations of coral bleaching in the central Saudi Arabian Red Sea during the 2015/2016 global coral bleaching event. Plos One 13:e0195814
Morris LA, Voolstra CR, Quigley KM, Bourne DG, Bay LK (2019) Nutrient availability and metabolism affect the stability of coral-Symbiodiniaceae symbioses. Trends Microbiol 27:678–689
NOAA Corl Reef Watch NCR (2019) NOAA Coral Reef Watch Version 3.1 Daily Global 5-km Satellite Coral Bleaching Degree Heating Week Product. NOAA Coral Reef Watch, College Park, Maryland, USA
Oksanen J, Blanchet FG, Friendly M, Kindt R, Legendre P, McGlinn D, Minchin PR, O’Hara RB, Simpson GL, Solymos P, Stevens MHH, Szoecs E, Wagner H (2019) vegan: Community Ecology PackageR package version 2.5-5
Palumbi SR, Barshis DJ, Traylor-Knowles N, Bay RA (2014) Mechanisms of reef coral resistance to future climate change. Science 344:895–898
Pettay DT, Wham DC, Smith RT, Iglesias-Prieto R, LaJeunesse TC (2015) Microbial invasion of the Caribbean by an Indo-Pacific coral zooxanthella. Proc Natl Acad Sci U S A 112:7513–7518
Quigley KM, Bay LK, Willis BL (2018) Leveraging new knowledge of Symbiodinium community regulation in corals for conservation and reef restoration. Mar Ecol Prog Ser 600:245–253
Quigley KM, Davies SW, Kenkel CD, Willis BL, Matz MV, Bay LK (2014) Deep-sequencing method for quantifying background abundances of Symbiodinium types: Exploring the rare Symbiodinium biosphere in reef-building corals. Plos One 9:e94297
R Core Team (2019). R: A language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria. URL https://www.R-project.org/
Rohwer F, Seguritan V, Azam F, Knowlton N (2002) Diversity and distribution of coral-associated bacteria. Mar Ecol Prog Ser 243:1–10
Roik A, Rothig T, Roder C, Ziegler M, Kremb SG, Voolstra CR (2016) Year-long monitoring of physico-chemical and biological variables provide a comparative baseline of coral reef functioning in the Central Red Sea. Plos One 11:e0163939
Rowan R, Knowlton N, Baker A, Jara J (1997) Landscape ecology of algal symbionts creates variation in episodes of coral bleaching. Nature 388:265–269
Sampayo EM, Ridgway T, Bongaerts P, Hoegh-Guldberg O (2008) Bleaching susceptibility and mortality of corals are determined by fine-scale differences in symbiont type. P Natl Acad Sci USA 105:10444–10449
Santos SR, Shearer TL, Hannes AR, Coffroth MA (2004) Fine-scale diversity and specificity in the most prevalent lineage of symbiotic dinoflagellates (Symbiodinium, Dinophyceae) of the Caribbean. Mol Ecol 13:459–469
Silverstein RN, Cunning R, Baker AC (2015) Change in algal symbiont communities after bleaching, not prior heat exposure, increases heat tolerance of reef corals. Global Change Biol 21:236–249
Smith EG, Ketchum RN, Burt JA (2017) Host specificity of Symbiodinium variants revealed by an ITS2 metahaplotype approach. ISME J 11:1500–1503
Spalding MD, Ravilious C, Green EP (2001) World Atlas of Coral Reefs. The University of California Press, Berkeley
Suggett DJ, Goyen S, Evenhuis C, Szabo M, Pettay DT, Warner ME, Ralph PJ (2015) Functional diversity of photobiological traits within the genus Symbiodinium appears to be governed by the interaction of cell size with cladal designation. New Phytol 208:370–381
Swain TD, Chandler J, Backman V, Marcelino L (2016) Consensus thermotolerance ranking for 110 Symbiodinium phylotypes: an exemplar utilization of a novel iterative partial-rank aggregation tool with broad application potential. Funct Ecol 31:172–183
Takahashi S, Whitney SM, Badger MR (2009) Different thermal sensitivity of the repair of photodamaged photosynthetic machinery in cultured Symbiodinium species. P Natl Acad Sci USA 106:3237–3242
Tchernov D, Gorbunov MY, de Vargas C, Yadav SN, Milligan AJ, Haggblom M, Falkowski PG (2004) Membrane lipids of symbiotic algae are diagnostic of sensitivity to thermal bleaching in corals. P Natl Acad Sci USA 101:13531–13535
Terraneo TI, Fusi M, Hume BCC, Arrigoni R, Voolstra CR, Benzoni F, Forsman ZH, Berumen ML (2019) Environmental latitudinal gradients and host-specificity shape Symbiodiniaceae distribution in Red Sea Porites corals. J Biogeogr 46(10):2323–2335
Teschima MM, Garrido A, Paris A, Nunes FLD, Zilberberg C (2019) Biogeography of the endosymbiotic dinoflagellates (Symbiodiniaceae) community associated with the brooding coral Favia gravida in the Atlantic Ocean. Plos One 14:e0215167
Thornhill DJ, Lajeunesse TC, Santos SR (2007) Measuring rDNA diversity in eukaryotic microbial systems: how intragenomic variation, pseudogenes, and PCR artifacts confound biodiversity estimates. Mol Ecol 16:5326–5340
Thornhill DJ, Lewis AM, Wham DC, LaJeunesse TC (2014) Host-specialist lineages dominate the adaptive radiation of reef coral endosymbionts. Evolution 68:352–367
Trench RK, Blank RJ (1987) Symbiodinium-Microadriaticum Freudenthal, Symbiodinium-Goreauii Sp-Nov, Symbiodinium-Kawagutii Sp-Nov and Symbiodinium-Pilosum Sp-Nov - Gymnodinioid Dinoflagellate symbionts of marine-invertebrates. J Phycol 23:469–481
UNEP-WCMC, WorldFish Centre, WRI, TNC (2018) Global distribution of coral reefs, compiled from multiple sources including the Millennium Coral Reef Mapping Project. Version 4.0, updated by UNEP-WCMC. Includes contributions from IMaRS- USF and IRD (2005), IMaRS-USF (2005) and Spalding et al. (2001). Cambridge (UK): UNEP World Conservation Monitoring Centre
Varasteh T, Shokri MR, Rajabi-Maham H, Behzadi S, Hume BC (2017) Symbiodinium thermophilum symbionts in Porites harrisoni and Cyphastrea microphthalma in the northern Persian Gulf, Iran. J Mar Biol Assoc United Kingdom 98:2067–2073
Ziegler M, Eguíluz VM, Duarte CM, Voolstra CR (2017) Rare symbionts may contribute to the resilience of coral–algal assemblages. ISME J 12:161–172
Acknowledgements
We would like to thank Roberto Arrigoni, Darren Cocker, Tullia Terraneo, Rodrigo Villalobos, Michael Wooster, Sara Wilson, and Emily Troyer for their assistance with laboratory and field work; the KAUST Bioscience Core Laboratory for assistance with sequencing; and the KAUST Coastal and Marine Resources Core Laboratory for assistance with field work. Funding for this project was provided by the King Abdullah University of Science and Technology (baseline funds to M.L.B. and C.R.V.).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
On behalf of all authors, the corresponding author states that there is no conflict of interest.
Additional information
Topic Editor Simon Davy
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Hume, B.C.C., Mejia-Restrepo, A., Voolstra, C.R. et al. Fine-scale delineation of Symbiodiniaceae genotypes on a previously bleached central Red Sea reef system demonstrates a prevalence of coral host-specific associations. Coral Reefs 39, 583–601 (2020). https://doi.org/10.1007/s00338-020-01917-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00338-020-01917-7