Abstract
The seeds of many legume species including soybean, Pongamia pinnata and the model legume Medicago truncatula store considerable oil, apart from protein, in their cotyledons. However, as a group, legume storage strategies are quite variable and provide opportunities for better understanding of carbon partitioning into different storage products. Legumes with their ability to fix nitrogen can also increase the sustainability of agricultural systems. This review integrates the cell biology, biochemistry and molecular biology of oil body biogenesis before considering biotechnology strategies to enhance oil body biosynthesis. Cellular aspects of packaging triacylglycerol (TAG) into oil bodies are emphasized. Enhancing seed oil content has successfully focused on the up-regulation of the TAG biosynthesis pathways using overexpression of enzymes such as diacylglycerol acyltransferase1 and transcription factors such as WRINKLE1 and LEAFY COTYLEDON1. While these strategies are central, decreasing carbon flow into other storage products and maximizing the packaging of oil bodies into the cytoplasm are other strategies that need further examination. Overall there is much potential for integrating carbon partitioning, up-regulation of fatty acid and TAG synthesis and oil body packaging, for enhancing oil levels. In addition to the potential for integrated strategies to improving oil yields, the capacity to modify fatty acid composition and use of oil bodies as platforms for the production of recombinant proteins in seed of transgenic legumes provide other opportunities for legume biotechnology.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Legume seeds store proteins, lipids and starch required for energy and growth upon germination. The proportion of these storage components varies according to species (Table 1; Fig. 1). The seed can thus be harvested to serve human nutrition, stock feed, biofuels and industrial application (Duranti and Gius 1997; Djemel et al. 2005; Gallardo et al. 2008). Legume seed can accumulate considerable oil aside from protein, and the oil content is largely dependent on legume species (Table 1). For example, soybean seed contains 20% oil (Clemente and Cahoon 2009). Pongamia pinnata (Pongamia) seed commonly stores 35% oil, which offers a potential source for biofuel use (Scott et al. 2008). Low oil is usually associated with high starch, but not necessarily (Table 1). As the world population is growing rapidly there is increasing pressure on food security, and seed oil production is part of this. In addition, biodiesel demand for future energy also relies on greater seed oil production. Therefore, improvement of seed oil production will be important in addressing these challenges (Roesler et al. 2016).
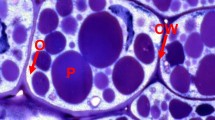
Oil bodies in three legumes: Pongamia, soybean and Medicago. Light microscope images of cotyledon storage cells of Pongamia (a), soybean (b) and M. truncatula (c). In Pongamia and soybean, oil bodies are white dots filling the cytoplasm, while in M. truncatula they are white borders ringing the protein bodies. Electron microscope images of oil bodies of Pongamia (d), soybean (e) and M. truncatula (f). a amyloplast, n nucleus, cw cell wall, p protein body, arrows point to individual oil bodies. Bars a–c 10 µm and d–f 1 µm
Legumes are second to cereals in agricultural importance, based on area harvested and total production (Gepts et al. 2005). Grain legumes provide about one-third of dietary protein nitrogen and vegetable oil for human consumption (Graham and Vance 2003). Intercropping and rotation of grain legumes with cereals or other non-leguminous crops can increase nitrogen use efficiency and enhance yield (Rose 2008; Siddique et al. 2012). It has been estimated that crop legumes contribute 21 Mt (megatonne, 106 tonnes, tonne is the metric ton) of N2 annually by legume–rhizobia symbioses (Herridge et al. 2008) and return 5–7 Mt of N2 to soil saving US$8–12 billion (Foyer et al. 2016). Further, synthetic nitrogen fertilizers are a major source of nitrous oxide emissions (Davidson 2009; Foyer et al. 2016). Nitrous oxide is the fourth most important anthropogenic greenhouse gas. It is also a reactant in the destruction of atmospheric ozone (Davidson 2009). In addition, it is argued that legumes have been underutilized, with a negative impact on human health and sustainable food production (Foyer et al. 2016).
In this review we first integrate the cell biology, biochemistry and molecular biology of oil body biogenesis. We then examine biotechnological strategies for modification of oil content. Enhancing seed oil content has to date particularly focused on the up-regulation of the oil biosynthesis pathways. While this strategy is important, decreasing carbon flow into other storage products and maximizing the packaging of oil bodies into the cytoplasm need further examination. The capacity to modify fatty acid composition and the use of oil bodies as platforms for the production of recombinant proteins in seed of transgenic legumes are also considered.
Oil bodies in legumes
Seed oil is stored as oil bodies (also known as lipid droplets) which are subcellular structures in the cytoplasm (Fig. 1). Oil bodies are generally considered to be circular to ovoid, with diameter varying between species, but within the range of 0.5–2.5 µm (Huang 1992; Tzen et al. 1993; Wang et al. 2012). However, as can be seen for Pongamia and soybean (Fig. 1) some oil body diameters can be 2–3 times larger than 2.5 µm. Oil bodies are filled with triacylglycerols (TAGs) enclosed by a monolayer of phospholipid (PL) embedded with integral membrane oleosin proteins (Huang 1992; Tzen et al. 1993; Tzen 2012). There are different isoforms of oleosin and two minor integral proteins, caleosin and steroleosin (Chen et al. 1999; Tzen 2012). The structure of oil bodies allow them to be maintained as small discrete structures (Fig. 1). In Medicago truncatula oil bodies are aligned around the protein bodies (Fig. 1f) and adjacent to the plasma membrane. In legumes with more lipids the oil bodies pack into the cytoplasm with different patterns of oil body sizes (Fig. 1d, e). It is possible that the arrangement of oil bodies adjacent to the plasma membrane may have a cellular protective effect during seed desiccation. Knockdown of oleosin in soybean, which leads to the formation of giant oil bodies, results in few, if any, viable cotyledon cells after hydration and germination (Schmidt and Herman 2008).
The oil body can be detected as early as the heart stage in Brassica napus (He and Wu 2009). The different oleosin isoforms, caleosin and steroleosin accumulate sequentially throughout seed development (Gallardo et al. 2016). Fatty acid accumulation starts from 8 days after pollination, corresponding to the heart stage in M. truncatula (Wang et al. 2012). While this review focuses on seed oil bodies, oil bodies can be present in many different tissues and organs (Gallardo et al. 2016).
Oil body biogenesis
The pathway from photosynthate to the final oil bodies is composed of three major parts: fatty acid biosynthesis, TAG assembly and oil body formation. As a consequence, oil body formation in developing seed is regulated at multiple steps.
Fatty acid synthesis
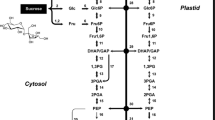
Fatty acids are synthesized de novo in seed plastids using the sucrose imported from photosynthesis (Fig. 2). The sucrose is first cleaved into glucose and fructose, followed by processing into pyruvate via glycolysis. While pyruvate may enter the plastid directly from the cytoplasm by a passive mechanism, or unknown translocator, there are other sources of pyruvate in soybean plastids (Hajduch et al. 2011; Gerrard Wheeler et al. 2016). Phosphoenol pyruvate (PEP) can be imported into the plastid via a PEP translocator and pyruvate formed via a plastid pyruvate kinase. Likely more important in soybean (Hajduch et al. 2011) is the production of malate from oxaloacetate derived from PEP. Malate is imported into the plastid via a dicarboxylic translocator and converted to pyruvate by a NADP malic enzyme (Hajduch et al. 2011; Gerrard Wheeler et al. 2016). In the plastid, pyruvate dehydrogenase converts pyruvate to CO2, which is recycled, and acetyl-CoA which is utilized for fatty acid biosynthesis (Fig. 2). The enzyme acetyl-CoA carboxylase (ACCase) is generally considered to catalyse the first reaction of the fatty acid biosynthetic pathway: the formation of malonyl-CoA from acetyl-CoA. Malonyl-CoA is used for the production of 16:0 ACP, 18:0 ACP and 18:1 ACP by the fatty acid (FA) synthase (Chapman and Ohlrogge 2012). Two carbon fragments are added to the growing FA chain, which is covalently attached to the acyl carrier protein (ACP). Free FAs are produced from 16:0 ACP, 18:0 ACP and 18:1 ACP by two acyl-ACP thioesterases (Ohlrogge and Browse 1995; Chapman and Ohlrogge 2012). The FAs are activated to CoA on the chloroplast outer envelope and 16:0 CoA, 18:0 CoA and 18:1 CoA are transferred to the endoplasmic reticulum (ER) for TAG assembly. The majority of commonly grown oil crops accumulate mainly C16 or C18 saturated and unsaturated fatty acids in their storage lipids.
Oil body biogenesis summary diagram. Biosynthesis of fatty acids and oil bodies. Information from Ohlrogge and Browse (1995), Chapman and Ohlrogge (2012), Tzen (2012), Bates et al. (2013), Bates (2016), Xu and Shanklin (2016), Pyc et al. (2017). PEP phosphoenolpyruvate, GA-3-P glyceraldehyde-3-phosphate, G6-P glucose-6-phosphate, ACCase acetyl-CoA carboxylase, FAS fatty acid synthase, ACP acyl carrier protein, FAs fatty acids, GPAT glycerol-3-phosphate acyltransferase, LPA lysophosphatidic acid, LPAAT lysophosphatidic acid acyltransferase, PA phosphatidic acid, PAP phosphatidic acid phosphatase, DAG diacylglycerol, DGAT diacylglycerol acyltransferase, PC phosphatidylcholine, PDCT PC:DAG cholinephosphotransferase, TAG triacylglycerol. The accumulation of TAGs occurs between the two unit membranes of the ER. This would place at least some reactions shown diagrammatically, within the ER membrane
TAG assembly
Lipids are stored in the seed in the form of TAGs, in which fatty acids are linked by ester bonds with the three available hydroxyl groups of glycerol (Ohlrogge and Browse 1995; Bates et al. 2013; Fig. 2). Fatty acids are transferred from acyl-CoA to the glycerol-3-phosphate backbone at the sn-1 position by glycerol-3-phosphate acyltransferase (GPAT) and the sn-2 position by the lysophosphatidic acid acyltransferase (LPAAT), yielding the central metabolite phosphatidic acid (PA). Phosphatidic acid is then dephosphorylated to diacylglycerol (DAG) and a third fatty acid is transferred to the free sn-3 position of DAG by diacylglycerol acyltransferase (DGAT). DAG can also be converted to phosphatidylcholine (PC) and flux through a PC pool, and then made available for TAG biosynthesis (Bates et al. 2013; Xu and Shanklin 2016). This last fatty acid transfer step for TAG biosynthesis in the sn-3 position, carried out by DGATs, is specific to TAG biosynthesis. There are two major distinctive DGATs (DGAT1 and DGAT2) located in the ER membrane (Bates et al. 2013). Their relative activity is species dependent. There is also an acyl-CoA-independent reaction catalysed by phospholipid:diacylglycerol acyltransferase (PDAT1), not shown in Fig. 2, which uses phospholipids as an acyl donor and DAG as an acyl acceptor to produce TAGs and lysophospholipids (Xu and Shanklin 2016).
Oil body assembly
Free TAGs undergo further packaging into oil bodies. As outlined above, the mature oil bodies in the cytosol are filled with a TAG matrix surrounded by a PL monolayer membrane embedded with the abundant oleosin protein of ~15–30 kDa (Chapman et al. 2012) and the minor caleosin and steroleosin proteins. There are two abundant oleosin isomers that may form heterodimers or heteromultimers on the surface of the oil bodies (Huang 1996). The current understanding is that oleosins are co-translationally inserted into the ER and associate with the TAGs which accumulate between the two phospholipid leaflets (Hsieh and Huang 2004; Shimada and Hara-Nishimura 2010; Chapman et al. 2012; Tzen 2012). The TAGs surrounded by the phospholipid monolayer, with the embedded proteins, eventually bud off to form the oil body (Figs. 1, 2). The models are supported by ultrastructure studies (Herman 2008), linking oleosin biosynthesis to the ER (Sarmiento et al. 1997) and the thermodynamics involved (Huang 1992). Because of their hydrophobicity, the TAGs synthesized in the ER can be sequestered between the two PL layers (Huang 1996). Models for the organization of the integral oil body proteins (oleosin, as well as caleosin and steroleosin) on the surface of the oil body can be found in the review by Tzen (2012) on integral proteins in plant oil bodies. Oleosin has an N-terminal amphipathic domain, a central lipophilic anchoring domain and a C-terminal amphipathic domain (Huang 1992; Tzen et al.1993; Tzen 2012). Around 20% of amino acid residues are immersed in the PL layer, 30% are located in the TAG matrix, and the remaining 50% are exposed to the cytosol (Tzen et al. 1993). Oleosin stabilizes the oil bodies and prevents coalescence of oil bodies by electronegative repulsion and steric hindrance (Tzen et al. 1992; Tzen and Huang 1998). The structural role of oleosins allows the oil bodies to be maintained as relatively small discrete organelles (Tzen 2012) that can be packed into the cytoplasm (Fig. 1). Even with the cytoplasmic compression during dehydration in late maturation, where oil bodies are forced into contact with each other, the organelles resist coalescence and remain as small individual bodies due to oleosins (Murphy 2001). The large surface area per unit of TAG also facilitates lipolysis during germination (Hsieh and Huang 2004).
Extensive phylogenetic studies have identified five oleosin lineages (Huang and Huang 2015). These are P (primitive; in algae, mosses and ferns), U (in all land plants), SL (seed low—molecular weight, in seeds of gymnosperms and angiosperms), SH (seed high molecular weight in seeds of angiosperms) and T (tapetum, in tapeta of Brassicaceae). Legumes have U, SL and SH oleosins (Huang and Huang 2016). In soybean, with 13 oleosin genes, 2 SL and 2 SH oleosin genes produce 90% of the oleosin. A role for different oleosin isoforms in oil body dynamics has been demonstrated using Arabidopsis mutants where not only oil body size, but also spatial distribution is influenced (Miquel et al. 2014). Miquel et al. also show that fusion of smaller oil bodies is a contributor to oil body growth. It seems plausible that the different oil body sizes and distribution are influenced by the way different oleosins are expressed. This could contribute to the differences seen in Fig. 1 between Pongamia and soybean oil body size distribution, and is worthy of further investigation. Another outstanding question in oil body assembly is how the composition difference between the unit phospholipid membrane and the cytosolic leaflet of the ER is generated (Miquel et al. 2014).
The biogenesis of oil bodies can occur in the absence of oleosins in other tissues and organs (Chapman et al. 2012). In the mesocarp of avocado and olive there are ‘oil bodies’ that likely lack oleosin and are subsequently very large, from 10 to 20 μm in diameter; while oil bodies in the seed contain oleosin and are 0.5–2.0 μm in diameter (Ross et al. 1993). Importantly, it is the small oil bodies in the seed that undergo desiccation and are then mobilized for germination (Murphy 2012). It does suggest that the oleosins are important in seed oil bodies to meet the storage, desiccation and germination requirements of the seed. In a more recent study on avocado mesocarp cells (Huang and Huang 2016), oleosin transcripts of a special phylogenic lineage (designated M) were observed in the mesocarp cells. Very small oil bodies (<0.5 μm) on the periphery of the cell in addition to the very large ones (>5.0 μm) were identified. The oleosin was mainly associated with the small oil bodies as determined by immuno-confocal laser scanning microscopy. Very large oil bodies may lack oleosins and be stabilized by the lipid-associated proteins LDAP1 and LDAP2 (Horn et al. 2013) which are similar to small rubber particle proteins. In green algae, oleosins have been detected but are not major proteins (Huang et al. 2013). The 28 kDa hydrophobic protein MLDP (‘Major Lipid Droplet Protein’) is the most abundant protein associated with oil bodies in green algae and shares no primary sequences with oleosin; MLDP, however, appears to influence oil body size and prevent fusion (Goold et al. 2015). In Chlamydomonas when MLDP synthesis is inhibited, oil body size increases (Goold et al. 2015). In addition to oleosins, there are a number of other proteins associated with plant oil bodies (Gallardo et al. 2016). It seems likely that this includes peripheral proteins associated with the surface structures of oil bodies for oil body mobilization during germination (Feussner et al. 2001; Jolivet et al. 2013).
While oleosin has a major influence on oil body size and distribution and maintains the integrity of the oil body in desiccation, seipen is another protein that is important in determining the number and size of oil bodies. Seipens in plants were discovered as homologues of animal and yeast seipens (Cai et al. 2015). Three seipens were found in Arabidopsis and there are three homologues in M. truncatula (Cai et al. 2015). Transient expression of the Arabidopsis genes in Nicotiana benthamiana showed that SEIPIN1 caused the accumulation of large oil bodies, while expression of SEIPIN2 and SEIPIN3 accumulated small oil bodies. SEIPEN1 has its highest expression in developing seeds (Cai et al. 2015). GFP-tagged SEIPENS localized to oil body-forming sites on the ER of N. benthamiana leaves, with the ER identified by the ER marker protein CFP-HDEL (Cai et al. 2015). The morphology of the ER after the co-labelling with GFP-tagged SEIPENS and the CFP-HDEL suggested that SEIPENS are involved in organizing sub-domains of the ER devoted to TAG synthesis and oil body assembly (Fig. 2).
Transcription factors and the regulation of oil body biogenesis
The transcriptional regulation of oil body biogenesis in legumes has not been as extensively investigated compared to Arabidopsis with its 36% seed oil storage (Li et al. 2006), availability of mutants and ease of transgenic analysis.
The transcription factor most directly linked to fatty acid biosynthesis is WRINKLED1 (WRI1), discovered from the isolation of the Arabidopsis wri1 mutant (Focks and Benning 1998). WRINKLED1 regulates genes encoding most of the key enzymes of the later stages of glycolysis and fatty acid biosynthesis (Focks and Benning 1998; Cernac and Benning 2004; Kim et al. 2016). Homologues of WRI1 are widely spread, from Arabidopsis to oil palm (Ma et al. 2013). Homologues are present in legumes (Ma et al. 2013) but have not been the subject of experimental work. It seems likely that in higher plants WRI1, or closely related genes, is the important regulator of fatty acid biosynthesis. WRI1 is a member of the AP2/ERBP transcription factor family (Cernac and Benning 2004) and binds to the AW box sequence [5′-[CnTnG](n)7[CG]-3′] of promoters of the target glycolysis and fatty acid biosynthesis genes (Maeo et al. 2009). Further investigations by Kim et al. (2016) have identified MEDIATOR15 (MED15), a subunit of the Mediator complex, as an interacting partner of WRI1. The work suggests that WRI1 targets the promoters of glycolysis-related and fatty acid biosynthesis genes and then recruits the mediator complex and RNA polymerase II, via MED15, to activate transcription. Another transcriptional regulator (Shi et al. 2012) implicated in the regulation of oil biosynthesis is GLABRA2 (GL2). This is discussed in the “Biotechnology—increasing oil production” section below.
Oil body biogenesis has been linked to the transcriptional regulators of embryo maturation, given that the production of seed storage reserves goes hand in hand with the maturation of the embryo (Kurdyukov et al. 2014). Four ‘master’ transcription regulators have been identified in Arabidopsis as controllers of seed maturation (Meinke et al. 1994; Gutierrez et al. 2007) together with LEAFY COTYLEDON1-like (L1L, Kwong et al. 2003). The four master regulators are LEAFY COTYLEDON1 (LEC1) (West et al. 1994; Lotan et al. 1998; Casson and Lindsey 2006), LEAFY COTYLEDON2 (LEC2) (Stone et al. 2001), FUSCA3 (FUS3) (Bäumlein et al. 1994; Parcy et al. 1997) and ABSCISIC ACID-INSENSITIVE3 (ABI3; Nambara et al. 1995; Parcy et al. 1997). LEC2, FUS3 and ABI3 belong to the B3 family of transcription factors, with B3 DNA-binding domains (Stone et al. 2001). LEC1 belongs to the CBF family of transcription factors and encodes a CCAAT box-binding HAP3 subunit (Lee et al. 2003). LIL is similar to LEC1 but encodes a different HAP3 subunit (Kwong et al. 2003).
A number of studies have directly implicated the master transcription factors in the regulation of oil biosynthesis. Mu et al. (2008) using LEC1-overexpressing Arabidopsis plants under the control of an estradiol-inducible promoter found that a number of enzyme-coding genes in the plastid fatty acid synthesis pathway were up-regulated, including three subunits of ACCase. Genes involved in glycolysis and lipid accumulation were also up-regulated; and oil bodies formed in vegetative tissues. LEC1 function was found to be partially dependent on ABI3, FUS3 and WRI1 (Mu et al. 2008). LEC1 and LEC2 act upstream of ABI3 and FUS3 (To et al. 2006; Wang et al. 2007). FUS3 (Wang et al. 2007) but not ABI3 (Mu et al. 2008) was able to directly stimulate oil biosynthesis genes when overexpressed in Arabidopsis (Wang et al. 2007). In B. napus both LEC1 and LIL when overexpressed act similar to LEC1 overexpression in Arabidopsis (Mu et al. 2008), consistent with similar functions of LEC1 and LIL (Kwong et al. 2003).
Using transgenic Arabidopsis plants with an inducible LEC2, it was shown that LEC2 is somewhat similar to LEC1 in its action. LEC2 activation triggered the accumulation of oil- and seed-specific mRNAs and induced LEC1, FUS3 and ABI3 (Santos Mendoza et al. 2005).
LEC2 and LEC1 (Mu et al. 2008) also control the expression of the AP2/ERFP transcription factor WRI1 which is essential for oil accumulation (Focks and Benning 1998; Cernac and Benning 2004; Baud et al. 2007). Though WRI1 acts downstream of LEC1 and LEC2, and they appear to regulate WRI1, binding to the WRI1 promoter by LEC1 and LEC2 has not been demonstrated (Marchive et al. 2014). It has recently been found that the transcription factor MYB89 is able to directly repress WRI1 (Li et al. 2017). WRI1 and related transcription factors have a key role in the controlled production of fatty acids (Focks and Benning 1998; Cernac and Benning 2004; Baud et al. 2007; To et al. 2012; Marchive et al. 2014).
In Legumes, the sequencing of a number of genomes, Lotus japonicus (Sato et al. 2008), soybean (Schmutz et al. 2010), M. truncatula (Young et al. 2011), chickpea (Varshney et al. 2013), lupin (Yang et al. 2013) and common bean (Schmutz et al. 2014), has enabled identification of orthologues of the legume transcription factors shown to regulate oil body biogenesis in Arabidopsis. In M. truncatula there is a range of proteomic (Gallardo et al. 2007; Thompson et al. 2009) and gene expression data (Gallardo et al. 2007; Udvardi et al. 2007; Verdier et al. 2008; Kurdyukov et al. 2014; Nolan et al. 2014) in relation to the master regulators of oil body biogenesis. There have not been overexpression studies that have been carried out in other species.
Biotechnology—increasing oil production
In addition to up-regulation of lipid biosynthesis, to maximize oil body production it is necessary to consider the partitioning of carbon into oil and packaging of the lipid into oil bodies of suitable size and cellular distribution.
Modifying carbon partitioning
Examination of Table 1 indicates the variable nature of carbon partitioning in legumes and offers the potential for investigating partitioning into different storage products. A number of legumes (e.g. soybean, Pongamia) are rich in oil and protein while others are rich in starch and protein (e.g. chickpea, faba bean), and in some cases protein is quite dominant (e.g. lupin). Where oil is relatively low (Table 1) there can be a trade-off with increased starch (e.g. chickpea) or fibre (e.g. lupin). A hierarchical comparative genomics analysis of ten legume genomes to produce multiple alignments of homologous genes is a new resource for legume biology, which can aid comparative studies (Wang et al. 2017). More than 850 oil synthesis-related genes were located in the peanut genomes and 1528 in soybean (Wang et al. 2017).
There has been experimentation on the effects of reducing starch on oil content. In peas, Weigelt et al. (2009) used RNAi to repress ADP-glucose pyrophosphorylase (AGP) to reduce starch. Reduced starch was associated with increased protein, lipid and sucrose. In mature seed, lipid increased from 40 to 58 mg g−1 and total N increased from 2.9 to 4.3%. There was, however, a yield penalty with seed weight reduced from 345 to 302 mg. There were some negative stress signalling effects but there were responses to circumvent these (Weigelt et al. 2009). In Arabidopsis, using RNAi against AGP was combined with overexpression of the WRI1 transcription factor (Sanjaya et al. 2011). This resulted in less starch and more hexoses and 5.8-fold more oil in vegetative tissues compared with reduced AGP alone or overexpressed WRI1 alone.
In considering carbon partitioning in seed, cell wall storage polysaccharides (mannans, xyloglucans and galactans) are important components (Buckeridge et al. 2000) which have received little attention in legumes. In legumes, mucilage (a galactomannan) is located in the endosperm cell wall (Edwards et al. 1999; Naoumkina et al. 2008). The main function of the endosperm cell walls appears to be storage, with the yield of galactomannan reaching more than 30% of the seed dry weight in many legume species (Buckeridge 2010). Some years ago, in a study based on 300 legume species, it was shown that there was an inverse relationship between legume mucilage and oil plus protein (Tookey and Jones 1965). We have also shown this inverse relationship between Medicago orbicularis with low oil (Tonnet and Snudden 1974) and high endosperm mucilage compared to M. truncatula (unpublished data).
In Arabidopsis, seed coat mucilage which is released from the cell wall on hydration forms a gel-like capsule surrounding the seed. The mucilage may mediate seed dispersal through adhesion or possibly facilitate seed hydration (Western 2012). Carbon partitioning into increased oil production can also be obtained in Arabidopsis by reducing the expression of the transcription factor GLABRA2 (GL2). Mutants of the GL2 transcription factor (Shi et al. 2012) have reduced mucilage on the seed surface and increased oil production. While mucilage is not present in the seed coat of legumes (e.g. M. truncatula; Verdier et al. 2013), as pointed out above, it is present in the remaining seed endosperm. Mucilage and other storage polysaccharides are potentially available to contribute to oil production, perhaps by down-regulating GL2 homologues.
Interestingly, in the study by Roesler et al. (2016) in soybean using improved DGAT variants, a 3.5 percentage point increase of oil and protein was accompanied by a 1.9 percentage point reduction of soluble sugars. Preliminary analysis also indicated a reduction of cell wall polysaccharides and likely the oligosaccharides raffinose and stachyose.
Investigations of the interaction between three of the master regulators of seed maturation LEC2, ABI3 and FUS3 in Arabidopsis provided some information on the partitioning between storage products using single, double and triple mutants and ectopic overexpression (Roscoe et al. 2015). It was concluded that LEC2 influences lipid and protein storage to a similar extent, while ABI3 increased protein relative to lipid and the converse for FUS3.
Metabolic engineering of enzymes involved in TAG biosynthesis
A number of enzyme-coding genes involved in fatty acid synthesis, glycerol synthesis and ultimately TAGs when overexpressed increase oil production (Weselake et al. 2009). These include genes involved in fatty acid synthesis (e.g. homomeric ACCase, Roesler et al. 1997), synthesis of the glycerol backbone (e.g. glycerol-3-phosphate dehydrogenase, Vigeolas et al. 2007), GPAT (Jain et al. 2000) and LPAAT (Zou et al. 1997) involved, respectively, in the first and second acyl-CoA-dependent acylation of the glycerol backbone. Most prominent has been DGAT (Fig. 2) which catalyses the final acyl-CoA-dependent acylation, transferring an acyl group from acyl-CoA to diacylglycerol. This last fatty acid transfer step for TAG biosynthesis in the sn-3 position, carried out by DGATs, is specific to TAG biosynthesis.
DGAT investigations have resulted in field trials demonstrating increased oil production in both canola (B. napus) and soybean. In Arabidopsis overexpression of seed-specific DGAT cDNA increased the oil content from 34 to 46% (Jako et al. 2001). In canola (B. napus) DGAT1 genes from Arabidopsis or B. napus increased oil content from 2.5 to 7% on an absolute dry matter basis, under field conditions (Taylor et al. 2009). In soybean a DGAT1 variant increased the percentage of seed oil by 3 percentage points (approximately from 19 to 22%) based on the T2 and T3 transgenic generations in field trials (Roesler et al. 2016). This increase in seed oil was an improvement on previous studies where the increase was 1.5 percentage points using a DGAT2 from the fungus Umbelopsis ramanniana optimized for expression in soybean (Lardizabal et al. 2008) or a DGAT1 from Sesamum indicum (Wang et al. 2014). The study by Roesler et al. (2016) used amino acid substitutions in soybean DGAT1. The amino acid substitutions were based on the amino acid composition of DGAT1 from the American hazelnut shrub Corylus americana. There are suggestions that DGAT1 overexpression is less sensitive to environmental stress and warrants further study in this context, given the sensitivity of oil production to abiotic stress (Singer et al. 2016).
In Camelina sativa it has been shown that DAG supply can be enhanced by co-expressing two Arabidopsis phospholipase Dζ genes (AtPLD ζ 1 and AtPLD ζ 2) which convert PC to DAG (Yang et al. 2017). This results in a 2–3% increase of TAGs in total seed biomass. Bates (2016) has argued that an enhanced understanding of the control of acyl flux through the lipid metabolic network is required for the best approaches to genetic engineering of oilseeds.
Liu et al. (2015) have been able to show that simultaneous overexpression of multiple TAG biosynthetic genes in a high-oil canola cultivar can increase oil content more than overexpression of a single gene. The simultaneously overexpressed genes were GPDH, GPAT, LPAAT and DGAT.
Transcription factors
The overexpression of the master regulators of seed maturation provide another approach to increasing seed oil content, particularly LEC1 and LEC2 which are able to regulate FUS3 and ABI3 as well as WRI1 (Weselake et al. 2009). As indicated above, when LEC1 is overexpressed in Arabidopsis it is able to regulate many genes involved in oil body biogenesis (Mu et al. 2008). This also leads to increases in the level of major fatty acid species and increases in fatty acid biosynthesis also occur when B. napus LEC1 and L1L are overexpressed in Arabidopsis. Tan et al. 2011 have shown that seed-specific expression of BnLEC1 and BnL1L using truncated napin A promoters in transgenic B. napus increased seed oil by 2–20%. Recent studies with BnLEC1 overexpressed constitutively in B. napus led to an increased seed oil content of 7–16% (Elahi et al. 2016).
Ectopic expression of LEC2 in Arabidopsis vegetative leaves can also induce storage oil in leaves (Santos Mendoza et al. 2005; Stone et al. 2008). Similar results were obtained with transgenic expression of FUS3 which quickly induced fatty acid synthesis in transgenic seedlings and mesophyll protoplasts (Wang et al. 2007). Clearly master TFs are important in approaches to enhance oil body synthesis. Both LEC1 and LEC2 act upstream of FUS3 and ABI3 (Wang et al. 2007; Weselake et al. 2009). It seems based on existing information that LEC1or LEC1L are the master regulators of choice for improved oil biosynthesis (Mu et al. 2008; Weselake et al. 2009; Tan et al. 2011; Elahi et al. 2016).
LEC1 and LEC2 both target the WRI1 transcription factor which has been shown to more directly target fatty acid synthesis. Therefore, it is not surprising that WRI1 overexpression enhances oil biosynthesis in Arabidopsis (Cernac and Benning 2004) and maize (Pouvreau et al. 2011) and is the current focus for enhancing oil biosynthesis (Marchive et al. 2014; Horn and Benning 2016). WRI1 has been a gene of choice in combination with other genes. Vanhercke et al. (2013) have demonstrated what they call a ‘push’ and ‘pull’ approach to obtain increased plant oil levels in N. benthamiana leaves. The ‘push’ involved using WRI1 to increase fatty acid levels and the ‘pull’ to increase TAG assembly using DGAT1. Co-expression of these two genes resulted in a synergistic effect. Given the success of DGAT1 overexpression in increasing soybean levels (Roesler et al. 2016) the additional use of WRI1 is attractive.
Promoting TAG packaging into oil bodies
The size of oil bodies is related to the ratio of oleosin to TAGs. A high ratio of oil to oleosin produces large oil bodies, while small oil bodies have a low ratio of oil to oleosin (Shimada and Hara-Nishimura 2010). This offers the potential to modify oil production by optimizing the oleosin level. A deficiency of oleosin results in an inhibition of oil content (Siloto et al. 2006). Down-regulation of oleosin in Arabidopsis leads to increased oil body size, due to coalescence, and decreased oil production (Siloto et al. 2006). A similar relationship between oleosin and final oil body size has been shown for soybean (Schmidt and Herman 2008). Suppression of soybean oleosin caused the formation of giant oil bodies, observed at the onset of dormancy, but final oil content was not reported. In canola, higher oil content is associated with higher oleosin (Hu et al. 2009) and large oil bodies are characteristic of the lower oil content. Rice seed oil can be increased by overexpression of soybean oleosin and there are more numerous and smaller oil bodies (Liu et al. 2013). Overexpression of DGAT1 and a stabilized cysteine [Cys]-oleosin in Arabidopsis leaves enabled the accumulation of TAG to 2.1% dry weight, 44-fold higher than wild type (Winichayakul et al. 2013). Transient co-expression of WRI1 and OLEOSIN1 in N. benthamiana leaves also increased TAG production (Zhai et al. 2017).
SEIPEN1 can also been used to influence oil content and oil body size. An increased seed oil content of up to 10% in Arabidopsis has been obtained by ectopic expression of SEIPEN1 and was accompanied by an increase in oil body size compared to wild type (Cai et al. 2015). An integral ER protein in animals called FAT STORAGE-INDUCING TRANSMEMBRANE PROTEIN 2 (FIT2) when co-expressed with LEC2 and DGAT2 increased oil content. This suggests FIT2 is present in plant cells and shows the value of ‘push’ (LEC2), ‘pull’ (DGAT2) and ‘protect’ (FIT2) strategies (Pyc et al. 2017).
Oleosins also provide other opportunities for other biotechnological strategies for oil bodies. As discussed by Horn and Benning (2016), it may be possible to use oleosins to interact with other proteins that could act as catalysts for reactions to broaden the engineering capacity of plants. It could be a useful way of fortifying seeds with various proteins for health or to enhance seed protein yields. Yi et al. (2015) used oleosin translational fusion technology (Van Rooijen and Moloney 1995a, b) to produce human growth factor 9 fused to oleosin in Arabidopsis cells.
Modifying fatty acid composition
In addition to oil production, specific fatty acids are best suited for specific purposes such as human health needs, industrialized products and biofuels (Clemente and Cahoon 2009; Haslam et al. 2013). There has been important progress in this area. In soybean, lines with high oleic acid levels have been produced by reducing expression of fatty acid desaturase 2 genes, which catalyse the conversion of the monounsaturated oleic acid to the polyunsaturated linoleic acid, using RNAi (Buhr et al. 2002; Wagner et al. 2011) or by mutations produced by TALENS (Haun et al. 2014; Demorest et al. 2016). The monounsaturated oleic acid is considered better for cardiovascular health than the polyunsaturated linoleic acid. Reduced linoleic acid improves oxidative stability without the production of trans-fatty acids (Clemente and Cahoon 2009). The omega-3 long-chain polyunsaturated fatty acids (LC-PUFAs) such as eicosapentaenoic acid (EPA) and docosahexaenoic (DHA) have been linked to reduced risk of cardiovascular disease and improved aspects of cognition and mental health (Haslam et al. 2013). These fatty acids are not known to be produced in higher plants and are present in fish oil (Haslam et al. 2013). Marine fish oil is considered to be insufficient to meet increasing demand for LC-PUFAs (Haslam et al. 2013; Horn and Benning 2016). In C. sativa up to 12% EPA and 14% DHA of total fatty acids have been obtained when co-synthesized and 31% when EPA alone was targeted (Ruiz-Lopez et al. 2014; Horn and Benning 2016). This was achieved by redirecting endogenous 18:3 through a series of desaturases and elongases from microalgae and fungi. The construct designed to accumulate EPA and DHA in the transgenics involved seven genes (Ruiz-Lopez et al. 2014). In soybean EPA levels of almost 20% of the total seed fatty acids have been produced (Clemente and Cahoon 2009). Long-chain w3 fatty acids and long-chain w6 fatty acids have been produced in transgenic Arabidopsis (Qi et al. 2004) and soybean (Sato et al. 2004). Similar to modifying fatty acids to improve health outcomes fatty acid modification can be used to optimize oil for industrial and biofuel industries. High-oleic and low-palmitic oils are more oxidatively stable than commodity soybean oils which results in reduced NOx emissions (Kinney and Clemente 2005; Santos et al. 2013).
Conclusions and future prospects
There has been successful utilization of the understanding of fatty acid and TAG biosynthesis to increase seed oil in legumes using overexpression of key genes. There are opportunities to build on this using the so-called ‘push’, ‘pull’ and ‘protect’ approach (Pyc et al. 2017). Current genomics in legumes makes it possible to readily locate orthologues based on studies in other species. The optimization of oil body assembly for maximum oil is not completely clear, e.g. see the different oil body size patterns between Pongamia, soybean and M. truncatula oil bodies (Fig. 1). However, both oleosins and seipens have important roles. In legumes, suitable experimental systems are required that can point to the most appropriate genes for stacking, for subsequent whole plant analysis. Such systems are available in the model legume M. truncatula, using transient leaf expression and somatic embryos (Picard et al. 2013; Song et al. 2013).
Regulation of carbon partitioning determines the mix of carbohydrate (sugars, starch, cell wall storage polysaccharides) and oil. Legumes with their diversity of seed protein, oil and carbohydrate offer potential for progressing understanding of the key determinants of storage partitioning. The large amount of variation in the legume family is a major resource for the future (Gresshoff et al. 2015). Improved understanding of partitioning will likely require gene knockdown to reduce carbon flow into some pathways, to promote oil biosynthesis. Knockdown can be facilitated by the utilization of CRISR-cas9 technology (Luo et al. 2016).
There are other opportunities with non-transgenic molecular breeding approaches (Chaudhary et al. 2015). The improved knowledge of oil body biogenesis, particularly in relation to oil body assembly, together with available genomic resources can facilitate development of selection markers. The current knowledge of biochemistry and molecular biology knowledge can be linked to the information derived from QTLs (Quantitative Trait Loci) and genome -wide association studies, available in legume crops such as soybean (Chaudhary et al. 2015). Meta-QTL analysis has been carried out on oil content in legumes (Qi et al. 2011; Zhaoming et al. 2017). Identification of transcript polymorphisms using integrated RNA-seq and bioinformatics is another approach to identify lines varying in oil composition (Goettel et al. 2014). CRISPR-cas9 gene-editing technology can be utilized where there are useful variants of known genes and regulatory regions (Luo et al. 2016).
Biotechnology to enhance oil production in legumes can be utilized not only in oil crops but for nutritional enhancement in other food legumes with low-lipid seed stores. While maximizing oil levels is a major biotechnological objective, there is also substantive potential in modifying fatty acids for different types of oil utilization as well as using oil bodies as a vehicle for seed protein enrichment.
Author contribution statement
YS and RJR designed and wrote the review; X-D Wang did the oil body cytology.
References
Bala M, Nag TN, Kumar S, Vyas M, Kumar A, Bhogal NS (2011) Proximate composition and fatty acid profile of Pongamia pinnata, a potential biodiesel crop. J Am Oil Chem Soc 88:559–562
Bates PD (2016) Understanding the control of acyl flux through the lipid metabolic network of plant oil biosynthesis. Biochim Biophys Acta Mol Cell Biol Lipids 1861:1214–1225
Bates PD, Stymne S, Ohlrogge J (2013) Biochemical pathways in seed oil synthesis. Curr Opin Plant Biol 16:358–364
Baud S, Santos-Mendoza M, To A, Harscoët E, Lepiniec L, Dubreucq B (2007) WRINKLED1 specifies the regulatory action of LEAFY COTYLEDON2 towards fatty acid metabolism during seed maturation in Arabidopsis. Plant J 50:825–838
Bäumlein H, Miséra S, Luerßen H, Kölle K, Horstmann C, Wobus U et al (1994) The FUS3 gene of Arabidopsis thaliana is a regulator of gene expression during late embryogenesis. Plant J 6:379–387
Buckeridge MS (2010) Seed cell wall storage polysaccharides: models to understand cell wall biosynthesis and degradation. Plant Physiol 154:1017–1023
Buckeridge MS, dos Santos HP, Tiné MAS (2000) Mobilisation of storage cell wall polysaccharides in seeds. Plant Physiol Biochem 38:141–156
Buhr T, Sato S, Ebrahim F, Xing A, Zhou Y, Mathiesen M et al (2002) Ribozyme termination of RNA transcripts down-regulate seed fatty acid genes in transgenic soybean. Plant J 30:155–163
Cai Y, Goodman JM, Pyc M, Mullen RT, Dyer JM, Chapman KD (2015) Arabidopsis SEIPIN proteins modulate triacylglycerol accumulation and influence lipid droplet proliferation. Plant Cell 27:2616–2636
Casson SA, Lindsey K (2006) The turnip mutant of Arabidopsis reveals that LEAFY COTYLEDON1 expression mediates the effects of auxin and sugars to promote embryonic cell identity. Plant Physiol 142:526–541
Cernac A, Benning C (2004) WRINKLED1 encodes an AP2/EREB domain protein involved in the control of storage compound biosynthesis in Arabidopsis. Plant J 40:575–585
Chapman KD, Ohlrogge JB (2012) Compartmentation of triacylglycerol accumulation in plants. J Biol Chem 287:2288–2294
Chapman KD, Dyer JM, Mullen RT (2012) Biogenesis and functions of lipid droplets in plants. Thematic review series: lipid droplet synthesis and metabolism from yeast to man. J Lipid Res 53:215–226
Chaudhary J, Patil GB, Sonah H, Deshmukh RK, Vuong TD, Valliyodan B et al (2015) Expanding omics resources for improvement of soybean seed composition traits. Front Plant Sci 6:1021
Chen JCF, Tsai CCY, Tzen JTC (1999) Cloning and secondary structure analysis of caleosin, a unique calcium-binding protein in oil bodies of plant seeds. Plant Cell Physiol 40:1079–1086
Clemente TE, Cahoon EB (2009) Soybean oil: genetic approaches for modification of functionality and total content. Plant Physiol 151:1030–1040
Dam S, Laursen BS, Ørnfelt JH, Jochimsen B, Stærfeldt HH, Friis C et al (2009) The proteome of seed development in the model legume Lotus japonicus. Plant Physiol 149:1325–1340
Davidson EA (2009) The contribution of manure and fertilizer nitrogen to atmospheric nitrous oxide since 1860. Nat Geosci 2:659–662
Demorest ZL, Coffman A, Baltes NJ, Stoddard TJ, Clasen BM, Luo S et al (2016) Direct stacking of sequence-specific nuclease-induced mutations to produce high oleic and low linolenic soybean oil. BMC Plant Biol 16:225
Djemel N, Guedon D, Lechevalier A, Salon C, Miquel M, Prosperi JM et al (2005) Development and composition of the seeds of nine genotypes of the Medicago truncatula species complex. Plant Physiol Biochem 43:557–566
Duranti M, Gius C (1997) Legume seeds: protein content and nutritional value. Field Crops Res 53:31–45
Edwards ME, Dickson CA, Chengappa S, Sidebottom C, Gidley MJ, Reid JS (1999) Molecular characterisation of a membrane-bound galactosyltransferase of plant cell wall matrix polysaccharide biosynthesis. Plant J 19:691–697
Elahi N, Duncan RW, Stasolla C (2016) Modification of oil and glucosinolate content in canola seeds with altered expression of Brassica napus LEAFY COTYLEDON1. Plant Physiol Biochem 100:52–63
Feussner I, Kühn H, Wasternack C (2001) Lipoxygenase-dependent degradation of storage lipids. Trends Plant Sci 6:268–273
Focks N, Benning C (1998) wrinkled1: a novel, low-seed-oil mutant of Arabidopsis with a deficiency in the seed-specific regulation of carbohydrate metabolism. Plant Physiol 118:91–101
Foyer CH, Lam HM, Nguyen HT, Siddique KH, Varshney RK, Colmer TD et al (2016) Neglecting legumes has compromised human health and sustainable food production. Nat Plants 2:16112
Gallardo K, Firnhaber C, Zuber H, Héricher D, Belghazi M, Henry C et al (2007) A combined proteome and transcriptome analysis of developing Medicago truncatula seeds. Evidence for metabolic specialization of maternal and filial tissues. Mol Cell Proteom 6:2165–2179
Gallardo K, Thompson R, Burstin J (2008) Reserve accumulation in legume seeds. C R Biol 331:755–762
Gallardo K, Jolivet P, Vernoud V, Canonge M, Larré C, Chardot T (2016) Storage cells—oil and protein bodies. In: Rose RJ (ed) Molecular cell biology of the growth and differentiation of plant cells. CRC Press, London, pp 362–381
Gepts P, Beavis WD, Brummer EC, Shoemaker RC, Stalker HT, Weeden NF et al (2005) Legumes as a model plant family. Genomics for food and feed report of the cross-legume advances through genomics conference. Plant Physiol 137:1228–1235
Gerrard Wheeler MC, Arias CL, Righini S, Badia MB, Andreo CS, Drincovich MF et al (2016) Differential contribution of malic enzymes during soybean and castor seeds maturation. PLoS One 11:e0158040
Goettel W, Xia E, Upchurch R, Wang ML, Chen P, An YQ (2014) Identification and characterization of transcript polymorphisms in soybean lines varying in oil composition and content. BMC Genom 15:299
Goold H, Beisson F, Peltier G, Li-Beisson Y (2015) Microalgal lipid droplets: composition, diversity, biogenesis and functions. Plant Cell Rep 34:545–555
Graham PH, Vance CP (2003) Legumes: importance and constraints to greater use. Plant Physiol 131:872–877
Gresshoff PM, Hayashi S, Biswas B, Mirzaei S, Indrasumunar A, Reid D et al (2015) The value of biodiversity in legume symbiotic nitrogen fixation and nodulation for biofuel and food production. J Plant Physiol 172:128–136
Guillon F, Champ MJ (2002) Carbohydrate fractions of legumes: uses in human nutrition and potential for health. Br J Nutr 88(S3):S293–S306
Gutierrez L, Van Wuytswinkel O, Castelain M, Bellini C (2007) Combined networks regulating seed maturation. Trends Plant Sci 12:294–300
Hajduch M, Matusova R, Houston NL, Thelen JJ (2011) Comparative proteomics of seed maturation in oilseeds reveals differences in intermediary metabolism. Proteomics 11:1619–1629
Haslam RP, Ruiz-Lopez N, Eastmond P, Moloney M, Sayanova O, Napier JA (2013) The modification of plant oil composition via metabolic engineering—better nutrition by design. Plant Biotech J 11:157–168
Haun W, Coffman A, Clasen BM, Demorest ZL, Lowy A, Ray E et al (2014) Improved soybean oil quality by targeted mutagenesis of the fatty acid desaturase 2 gene family. Plant Biotechnol J 12:934–940
He YQ, Wu Y (2009) Oil body biogenesis during Brassica napus embryogenesis. J Integr Plant Biol 51:792–799
Herman EM (2008) Endoplasmic reticulum bodies: solving the insoluble. Curr Opin Plant Biol 11:672–679
Herridge DF, Peoples MB, Boddey RM (2008) Global inputs of biological nitrogen fixation in agricultural systems. Plant and Soil 311:1–8
Horn PJ, Benning C (2016) The plant lipidome in human and environmental health. Science 353:1228–1232
Horn PJ, James CN, Gidda SK, Kilaru A, Dyer JM, Mullen RT et al (2013) Identification of a new class of lipid droplet-associated proteins in plants. Plant Physiol 162:1926–1936
Hsieh K, Huang AH (2004) Endoplasmic reticulum, oleosins, and oils in seeds and tapetum cells. Plant Physiol 136:3427–3434
Hu Z, Wang X, Zhan G, Liu G, Hua W, Wang H (2009) Unusually large oil bodies are highly correlated with lower oil content in Brassica napus. Plant Cell Rep 28:541–549
Huang AHC (1992) Oil bodies and oleosins in seeds. Annu Rev Plant Physiol Mol Biol 43:177–200
Huang AHC (1996) Oleosins and oil bodies in seeds and other organs. Plant Physiol 10:1055–1061
Huang MD, Huang AH (2015) Bioinformatics reveal five lineages of oleosins and the mechanism of lineage evolution related to structure/function from green algae to seed plants. Plant Physiol 169:453–470
Huang MD, Huang AH (2016) Subcellular lipid droplets in vanilla leaf epidermis and avocado mesocarp are coated with oleosins of distinct phylogenic lineages. Plant Physiol 171:1867–1878
Huang J, Schols HA, van Soest JJ, Jin Z, Sulmann E, Voragen AG (2007) Physicochemical properties and amylopectin chain profiles of cowpea, chickpea and yellow pea starches. Food Chem 101:1338–1345
Huang N-L, Huang M-D, Chen T-LL, Huang AHC (2013) Oleosin of subcellular lipid droplets evolved in green algae. Plant Physiol 161:1862–1874
Jain RK, Coffey M, Lai K, Kumar A, MacKenzie SL (2000) Enhancement of seed oil content by expression of glycerol-3-phosphate acyltransferase genes. Biochem Soc Trans 28:958–961
Jako C, Kumar A, Wei YD, Zou JT, Barton DL, Giblin EM et al (2001) Seed-specific over-expression of an Arabidopsis cDNA encoding a diacylglycerol acyltransferase enhances seed oil content and seed weight. Plant Physiol 126:861–874
Jolivet P, Acevedo F, Boulard C, d’Andréa S, Faure JD, Kohli A et al (2013) Crop seed oil bodies: from challenges in protein identification to an emerging picture of the oil body proteome. Proteomics 13:1836–1849
Kim MJ, Jang IC, Chua NH (2016) MED15 subunit mediates activation of downstream lipid-related genes by Arabidopsis WRINKLED1. Plant Physiol 171:1951–1964
Kinney AJ, Clemente TE (2005) Modifying soybean oil for enhanced performance in biodiesel blends. Fuel Process Technol 86:1137–1147
Kurdyukov S, Song Y, Sheahan MB, Rose RJ (2014) Transcriptional regulation of early embryo development in the model legume Medicago truncatula. Plant Cell Rep 33:349–362
Kwong RW, Bui AQ, Lee H, Kwong LW, Fischer RL, Goldberg RB et al (2003) LEAFY COTYLEDON1-LIKE defines a class of regulators essential for embryo development. Plant Cell 15:5–18
Lardizabal K, Effertz R, Levering C, Mai J, Pedroso MC, Jury T et al (2008) Expression of Umbelopsis ramanniana DGAT2A in seed increases oil in soybean. Plant Physiol 148:89–96
Lee H, Fischer RL, Goldberg RB, Harada JJ (2003) Arabidopsis LEAFY COTYLEDON1 represents a functionally specialized subunit of the CCAAT binding transcription factor. Proc Natl Acad Sci 100:2152–2156
Li Y, Beisson F, Pollard M, Ohlrogge J (2006) Oil content of Arabidopsis seeds: the influence of seed anatomy, light and plant-to-plant variation. Phytochemistry 67:904–915
Li D, Jin C, Duan S, Zhu Y, Qi S, Liu K et al (2017) MYB89 Transcription factor represses seed oil accumulation. Plant Physiol 173:1211–1215
Liu WX, Liu HL, Qu LQ (2013) Embryo-specific expression of soybean oleosin altered oil body morphogenesis and increased lipid content in transgenic rice seeds. Theor Appl Genet 126:2289–2297
Liu F, Xia Y, Wu L, Fu D, Hayward A, Luo J et al (2015) Enhanced seed oil content by overexpressing genes related to triacylglyceride synthesis. Gene 557:163–171
Lotan T, Ohto MA, Yee KM, West MA, Lo R, Kwong RW et al (1998) Arabidopsis LEAFY COTYLEDON1 is sufficient to induce embryo development in vegetative cells. Cell 93:1195–1205
Luo M, Gilbert B, Ayliffe M (2016) Applications of CRISPR/Cas9 technology for targeted mutagenesis, gene replacement and stacking of genes in higher plants. Plant Cell Rep 35:1439–1450
Ma W, Kong Q, Arondel V, Kilaru A, Bates PD, Thrower NA et al (2013) WRINKLED1, a ubiquitous regulator in oil accumulating tissues from Arabidopsis embryos to oil palm mesocarp. PLoS One 8:e68887
Maeo K, Tokuda T, Ayame A, Mitsui N, Kawai T, Tsukagoshi H et al (2009) An AP2-type transcription factor, WRINKLED1, of Arabidopsis thaliana binds to the AW-box sequence conserved among proximal upstream regions of genes involved in fatty acid synthesis. Plant J 60:476–487
Marchive C, Nikovics K, To A, Lepiniec L, Baud S (2014) Transcriptional regulation of fatty acid production in higher plants: molecular bases and biotechnological outcomes. Eur J Lipid Sci Technol 116:1332–1343
Meinke DW, Franzmann LH, Nickle TC, Yeung EC (1994) Leafy cotyledon mutants of Arabidopsis. Plant Cell 6:1049–1064
Miquel M, Trigui G, d’Andréa S, Kelemen Z, Baud S, Berger A et al (2014) Specialization of oleosins in oil body dynamics during seed development in Arabidopsis seeds. Plant Physiol 164:1866–1878
Mohamed AI, Bhardwaj HL, Rangappa M, Hoggard G (1995) Seed characteristics and nutrient composition of selected beans (Phaseolus vulgaris L.) with different ozone tolerance. Plant Foods Hum Nutr 47:29–38
Mu J, Tan H, Zheng Q, Fu F, Liang Y, Zhang J et al (2008) LEAFY COTYLEDON1 is a key regulator of fatty acid biosynthesis in Arabidopsis. Plant Physiol 148:1042–1054
Murphy DJ (2001) The biogenesis and functions of lipid bodies in animals, plants and microorganisms. Prog Lipid Res 40:325–438
Murphy DJ (2012) The dynamic roles of intracellular lipid droplets: from archaea to mammals. Protoplasma 249:541–585
Nambara E, McCourt P, Naito S (1995) A regulatory role for the ABI3 gene in the establishment of embryo maturation in Arabidopsis thaliana. Development 121:629–636
Naoumkina M, Vaghchhipawala S, Tang Y, Ben Y, Powell RJ, Dixon RA (2008) Metabolic and genetic perturbations accompany the modification of galactomannan in seeds of Medicago truncatula expressing mannan synthase from guar (Cyamopsis tetragonoloba L.). Plant Biotechnol J 6:619–631
Nolan KE, Song Y, Liao S, Saeed NA, Zhang X, Rose RJ (2014) An unusual abscisic acid and gibberellic acid synergism increases somatic embryogenesis, facilitates its genetic analysis and improves transformation in Medicago truncatula. PLoS One 9:e99908
Ohlrogge JB, Browse J (1995) Lipid biosynthesis. Plant Cell 7:957–970
Parcy F, Valon C, Kohara A, Miséra S, Giraudat J (1997) The ABSCISIC ACID-INSENSITIVE3, FUSCA3, and LEAFY COTYLEDON1 loci act in concert to control multiple aspects of Arabidopsis seed development. Plant Cell 9:1265–1277
Picard K, Lee R, Hellens R, Macknight R (2013) Transient gene expression in Medicago truncatula. In: Rose RJ (ed) Legume genomics. Methods and protocols, vol 1069. Methods in molecular biology. Humana Press, Springer, New York, pp 215–226
Pickett TA (1950) Composition of developing peanut seed. Plant Physiol 25:210–224
Pouvreau B, Baud S, Vernoud V, Morin V, Py C, Gendrot G et al (2011) Duplicate maize Wrinkled1 transcription factors activate target genes involved in seed oil biosynthesis. Plant Physiol 156:674–686
Pyc M, Cai Y, Greer MS, Yurchenko O, Chapman KD, Dyer JM et al (2017) Turning over a new leaf in lipid droplet biology. Trends Plant Sci 22:596–609
Qi B, Fraser T, Mugford S, Dobson G, Sayanova O, Butler J et al (2004) Production of very long chain polyunsaturated omega-3 and omega-6 fatty acids in plants. Nat Biotechnol 22:739–745
Qi ZM, Wu Q, Han X, Sun YN, Du XY, Liu CY et al (2011) Soybean oil content QTL mapping and integrating with meta-analysis method for mining genes. Euphytica 179:499–514
Rachwa-Rosiak D, Nebesny E, Budryn G (2015) Chickpeas—composition, nutritional value, health benefits, application to bread and snacks: a review. Crit Rev Food Sci Nutr 55:1137–1145
Roesler K, Shintani D, Savage L, Boddupalli S, Ohlrogge J (1997) Targeting of the Arabidopsis homomeric acetyl-coenzyme A carboxylase to plastids of rapeseeds. Plant Physiol 113:75–81
Roesler K, Shen B, Bermudez E, Li C, Hunt J, Damude HG et al (2016) An improved variant of soybean type 1 diacylglycerol acyltransferase increases the oil content and decreases the soluble carbohydrate content of soybeans. Plant Physiol 171:878–893
Roscoe TT, Guilleminot J, Bessoule JJ, Berger F, Devic M (2015) Complementation of seed maturation phenotypes by ectopic expression of ABSCISIC ACID INSENSITIVE3, FUSCA3 and LEAFY COTYLEDON2 in Arabidopsis. Plant Cell Physiol 56:1215–1228
Rose RJ (2008) Medicago truncatula as a model for understanding plant interactions with other organisms, plant development and stress biology: past, present and future. Funct Plant Biol 35:253–264
Ross JH, Sanchez J, Millan F, Murphy DJ (1993) Differential presence of oleosins in oleogenic seed and mesocarp tissues in olive (Olea europaea) and avocado (Persea americana). Plant Sci 93:203–210
Ruiz-Lopez N, Haslam RP, Napier JA, Sayanova O (2014) Successful high-level accumulation of fish oil omega-3 long-chain polyunsaturated fatty acids in a transgenic oilseed crop. Plant J 77:198–208
Sanjaya Durrett TP, Weise SE, Benning C (2011) Increasing the energy density of vegetative tissues by diverting carbon from starch to oil biosynthesis in transgenic Arabidopsis. Plant Biotechnol J 9:874–883
Santos Mendoza M, Dubreucq B, Miquel M, Caboche M, Lepiniec L (2005) LEAFY COTYLEDON 2 activation is sufficient to trigger the accumulation of oil and seed specific mRNAs in Arabidopsis leaves. FEBS Lett 579:4666–4670
Santos EM, Piovesan ND, de Barros EG, Moreira MA (2013) Low linolenic soybeans for biodiesel: characteristics, performance and advantages. Fuel 104:861–864
Sarmiento C, Ross JH, Herman E, Murphy DJ (1997) Expression and subcellular targeting of a soybean oleosin in transgenic rapeseed. Implications for the mechanism of oil-body formation in seeds. Plant J 11:783–796
Sato S, Xing A, Ye X, Schweiger B, Kinney A, Graef G et al (2004) Production of γ-linolenic acid and stearidonic acid in seeds of marker-free transgenic soybean. Crop Sci 44:646–652
Sato S, Nakamura Y, Kaneko T, Asamizu E, Kato T, Nakao M et al (2008) Genome structure of the legume, Lotus japonicus. DNA Res 15:227–239
Schmidt MA, Herman EM (2008) Suppression of soybean oleosin produces micro-oil bodies that aggregate into oil body/ER complexes. Mol Plant 1:910–924
Schmutz J, Cannon SB, Schlueter J, Ma J, Mitros T, Nelson W et al (2010) Genome sequence of the palaeopolyploid soybean. Nature 463:178–183
Schmutz J, McClean PE, Mamidi S, Wu GA, Cannon SB, Grimwood J et al (2014) A reference genome for common bean and genome-wide analysis of dual domestications. Nat Genet 46:707–713
Scott PT, Pregelj L, Chen N, Hadler JS, Djordjevic MA, Gresshoff PM (2008) Pongamia pinnata: an untapped resource for the biofuels industry of the future. Bioenergy Res 1:2–11
Shi L, Katavic V, Yu Y, Kunst L, Haughn G (2012) Arabidopsis glabra2 mutant seeds deficient in mucilage biosynthesis produce more oil. Plant J 69:37–46
Shimada TL, Hara-Nishimura I (2010) Oil-body-membrane proteins and their physiological functions in plants. Biol Pharm Bull 33:360–363
Siddique KH, Johansen C, Turner NC, Jeuffroy MH, Hashem A et al (2012) Innovations in agronomy for food legumes. A review. Agron Sustain Dev 32:45–64
Siloto RM, Findlay K, Lopez-Villalobos A, Yeung EC, Nykiforuk CL, Moloney MM (2006) The accumulation of oleosins determines the size of seed oil bodies in Arabidopsis. Plant Cell 18:1961–1974
Singer SD, Zou J, Weselake RJ (2016) Abiotic factors influence plant storage lipid accumulation and composition. Plant Sci 243:1–9
Song Y, Nolan KE, Rose RJ (2013) Stable transformation of Medicago truncatula cv. Jemalong for gene analysis. In: Rose RJ (ed) Legume genomics Methods and protocols, vol 1069. Methods in Molecular Biology. Humana Press, Springer, New York, pp 203–214
Stone SL, Kwong LW, Yee KM, Pelletier J, Lepiniec L, Fischer RL et al (2001) LEAFY COTYLEDON2 encodes a B3 domain transcription factor that induces embryo development. Proc Natl Acad Sci 98:11806–11811
Stone SL, Braybrook SA, Paula SL, Kwong LW, Meuser J, Pelletier J et al (2008) Arabidopsis LEAFY COTYLEDON2 induces maturation traits and auxin activity: implications for somatic embryogenesis. Proc Natl Acad Sci USA 105:3151–3156
Tan H, Yang X, Zhang F, Zheng X, Qu C, Mu J et al (2011) Enhanced seed oil production in canola by conditional expression of Brassica napus LEAFY COTYLEDON1 and LEC1-LIKE in developing seeds. Plant Physiol 156:1577–1588
Taylor DC, Zhang Y, Kumar A, Francis T, Giblin EM, Barton DL et al (2009) Molecular modification of triacylglycerol accumulation by over-expression of DGAT1 to produce canola with increased seed oil content under field conditions. Botany 87:533–543
Thompson R, Burstin J, Gallardo K (2009) Post-genomics studies of developmental processes in legume seeds. Plant Physiol 151:1023–1029
To A, Valon C, Savino G, Guilleminot J, Devic M, Giraudat J, Parcy F (2006) A network of local and redundant gene regulation governs Arabidopsis seed maturation. Plant Cell 18:1642–1651
To A, Joubes J, Barthole G, Lecureuil A, Scagnelli A, Jasinski S, Lepiniec L, Baud S (2012) WRINKLED transcription factors orchestrate tissue-specific regulation of fatty acid biosynthesis in arabidopsis. Plant Cell 24(12):5007–5023
Tonnet M, Snudden P (1974) Oil and protein content of the seeds of some pasture legumes. Aust J Agric Res 25:767–774
Tookey HL, Jones Q (1965) New sources of water-soluble seed gums. Econ Bot 19:165–174
Tzen JTC (2012) Integral proteins in plant oil bodies. ISRN Bot 2012:Article ID 173954
Tzen JTC, Huang AHC (1998) Surface structure and properties of plant seed oil bodies. J Cell Biol 117:327–335
Tzen JTC, Lie GC, Huang AH (1992) Characterization of the charged components and their topology on the surface of plant seed oil bodies. J Biol Chem 267:15626–15634
Tzen JTC, Cao YZ, Laurent P, Ratnayake C, Huang AHC (1993) Lipids, proteins, and structure of seed oil bodies from diverse species. Plant Physiol 101:267–276
Udvardi MK, Kakar K, Wandrey M, Montanari O, Murray J, Andriankaja A et al (2007) Legume transcription factors: global regulators of plant development and response to the environment. Plant Physiol 144:538–549
Van Rooijen GJ, Moloney MM (1995a) Structural requirements of oleosin domains for subcellular targeting to the oilbody. Plant Physiol 109:1353–1361
Van Rooijen GJ, Moloney MM (1995b) Plant seed oil-bodies as carriers for foreign proteins. Biotechnology 13:72–77
Vanhercke T, El Tahchy A, Shrestha P, Zhou XR, Singh SP, Petrie JR (2013) Synergistic effect of WRI1 and DGAT1 coexpression on triacylglycerol biosynthesis in plants. FEBS Lett 587:364–369
Varshney RK, Song C, Saxena RK, Azam S, Yu S, Sharpe AG et al (2013) Draft genome sequence of chickpea (Cicer arietinum) provides a resource for trait improvement. Nat Biotechnol 31:240–246
Verdier J, Kakar K, Gallardo K, Le Signor C, Aubert G, Schlereth A et al (2008) Gene expression profiling of M. truncatula transcription factors identifies putative regulators of grain legume seed filling. Plant Mol Biol 67:567–580
Verdier J, Dessaint F, Schneider C, Abirached-Darmency M (2013) A combined histology and transcriptome analysis unravels novel questions on Medicago truncatula seed coat. J Exp Bot 64:459–470
Vigeolas H, Waldeck P, Zank T, Geigenberger P (2007) Increasing seed oil content in oil-seed rape (Brassica napus L.) by over-expression of a yeast glycerol-3-phosphate dehydrogenase under the control of a seed-specific promoter. Plant Biotechnol J 5:431–441
Wagner N, Mroczka A, Roberts PD, Schreckengost W, Voelker T (2011) RNAi trigger fragment truncation attenuates soybean FAD2-1 transcript suppression and yields intermediate oil phenotypes. Plant Biotechnol J 9:723–728
Wang H, Guo J, Lambert KN, Lin Y (2007) Developmental control of Arabidopsis seed oil biosynthesis. Planta 226:773–783
Wang X-D, Song Y, Sheahan MB, Garg ML, Rose RJ (2012) From embryo sac to oil and protein bodies: embryo development in the model legume Medicago truncatula. New Phytol 193:327–338
Wang Z, Huang W, Chang J, Sebastian A, Li Y, Li H et al (2014) Overexpression of SiDGAT1, a gene encoding acyl-CoA: diacylglycerol acyltransferase from Sesamum indicum L. increases oil content in transgenic Arabidopsis and soybean. Plant Cell Tissue Organ Cult (PCTOC) 119:399–410
Wang J, Sun P, Li Y, Liu Y, Yu J, Ma X et al (2017) Hierarchically aligning 10 legume genomes establishes a family-level genomics platform. Plant Physiol 174:284–300
Weigelt K, Küster H, Rutten T, Fait A, Fernie AR, Miersch O et al (2009) ADP-glucose pyrophosphorylase-deficient pea embryos reveal specific transcriptional and metabolic changes of carbon-nitrogen metabolism and stress responses. Plant Physiol 149:395–411
Weselake RJ, Taylor DC, Rahman MH, Shah S, Laroche A, McVetty PB et al (2009) Increasing the flow of carbon into seed oil. Biotechnol Adv 27:866–878
West MA, Yee KM, Danao J, Zimmerman JL, Fischer RL, Goldberg RB et al (1994) LEAFY COTYLEDON1 is an essential regulator of late embryogenesis and cotyledon identity in Arabidopsis. Plant Cell 6:1731–1745
Western TL (2012) The sticky tale of seed coat mucilages: production, genetics, and role in seed germination and dispersal. Seed Sci Res 22:1–25
Winichayakul S, Scott RW, Roldan M, Hatier JH, Livingston S, Cookson R et al (2013) In vivo packaging of triacylglycerols enhances Arabidopsis leaf biomass and energy density. Plant Physiol 162:626–639
Xu C, Shanklin J (2016) Triacylglycerol metabolism, function, and accumulation in plant vegetative tissues. Annu Rev Plant Biol 67:179–206
Yang H, Tao Y, Zheng Z, Zhang Q, Zhou G, Sweetingham MW et al (2013) Draft genome sequence, and a sequence-defined genetic linkage map of the legume crop species Lupinus angustifolius L. PLoS One 8:e64799
Yang W, Wang G, Li J, Bates PD, Wang X, Allen DK (2017) Phospholipase Dζ enhances diacylglycerol flux into triacylglycerol. Plant Physiol 174:110–123
Yi S, Yang J, Huang J, Guan L, Du L, Guo Y et al (2015) Expression of bioactive recombinant human fibroblast growth factor 9 in oil bodies of Arabidopsis thaliana. Protein Expr Purif 116:127–132
Young ND, Debellé F, Oldroyd GE, Geurts R, Cannon SB, Udvardi MK et al (2011) The Medicago genome provides insight into the evolution of rhizobial symbioses. Nature 480:520–524
Zhai Z, Liu H, Shanklin J (2017) Phosphorylation of WRINKLED1 by KIN10 results in its proteasomal degradation, providing a link between energy homeostasis and lipid biosynthesis. Plant Cell 29:871–889
Zhaoming Q, Xiaoying Z, Huidong Q, Dawei X, Xue H, Hongwei J et al (2017) Identification and validation of major QTLs and epistatic interactions for seed oil content in soybeans under multiple environments based on a high-density map. Euphytica 213:162
Zou J, Katavic V, Giblin EM, Barton DL, MacKenzie SL, Keller WA et al (1997) Modification of seed oil content and acyl composition in the Brassicaceae by expression of a yeast sn-2 acyltransferase gene. Plant Cell 9:909–923
Acknowledgements
This research was supported by an Australian Research Council (ARC) Grant (CEO348212). We thank Prof. Peter Gresshoff for supplying the Pongamia seed.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare they have no conflict of interest.
Additional information
Communicated by Dr. Neal Stewart.
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Song, Y., Wang, XD. & Rose, R.J. Oil body biogenesis and biotechnology in legume seeds. Plant Cell Rep 36, 1519–1532 (2017). https://doi.org/10.1007/s00299-017-2201-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00299-017-2201-5