Abstract
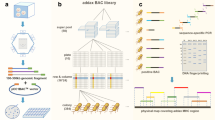
A goat Bacterial Artificial Chromosome (BAC) library of 61,440 independent clones was constructed and characterized. The average size of the inserts was estimated at 153 kilobases by analyzing almost 500 clones using Not1 digestion followed by FIGE (Field Inverted Gel Electrophoresis) analysis. The library represents about three genome equivalents, which yields a theoretical probability of 0.95 of isolating a particular DNA sequence. After individual growth, the clones were arrayed in 40 superpools, which were organized in three dimension pools. A rapid technique for pool DNA preparation by microwave treatment was set up. This technique was compatible with PCR analysis. Primer pairs from 166 sequences (microsatellites, coding sequences from goat, and conserved Expressed Sequence Tags (ESTs) from humans) enabled the library to be successfully searched in 165 cases, with an average of 3.52 positive superpools. Only one sequence could not be found. The degree of chimerism was evaluated by FISH analysis with DNA from over 110 clones and was estimated at 4%. This BAC library will constitute an invaluable tool for positional cloning in ruminants, as well as for more general comparative mapping studies in mammals.
Similar content being viewed by others
References
Andersson L, Archibald A, Ashburner M, Audun S, Barendse W, Bitgood J, Bottema C, Broad T, Brown S, Burt D, Charlier C, Copeland N, Davis S, Davisson M, Edwards J, Eggen A, Elgar G, Eppig JT, Franklin I, Grewe P, Gill T, Graves JAM, Hawken R, Hetzel J, Hilyard A, Jacob H, Jaswinska L, Jenkins N, Kunz H, Levan G, Lie O, Lyons L, Maccarone P, Mellersh C, Montgomery G, Moore S, Moran C, Morizot D, Neff M, Nicholas F, Obrien S, Parsons Y, Peters J, Postlethwait J, Raymond M, Rothschild M, Schook L, Sugimoto Y, Szpirer C, et al. (1996) Comparative genome organization of vertebrates. Mamm Genome 7, 717–734
Barendse W, Vaiman D, Kemp S, Sugimoto Y, Armitage SM, Williams J, Sun S, Eggen A, Agaba M, Aleyasin A, Band M, Bishop M, Buitkamp J, Byrne K, Collins F, Cooper L, Coupettiers W, Denis B, Drinkwater R, Easterday K, Ennis S, Erhardt G, Ferretti L, Gao Q, Georges M, Gurung R, Harlizius B, Hawkins G, Hetzel DJS, Hirano T, Joergensen C, Kessler M, Kirkpatrick B, Konfortov B, Kuhn C, Lenstra H, Leveziel H, Lewin H, Leyhe B, Li L, Martin Burriel I, McGraw RA, Miller JR, Moody D, Moore S, Nakane S, Nijman I, Olsaker I, Pomp D, Rando A, Ron M, Shalom A, Soller M, Teale A, Thieven U, Vage D, Varvio S, Velmala RVJ, Weikard R, Woodside C, Womack J, Zanotti M, Zaragoza P (1997) A medium-density genetic linkage map of the bovine genome. Mamm Genome 8, 21–28
Beckmann JS, Weber, JL (1992) Survey of human and rat microsatellites. Genomics 12, 627–631
Bellané-Chantelot C, Lacroix B, Ougen P, Billault A, Beaufils S, Bertrand S, Georges I, Gilbert F, Gros I, Lucotte G, Susini L, Codani J-J, Gesnouin P, Pook S, Vayssex G, Lu-Kuo J, Ried T, Ward D, Chumakov I, Le Paslier D, Barillot E, Cohen D (1992) Mapping the whole human genome by fingerprinting yeast artificial chromosomes. Cell 70, 1059–1068
Broom MF, Hill DF (1994) Construction of a large-insert yeast-artificial chromosome library from sheep DNA. Mamm Genome 5, 817–819
Cai L, Taylor JF, Wing RA, Gallagher DS, Woo S-S, Davis SK (1995) Construction and characterization of a bovine bacterial artificial chromosome library. Genomics 29, 413–425
Cai L, Schalkwyk LC, Schoeberlein-Stehli A, Zee RYL, Smith A, Haaf T, Georges M, Lehrach H, Lindpaintner K (1997) Construction and characterization of a 10-genome equivalent yeast artificial chromosome library for the laboratory rat Rattus norvegicus. Genomics 39, 385–392
Chowdhary BP, Frönicke L, Gustavsson I, Scherthan H (1996) Comparative analysis of cattle and human genomes: detection of ZOO-FISH and gene mapping-based chromosomal homologies. Mamm Genome 7, 297–302
Crawford AM, Dodds KG, Pierson CA, Ede AJ, Montgomery GW, Garmonsway HG, Beattie AE, Davies K, Maddox JF, Kappes SW, Stone RT, Nguyen TC, Penty JM, Lord EA, Broom JE, Buitkamp J, Schwenger W, Epplen JT, Matthew P, Matthews ME, Hulme DJ, Beh KJ, McGraw RA, Beattie CW (1995) An autosomal genetic linkage map of the sheep genome. Genetics 140, 703–724
Cribiu EP, Matejka M, Denis B, Malher X (1988) Etude chromosomique d’un hybride chèvre x mouton fertile. Genet Sel Evol 20, 379–386
Georges M, Drinkwater R, King T, Mishra A, Moore SS, Nielsen D, Sargeant LS, Sorensen A, Steele MR, Zhao X, Womack JE, Hetzel J (1993) Microsatellite mapping of a gene affecting horn development in Bos taurus. Nature Genet 4, 206–210
Goureau A, Yerle M, Schmitz A, Riquet J, Milan D, Pinton P, Frelat G, Gellin J (1996) Human and porcine correspondence of chromosome segments using bidirectional chromosome painting. Genomics 36, 252–262
Guillemot E, Gary F, Berland HM, Durand V, Darré R, Cribiu EP (1991) Cytogenetic investigation in Saanen and Alpine artificial insemination bucks. Identification of a Robertsonian translocation. Genet Sel Evol 23, 449–154
Gyapay G, Schmitt K, Fizames C, Jones H, Vega-Czarny N, Spillett D, Muselet D, Prud’Homme J-F, Dib C, Auffray C, Morissette J, Weissenbach J, Goodfellow PN (1996) A radiation hybrid map of the human genome. Hum Mol Genet 5, 339–346
Hanahan D (1983) Studies on transformation of Escherichia coli with plasmids. J Mol Biol 166, 557–580
Hayes H (1995) Chromosome painting with human chromosome specific DNA libraries reveals the extent and the distribution of conserved segments in bovine chromosomes. Cytogenet Cell Genet 71, 168–174
Kim UJ, Birren BW, Slepak T, Mancino V, Boysen C, Kang HL, Simon MI, Shizuya H (1996) Construction and characterization of a human bacterial artificial chromosome library. Genomics 34, 213–218
Klein R, Wolf ED, Wu R, Sanford JC (1992) High-velocity microprojectiles for delivering nucleic acids into living cells. Biotechnology 24, 384–386
Kondo Y, Mori M, Kuramoto T, Yamada J, Beckmann JS, Simon-Chazottes D, Montagutelli X, Guénet J-L, Serikawa T (1993) DNA segments mapped by reciprocal use of microsatellite primers between mouse and rat. Mamm Genome 4, 571–576
Larsen F, Gundersen G, Lopez R, Prydz H (1992) CpG islands as gene markers in the human genome. Genomics 13, 1095–1107
Le Provost F, Lepingle A, Martin P (1996) A survey of the goat genome transcribed in the lactating mammary gland. Mamm Genome 7, 657–666
Libert F, Lefort A, Okimoto R, Womack J, Georges M (1993) Construction of a bovine genomic library of large yeast artificial chromosome clones. Genomics 270–276.
Lindsay S, Bird AP (1987) Use of restriction enzymes to detect potential gene in mammalian DNA. Nature 327, 336–338
Lyons LA, Laughlin TF, Copeland NG, Jenkins NA, Womack JE, O’Brien SJ (1997) Comparative anchor tagged sequences (CATS) for integrative mapping of mammalian genomes. Nature Genet 15, 47–56
Moore SS, Sargeant LL, King TJ, Mattick JS, Georges M, Hetzel DJS (1991) The conservation of dinucleotide microsatellites among mammalian genomes allows the use of heterologous PCR primer pairs in closely related species. Genomics 10, 654–660
Moore SS, Byrne K, Berger KT, Barendse W, McCarthy F, Womack JE, Hetzel DJS (1994) Characterization of 65 bovine microsatellites. Mamm Genome 5, 84–90
O’Brien SJ, Womack JE, Lyons LA, Moore KJ, Jenkins NA, Copeland NG (1993) Anchored reference loci for comparative genome mapping in mammals. Nature Genetics 3, 103–112
Pépin L, Amigues Y, Lépingle A, Berthier J-L, Bensaïd A, Vaiman D (1995) Sequence conservation of microsatellites between cattle (Bos taurus) and goat (Capra hircus), and related species. Examples of use in parentage testing and phylogeny analysis. Heredity 74, 53–61
Rogel-Gaillard C, Bourgeaux N, Save J-C, Renard C, Coullin P, Pinton P, Yerle M, Vaiman M, Chardon P (1997) Construction of a swine YAC library allowing an efficient recovery of unique and centromeric repeated sequences. Mamm Genome 8, 186–192
Schlötterer C, Tautz D (1992) Slippage synthesis of simple sequence DNA. Nucl Acids Res 20, 211–215
Shizuya H, Birren B, Kim U-J, Mancino V, Slepak T, Tachiri Y, Simon M (1992) Cloning and stable expression of 300-kilobase-pair fragments of human DNA in Escherischia coli using an F-factor-based vector. Proc Natl Acad Sci USA 89, 8794–8797
Smith TPL, Alexander LJ, Sonstegard TS, Yoo J, Beattie CW, Broom MF (1996) Construction and characterization of a large insert bovine YAC library with five-fold genomic coverage. Mamm Genome 7, 155–156
Solinas-Toldo S, Lengauer C, Fries R (1995) Comparative genome map of human and cattle. Genomics 27, 489–496
Steffen PJ (1992) Systematische Suche nach hochpolymorphen Markerloci im Rindergenom Zürich.
Stevanovic M, Zuffardi O, Collignon J, Lovell-Badge R, Goodfellow P (1994) The cDNA sequence and chromosomal location of the human SOX2 gene. Mamm Genome 5, 640–642
Stubbs L, Carver EA, Shannon ME, Kim J, Geisler J, Generoso EE, Stanford BG, Dunn WC, Mohrenweiser H, Zimmermann W, Watt SM, Ashworth LK (1996) Detailed comparative map of human chromosome 19q and related regions of the mouse genome. Genomics 35, 499–508
Sun HS, Cai L, Davis SK, Taylor JF, Doud LK, Bishop MD, Hayes H, Barendse W, Vaiman D, McGraw RA, Hirano T, Sugimoto Y, Kirkpatrick BW (1997) Comparative linkage mapping of human chromosome 13 and bovine chromosome 12. Genomics 39, 47–54
Vaiman D, Koutita O, Oustry A, Elsen J-M, Manfredi E, Fellous M, Cribiu EP (1996a) Genetic mapping of the autosomal region involved in XX sex-reversal and horn development in goats. Mamm Genome 7, 133–137
Vaiman D, Mercier D, Moazami-Goudarzi K, Eggen A, Ciampolini R, Lépingle A, Velmala R, Kaukinen J, Varvio S-L, Martin P, Levéziel H, Guérin G (1994) A set of 99 cattle microsatellites: characterization, synteny mapping and polymorphism. Mamm Genome 5, 288–297
Vaiman D, Pailhoux E, Schibler L, Oustry A, Chaffaux S, Cotinot C, Fellous M, Cribiu EP (1997) Genetic mapping of the polled/intersex locus (PIS) in goats. Theriogenology 47, 103–109
Vaiman D, Schibler L, Bourgeois F, Oustry A, Amigues Y, Cribiu EP (1996b) A genetic linkage map of the male goat genome. Genetics 144, 279–305
Wang GL, Holsten TE, Song WY, Wang HP, Ronald PC (1995) Construction of a rice bacterial artificial chromosome library and identification of clones linked to the Xa-21 disease resistance locus. Plant J 7, 525–533
Woo S-S, Jiang J, Gill BS, Paterson AH, Wing RA (1994) Construction and characterization of a bacterial artificial chromosome library of Sorghum bicolor. Nucl Acids Res 22, 4922–4931
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Schibler, L., Vaiman, D., Oustry, A. et al. Construction and extensive characterization of a goat Bacterial Artificial Chromosome library with threefold genome coverage. Mammalian Genome 9, 119–124 (1998). https://doi.org/10.1007/s003359900701
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/s003359900701