Abstract
Objectives
The aim of this study was to assess the potential of machine learning based on B-mode, shear-wave elastography (SWE), and dynamic contrast-enhanced ultrasound (DCE-US) radiomics for the localization of prostate cancer (PCa) lesions using transrectal ultrasound.
Methods
This study was approved by the institutional review board and comprised 50 men with biopsy-confirmed PCa that were referred for radical prostatectomy. Prior to surgery, patients received transrectal ultrasound (TRUS), SWE, and DCE-US for three imaging planes. The images were automatically segmented and registered. First, model-based features related to contrast perfusion and dispersion were extracted from the DCE-US videos. Subsequently, radiomics were retrieved from all modalities. Machine learning was applied through a random forest classification algorithm, using the co-registered histopathology from the radical prostatectomy specimens as a reference to draw benign and malignant regions of interest. To avoid overfitting, the performance of the multiparametric classifier was assessed through leave-one-patient-out cross-validation.
Results
The multiparametric classifier reached a region-wise area under the receiver operating characteristics curve (ROC-AUC) of 0.75 and 0.90 for PCa and Gleason > 3 + 4 significant PCa, respectively, thereby outperforming the best-performing single parameter (i.e., contrast velocity) yielding ROC-AUCs of 0.69 and 0.76, respectively. Machine learning revealed that combinations between perfusion-, dispersion-, and elasticity-related features were favored.
Conclusions
In this paper, technical feasibility of multiparametric machine learning to improve upon single US modalities for the localization of PCa has been demonstrated. Extended datasets for training and testing may establish the clinical value of automatic multiparametric US classification in the early diagnosis of PCa.
Key Points
• Combination of B-mode ultrasound, shear-wave elastography, and contrast ultrasound radiomics through machine learning is technically feasible.
• Multiparametric ultrasound demonstrated a higher prostate cancer localization ability than single ultrasound modalities.
• Computer-aided multiparametric ultrasound could help clinicians in biopsy targeting.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
With more than an estimated 164,000 new diagnoses in the USA [1] and almost 450,000 in Europe [2], prostate cancer (PCa) remains the most frequently occurring non-skin malignancy in Western men in 2018. Unfortunately, after prostate-specific antigen (PSA) serum level testing and/or digital rectal examination, the standard diagnostic approach strongly relies on a 10- to 12-core systematic biopsy [3]. Aside from complications associated with this procedure [4], high levels of underdiagnoses and overtreatment have been reported [5]. Given the strong clinical demand for reliable imaging that enables targeted biopsy, recent years have shown promising advances in multiparametric magnetic resonance imaging (mpMRI). Whereas individual modalities of MRI are not considered sufficiently accurate in PCa diagnosis, mpMRI leverages the combination of these modalities through scoring according to the Prostate Imaging Reporting and Data System (PI-RADS) [6]. The 2019 guidelines of the European Association of Urology recommend the use of a pre-biopsy mpMRI in the diagnostic pathway. However, aside from some inherent limitations of MRI (e.g., its high cost, limited availability, and impracticality for bedside use), such scoring systems are known to exhibit a slow learning curve and are at risk of high operator disagreement [7].
Another potential candidate for PCa imaging is ultrasound (US), which is cost-effective, widely available, and practical. Even though US modalities such as shear-wave elastography (SWE) and dynamic contrast-enhanced ultrasound (DCE-US) have shown promising results, targeted biopsy with US techniques still is not superior over systematic biopsy [8]. However, to date, a multiparametric US approach has been scarcely investigated [9]. The rationale for a multiparametric approach (i.e., combining information from complementary biomarkers such as tissue texture, elasticity or perfusion to image a notoriously multifocal and heterogeneous disease like PCa [10]) applies to both MRI and US. On top of that, the use of quantitative features known as radiomics is gaining attention [11]. Radiomics quantifies the spatial representation of tissue in an image such as heterogeneity or asymmetric enhancement by locally extracting textural and statistical features from the (parametric) images. In this work, we strived to combine the information from different modalities as well as their radiomics for image-based diagnosis of PCa. To examine the potential of such an approach, we employed machine learning technology by means of a random forest to optimally combine the underlying parameters. A random forest forms the core of a computer-aided diagnosis algorithm that combines all information into a single multiparametric image for the clinician to review [12].
In the classifier, inputs from B-mode US, SWE, and DCE-US are considered [13]. Although B-mode US by itself is not a suitable option for PCa imaging, biopsy guidelines highly recommend targeting of suspicious hypoechoic lesions [5]. As for SWE, tissue stiffness is regarded as a strong indicator of malignancy [14]. Recent studies have demonstrated its usefulness for the detection of PCa [15,16,17]. DCE-US, in which contrast agents are employed to visualize the vascularity, allows the assessment of tissue perfusion and contrast dispersion [18]. In fact, it was shown that quantification of the contrast agent kinetics by contrast ultrasound dispersion imaging (CUDI) allows the estimation of parameters reflecting the characteristics of angiogenic (micro)vasculature [19,20,21]. Whereas DCE-US images primarily represent vascular tissue characteristics, SWE images are related to the cell density and collagen deposition in the tissue [22, 23]. Therefore, being complementary in nature, it can be hypothesized that their combination leads to an increased diagnostic potential. In a recently published study, perfusion- and dispersion-related DCE-US parameters were already successfully combined in a machine learning approach [24].
This work validates a proposed random forest–based classifier in a leave-one-patient-out fashion, both pixel-wise and region-wise. Furthermore, the correlations among different features were investigated and their individual and combined importance for the localization of (clinically significant) PCa was evaluated.
Materials and methods
Data acquisition
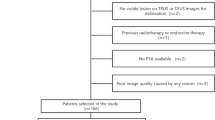
At the Martini Clinic Prostate Cancer Centre (University Hospital Hamburg Eppendorf, Germany), 50 men with biopsy-confirmed PCa referred for radical prostatectomy (RP) underwent a multiparametric US procedure. Only patients with a PSA level below 20 ng/mL, a prostate volume of < 80 mL, and no indication of extracapsular invasion were included in the study; patients with contra-indication for DCE-US or previous PCa therapy were excluded. Institutional review board approval was acquired and all participants signed an informed consent. Of them, 48 men underwent RP and were included in the study. The patient characteristics are listed in Table 1. Each patient received a B-mode, SWE, and 2-min DCE-US recording of the apex, mid, and base section of the gland. The examinations were performed manually, with an Aixplorer® ultrasound scanner (SuperSonic Imagine) equipped with a SE12-3 endocavity probe. For the DCE-US recordings, a 2.4-mL bolus of SonoVue® (Bracco) was intravenously administered. DCE-US was performed in “Gen” contrast-specific, low-mechanical-index mode; SWE images were obtained with minimal pre-compression and after a few-second stabilization period. This work is related to a clinical trial on multiparametric ultrasound (i.e., under registration number NCT03091231) and more information on the clinical workflow can be found in a previously published protocol paper [25].
Histopathological examination
After resection, the RP specimens were histopathologically examined. The annotated PCa regions were used to reconstruct a 3D model of the prostate and its lesions [26]. This model was subsequently digitally cross-sectioned at the apex, mid gland, and base to be matched to the imaging planes, allowing for direct US histopathology comparison [27]. Taking into account registration inaccuracy, a maximum of one unambiguously malignant and one unambiguously benign region of interest (ROI) were delineated in the B-mode image to serve as labeled ground truth for training and validation. The ROIs were drawn such that the number of malignant and the number of benign pixels, as well as those originating in the peripheral zone (PZ) and transition zone (TZ), were in balance.
Algorithm structure
An overview of the proposed method is shown in Fig. 1 and comprises (A) prostate segmentation, (B) data registration, (C) feature extraction, and (D) multiparametric classification. Testing and validation of the model is discussed in the last section of this “Materials and methods” section.
Prostate segmentation
Firstly, the prostate is located and delineated in each modality. To this end, we employed an automated deep learning–based TRUS segmentation algorithm on the side-view fundamental B-mode images of both the SWE and DCE-US acquisition [28, 29]. For DCE-US, the prostate position during wash-in (i.e., at 30 s) was used as a reference. Automatically, the prostate images were also zonally segmented, labeling pixels belonging to either the PZ or TZ for further use in the classification algorithm. The deep learning–based segmentations were checked by both an engineer and a urologist with 4 years of experience in TRUS imaging. Aside from prostate segmentation, a detection algorithm was designed to outline calcifications in the B-mode images. Calcifications were identified by high-valued regions in the fundamental-mode image after convolution with 2D Gaussian kernels having an empirically chosen standard deviation of ~ 0.6 mm and ~ 1.8 mm, thus detecting hyperechoic spots with diameters of approximately 1.2 and 3.6 mm. The purpose of calcification detection was to prevent false-positive readings due to elevated stiffness of calcified regions.
Data registration
As the proposed method aims at pixel-specific classification, a pixel-to-pixel match between the different US modalities is required. Again, the 30-s fundamental view image of the DCE-US recording was chosen as a reference. The SWE data were elastically registered to this image based on the segmented contours. Moreover, motion compensation was applied to the DCE-US video by rigid registration of every 5th frame to the reference position; the registration of the intermediate frames was performed by interpolating the translation-rotation matrix.
Imaging feature extraction
The proposed classifier includes a two-step feature extraction. First, model-based blood flow features are retrieved from the DCE-US imaging. The model-based feature extraction serves two purposes: on the one hand, physically meaningful parameters with known correlation to PCa are estimated and, on the other hand, the dimensionality of the DCE-US is reduced to 2D, matching the SWE and grayscale image prior to texture analysis. Secondly, radiomic features are extracted from the resulting model-based feature maps as well as the SWE and grayscale image.
The model-based feature extraction was based on CUDI, a family of quantification methods that estimate underlying physical quantities of a DCE-US recording related to perfusion and dispersion [19,20,21, 30, 31]. A total of 12 DCE-US features were extracted for every pixel, which are listed in Table 2. In CUDI, the spreading of contrast through the prostate is regarded as a convective-dispersive process, which can be quantified by assessing the evolution of contrast over time. The contrast velocity (v), dispersion (D), and Péclet (Pe) number were estimated through local system identification [21]. Alternatively, the local degree of dispersion can also be quantified by the similarity in contrast behavior among pixels. This was quantified either by spatiotemporal correlation (r) [30, 31] or spectral coherence (ρ) [20]. In addition, we fitted the contrast curves in a single pixel by a modified local density random walk model, enabling us to estimate the mean transit time (μ), the dispersion-related parameter (κ), and the area under the contrast curve (α) [19]. Finally, also heuristic parameters such as the wash-in time (WIT), appearance time (AT), peak intensity (PI), and peak time (PT) were extracted.
The rationale for the use of radiomic features is that not only pixel values but also local spiculation, heterogeneity, and granularity are widely considered as important biomarkers of cancer. Moreover, asymmetric patterns in perfusion or elasticity regardless of the pixel values are also seen as indicative of malignancy. To take into account intra-prostate asymmetry, as well as relatively high parameter values, we introduced the parameter value relative to the median parameter value per image as a feature. Likewise, to quantify parameter heterogeneity, we extracted the entropy of the parameter distribution in a circular kernel around the pixel of interest. A multiscale approach was adopted, using heuristic kernel radii of ~ 1 mm, ~ 2 mm, and ~ 3 mm. In addition, the parameter variance was calculated in a ~ 2-mm kernel.
Automated multiparametric combination
Multiparametric combination of the features was achieved through machine learning based on a random forest algorithm. A random forest is an ensemble of independently trained decision trees, which vote together on the final classification score [32]. Having a branch-like structure of decision nodes, single-classification trees classify a sample by a series of decisions based on the input variables. Node by node, the tree structure is grown by evaluating for which feature (a subset of) the labeled training instances can be most effectively separated in terms of their class. Subsequently, the robustness of a random forest is established by growing each tree using another random subset of the training samples [33].
In this work, we enforced the first split to be based on the zonal location (either PZ or TZ), as it is established that tissue stiffness [34] and the influx of contrast agents [35] differ substantially between zones. Then, a random forest was grown consisting of 1000 trees using 1/1000th of the training set with replacement. To promote generalizability, six random training patients were completely discarded prior to growing each tree. The cross-entropy of labels within the nodes was adopted as the splitting criterion and the tree depth was at most 50 nodes. Pixels containing calcifications were omitted in the training phase as they might obscure the underlying tissue type. The final multiparametric score, ranging from − 1 to 1, was defined by the ratio between the number of malignant and benign classifications among the trees in the random forest. After classification, outliers were removed from the multiparametric images by assigning the median multiparametric score in a 15-pixel region (~ 2.5 mm), corresponding to approximately half the radius of clinically significant PCa [36].
Validation and statistical analysis
The classifier was validated in a leave-one-patient-out cross-validation procedure, in which each patient is tested using 1 classifier that is trained on the data of the remaining patients. The performance was assessed by computing the area under the receiver operating characteristics curve (ROC-AUC) of the parameter values or the multiparametric score, both in a pixel-wise and a region-wise fashion. In the latter approach, a ROI was characterized by its mean parameter value or multiparametric score. Differences between distributions were statistically assessed with a Wilcoxon rank sum test [37]. Throughout this work, p values of < 0.05 and < 0.005 are defined to describe significant and highly significant differences among groups, respectively.
Results
Correlation among radiomics
Figure 2 depicts the correlations between radiomics in a correlation matrix. Strong positive and negative correlations are color-coded in red and blue, respectively. Features from the same analysis typically exhibit high correlation. In addition, especially μ and WIT as well as α and PI seem related. The low correlation between Young’s modulus (E), gray levels (G), and DCE-US features is an indication that B-mode, SWE, and DCE-US are indeed complementary.
Classification performance
Figure 3 illustrates the power of the proposed multiparametric analysis, showing a number of single-parametric maps alongside the multiparametric image obtained in a patient with a left-apical 4 + 5 = 9 tumor. The segmentation, zonal boundary, detected calcifications, and annotated ROI locations are indicated as well. The single- and multiparametric results across the entire dataset are presented in Table 2. Multiparametric radiomic-based classification yields a region-wise ROC-AUC of 0.75 and 0.90 for PCa and significant PCa versus benign regions, respectively. In our dataset, binary ROI classification (i.e., with a positive multiparametric score referring to malignancy and a negative to benign tissue) would lead to 32 (27%) benign regions erroneously classified as sPCa and 1 (3.3%) sPCa lesion as benign.
Image plane example, showing the B-mode (a), Young’s modulus (SWE) (b), Péclet number (c), spatiotemporal correlation (d), dispersion-related parameter (e), wash-in time (f), velocity (g), velocity relative to image median (h), 2-mm entropy of velocity (i), and resulting multiparametric map (j). In each map, the prostate and zonal segmentations are depicted in white, the calcifications are encircled in blue, and histopathologically confirmed malignant and benign ROIs are indicated in red and green, respectively
In comparison, applying the classifier on only the radiomic features for the best-performing DCE-US parameter (i.e., contrast velocity, v) resulted in a region-wise ROC-AUC of 0.71 and 0.84 for PCa and sPCa, respectively. The classification performance can thus be partly attributed to the use of radiomics and partly to multiparametric combination. With the use of non-contrast features, only region-wise ROC-AUCs of 0.58 for PCa and 0.65 for sPCa were achieved.
Feature relevance
The relative relevance of different parameters is assessed by examining which parameters are selected for the first, second, etc. decision nodes in the trees of the random forest. Figure 4 presents the most prominent parameters. Based on this data, it seems that in particular the combination between v, r, and E is favored. In the TZ, also ρ and PT are relevant parameters. In terms of radiomics, mostly the parametric value itself and the large-kernel entropy are selected.
Relation to cancer grade
To assess the degree to which parameters and the multiparametric score correspond to cancer aggressiveness, Fig. 5 shows how the mean values per ROI are distributed for different Gleason groups and prostate zones (i.e., PZ and TZ). Both SWE-derived Young’s modulus and the best-performing DCE-US parameter (i.e., v) are depicted alongside the final multiparametric score. Significant and highly significant differences are indicated with a single asterisk and double asterisks, respectively. It should be emphasized that healthy TZ tissue is generally stiffer than PZ tissue [34], as evidenced in Fig. 5, hampering the analysis of TZ and PZ regions as a single group.
Overview of the parameter values and classifier score for the velocity (a), Young’s modulus (E) (b), and the multiparametric classifier score (c). Individual regions are represented by a bullet. The violin plots represent the group distribution in the PZ (left, blue) and TZ (right, red). Significant and highly significant differences according to a Wilcoxon rank sum test are indicated with a single asterisk and double asterisks
Discussion
In this work, we report on the development of a random forest–based classifier for multiparametric classification of PCa based on co-registered B-mode, shear-wave elastography, and contrast-enhanced ultrasound. Aside from model-based parameters, radiomics are introduced in the classifier framework to extract additional information from the parametric maps. The ROC-AUC, validated in a leave-one-patient-out cross-validation fashion, shows a region-wise improvement from 0.76 of the best-performing individual parameter, v, to a multiparametric 0.90 for significant Gleason > 3 + 4 PCa. A similar improvement is achieved using a pixel-wise approach. The improvement is partly the result of the radiomic extraction and partly of the multiparametric combination.
The random forest classifier is a powerful tool for classification that allows for the integration of a large range of (radiomic) features and generates an intuitive multiparametric score. The frequency at which parameters are being selected for classification (see Fig. 4) substantiates the multiparametric hypothesis, favoring a combination of a perfusion-related (i.e., v, PT), dispersion-related (i.e., r, ρ), and elastographic (i.e., E) parameters. This is in line with earlier work that only included DCE-US parameters, reporting that model-based parameters that are related to different underlying biomarkers combine most effectively [24]. In addition, the selected parameters differ substantially between the PZ and TZ. This might be due to the anatomic or physiological differences between zones; however, it could also be a result of less robust parameter estimation farther away from the probe (i.e., in the TZ), due to the increasing impact of attenuation and shadowing. This stresses the need for adequate zonal segmentation in the proposed framework, here obtained through deep learning [28, 29].
The multiparametric score is shown to scale with tumor Gleason grade, with significant differences between benign, insignificant, and significant disease. Several definitions of clinically significant prostate cancer are used in the literature; due to the limited amount of 3 + 3 disease in this RP-validated study and the distinction between Gleason 3 + 4 = 7 and 4 + 3 = 7 being strongly associated with PCa prognosis [38], we report on both the identification of ≥ 3 + 3 and > 3 + 4 PCa. Our results (Fig. 5) show no significant difference between 3 + 4 PCa and the small group of 3 + 3 PCa. This might be partly explained due to a bias in the 3 + 3 group, with tumors being disproportionately large for clinicians to decide upon RP as a treatment instead of active surveillance and, thus, for inclusion in the presented study.
Furthermore, only a few radiomic features were introduced in this research. Many more have been proposed in the literature, including morphological, intensity-based, texture-based, and statistics-based features [39,40,41,42,43]. Alternatively, novel model-based features could be considered. For example, velocity vector field entropy [44] and viscoelasticity [23, 45] have shown promise as markers for PCa in DCE-US and SWE. Even though the quality of the input images remains dependent on the operator recording the SWE and CEUS acquisitions [46, 47], the use of automatically generated single multiparametric images might reduce interobserver variability compared with cognitive reading of a large ensemble of parametric maps.
Compared with other research, a meta-analysis of SWE and DCE-US has reported ROC-AUCs of 0.90 [15] to 0.91 [16] and 0.83 [48], respectively. It should be emphasized that these results are based on systematic biopsy, known for its systematic and random errors [49], as a reference standard. Furthermore, these scores were obtained by cognitive reading of images and videos rather than an automatic pixel value–based approach like in this study, which might hamper comparison. The value of multiparametric images for cognitive reading, either as stand-alone tool or combined with the source images, remains to be investigated.
Despite the performance gains obtained using the proposed method, some malignant ROIs were still missed and some benign ROIs were wrongly classified as malignant by the algorithm. In the future, immunohistochemical techniques [50, 51] might elucidate more on the nature of the false readings. There are indications that (co-occurring) prostatitis or BPH might be responsible for false positives, as these diseases are known to also promote angiogenesis [52, 53]. Qualitative inspection of the false negatives revealed that these were indeed invisible to the naked eye on all US modalities. Future analysis of tumors that are missed on all US imaging modalities might potentially direct us towards new parameters or radiomic features contributing to the multiparametric classification.
Furthermore, this study was conducted in a single center, where a dataset of 50 patients presenting biopsy-proven PCa was collected in order to have RP specimens as histopathological ground truth for PCa localization. As a result, ROC-AUCs were calculated for the separation of benign and malignant ROIs. Prospective, multicenter, targeted biopsy-based studies might eventually confirm the diagnostic value of the machine learning classification presented in this work in a more varied patient group [54]. Another limitation is the 2D nature of this approach, requiring the acquisition of three planes per patient for every modality. In a clinical setting, the use of more planes per patient would reduce the risk of missing out-of-plane tumors at the cost of an increased procedure time. However, as 3D SWE and DCE-US have recently been introduced [55, 56], expansion to three dimensions can be envisaged.
In conclusion, we demonstrated the feasibility of a multiparametric classifier to improve upon single US modalities for the localization of PCa. This is in line with recently published work on multiparametric US for the identification of malignant and benign breast lesions [57]. We aim to further extend the dataset, so that the classification approach can be expanded to more radiomics and features. Once the performance is consolidated, we believe that a three-dimensional approach might bring clinical adoption closer within reach.
Abbreviations
- CUDI:
-
Contrast ultrasound dispersion imaging
- DCE-US:
-
Dynamic contrast-enhanced ultrasound
- MRI:
-
Magnetic resonance imaging
- PCa:
-
Prostate cancer
- PI-RADS:
-
Prostate imaging reporting and data system
- PSA:
-
Prostate-specific antigen
- PZ:
-
Peripheral zone
- ROC-AUC:
-
Area under the receiver operating characteristics curve
- ROI:
-
Region of interest
- RP:
-
Radical prostatectomy
- SWE:
-
Shear-wave elastography
- TZ:
-
Transition zone
- US:
-
Ultrasound
References
Siegel RL, Miller KD, Jemal A (2018) Cancer statistics, 2018. CA Cancer J Clin 68:7–30
Ferlay J, Colombet M, Soerjomataram I et al (2018) Cancer incidence and mortality patterns in Europe: estimates for 40 countries and 25 major cancers in 2018. Eur J Cancer 103:356–387
Mottet N, Bellmunt J, Bolla M et al (2017) EAU-ESTRO-SIOG guidelines on prostate cancer. Part 1: screening, diagnosis, and local treatment with curative intent. Eur Urol 71:618–629. https://doi.org/10.1016/j.eururo.2016.08.003
Loeb S, Vellekoop A, Ahmed HU et al (2013) Systematic review of complications of prostate biopsy. Eur Urol 64:876–892. https://doi.org/10.1016/j.eururo.2013.05.049
Ukimura O, Coleman JA, de la Taille A et al (2013) Contemporary role of systematic prostate biopsies: indications, techniques, and implications for patient care. Eur Urol 63:214–230
Weinreb JC, Barentsz JO, Choyke PL (2016) PIRADS prostate imaging - reporting and data system: 2015, version 2. Eur Urol 69:16–40
Smith CP, Harmon SA, Barrett T et al (2018) Intra- and interreader reproducibility of PI-RADSv2: a multireader study. J Magn Reson Imaging. https://doi.org/10.1002/jmri.26555
van Hove A, Savoie PH, Maurin C et al (2014) Comparison of image-guided targeted biopsies versus systematic randomized biopsies in the detection of prostate cancer: a systematic literature review of well-designed studies. World J Urol 32:847–858
Postema A, Mischi M, de la Rosette J, Wijkstra H (2015) Multiparametric ultrasound in the detection of prostate cancer: a systematic review. World J Urol 33:1651–1659
Andreoiu M, Cheng L (2010) Multifocal prostate cancer: biologic, prognostic, and therapeutic implications. Hum Pathol 41:781–793. https://doi.org/10.1016/j.humpath.2010.02.011
Gillies RJ, Kinahan PE, Hricak H (2015) Radiomics: images are more than pictures, they are data. Radiology 278:563–577. https://doi.org/10.1148/radiol.2015151169
Wang S, Summers RM (2012) Machine learning and radiology. Med Image Anal 16:933–951. https://doi.org/10.1016/j.media.2012.02.005
Klibanov AL, Hossack JA (2015) Ultrasound in radiology: from anatomic, functional, molecular imaging to drug delivery and image-guided therapy. Investig Radiol 50:657–670
Correas J-M, Tissier A-M, Khairoune A et al (2013) Ultrasound elastography of the prostate: state of the art. Diagn Interv Imaging 94:551–560
Woo S, Suh CH, Kim SY, Cho JY, Kim SH (2017) Shear-wave elastography for detection of prostate cancer: a systematic review and diagnostic meta-analysis. AJR Am J Roentgenol 209:806–814. https://doi.org/10.2214/AJR.17.18056
Sang L, Wang XM, Xu DY, Cai YF (2017) Accuracy of shear wave elastography for the diagnosis of prostate cancer: a meta-analysis. Sci Rep 7:1949. https://doi.org/10.1038/s41598-017-02187-0
Boehm K, Salomon G, Beyer B et al (2015) Shear wave elastography for localization of prostate cancer lesions and assessment of elasticity thresholds: implications for targeted biopsies and active surveillance protocols. J Urol 193:794–800
Halpern EJ, McCue PA, Aksnes AK, Hagen EK, Frauscher F, Gomella LG (2002) Contrast-enhanced US of the prostate with Sonazoid: comparison with whole-mount prostatectomy specimens in 12 patients. Radiology 222:361–366. https://doi.org/10.1148/radiol.2222010582
Kuenen MP, Mischi M, Wijkstra H (2011) Contrast-ultrasound diffusion imaging for localization of prostate cancer. IEEE Trans Med Imaging 30:1493
Mischi M, Kuenen MPJ, Wijkstra H (2012) Angiogenesis imaging by spatiotemporal analysis of ultrasound contrast agent dispersion kinetics. IEEE Trans Ultrason Ferroelectr Freq Control 59:621–629. https://doi.org/10.1109/TUFFC.2012.2241
van Sloun RJ, Demi L, Postema AW, de la Rosette JJ, Wijkstra H, Mischi M (2017) Ultrasound-contrast-agent dispersion and velocity imaging for prostate cancer localization. Med Image Anal 35:610–619. https://doi.org/10.1016/j.media.2016.09.010
Pallwein L, Mitterberger M, Gradl J et al (2007) Value of contrastenhanced ultrasound and elastography in imaging of prostate cancer. Curr Opin Urol 17:39–47. https://doi.org/10.1097/MOU.0b013e328011b85c
Hoyt K, Castaneda B, Zhang M et al (2008) Tissue elasticity properties as biomarkers for prostate cancer. Cancer Biomark 4:213–225
Wildeboer RR, Postema AW, Demi L et al (2017) Multiparametric dynamic contrast-enhanced ultrasound imaging of prostate cancer. Eur Radiol 27:3226–3234
Mannaerts CK, Wildeboer RR, Postema AW et al (2018) Multiparametric ultrasound: evaluation of grayscale, shear wave elastography and contrast-enhanced ultrasound for prostate cancer detection and localization in correlation to radical prostatectomy specimens. BMC Urol 18:98
Wildeboer RR, Schalk SG, Demi L et al (2017) Three-dimensional histopathological reconstruction as a reliable ground truth for prostate cancer studies. Biomed Phys Eng Express 3:35014
Wildeboer RR, van Sloun RJG, Postema AW et al (2018) Accurate validation of ultrasound imaging of prostate cancer: a review of challenges in registration of imaging and histopathology. J Ultrasound 21:197–207. https://doi.org/10.1007/s40477-018-0311-8
van Sloun RJG, Wildeboer RR, Postema AW et al (2018) Zonal segmentation in transrectal ultrasound images of the prostate through deep learning. 2018 IEEE Int Ultrason Symp Proc
van Sloun RJG, Wildeboer RR, Mannaerts CK et al (2019) Deep learning for real-time, automatic, and scanner-adapted prostate (zone) segmentation of transrectal ultrasound, for example,magnetic resonance imaging–transrectal ultrasound fusion prostate biopsy. Eur Urol Focus. https://doi.org/10.1016/j.euf.2019.04.009
Kuenen M, Saidov T, Wijkstra H, Mischi M (2013) Contrastultrasound dispersion imaging for prostate cancer localization by improved spatiotemporal similarity analysis. Ultrasound Med Biol 39:1631
Kuenen MPJ, Saidov TA, Wijkstra H et al (2013) Spatiotemporal correlation of ultrasound contrast agent dilution curves for angiogenesis localization by dispersion imaging. IEEE Trans Ultrason Ferroelectr Freq Control 60:2665–2669. https://doi.org/10.1109/TUFFC.2013.2865
Breiman L (1996) Bagging predictors. Mach Learn 24:123–140
Geurts P, Irrthum A, Wehenkel L (2009) Supervised learning with decision tree-based methods in computational and systems biology. Mol BioSyst 5:1593–1605. https://doi.org/10.1039/B907946G
Rouvière O, Melodelima C, Hoang Dinh A et al (2017) Stiffness of benign and malignant prostate tissue measured by shear-wave elastography: a preliminary study. Eur Radiol 27:1858–1866
Halpern EJ (2006) Contrast-enhanced ultrasound imaging of prostate cancer. Rev Urol 8:S29
Stamey TA, Freiha FS, McNeal JE et al (1993) Localized prostate cancer. Relationship of tumor volume to clinical significance for treatment of prostate cancer. Cancer 71:933–938
Fay MP, Proschan MA (2010) Wilcoxon-Mann-Whitney or t-test? On assumptions for hypothesis tests and multiple interpretations of decision rules. Stat Surv 4:1–39. https://doi.org/10.1214/09-SS051
Stark JR, Perner S, Stampfer MJ et al (2009) Gleason score and lethal prostate cancer: does 3 + 4 = 4 + 3? J Clin Oncol 27:3459–3464. https://doi.org/10.1200/JCO.2008.20.4669
Smith CP, Czarniecki M, Mehralivand S et al (2018) Radiomics and radiogenomics of prostate cancer. Abdom Radiol (NY) 44:2021–2029
Rohrbach D, Wodlinger B, Wen J et al (2018) High-frequency quantitative ultrasound for imaging prostate cancer using a novel micro-ultrasound scanner. Ultrasound Med Biol 44:1341–1354. https://doi.org/10.1016/j.ultrasmedbio.2018.02.014
Guo Y, Hu Y, Qiao M et al (2018) Radiomics analysis on ultrasound for prediction of biologic behavior in breast invasive ductal carcinoma. Clin Breast Cancer 18:e335–e344
Coroller TP, Agrawal V, Narayan V et al (2016) Radiomic phenotype features predict pathological response in non-small cell lung cancer. Radiother Oncol 119:480–486. https://doi.org/10.1016/j.radonc.2016.04.004
Ingrisch M, Schneider MJ, Nörenberg D et al (2017) Radiomic analysis reveals prognostic information in T1-weighted baseline magnetic resonance imaging in patients with glioblastoma. Investig Radiol 52:360–366
van Sloun RJ, Demi L, Postema AW, de la Rosette JJ, Wijkstra H, Mischi M (2017) Entropy of ultrasound-contrast-agent velocity fields for angiogenesis imaging in prostate cancer. IEEE Trans Med Imaging 36:826–837
van Sloun RJ, Wildeboer RR, Wijkstra H, Mischi M (2017) Viscoelasticity mapping by identification of local shear wave dynamics. IEEE Trans Ultrason Ferroelectr Freq Control 64:1666–1673. https://doi.org/10.1109/TUFFC.2017.2743231
Harvey H, Morgan V, Fromageau J et al (2018) Ultrasound shear wave elastography of the normal prostate: interobserver reproducibility and comparison with functional magnetic resonance tissue characteristics. Ultrason Imaging 40:158–170. https://doi.org/10.1177/0161734618754487
Fröhlich E, Muller R, Cui X-W et al (2015) Dynamic contrastenhanced ultrasound for quantification of tissue perfusion. J Ultrasound Med 34:179–196
Li Y, Tang J, Fei X, Gao Y (2013) Diagnostic performance of contrast enhanced ultrasound in patients with prostate cancer: a meta-analysis. Acad Radiol 20:156–164. https://doi.org/10.1016/j.acra.2012.09.018
Ahmed HU, El-Shater Bosaily A, Brown LC et al (2017) Diagnostic accuracy of multi-parametric MRI and TRUS biopsy in prostate cancer (PROMIS): a paired validating confirmatory study. Lancet 389:815–822
Marien KM, Croons V, Waumans Y et al (2016) Development and validation of a histological method to measure microvessel density in whole-slide images of cancer tissue. PLoS One 11:e0161496
Ferrer FA, Miller LJ, Andrawis RI et al (1998) Angiogenesis and prostate cancer: in vivo and in vitro expression of angiogenesis factors by prostate cancer cells. Urology 51:161–167. https://doi.org/10.1016/S0090-4295(97)00491-3
Shih S-J, Dall’Era MA, Westphal JR et al (2003) Elements regulating angiogenesis and correlative microvessel density in benign hyperplastic and malignant prostate tissue. Prostate Cancer Prostatic Dis 6:131–137
Sandhu JS (2008) Prostate cancer and chronic prostatitis. Curr Urol Rep 9:328–332
Postema AW, Scheltema MJV, Mannaerts CK et al (2017) The prostate cancer detection rates of CEUS-targeted versus MRI-targeted versus systematic TRUS-guided biopsies in biopsy-naïve men: a prospective, comparative clinical trial using the same patients. BMC Urol 17:27. https://doi.org/10.1186/s12894-017-0213-7
Shoji S, Hashimoto A, Nakamura T et al (2018) Novel application of three-dimensional shear wave elastography in the detection of clinically significant prostate cancer. Biomed Rep 8:373–377. https://doi.org/10.3892/br.2018.1059
Schalk SG, Huang J, Li J et al (2018) 3-D quantitative dynamic contrast ultrasound for prostate cancer localization. Ultrasound Med Biol 44:807–814. https://doi.org/10.1016/j.ultrasmedbio.2017.12.005
Kapetas P, Clauser P, Woitek R et al (2019) Quantitative multiparametric breast ultrasound: application of contrastenhanced ultrasound and elastography leads to an improved differentiation of benign and malignant lesions. Investig Radiol 54:257–264
Funding
This study has received funding from the Dutch Cancer Society (#UVA2013-5941), a European Research Council Starting Grant (#280209), and was performed within the framework of the IMPULS2-program within the Eindhoven University of Technology in collaboration with Philips. The authors would like to acknowledge SuperSonic Imagine (Aix-en-Provence, France) for providing the Aixplorer® Ultrasound platform and technical support and NVIDIA Corporation for granting the Titan XP graphics processing unit.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Guarantor
The scientific guarantor of this publication is Massimo Mischi.
Conflict of interest
The authors of this manuscript declare relationships with the following companies: Philips.
Statistics and biometry
No complex statistical methods were necessary for this paper.
Informed consent
Written informed consent was obtained from all subjects (patients) in this study.
Ethical approval
Institutional Review Board approval was obtained.
Methodology
• retrospective
• diagnostic or prognostic study
• performed at one institution
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Wildeboer, R.R., Mannaerts, C.K., van Sloun, R.J.G. et al. Automated multiparametric localization of prostate cancer based on B-mode, shear-wave elastography, and contrast-enhanced ultrasound radiomics. Eur Radiol 30, 806–815 (2020). https://doi.org/10.1007/s00330-019-06436-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00330-019-06436-w