Abstract
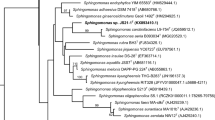
Using antagonistic bacterium is an effective method to control plant disease by fungal pathogens. An aerobic bacterium designated SJ-25, capable of suppressing Fusarium graminearum, Exserohilum turcicum, Pythium aphanidermatum, and Cochliobolus sativus, was isolated from farmland soil. The phylogenetic analysis revealed that strain SJ-25 belongs to the species of Sphingobacterium psychroaquaticum. The genome of strain SJ-25 consists of a 4,396,535-bp chromosome with a G+C content of 41.7 mol%; including 3696 CDS, 64 tRNA genes and six rRNA operons. Genomic analysis revealed that its genome contains multiple genes responsible for biosynthesis of siderophore, methyl 4-hydroxybenzoate, chitinase, giving strain SJ-25 the antagonistic ability on fungi pathogen. Strain SJ-25 harbors sets genes responsible for production of 2, 3-butanediol and salicylic acid, which could elicit the induced systemic resistance of the host plant. This genome sequence could be used as a basis material for further exploration of antagonistic mechanisms on fungi, widening our understanding of the ecological role of the genus Sphingobacterium in farmland ecosystem.




Similar content being viewed by others
References
Muller R, Shinn C, Waldvogel AM, Oehlmann J, Ribeiro R, Moreira-Santos M (2019) Long-term effects of the fungicide pyrimethanil on aquatic primary producers in macrophyte-dominated outdoor mesocosms in two European ecoregions. Sci Total Environ 665:982–994
Haas D, Defago G (2005) Biological control of soil-borne pathogens by fluorescent pseudomonads. Nat Rev Microbiol 3:307–319
Han GZ (2019) Origin and evolution of the plant immune system. New Phytol 222:70–83
Ma J-P, Wang Z, Lu P, Wang H-J, Ali SW, Li S-P, Huang X (2010) Biodegradation of the sulfonylurea herbicide chlorimuron-ethyl by the strain Pseudomonas sp. LW3. FEMS Microbiol Lett 296:203–209
Ito S, Yazawa S, Nakagawa Y, Sasaki Y, Yajima S (2015) Effects of alkyl parabens on plant pathogenic fungi. Bioorg Med Chem Lett 25:1774–1777
Hider RC, Kong X (2010) Chemistry and biology of siderophores. Nat Product Rep 27:637–657
Arrebola E, Tienda S, Vida C, de Vicente A, Cazorla FM (2019) Fitness features involved in the biocontrol interaction of pseudomonas chlororaphis with host plants: the case study of PcPCL1606. Front Microbiol 10:719
Sarwar A, Latif Z, Zhang S, Hao J, Bechthold A (2019) A potential biocontrol agent Streptomyces violaceusniger AC12AB for managing potato common scab. Front Microbiol 10:202
Yin CP, Jin LP, Li S, Xu X, Zhang YL (2019) Diversity and antagonistic potential of Actinobacteria from the fungus-growing termite Odontotermes formosanus. 3 Biotech 9:45
Li YG, Wang RT, Liu JX, Xu LK, Ji PS, Sun L, Pan HY, Jiang BW, Li LR (2019) Identification of a biocontrol agent Bacillus vallismortis BV23 and assessment of effects of its metabolites on Fusarium graminearum causing corn stalk rot. Biocontrol Sci Technol 29:263–275
Sun JQ, Liu M, Wang XY, Xu L, Wu XL (2015) Sphingobacterium suaedae sp. nov., isolated from the rhizosphere soil of Suaeda corniculata. Int J Syst Evol Microbiol 65:4508–4513
Xu L, Sun J-Q, Wang L-J, Gao Z-W, Sun L-Z, Wu X-L (2017) Sphingobacterium alkalisoli sp. nov, isolated from a saline-alkaline soil. Int J Syst Evol Microbiol 67:1943–1948
Ahmed I, Ehsan M, Sin Y, Paek J, Khalid N, Hayat R, Chang YH (2014) Sphingobacterium pakistanensis sp. nov, a novel plant growth promoting rhizobacteria isolated from rhizosphere of Vigna mungo. Antonie Van Leeuwenhoek 105:325–333
Mehnaz S, Weselowski B, Lazarovits G (2007) Sphingobacterium canadense sp. nov., an isolate from corn roots. Syst Appl Microbiol 30:519–524
Matsuda Y, Iida Y, Shinogi T, Kakutani K, Nonomura T, Toyoda H (2001) In vitro suppression of mycelial growth of Fusarium oxysporum by extracellular chitosanase of Sphingobacterium multivorum and cloning of the chitosanase gene csnSM1. J Gen Plant Pathol 67:318–324
Smibert RM, Krieg NR (1994) Phenotypic characterization. In Methods for general and molecular bacteriology. American Society for Microbiology, Washington, DC
Schwyn B, Neilands JB (1987) Universal chemical assay for the detection and determination of siderophores. Anal Biochem 160:47–56
Dong XZ, Cai MY (2001) Determinative manual for routine bacteriology. Scientific Press, Beijing
Tamura K, Stecher G, Peterson D, Filipski A, Kumar S (2013) MEGA6: Molecular evolutionary genetics analysis version 6.0. Mol Biol Evol 30:2725–2729
Sun JQ, Wang XY, Wang LJ, Xu L, Liu M, Wu XL (2016) Saccharibacillus deserti sp. nov, isolated from desert soil. Int J Syst Evol Microbiol 66:623–627
Aziz R, Bartels D, Best A, DeJongh M, Disz T, Edwards R, Formsma K, Gerdes S, Glass E, Kubal M et al (2008) The RAST server: rapid annotations using subsystems technology. BMC Genomics 9:75
Zhou Y, Liang Y, Lynch K, Dennis JJ, Wishart DS (2011) PHAST: a fast phage search tool. Nucl Acids Res 39:347–352
Richter M, Rosselló-Móra R, Glöckner FO, Peplies J (2016) JSpeciesWS: a web server for prokaryotic species circumscription based on pairwise genome comparison. Bioinformatics 32:929–931
Darling AE, Mau B, Perna NT (2010) progressiveMauve: multiple genome alignment with gene gain, loss and rearrangement. PLoS ONE 25:e11147
Acknowledgements
The reported study was supported by High-Level Talent Start-Up Research Project of Inner Mongolia University (Nos. 21800–5185133).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors have declared there was no conflict of interest.
Ethical Approval
This article doesn’t contain any study with human participants or animals performed by any of the authors.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Xu, L., Zhang, H., Xing, YT. et al. Complete Genome Sequence of Sphingobacterium psychroaquaticum Strain SJ-25, an Aerobic Bacterium Capable of Suppressing Fungal Pathogens. Curr Microbiol 77, 115–122 (2020). https://doi.org/10.1007/s00284-019-01789-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00284-019-01789-3