Abstract
Neuraminic acid synthases are an important yet underexplored group of enzymes. Thus, in this research, we performed a detailed kinetic and stability analysis and a comparison of previously known neuraminic acid synthase from Neisseria meningitidis, and a novel enzyme, PNH5, obtained from a metagenomic library. A systematic analysis revealed a high level of similarity of PNH5 to other known neuraminic acid synthases, except for its pH optimum, which was found to be at 5.5 for the novel enzyme. This is the first reported enzyme from this family that prefers an acidic pH value. The effect of different metal cofactors on enzyme activity, i.e. Co2+, Mn2+ and Mg2+, was studied systematically. The kinetics of neuraminic acid synthesis was completely elucidated, and an appropriate kinetic model was proposed. Enzyme stability study revealed that the purified enzyme exhibits changes in its structure during time as observed by differential light scattering, which cause a drop in its activity and protein concentration. The operational enzyme stability for the neuraminic acid synthase from N. meningitidis is excellent, where no activity drop was observed during the batch reactor experiments. In the case of PNH5, some activity drop was observed at higher concentration of substrates. The obtained results present a solid platform for the future application of these enzymes in the synthesis of sialic acids.
Key points
• A novel neuraminic acid synthase was characterized.
• The effect of cofactors on NeuS activity was elucidated.
• Kinetic and stability characterization of two neuraminic acid synthases was performed.
Graphical Abstract

Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Sialic acid synthases are an underexplored family of enzymes, especially from the perspective of their application potential. Still, their role and function in physiological processes are more or less well defined. Prominent members of this family are N-acetylneuraminic acid synthases (NeuSs) that catalyse the formation of N-acetylneuraminic acid (Neu5Ac) by an aldol-like condensation of N-acetylmannosamine (ManNAc) and phosphoenolpyruvate (PEP) (Scheme S1). NeuS enzymes represent attractive targets for protein engineering, in order to alter their properties and provide easy irreversible enzymatic routes towards a wide range of sialic acid analogues (Knorst 1999; Suryanti et al. 2003; Schnellbächer 2017). The latter are very interesting compounds with a potential to serve as novel chemotherapeutic agents against bacterial and viral infections (Koketsu et al. 1997; Gunawan et al. 2005; Mertsch et al. 2020b; Mertsch et al. 2020b; Heida et al. 2021), as well as cancer and immunological disorders (Suryanti et al. 2003; Keil et al. 2022). Sialic acids, a structurally diverse group of more than 50 distinct negatively charged α-keto acids, are widespread in viruses, mammalian cells, and microorganisms controlling a variety of biological functions, such as development, recognition, cell signalling, cell–cell interactions, and adhesion (Varki 1993, 1997; Traving and Schauer 1998; He et al. 2011; Yi et al. 2013; Fessner et al. 2014; Schnaar et al. 2014; Colley et al. 2014). They are terminal components of glycoproteins and glycolipids (Mertsch et al. 2020a). During the embryonic development of vertebrates, neural cell adhesion molecules are sialylated, which allows neural tissue to develop properly (Cunningham et al. 1983). Additionally, in certain cancers, sialylation of the cell surface is correlated with tumorigenesis and metastasis suggesting that sialic acids have a significant influence on human morbidity (Fukuda 1996; Gorelik et al. 2001; Yi et al. 2013). Among the sialic acids in nature, Neu5Ac is the most common and best studied sialic acid (Yi et al. 2013; Mertsch et al. 2020a), which due to their biological profiles are of potential importance for commercial application, i.e. as pharmaceutical precursors for antiviral drugs (e.g. Zanamivir (Zhang et al. 2010; Mertsch et al. 2020b)), food supplements and drug delivery systems nanocarrier stabilizers (Gurung et al. 2013). Since relatively simple structural modifications of sialic acids may significantly change their biological activities (von ltzstein and Thomson 1997), it can be expected that the synthesis of novel derivatives will have a strong potential of bringing innovative medicines to the market (Kiefel and Von Itzstein 2002). Although there are chemical methods for the preparation of Neu5Ac analogues (Warwel and Fessner 2000), sustainable and effective enzymatic methods of preparation will be of great interest for the synthesis and production of pharmaceutically active compounds based on Neu5Ac. Some research was already done on this topic (Mahmoudian et al. 1997; Knorst 1999; Schnellbächer 2017).
Current sources of microbial neuraminic acid synthases are rather scarce. They were first discovered in pathogenic microorganisms, and the first enzyme was isolated and purified from Neisseria meningitidis (Blacklow and Warren 1962). Later, the enzyme was isolated and purified to homogeneity from Escherichia coli K1 (Vann et al. 1997) and then found in Campylobacter jejuni (Linton et al. 2000) and Streptococcus agalactiae (Suryanti et al. 2003). Also, enzymes from the fish pathogens Aliivibrio salmonicida (Gurung et al. 2013) and Moritella viscosa (Berg et al. 2015) were discovered, of which the latter is the first cold-adapted neuraminic acid synthase. The first enzyme from a non-pathogenic source was discovered in Idiomarina loihiensis, a microorganism found in a submarine volcano in Hawaii (García García et al. 2015). Studies up to 2002 were mostly limited to genetic context and detection of activity (Hwang et al. 2002). In most studies, only a basic kinetic analysis of enzyme activity was performed, meaning that the Michaelis constants for PEP and ManNAc were estimated, as well as the maximum reaction rates, and/or catalytic constants (Table S1). It should be also noted here that different assays were used to determine enzyme activity. As these enzymes are cofactor dependent, different divalent metal ions were tested, and for most of them, divalent manganese, magnesium, and cobalt were found to be the most appropriate choices. The first structural study by CD spectroscopy and MALDI-TOF was performed for EcNeuS, along with chemical cross-linking studies (Hwang et al. 2002). The authors found that the C-terminal region of the protein has a significant role in catalytic activity and conformational integrity. Sundaram et al. (2004) performed the first stereochemical analysis with CjNeuS. They also concluded that the purified enzyme was unstable and performed their research with a partially purified enzyme. Suryanti et al. (2003) investigated SaNeuS and suggested that arginine residues present in the active site are involved in substrate recognition, Arg 305 being the only totally conserved. The crystal structure of neuraminic acid synthase was elucidated for the N. meningitidis enzyme (Gunawan et al. 2005), and the mechanism of the action was clarified as well (Gunawan et al. 2005; Liu et al. 2009). Among the neuraminic acid synthases reported so far, most of them come from mesophilic origin and one is from a psychrophilic organism (M. viscosa).
We investigated the natural reaction catalysed by NmNeuS and a novel enzyme from the Prozomix Ltd panel named PNH5, identified as a putative NeuS homolog through sequence analysis using the bioinformatics platform ProzOMIGO and explored the kinetics of the synthesis of Neu5Ac starting from ManNAc and PEP (Scheme S1) as a model system. This reaction represents a good potential for the synthesis of sialic acid analogues. Thus, this research can be viewed to establish a platform that reveals possible bottlenecks in the reaction system and offers potential solutions. Therefore, we were focused on understanding the way this enzyme performs in a synthesis reactor, paying attention to the process, i.e. the reaction kinetics and operational stability of the enzyme, as well as the stability of the enzyme in general. Given the novelty of PNH5, we conducted bioinformatic analyses to elucidate its structural characteristics via homology modelling and establish its similarity to known NeuS enzymes. With the application of protein engineering on NeuS, native enzymes could be modified to accommodate various non-native substrates and provide novel compounds with pharmaceutical potential. Initial research into that direction has already been conducted (Joseph et al. 2013).
Materials and methods
Chemicals and plasmids
ManNAc was purchased from Glentham Life Sciences, UK. Neu5Ac, MgCl2, CoCl2, MnCl2, imidazole, o-benzylhydroxylamine hydrochloride (BnONH2·HCl), and pyridine were obtained from Acros Organics, Belgium. The neuB gene (GenBank: EFM03356.1) containing pET19b for NmNeuS and pnh5 gene (GenBank: PP887683) containing pET28a equivalent for PNH5 expression vectors in BL21 (DE3) cells were provided from a TU Darmstadt collection (Schnellbächer 2017) and Prozomix Ltd, UK, respectively. Trifluoroacetic acid and Tris buffer were from Fisher Bioreagents, USA. Acetonitrile was from VWR, USA. Synergi™ Fusion-RP (80 Å, 4 μm, 4.6 × 250 mm) HPLC column was from Phenomenex, USA. The methanol gradient grade was from J.T. Baker, USA.
Quantification of compounds by HPLC
The samples from the reaction mixture were analysed by HPLC equipped with a PDA detector at 215 nm (Shimadzu, Japan), and the analysis was performed at 50 °C. A gradient method was applied for the analysis with two mobile phases, i.e. mobile phase A: 0.1% (v/v) trifluoroacetic acid in ultra-pure water and mobile phase B: mixture of acetonitrile and water in a volume ratio 80:20 with the addition of 0.1% (v/v) trifluoroacetic acid. The gradient was programmed as follows: 10–34% B over 8 min, 34–37% B over 3 min, 37% B for 5 min, 37–50% B over 4 min, 50–100% B over 3 min, 100% B for 2 min, 100–10% B over 4 min, and 10% B for 3 min. The flow rate was 0.8 mL min−1.
ManNAc, PEP, and Neu5Ac were derivatised before the analysis with BnONH2·HCl (Garrabou et al. 2009; Česnik et al. 2019) by using the following modified procedure. A 10 μL of the reaction sample was mixed with 50 μL of the derivatisation solution (130 mM BnONH2·HCl in pyridine, methanol and water mixture containing 66, 30, and 4% v/v, respectively) and incubated at 50 °C and 900 rpm for 60 min. Then, 500 μL MeOH was added, and samples were centrifuged for 5 min at 14 000 rpm. Then, they were filtered through 0.22 μm hydrophobic PTFE filters and analysed by using the method described above. The linearity of the calibration curve was up to 25 and 15 mM for ManNAc and Neu5Ac, respectively. The retention times of PEP, Neu5Ac, and ManNAc derivatives were 3.7, 11.4, and 12.4 min, respectively (Figs. S1–S3).
Purification of NmNeuS and PNH5
E. coli BL21 (DE3) cells transformed with the neuB gene from Neisseria meningitidis (GenBank: EFM03356.1) (Bateman et al. 2021) containing recombinant plasmid (pET19b-Nme_neuB) were used for the overexpression of the protein of interest. A 5 µL of glycerol stock solution (OD600 = ~ 1.5) was inoculated into 10 mL of LB medium containing ampicillin (100 µg mL−1) and incubated overnight at 37 °C and 180 rpm. A 4 mL from the starter culture was inoculated into 400 mL of Prozomix® modified TBp medium containing ampicillin (100 µg mL−1) and incubated at 26 °C and 280 rpm for 22 h. After cultivation, the cells were harvested by centrifugation at 4000 rpm for 15 min. The cell pellets were resuspended in buffer A (20 mM sodium phosphate, 500 mM NaCl, 30 mM imidazole pH 7.5, 1.0 mg mL−1 lysozyme) and incubated on ice for 30 min. The resuspended cells were disrupted by sonication and insoluble cell debris and protein aggregates were removed by centrifugation (14,000 rpm, 40 min). The supernatant was filtered through a 0.45-µm filter and then applied to a 5 mL HisTrap™ HP column (GE Healthcare, USA), which was pre-equilibrated with buffer A. The column was washed with 25 mL of buffer A and the NmNeuS was eluted with buffer A containing different imidazole concentrations. The collected fractions were analysed by SDS-PAGE (Laemmli 1970) to identify fractions containing pure protein. For PNH5, cell-free extract (CFE) was mixed with buffer A, and the obtained lysate was clarified by centrifugation (14,000 rpm, 40 min). The supernatant was filtered through a 0.45-µm filter, and then the same procedure was used for the purification. The collected fractions were subjected to dilution up to a final volume of 50 mL with the reaction buffers (150 mM sodium phosphate pH 7.5 and sodium acetate pH 5.5 for NmNeuS and PNH5, respectively) containing 5 mM ethylenediaminetetraacetic acid (EDTA) for the removal of Ni2+ ions leaked from the HisTrap™ HP column during the purification process and other metals present. Enzyme solutions were concentrated with ultracentrifuge tubes (MM Amicon Ultracel-10 K). Subsequently, the concentrated enzyme samples were diluted and washed three times with reaction buffers aimed at the complete elimination of residual EDTA (Mónico et al. 2017). The concentrated enzyme solutions were again centrifuged for the removal of aggregates (14,000 rpm, 30 min). This process yielded 12 mL of NmNeuS at a concentration of 1.21 mg/mL and 10 mL of PNH5 at a concentration of 1.14 mg mL−1, with specific activities of 9.52 U mg−1 and 3.76 U mg−1, respectively.
Activity assay for the determination of neuraminic acid synthase activity
Neuraminic acid synthase (NeuS) activity was determined from the change in concentration of Neu5Ac formed in the metal-dependent condensation of ManNAc and PEP (Scheme S1). For the first activity screening, reactions were performed in sodium phosphate acid buffer (150 mM, pH 7.5). Subsequently, optimum reaction buffers were employed for each enzyme in the ensuing experimental procedures.
For the screening assays, the solution containing 5 mM PEP, 5 mM ManNAc, and 0.05 mM Co2+ was prepared in buffer in a total volume of 2 mL. The reactor was thermostated at 37 °C and shaked at 900 rpm. The reaction was started by the addition of 50 μL of 0.5 mg mL−1 stock solution of pure NeuS. Six samples were taken from the reaction mixture at specific time intervals and analysed as described above. Sampling was done for approximately 25 min, and from the linear slope of the curve, representing the change of Neu5Ac concentration in time, specific enzyme activity is calculated (Eq. 1) (S.A. specific enzyme activity, U mg−1; c molar concentration, mM; Vr reactor volume, mL; VNeuS enzyme volume, mL; γNeuS mass concentration, mg mL–1):
One unit of NmNeuS and PNH5 activity was defined as the amount of enzyme needed to catalyse the formation of 1 µmol Neu5Ac per minute at 37 °C in 150 mM sodium phosphate buffer, pH 7.5 for NmNeuS and 150 mM NaOAc buffer, and pH 5.5 for PNH5, respectively.
Screening of putative PNHs from Prozomix
A metagenomic library of wild-type PNHs sourced from diverse locations within the United Kingdom (Marshall et al. 2021) was sought to evaluate their potential for the aldol-like condensation of ManNAc and PEP to yield Neu5Ac. Reaction mixtures containing initial concentrations of 5 mM PEP, 5 mM ManNAc, and 1 mM Co2+ were prepared in 150 mM sodium phosphate pH 7.5 in a total volume of 950 μL. Reactors were thermostated at 37 °C and shaked at 900 rpm. The reactions were initiated by the addition of 50 μL of 4 mg mL−1 of CFEs. Then, 10 μL of the initial and 24-h samples were taken from the reaction mixture and analysed as described previously to elucidate PNHs activity in the synthesis of Neu5Ac. Subsequently, the PNHs yielding Neu5Ac underwent further investigation aimed at determining their specific activity rates in the synthesis of Neu5Ac. This evaluation was conducted by employing the activity assay described in the aforementioned section with the concentration of PNH in the assay of 0.2 mg mL−1.
Bioinformatic analysis and molecular modelling PNH5
To gain a comprehensive insight into the structure–function relationship of PNH5, bioinformatic analyses have been utilized. Amino acid sequences of NeuSs from UniProt (Bateman et al. 2021) were aligned with Clustal Omega (Sievers et al. 2011) (UniProt accession number for each NeuS amino acid sequence; Neisseria meningitidis serogroup B, Q7DDU0; Aeromonas caviae, Q9R9S2; Aliivibrio salmonicida, B6EHB9; Campylobacter jejuni, Q7BC41; Edwardsiella ictaluri, C5BCQ8; Escherichia coli, Q46675; Moritella viscosa, A0A090IGE4; Pseudomonas aeruginosa, Q8KH52; Streptococcus sp. HMSC069D09, A0A1F0CHD3). The graphical output of the multiple sequence alignment of NeuSs was prepared by using ESPRIPT 3.0 server (Robert and Gouet 2014) and NmNeuS (PDB ID: 1XUZ) as secondary structure information. The homology model of PNH5 was created using AlphaFold2 based on the published structures (PDB ID: 1VLI, 1VS1, 1WVO, 2WQP, 3G8R, 3UZJ, 4IPI, 4IPJ, 6NCS, 6PPW, 8H2C) (Berman et al. 2003; Eastman et al. 2017; Mirdita et al. 2017, 2019, 2022; Mitchell et al. 2019; Steinegger et al. 2019; van Kempen et al. 2023). The theoretical molecular weights of NmNeuS and PNH5 were calculated via Expasy compute pI/Mw tool (Gasteiger et al. 2005). The graphics were done by PyMOL (Molecular Graphics System, Version 2.0 Schrodinger, LLC).
The influence of buffer and pH on enzyme activity
The specific activities of NmNeuS and PNH5 were measured in 150 mM buffers; sodium acetate for pH 5–5.5, sodium phosphate for pH 6–8, Tris–HCl for pH 8.5, and glycine–NaOH for pH 9–10. The effect of pH on the activity of NmNeuS was determined by incubating the reaction mixture (1 mL total volume) with selected buffers containing initial 5 mM concentrations of ManNAc and PEP, and 1 mM MnCl2 for NmNeuS and 10 mM concentrations of ManNAc and PEP, and 0.05 mM CoCl2 for PNH5, respectively. The reactors (1.5 mL Eppendorf tubes) were thermostated at 37 °C and shaked at 900 rpm. Sampling and analysis were done as described previously. The concentrations of NmNeuS and PNH5 in the assay were 0.0125 and 0.023 mg mL−1, respectively.
Determination of enzyme kinetics
The enzyme kinetics were determined by using the initial reaction rate method. The influence of concentrations of all reaction compounds on the specific enzyme activity was evaluated, i.e. ManNAc, PEP, Neu5Ac, Co2+, Mn2+, and Mg2+. The enzyme activity was calculated according to Eq. (1) based on the change in product concentration in time. During these measurements, the substrate conversion was always below 10%. The experimental conditions for each data series are presented in the legends below the corresponding figures. It needs to be noted here that the effect of concentrations of each compound was investigated independently, which means that all the other reaction conditions were kept constant. For example, when the influence of ManNAc concentration on enzyme activity was evaluated, all other concentrations were kept constant, i.e. PEP and Co2+. Experiments were performed at 37 °C on a shaker at 900 rpm. The volume of the reaction mixture was 1 mL, and the total sampling volume was always below 10%. The experiments were carried out in 1.5 mL Eppendorf tubes.
NmNeuS and PNH5 storage stability
The evaluation of the enzyme storage stability was conducted through the integration of three methodologies, i.e. the assessment of enzyme activity using the activity assay described above, by following protein concentration using the Bradford method (Bradford 1976) and the quantification of protein size distribution through particle size distribution analysis (PSD). During storage, changes in protein structure occur and can lead to deactivation. These were followed by the dynamic light scattering (DLS) method. DLS is a highly sensitive technique capable of detecting particles ranging from sub-nanometre sizes up to several micrometres, depending on the specific instrument, which makes it suitable for detecting solvated protein molecules in homogeneous mixtures, as well as larger, insoluble aggregates. The characterization of particle distribution with varying hydrodynamic diameters (Dh) was achieved through the utilization of the number size distribution (NSD), the volume size distribution (VSD), and the intensity size distribution (ISD). NSD demonstrates the number of particles, serving as an indicator of relative protein concentration, and VSD demonstrates the volume that particles occupy. On the other hand, IDS represents the amount of light scattered by the particles in the sample, highlighting the presence of larger particles, which makes it particularly suitable for the early identification of aggregate formation. Increases in the size of protein molecules, exceeding those observed in enzyme crystal structures, are commonly ascribed to protein aggregation, a phenomenon detectable by monitoring PSD (Stetefeld et al. 2016; Milčić et al. 2022). Distributions were monitored by DLS on a Zetasizer Ultra (Malvern Panalytical, Malvern, United Kingdom). Pure NmNeuS and PNH5 enzyme stocks, stored at 4 °C in 150 mM of sodium phosphate buffer pH 7.5 (NmNeuS and PNH5) and 150 mM sodium acetate pH 5.5 (PNH5), were analysed during approximately 50 days of storage. Bovine serum albumin (BSA), well-characterized protein in terms of size and distribution, was used as a standard, while each measurement of NeuS enzymes samples was preceded by control measurement with a buffer solution same as sample matrix. Each measurement was performed in triplicate with a temperature equilibration time of 120 s at 25 °C.
The influence of reaction conditions on enzymatic synthesis of Neu5Ac in batch reactor experiments
Two series of batch reactor experiments were conducted at different reaction conditions, i.e. substrates and enzyme concentration, one with NmNeuS, and the other with PNH5. During these experiments, the concentration of the reaction compounds was followed, as well as enzyme activity to evaluate the enzyme operational stability. Reactions of 15 mL containing ca. 10, 20, 50, and 100 mM equimolar concentrations of ManNAc and PEP and 0.05 mM of Co2+ were initiated in 15-mL centrifuge tubes by the addition of 0.021 and 0.053 mg mL−1 of NmNeuS and PNH5, respectively. Designated concentrations represent the amount of enzyme in the reaction mixture, i.e. reactor. They correspond to activities of 0.34 and 0.30 U mL−1 of NmNeuS and PNH5, respectively. Subsequently, the reaction mixtures were distributed in 1 mL aliquots among 15 conical 1.5-mL Eppendorf tubes for the facilitation of sample collection. They were placed on a shaker at 900 rpm and 37 °C. At predetermined time points, 10 µL of the reaction mixture was taken from each aliquot for the quantification of substrate and product concentrations. Then, 0.99 mL reaction mixture from each aliquot was used to isolate the enzyme and subsequently separate it from the other components of the reaction mixture via ultracentrifugation in Amicon™ Ultra-0.5 centrifugal unit (Merck, USA; 4 °C, 14 000 rpm, 5 min). After the first round of ultracentrifugation, the concentrated reaction sample underwent successive washes with reaction buffers (450 µL × 2 times) to eliminate all components except for the enzyme. Eventually, the obtained enzyme samples (100 µL) were employed to initiate the activity assays (NmNeuS (10 mM PEP and ManNAc, 0.05 mM Co2+, 990 µL total reaction volume); PNH5 (20 mM PEP and ManNAc, 0.05 mM Co2+, 1050 µL total reaction volume in 1.5 mL conical Eppendorf tubes)). In these experiments, the concentrations of compounds were followed by HPLC as described before and the activities of NmNeuS, and PNH5 were calculated according to Eq. (1).
Data handling and development of the mathematical model
The experimental data on the dependence of the specific enzyme activity on the concentration of reaction compounds was used to estimate the kinetic constants. For this purpose, program package SCIENTIST and non-linear regression analysis were used. Simplex and Least squares methods implemented in SCIENTIST were used for the estimation of the kinetic parameters. The same software was used for the simulation of data.
The reaction rate (r) of the aldol-like condensation of ManNAc and PEP catalysed by NmNeuS was described by double substrate Michaelis–Menten kinetics (Eq. (2)), whereas for PNH5, the inhibition by PEP was included in the model (Eq. (3)). In both equations, there is a factor f included (Eq. (4)). It defines the influence of cofactor Co2+ concentration on the reaction rate. Namely, it is known that enzyme activity is dependent upon the concentration of the cofactor. It is however put in a separate equation because the cofactor is not spent in the reactor, but it affects the reaction rate.
Equations (5)–(7) represent the mass balance equation for ManNAc, PEP, and Neu5Ac in the batch reactor, which were used for the reaction simulation.
If operational stability decay occurs, it was assumed that it will occur by the kinetic model of the first order according to Eq. (8):
Results
In this work, the biocatalytic synthesis of Neu5Ac was investigated via aldol-like addition of ManNAc and PEP. For this purpose, we investigated the previously known enzyme from Neisseria meningitidis and evaluated a panel of putative enzymes from Prozomix Ltd. After the initial screening and the selection of the best candidate, we performed a preliminary characterization of the new enzyme, followed by thorough kinetic analysis for both enzymes, revealing the differences and similarities between them. Further, we conducted a computational study to model the molecular structure of PNH5 and its similarity to the currently known enzymes. We also focused on enzyme stability at both storage conditions and reactor conditions. A kinetic model of the synthesis reaction was developed and validated in the batch reactor.
Screening of putative homologues of neuraminic acid synthases
A panel of 13 putative homologues of NeuS from Prozomix Ltd (PNHs) was screened in the synthesis of Neu5Ac to find the best representative for further characterization according to the procedure described in the Experimental part. First, the enzymes were compared according to Neu5Ac production efficiencies (Fig. 1a). From these results, 5 hits, i.e. PNH2, PNH3, PNH5, PNH6, and PNH11, were further characterized by comparing their specific activities (Fig. 1b) in the targeted reaction. The best activity was obtained with PNH5, which was further characterized as a representative of the panel.
a PNH panel screening in Neu5Ac synthesis via metal-dependent condensation of ManNAc and PEP catalysed by PNHs (cManNAc = 5 mM, cPEP = 5 mM, cCo2+ = 1 mM, γPNH = 0.2 mg mL−1, 150 mM HEPES buffer pH 7.5, 24 h, T = 37 °C). b Comparison of specific activities of the best selected PNHs from (a) (cManNAc = 10 mM, cPEP = 10 mM, cCo2+ = 0.05 mM, γPNH = 0.2 mg mL.−1, 150 mM sodium phosphate buffer pH 7.5, T = 37 °C)
Bioinformatic analysis and molecular modelling of PNH5
As PNH5 showed the highest Neu5Ac production efficiency among the putative homologues, bioinformatic analyses were used to obtain a comprehensive insight into its structure–function relationship. The subsequent section aims to elucidate and interpret the implications of these analyses. The theoretical molecular weights of NmNeuS and PNH5 were calculated based on amino acid sequences as 38,347 and 38,377 Da, respectively. The accurate molecular weight of NmNeuS was determined with ESI–MS as 38,649.8 Da (Hao et al. 2005). Therefore, the determined molar masses exhibit a close concordance with the computed values.
Fig. S4 illustrates a structural alignment depicting the sequence comparison of various NeuS homologs. Most residues associated with secondary structure elements exhibit either identical matches or preservation of similar physiochemical properties. Across different NeuB sequences, the amino acids crucial for ligand binding in NmNeuS and PNH5 (Mn2+, PEP, and ManNAc) remain conserved. PNH5 exhibits the highest sequence identity with NeuS from Moritella viscosa (62.43%).
Structure similarity of NmNeuS and PNH5
Up to now, NmNeuS is the only NeuS enzyme for which a crystal structure has been solved (Gunawan et al. 2005; Liu et al. 2009). The substrate binding situation can be gathered from a crystal structure of NmNeuS that contains Mn2+, PEP, and a reduced non-reactive analogue of ManNAc (rManNAc) (Gunawan et al. 2005). In addition, NmNeuS has been crystallized with an analogue of the reaction intermediate (Liu et al. 2009). As seen in Fig. 2a, PNH5 has been identified as a domain-swapped homodimer. The monomers consist of a (β/α)8 catalytic barrel at the N-terminus coupled to an antifreeze protein-like (AFPL) domain at the C-terminus via an extended flexible linker region (Gunawan et al. 2005). The catalytic barrel of one monomer is capped by the AFPL domain on the other monomer during dimer assembly. As a result of the homology searches against type III antifreeze proteins, the AFPL domain had previously been identified and was suggested to function in polar sugar-substrate binding (Baardsnes and Davies 2001). It has since been demonstrated that the AFPL domain plays an essential role in the catalysis of NmNeuS by contributing active site residues, which provide the catalytic orientation of the ManNAc within the active site (Joseph et al. 2013, 2014).
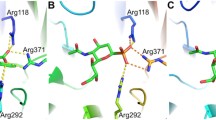
a Homology model of PNH5 dimer created with AlphaFold2 based on the published structures. PNH5’s Chain A is coloured grey, and Chain B is coloured by domain. The catalytic domain of PNH5 chain B is shown in green (residues Met1-Gly271), the linker domain in red (residues Ser272-Val281), and the AFPL domain (residues Thr282-Glu346) in yellow. PNH5 chain B monomer is superimposed with the structure of NmNeuS monomer coloured blue. rManNAc is coloured cyan, PEP orange, and Mn2+ magenta. b PNH5 active site. The N-terminal catalytic domain, linker region, and AFPL residues are coloured in blue, cyan, and magenta, respectively. rManNAc, PEP, and Mn+2 are coloured in green, orange, and pink, respectively
The corresponding model of PNH5’s active site architecture with bound Mn2+, PEP, and rManNAc is illustrated in Fig. 2b. The N-terminal domain of PNH5 contributes most of the catalytic residues. Single residues Phe285 and conserved the Arg311 are contributed by the AFPL domain. Arg311 interacts directly with the N-acetyl carbonyl of rManNAc in NmNeuS, while Phe285 forms part of the hydrophobic pocket. Phe285 is present in some members among the bacterial NeuS enzymes with AFPL domain, whereas Arg311 is completely conserved. Additionally, Arg311 is not present in AFPL domains of the mammalian Neu5Ac-9-phosphate synthase (Gunawan et al. 2005).
Furthermore, ManNAc interacts with Asp245, Gln53, and Tyr184 within the fully conserved catalytic residues besides Arg314. The C-6 hydroxyl of ManNAc interacts with Asn72, which contributes to the β2-α2 loop of the catalytic domain. While Asn72 is fully conserved in NeuS enzymes, it is not present in other sialic acid synthase enzymes. The phosphate group of PEP interacts via H-bonds with Ser130, Ser152, and Ser211 residues, which are fully conserved among the bacterial sialic acid synthases and mammalian Neu5Ac-9PSs. Additionally, the phosphate group of PEP also interacts with Asn182 which is not completely conserved among the sialic acid synthase family. The carboxylate group of PEP is interacting with the conserved Lys51 and Lys127 residues. While Thr108’s reactive group is close to the PEP carboxylate, it is not within the hydrogen bonding range. Additionally, among the sialic acid synthase family, the residue is conserved as either Ser or Thr (Gunawan et al. 2005).
Imidazole chains of His213 and His234, which are fully conserved among both bacterial and mammalian sialic acid synthases, form interactions with Mn2+ at the active site owing to metal’s octahedral geometry, indicating that the metal dependency is a key evolutionary characteristic of these enzymes (Gunawan et al. 2005).
The detailed analysis of PNH5’s active site, encompassing interactions with Mn2+, PEP, and ManNAc, underscores the significance of specific residues, such as Arg311 and Phe285, in catalytic orientation. Moreover, the conserved nature of key residues, including Asn72 and imidazole chains of His213 and His234, across bacterial and mammalian sialic acid synthases highlights the evolutionary importance of metal dependency in these enzymes. These findings not only enhance our understanding of PNH5’s catalytic mechanism as a NeuS but also provide a foundation for further exploration into the broader functions and potential applications of NeuS enzymes in the context of sialic acid biosynthesis.
pH and temperature dependence of enzyme activity
The influence of pH on the activity of NmNeuS and PNH5 is presented in Fig. 3. While the NmNeuS shows an optimum activity at pH 7.5 (Fig. 3a) just like reported by Hao et al. (2005), PNH5 prefers acidic pH value and exhibits its maximum activity at pH 5.5 (Fig. 3b). This is untypical for the NeuS homologs found so far and is reported for the first time for this enzyme family.
Effect of pH on (a) NmNeuS activity (cManNAc = 5 mM, cPEP = 5 mM, cMn2+ = 1 mM, γNeuS = 0.0125 mg mL−1, 150 mM buffer, 37 °C, 100% activity corresponds to 10.62 U mg−1) and (b) PNH5 activity (cManNAc = 10 mM, cPEP = 10 mM, cCo2+ = 0.05 mM, γPNH5 = 0.023 mg mL−1, 37 °C, 150 mM buffer, 100% activity corresponds to 5.02 U mg−1). Legend: black circle, sodium acetate buffer; black square, sodium phosphate buffer; black triangle, Tris–HCl buffer; black triangle down, glycine–NaOH
The effect of different buffers was evaluated for NmNeuS, and the results are presented in Fig. S5 showing an optimum activity in HEPES buffer. The choice of pH 7.6 was made to investigate the impact of the buffer on the activity of NmNeuS. Additionally, this pH value was selected to accommodate the inclusion of bicine buffer in the experimental buffer set. For practical reasons, sodium phosphate buffer pH 7.5 showing 91% of maximum activity was used in further characterization in the case of NmNeuS, whereas sodium acetate buffer pH 5.5 was selected for PNH5.
The influence of temperature on the activity of both enzymes is presented in Fig. 4. Both enzymes show an optimum activity at 50 °C. Considering the temperature profiles, 37 °C was chosen for subsequent experiments as the enzyme activity at this temperature is satisfactory, while stability is presumably higher than at the temperature optimum.
The influence of cofactor on the activity of NmNeuS and PNH5
The results presented in Fig. 5. indicate that the influence of different concentrations of all three cofactors can be described by Michaelis–Menten kinetics for both NmNeuS and PNH5, respectively. Additionally, both enzymes exhibit the maximum activity in the presence of Co2+ ions as cofactor (Fig. 5c and 5f), which was highlighted in the work of Hao et al. (2005) for NmNeuS. The experimental data (Fig. 5) were used to estimate the values of the kinetic parameters presented in Table 1. It can be concluded from the kinetic parameters that Mg2+, even though it was often used with NmNeuS (Blacklow and Warren 1962; Hao et al. 2005), is not a good choice because it suffers from the highest value of the apparent Michaelis constant. When it comes to PNH5, Mg2+ is by far the worst cofactor, considering that the estimated maximum reaction rate is only 0.11 U mg−1. Both enzymes exhibit high affinity towards Co2+ shown by low apparent Km values in Table 1. This means that a very low cobalt concentration is necessary to obtain maximum reaction rate, i.e. specific activity.
Dependence of specific enzyme activity on the concentrations of metal cofactors for NmNeuS (a–c cManNAc = 10 mM, cPEP = 10 mM, 150 sodium phosphate buffer pH 7.5, 37 °C) and PNH5 (d–f cManNAc = 20 mM, cPEP = 20 mM, 150 sodium acetate buffer pH 5.5, 37 °C, γPNH5 = 0.300 mg mL−1): a Mg2+ (γNeuS = 0.081 mg mL−1), b Mn2+ (γNeuS = 0.330 mg mL−1), c Co2+ (γNeuS = 0.33 mg mL−1), d Mg2+, e Mn2+, and f Co2+
Kinetics of aldol-like condensation of ManNAc and PEP catalysed by NmNeuS and PNH5
Having identified Co2+ as the optimum cofactor, a thorough investigation of the reaction kinetics was conducted for both enzymes. This means that the influence of ManNAc, co-substrate PEP, and Neu5Ac concentrations on the specific enzyme activity was evaluated. The results presented in Fig. 6. show that for NmNeuS, both ManNAc (Fig. 6a) and PEP (Fig. 6b) behave according to Michaelis–Menten kinetics. It appears that the product, Neu5Ac, does not inhibit the enzyme (Fig. 6c) in the investigated concentration range. Very similar results were obtained with PNH5 (Fig. 6d) except for the influence of PEP on the enzyme activity (Fig. 6e), where it was found that it mildly inhibits the enzyme as substrate. Kinetic parameters were estimated by using double substrate Michaelis–Menten kinetics from the data presented in Fig. 6. and are presented in Table 1.
Dependence of specific enzyme NmNeuS activity (a–c cCo2+ = 0.05 mM, γNeuS = 0.016 mg mL−1, 150 mM sodium phosphate buffer pH 7.5) and PNH5 activity (d–f cCo2+ = 0.2 mM, γPNH5 = 0.015 mg mL.−1, 150 mM sodium acetate buffer pH 5.5) on the concentration of (a) ManNAc (cPEP = 10 mM), b PEP (cManNAc = 20 mM), c Neu5Ac (cManNAc = 10 mM, cPEP = 10 mM), d ManNAc (cPEP = 10 mM), e PEP (cManNAc = 20 mM), and f Neu5Ac (cManNAc = 20 mM, cPEP = 20 mM)
Kinetic model of aldol-like addition of ManNAc and PEP
A kinetic model for the aldol-like addition of ManNAc and PEP catalysed by NeuS has not been reported yet to the best of our knowledge. We believe that such research is of great interest considering that modelling holds numerous benefits, such as the ability to simulate different reaction scenarios in silico. Simulations enable choosing the best reactor mode and in combination with selection of downstream processing methods make a great base for early stage process cost estimation. This enables comparison of different synthesis methods and process schemes at an early stage of process development. Additionally, it enables the comparison of the estimated product price in the investigated process with the prices on the market. Modelling also enables making assumptions and estimations about the effect of the process on the environment. Most of these benefits were demonstrated before for other reaction systems (Findrik Blažević et al. 2021; Sudar et al. 2021; Česnik Katulić et al. 2021). Based on the presented experimental data (Fig. 6.), a kinetic model was developed (Eqs. (2) and (3)) for the synthesis of Neu5Ac catalysed by NmNeuS and PNH5, respectively. In these equations, r1 and r2 present the reaction rate of Neu5Ac formation. Mass balance equations for ManNAc, PEP, and Neu5Ac in the batch reactor are presented by the Eqs. (5)–(7), respectively. They are vital for the validation of the mathematical model, which was done in the batch system in this case and will be discussed later.
Storage stability of NeuS and PNH5
We followed the activity and protein concentration of NmNeuS and PNH5 for 50 days to assess the stability of enzymes during storage. The results presented in Fig. 7. show that NmNeuS retains around 50% of its initial activity after 50 days (Fig. 7a) and forms a kind of equilibrium then, whereas PNH5 loses all its activity during the same period. As far as the protein concentration is concerned, it remains more or less constant for NmNeuS, whereas for PNH5, it follows the drop of enzyme activity (Fig. S6, Fig. 7b). It needs to be noted that the data for PNH5 were measured at pH 7.5 (Fig. S6), as well as 5.5 (Fig. 7b). While the enzyme shows poor stability at pH 7.5, it was shown that the stability is significantly improved at pH 5.5 (Fig. 7b), which is the optimum pH for enzyme activity.
Storage stability presented as the change of specific activity and protein concentration during 50 days for a NmNeuS (150 mM sodium phosphate buffer, pH 7.5, cManNAc = 10 mM, cPEP = 10 mM, cCo2+ = 0.05 mM, γNmNeuS = 0.048 mg mL−1) and b PNH5 (150 mM sodium acetate buffer pH 5.5, cManNAc = 20 mM, cPEP = 20 mM, cCo2+ = 0.05 mM, γPNH5 = 0.035 mg mL.−1, 37 °C)
To assess what happens with the protein molecules for 50 days, the DLS technique was performed. Namely, the changes in the enzyme performance during time can be attributed to the changes in its structure. As the enzyme was incubated in the same medium throughout the experiment, the surrounding conditions remained constant, meaning that any changes in the size distribution profile of the enzyme can be attributed solely to structural changes over time, such as enzyme degradation and consequent aggregation during storage. Since both enzymes displayed activity loss to some extent, DLS was performed to determine whether the stability decline could be attributed to the changes in protein structure and, consequently, aggregation. Thus, it makes sense to evaluate what is happening to protein size distributions, namely NSD, VSD, and ISD. The distribution of the particles with different hydrodynamic diameters is presented in Fig. 8. In the initial phase immediately after purification, the particle sizes of NmNeuS were in the range of 10 nm which corresponds to the size of the crystal dimer designated as 58.766 × 76.147 × 77.482 Å (PDB ID: 1XUU (Berman et al. 2003; Gunawan et al. 2005)). Such monomodal and narrow distribution around the size of the crystal protein indicates the existence of the active form of the enzyme in homogeneous mixture. Subsequently, following the initial stage of the purification process, a similar trend was noted with PNH5 particles, whose dimensions fell within the 10-nm range, aligning with the calculated dimensions, 123.8 × 49.1 × 48.7 Å, derived from an Inertia Axis Aligned Bounding Box (IABB, Draw Protein Dimensions Script (Schrödinger 2015)). Furthermore, it can be concluded from the presented data for this enzyme that the size of the protein molecules is approximately the same as that for NmNeuS (Figs. 8 a and b and 9 c and d, respectively). The similarity in dimensional characteristics between NmNeuS and PNH5 is evident in the superimposed 3D models illustrated in Fig. 2a. These results indicate a higher protein integrity of NmNeuS than PNH5 at these conditions. Even after 39 days, NmNeuS does not show visible aggregation when NSD and VSD are studied, while for PNH5, significant protein aggregation occurred after 39 days in both buffers, which is evident from the shift of the peak at the particle size of 100–1000 nm. These data at pH 7.5 correlate well with the protein concentration and specific activity data presented in Fig. S6. According to NSD, PNH5 at pH 5.5 appears to be in completely aggregated form after 49 days (Fig. 8b1), although activity measurements show a retention of about 50% of the initial value. This could, for example, be attributed to the change of surface charge of larger aggregates, which at pH 5.5 attract proteins in their active form through electrostatic interactions, potentially complicating their differentiation during DLS measurements. To inspect this further, the sample was centrifuged after 61 days, and the activity, protein concentration and PSD in the supernatant were measured. After separation by centrifugation, the concentration of soluble proteins in the supernatant was 0.297 mg/mL with specific activity 4.80 U/mg. PSD measurements also confirmed the existence of active proteins around 10 nm, which were not detectable by the same method before centrifugation (Fig. S7).
Size distributions of NmNeuS and PNH5 in buffer solution during 39–49 days of storage according to the number (a NmNeuS, b–b1 PNH5), volume (c NmNeuS, d–d1 PNH5, and intensity (e NmNeuS, f–f1 PNH5) (γEnzyme = 1 mg mL.−1; 150 mM sodium phosphate buffer pH 7.5 for (a), (b), (c), (d), (e), and (f); 50 mM sodium acetate pH 5.5 for (b1), (d1), and (f1); 4 °C)
Synthesis of Neu5Ac catalysed by NmNeuS (150 mM sodium phosphate buffer pH 7.5, 37 °C, γNeuS = 0.021 mg mL−1, cCo2+ = 0.05 mM) and PNH5 (150 mM sodium acetate buffer pH 5.5, 37 °C, γPNH5 = 0.053 mg mL−1, cCo2+ = 0.05 mM) in the batch reactor. a cManNAc = 83.79 mM, cPEP = 100.00 mM. a1 Operational stability of NmNeuS during the experiment presented in (a). b cManNAc = 47.90 mM, cPEP = 50.00 mM. b1 Operational stability of PNH5 during the experiment presented in (b). Legend: a–b black circles, ManNAc; white circles, Neu5Ac; black line, simulation. a1–b1 black circles, relative activity of NmNeuS (a1) and PNH5 (b1) during the experiments; black line, model simulation
Figure 8e–f1 present the enzyme particle ISD. Since this type of distribution weights particles according to the intensity of the signal, it is convenient for early identification of protein aggregation. These data show that even though the majority of particles correspond to the size of the protein crystal as elaborated in Fig. 8a–d1, both enzymes are prone to aggregation to some extent. For NmNeuS, it is less obvious from other data, but Fig. 8e shows that immediately after purification, there are particles in the range of 100 nm (black line). After 39 days, there are particles in the higher size ranges as well. This is more pronounced for PNH5 (Fig. 8 f and f1) with signals of relatively high intensity for particle sizes up to 1000 nm. This corresponds well to other provided experimental data, i.e. protein concentration and activity. Since after 20 days both enzymes at pH 7.5 retain approximately 50% of the initial activity (Fig. 7a and Fig. S6), and PNH5 around 65% at pH 5.5 (Fig. 8b), and the particle size distribution does not detect the formation of the same number of aggregates, it can be concluded that aggregation is not the only and predominant reason for loss of stability. However, the DLS method does not rule out the loss of activity due to other conformational changes such as dimer dissociation into monomers. The size difference between these units is not sufficient for the method resolution, as the factor of 3 is a prerequisite for the successful detection of definite change in size by this experimental method. In the case of PNH5 at pH 7.5, the drop in protein concentration after 50 days (Fig. S6) can be attributed to the formation of water-insoluble protein aggregates, as clearly confirmed by DLS.
Mathematical model simulations and the stability of enzyme during the reaction
The mathematical model of Neu5Ac synthesis was validated in a series of batch reactor experiments (Eqs. (2) and (4)–(7)). The synthesis of Neu5Ac catalysed by NeuS is a double substrate reaction that starts from ManNAc and PEP. The reaction therefore requires PEP in stoichiometric amounts as co-substrate and an addition of metal cofactor. Co2+ was found to be the best choice and was therefore used in further experiments. The first series of experiments was performed with NmNeuS and is presented in Fig. 9 a and a1 and Fig. S8. During the experiments, enzyme activity was evaluated, and the results are also presented in Fig. 9a1 and Fig. S8 a1 and b1. The operational stability is an enzyme property crucial for its application. It was evaluated by following enzyme activity in a series of batch reactor experiments.
Figure 9a and Fig. S8a–c show the change of concentrations of ManNAc and Neu5Ac during the experiment for the reaction catalysed by NmNeuS, whereas Fig. 9a1and Fig. S8a1–b1 show the change in the enzyme activity during the experiments. In all the cases, it can be observed that the mathematical model describes the data well and that the enzyme activity is practically constant implying that NmNeuS has good operational stability. Batch reactor experiments at different reaction conditions were conducted with PNH5 as well and are presented in Fig. 9b and Fig. S9. The mathematical model (Eqs. (3) and (4)–(7)) also described the data well. Thus, it can be verified for both enzymes that product inhibition is not present as assumed from the limited concentration range measurements. The experiment presented in Fig. S9b1 shows that the enzyme operational stability is good at the reaction conditions. Figure 9b1 shows that a slow enzyme activity drop can be observed in case of the experiment conducted at 50 mM of substrate. The operational stability decay rate constant of the first order was estimated to be kd = 9.6 ⋅ 10–5 ± 9.2 ⋅ 10–6 min−1. This value corresponds to 120.34 h enzyme half-life time. In the case of the experiment presented in Fig. S9c, the operational stability decay rate constant was estimated directly from the concentration vs. time data. It was found to be 0.0015 ± 0.0001 min−1, corresponding to the enzyme half-life time of 7.7 h. Thus, it appears that for this enzyme, an increase of substrate concentration also means a decrease in enzyme operational stability. Statistical output of the mathematical model, along with the explanation of its meaning is presented in Table S3.
Discussion
On the effect of pH on NeuS activity
A wide range of pH values have showed compatibility with NeuS activity from different organisms (Table S1), but the pH optimums reported were always in the same range between 7.0 and 8.3. For comparison, at the respective pH values, PNH5 displays only around 20% of its maximum activity observed at 5.5. At pH 5.0, PNH5 activity drops by 50%, whereas a complete activity loss is observed at pH 4.0. However, while pH 7.0 is an optimum for SaNeuS activity, AsNeuS was inactive at the same pH (Suryanti et al. 2003; Gurung et al. 2013). Furthermore, AsNeuB becomes active from its inactive form when pH is increased from 7.0 to 7.5, in accordance with NmNeuS activity (Blacklow and Warren 1962; Hao et al. 2005; Gurung et al. 2013). As the mechanistic studies have shown that the metal ions are vital for bacterial NeuS catalysis, a change in the protonation states of the metal ion coordinating the active site histidines (His215 and His236) could explain the increase in the activity (Gurung et al. 2013; Berg et al. 2015).
On the influence of cofactors on the activity of NmNeuS and PNH5 and enzyme kinetics
As stated before, in the relevant literature, several divalent metal cofactors were tested with NeuS homologues, i.e. magnesium, manganese, cobalt, calcium, and others (Blacklow and Warren 1962; Vann et al. 1997; Linton et al. 2000; Suryanti et al. 2003; Sundaram et al. 2004; Gunawan et al. 2005; Hao et al. 2005; Liu et al. 2009; Gurung et al. 2013; García García et al. 2015; Berg et al. 2015). For example, Hao et al. (2005) reported that the activity of NmNeuS was around 2.7-fold higher in the presence of cobalt than magnesium ions, while it was around 1.15-fold better in the presence of manganese than magnesium ions. Komaki et al. (1997) reported that NeuS from E. coli K1-M12 does not need a metal cofactor for enzyme activity. In most of these reports, some research on the cofactor effect was done. However, systematic studies on the effect of cofactor concentration on enzyme activity had not been reported so far. Such experiments are important to find the optimum conditions for enzyme activity in the reactor for producing sialic acid analogues. Additionally, none of the authors so far stated that they considered the metal cofactors already present in the enzyme samples after the purification. Therefore, we believe that Komaki et al. (1997) may have falsely reported that their enzyme needs no cofactor for activity. As a major difference to research done so far, all our measurements were performed with purified enzyme, from which the metal cofactors were previously removed by the EDTA treatment (Mónico et al. 2017). An ICP-MS analysis was performed to prove the absence of metals, which could affect enzyme activity (Table S2). We could detect only Mn2+ and Ni2+ in the purified enzyme samples, corresponding to the native cofactor and the affinity purification reagent. Enzyme preparations containing no metal after EDTA treatment did not exhibit activity in the synthesis of Neu5Ac. During enzyme production and purification, various metal ions present in solution may bind to the enzyme and affect the measured enzyme activity. If the bound metal had not been removed before, it would be difficult to judge the effect on enzyme activity of systematically adding different metal cofactors and their concentration, when the enzymes independently showed dubious activity even in the absence of added metal ions.
While some kinetic parameters can be found in the literature for NmNeuS, as well as for other enzymes (Table S1), the data is scarce and not measured in a systematic fashion. In addition, different methods and assays were applied. While Hao et al. (2005) reported kcat value of 49.5 min−1 for this enzyme, in our case, it is estimated to be significantly higher at 619.41 ± 11. 81 min−1. It needs to be noted here that we used broader concentration ranges of substrate and co-substrate in our measurements and non-linear regression analysis to estimate the values of the kinetic parameters. Additionally, the Michaelis constant for ManNAc was estimated to be almost tenfold lower in our case, and the value of kcat/Km was 7644.4 s−1 M−1. These differences come from the experimental and data handling methodology. Namely, Hao et al. (2005) applied a colorimetric TBA assay and used 1 mM Mg2+ as a cofactor, even though they also concluded that Co2+ yields a three-fold better enzyme activity. Additionally, they used different buffers and a rather narrow concentration range of ManNAc only up to 3 mM, which is significantly lower than the Michaelis constant they estimated for NmNeuS.
As far as PNH5 is concerned, the estimated Michaelis constants for PEP and ManNAc (Table 1) are quite low and comparable to NmNeuS meaning that ManNAc is an excellent substrate for this enzyme, and a good selection if, for some chemical reason, acidic pH is required. For example, this could be interesting for downstream processing purposes without the need for media exchange, possibly leading to a more profitable process, because Furuhata (2004) reported that sialic acids crystallize from water–acetic acid mixtures (Furuhata 2004).
The analysis of the molecular structure of PNH5 revealed that the molecular weight of the enzyme to be 38.4 kDa, kcat was calculated to be 219.01 ± 18.05 min−1 and kcat/Km was estimated to be 6755 s−1 M−1.
On the storage stability of NeuS and PNH5
Enzyme stability is an important feature of every industrial biocatalyst. It is therefore important to evaluate it and improve it if needed. As far as NeuS homologues are concerned, no systematic study of enzyme stability was reported so far to the best of our knowledge. Hwang et al. (2002) identified site-specific cleavage prone to proteolysis identified at Lys280 residue for EcNeuS and concluded that the purified enzyme, identified as tetramer (160 kDa), was not susceptible to cleavage. They described the degradation of monomer (size 40 kDa) to two different fractions, e.g. 33 and 7 kDa, and suspected that in that manner, the tetramer turns to trimer and loses the activity. Suryanti et al. (2003) reported that SaNeuS was prone to proteolysis during purification. Sundaram et al. (2004) reported that pure CjNeuS was not very stable, and for that reason, they conducted experiments with partially purified enzyme. Hao et al. (2005) reported that the recombinant NmNeuS is stable when stored at − 20 °C. The cold-adapted MvNeuS (Berg et al. 2015) was found to be stable for 3–4 weeks at 4 °C, but readily aggregated at room temperature. They performed a calorimetric study, which showed that the enzyme unfolds with a two-step transition, i.e. two peaks indicated that the dimer first separates into monomers (1st peak) and that monomers deactivate at higher temperatures (2nd peak).
From the presented literature overview, it is evident that the data is insufficient to understand what is going on with these enzymes in the solution, and that is why we combined enzyme activity and protein concentration measurements with DLS technique to elucidate the situation.
On the application of the mathematical model and the operational stability of enzymes
When discussing the optimal reaction conditions and design, one must have a deep insight into the kinetic behaviour of an enzyme. In case of NmNeuS, we can see that the reaction kinetics is very simple double substrate Michaelis–Menten kinetics. In these cases, the selection of a reactor mode is less difficult. If an enzyme is stable, such as in the NmNeuS case, any reactor will be applicable and good. However, if we aim towards high volumetric productivity, a continuous reactor mode would be the best option ensuring constant product quality (Fig. S10 for NmNeuS). In order to ensure good enzyme reusability, an ultrafiltration membrane reactor can be applied (Sudar et al. 2013; Valinger et al. 2014). However, the limitation of Neu5Ac solubility means that the highest substrate concentration should be in the range of 160 mM. Alternative continuous systems would require enzyme immobilization. In case of PNH5, PEP inhibition is mild, and does not really affect the outcome. However, since enzyme stability seems to be a potential issue, enzyme immobilization could be considered for further application. Instead, discontinuous reactor modes can also be applied. The results collected so far represent an excellent platform for the application of NeuS in the synthesis of unnatural sugars, while it is expected that the findings presented in this work can be generalized on other potential reactions catalysed by these enzymes. Our current research is directed toward that area.
Data availability
The data that supports the findings of this study are available in the supplementary material of this article.
References
Baardsnes J, Davies PL (2001) Sialic acid synthase: the origin of fish type III antifreeze protein? Trends Biochem Sci 26:468–469. https://doi.org/10.1016/S0968-0004(01)01879-5
Bateman A, Martin MJ, Orchard S, Magrane M, Agivetova R, Ahmad S, Alpi E, Bowler-Barnett EH, Britto R, Bursteinas B, Bye-A-Jee H, Coetzee R, Cukura A, da Silva A, Denny P, Dogan T, Ebenezer TG, Fan J, Castro LG, Garmiri P, Georghiou G, Gonzales L, Hatton-Ellis E, Hussein A, Ignatchenko A, Insana G, Ishtiaq R, Jokinen P, Joshi V, Jyothi D, Lock A, Lopez R, Luciani A, Luo J, Lussi Y, MacDougall A, Madeira F, Mahmoudy M, Menchi M, Mishra A, Moulang K, Nightingale A, Oliveira CS, Pundir S, Qi G, Raj S, Rice D, Lopez MR, Saidi R, Sampson J, Sawford T, Speretta E, Turner E, Tyagi N, Vasudev P, Volynkin V, Warner K, Watkins X, Zaru R, Zellner H, Bridge A, Poux S, Redaschi N, Aimo L, Argoud-Puy G, Auchincloss A, Axelsen K, Bansal P, Baratin D, Blatter MC, Bolleman J, Boutet E, Breuza L, Casals-Casas C, de Castro E, Echioukh KC, Coudert E, Cuche B, Doche M, Dornevil D, Estreicher A, Famiglietti ML, Feuermann M, Gasteiger E, Gehant S, Gerritsen V, Gos A, Gruaz-Gumowski N, Hinz U, Hulo C, Hyka-Nouspikel N, Jungo F, Keller G, Kerhornou A, Lara V, Le Mercier P, Lieberherr D, Lombardot T, Martin X, Masson P, Morgat A, Neto TB, Paesano S, Pedruzzi I, Pilbout S, Pourcel L, Pozzato M, Pruess M, Rivoire C, Sigrist C, Sonesson K, Stutz A, Sundaram S, Tognolli M, Verbregue L, Wu CH, Arighi CN, Arminski L, Chen C, Chen Y, Garavelli JS, Huang H, Laiho K, McGarvey P, Natale DA, Ross K, Vinayaka CR, Wang Q, Wang Y, Yeh LS, Zhang J, Ruch P, Teodoro D (2021) UniProt: the universal protein knowledgebase in 2021. Nucleic Acids Res 49:D480–D489. https://doi.org/10.1093/nar/gkaa1100
Berg TO, Gurung MK, Altermark B, Smalås AO, Ræder ILU (2015) Characterization of the N-acetylneuraminic acid synthase (NeuB) from the psychrophilic fish pathogen Moritella viscosa. Carbohyd Res 402:133–145
Berman H, Henrick K, Nakamura H (2003) Announcing the worldwide Protein Data Bank. Nat Struct Biol 10:980
Blacklow RS, Warren L (1962) Biosynthesis of sialic acids by Neisseria meningitidis. J Biol Chem 237:3520–3526. https://doi.org/10.1016/s0021-9258(19)70850-3
Bradford M (1976) A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal Biochem 72:248–254. https://doi.org/10.1006/abio.1976.9999
Česnik Katulić M, Sudar M, Hernández K, Qi Y, Charnock SJ, Vasić-Rački Đ, Clapés P, Findrik Blažević Z (2021) Cascade synthesis of l -homoserine catalyzed by lyophilized whole cells containing transaminase and aldolase activities: the mathematical modeling approach. Ind Eng Chem Res 60:13846–13858. https://doi.org/10.1021/acs.iecr.1c02343
Česnik M, Sudar M, Roldan R, Hernandez K, Parella T, Clapés P, Charnock S, Vasić-Rački Đ, Findrik Blažević Z (2019) Model-based optimization of the enzymatic aldol addition of propanal to formaldehyde: a first step towards enzymatic synthesis of 3-hydroxybutyric acid. Chem Eng Res Des 150:140–152. https://doi.org/10.1016/j.cherd.2019.06.025
Colley KJ, Kitajima K, Sato C (2014) Polysialic acid: biosynthesis, novel functions and applications. Crit Rev Biochem Mol Biol 49:498–532. https://doi.org/10.3109/10409238.2014.976606
Cunningham BA, Hoffman S, Rutishauser U, Hemperly JJ, Edelman GM (1983) Molecular topography of the neural cell adhesion molecule N-CAM: surface orientation and location of sialic acid-rich and binding regions. PNAS 80:3116–3120. https://doi.org/10.1073/pnas.80.10.3116
Eastman P, Swails J, Chodera JD, McGibbon RT, Zhao Y, Beauchamp KA, Wang L-P, Simmonett AC, Harrigan MP, Stern CD, Wiewiora RP, Brooks BR, Pande VS (2017) OpenMM 7: Rapid development of high performance algorithms for molecular dynamics. PLOS Comput Biol 13. https://doi.org/10.1371/journal.pcbi.1005659
Fessner WD, He N, Yi D, Unruh P, Knorst M (2014) Enzymatic Generation of Sialoconjugate Diversity. In: Riva S, Fessner W-D (eds) Cascade biocatalysis: integrating stereoselective and environmentally friendly reactions. Wiley-VCH Verlag GmbH & Co, KGaA, pp 361–392
Findrik Blažević Z, Milčić N, Sudar M, Majerić Elenkov M (2021) Halohydrin dehalogenases and their potential in industrial application – a viewpoint of enzyme reaction engineering. Adv Synth Catal 363:388–410. https://doi.org/10.1002/adsc.202000984
Fukuda M (1996) Possible roles of tumor-associated carbohydrate antigens. Cancer Res 56:2237–2244
Furuhata K (2004) Chemistry of N-acetylneuraminic acid (Neu5Ac). Trends Glycosci Glyc 16:143–169
García García MI, Lau K, Von Itzstein M, García Carmona F, Sánchez Ferrer Á (2015) Molecular characterization of a new N-acetylneuraminate synthase (NeuB1) from Idiomarina loihiensis. Glycobiol 25:115–123. https://doi.org/10.1093/glycob/cwu096
Garrabou X, Castillo JA, Guérard-Hélaine C, Parella T, Joglar J, Lemaire M, Clapés P (2009) Asymmetric self- and cross-aldol reactions of glycolaldehyde catalyzed by D-fructose-6-phosphate aldolase. Angew Chem Int Ed 48:5521–5525. https://doi.org/10.1002/anie.200902065
Gasteiger E, Hoogland C, Gattiker A, Duvaud S, Wilkins MR, Appel RD, Bairoch A (2005) Protein identification and analysis tools on the ExPASy server. In: Walker JM (ed) The Proteomics Protocols Handbook. Humana Press, Totowa, NJ, pp 571–607
Gorelik E, Galili U, Raz A (2001) On the role of cell surface carbohydrates and their binding proteins (lectins) in tumor metastasis. Cancer Metastasis Rev 20:245–277
Gunawan J, Simard D, Gilbert M, Lovering AL, Wakarchuk WW, Tanner ME, Strynadka NCJ (2005) Structural and mechanistic analysis of sialic acid synthase NeuB from Neisseria meningitidis in complex with Mn2+, phosphoenolpyruvate, and N-acetylmannosaminitol. J Biol Chem 280:3555–3563. https://doi.org/10.1074/jbc.M411942200
Gurung MK, Ræder IL, Altermark B, Smalås AO (2013) Characterization of the sialic acid synthase from Aliivibrio salmonicida suggests a novel pathway for bacterial synthesis of 7-O-acetylated sialic acids. Glycobiol 23:806–819. https://doi.org/10.1093/glycob/cwt018
Hao J, Balagurumoorthy P, Sarilla S, Sundaramoorthy M (2005) Cloning, expression, and characterization of sialic acid synthases. Biochem Biophys Res Comm 338:1507–1514. https://doi.org/10.1016/j.bbrc.2005.10.113
He N, Yi D, Fessner W (2011) Flexibility of substrate binding of cytosine-5′-monophosphate- n -acetylneuraminate synthetase (CMP-sialate synthetase) from Neisseria meningitidis: an enabling catalyst for the synthesis of neo-sialoconjugates. Adv Synth Catal 353:2384–2398. https://doi.org/10.1002/adsc.201100412
Heida R, Bhide YC, Gasbarri M, Kocabiyik Ö, Stellacci F, Huckriede ALW, Hinrichs WLJ, Frijlink HW (2021) Advances in the development of entry inhibitors for sialic-acid-targeting viruses. Drug Discov Today 26:122–137. https://doi.org/10.1016/j.drudis.2020.10.009
Hwang TS, Hung CH, Teo CF, Chen GT, Chang LS, Chen SF, Chen YJ, Lin CH (2002) Structural characterization of Escherichia coli sialic acid synthase. Biochem Biophys Res Commun 295:167–173. https://doi.org/10.1016/S0006-291X(02)00620-4
Joseph DDA, Jiao W, Kessans SA, Parker EJ (2014) Substrate-mediated control of the conformation of an ancillary domain delivers a competent catalytic site for N-acetylneuraminic acid synthase. Proteins: Structure. Function and Bioinformatics 82:2054–2066. https://doi.org/10.1002/prot.24558
Joseph DDA, Jiao W, Parker EJ (2013) Arg314 Is essential for catalysis by N-acetyl neuraminic acid synthase from Neisseria meningitidis. Biochem 52:2609–2619. https://doi.org/10.1021/bi400062c
Keil JM, Rafn GR, Turan IM, Aljohani MA, Sahebjam-Atabaki R, Sun X-L (2022) Sialidase inhibitors with different mechanisms. J Med Chem 65:13574–13593. https://doi.org/10.1021/acs.jmedchem.2c01258
Kiefel MJ, Von Itzstein M (2002) Recent advances in the synthesis of sialic acid derivatives and sialylmimetics as biological probes. Chem Rev 102:471–490. https://doi.org/10.1021/cr000414a
Knorst M (1999) Neue Enzyme zur Synthese von sialinsäurehaltigen Oligosacchariden. Aachen Techn Hochsch Diss
Koketsu M, Nitoda T, Sugino H, Juneja LR, Kim M, Yamamoto T, Abe N, Kajimoto T, Wong C-H (1997) Synthesis of a novel sialic acid derivative (sialylphospholipid) as an antirotaviral agent. J Med Chem 40:3332–3335. https://doi.org/10.1021/jm9701280
Komaki E, Otha Y, Tsukada Y (1997) Purification and characterization of n-acetylneuraminate synthase from Escherichia coli kl-m12. Biosci, Biotechnol Biochem 61:2046–2050. https://doi.org/10.1271/bbb.61.2046
Laemmli UK (1970) Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature 227:680–685. https://doi.org/10.1038/227680a0
Linton D, Karlyshev AV, Hitchen PG, Morris HR, Dell A, Gregson NA, Wren BW (2000) Multiple N-acetyl neuraminic acid synthetase (neuB) genes in Campylobacter jejuni: Identification and characterization of the gene involved in sialylation of lipo-oligosaccharide. Mol Microbiol 35:1120–1134. https://doi.org/10.1046/j.1365-2958.2000.01780.x
Liu F, Lee HJ, Strynadka NCJ, Tanner ME (2009) Inhibition of Neisseria meningitidis sialic acid synthase by a tetrahedral intermediate analogue. Biochem 48:9194–9201. https://doi.org/10.1021/bi9012758
Mahmoudian M, Noble D, Drake CS, Middleton RF, Montgomery DS, Piercey JE, Ramlakhan D, Todd M, Dawson MJ (1997) An efficient process for production of N-acetylneuraminic acid using N-acetylneuraminic acid aldolase. Enzyme Microb Technol 20:393–400. https://doi.org/10.1016/S0141-0229(96)00180-9
Marshall JR, Yao P, Montgomery SL, Finnigan JD, Thorpe TW, Palmer RB, Mangas-Sanchez J, Duncan RAM, Heath RS, Graham KM, Cook DJ, Charnock SJ, Turner NJ (2021) Screening and characterization of a diverse panel of metagenomic imine reductases for biocatalytic reductive amination. Nat Chem 13:140–148. https://doi.org/10.1038/s41557-020-00606-w
Mertsch A, He N, Yi D, Kickstein M, Fessner WD (2020) An α2,3-Sialyltransferase from Photobacterium phosphoreum with Broad Substrate Scope: Controlling Hydrolytic Activity by Directed Evolution. Chem Eur J 26:11614–11624
Mertsch A, Poschenrieder S, Fessner WD (2020) Semi-synthetic sialic acid probes for challenging the substrate promiscuity of enzymes in the sialoconjugation pathway. Adv Synth Catal 362:5485–5495. https://doi.org/10.1002/adsc.202000859
Milčić N, Stepanić V, Crnolatac I, Findrik Blažević Z, Brkljača Z, Majerić Elenkov M (2022) Inhibitory effect of DMSO on halohydrin dehalogenase: experimental and computational insights into the influence of an organic co-solvent on the structural and catalytic properties of a biocatalyst. Chem Eur J 28:e202201923. https://doi.org/10.1002/chem.202201923
Mirdita M, Schütze K, Moriwaki Y, Heo L, Ovchinnikov S, Steinegger M (2022) ColabFold: making protein folding accessible to all. Nat Methods 19:679–682. https://doi.org/10.1038/s41592-022-01488-1
Mirdita M, Steinegger M, Söding J (2019) MMseqs2 desktop and local web server app for fast, interactive sequence searches. Bioinformatics 35:2856–2858. https://doi.org/10.1093/bioinformatics/bty1057
Mirdita M, von den Driesch L, Galiez C, Martin MJ, Söding J, Steinegger M (2017) Uniclust databases of clustered and deeply annotated protein sequences and alignments. Nucleic Acids Res 45:D170–D176. https://doi.org/10.1093/nar/gkw1081
Mitchell AL, Almeida A, Beracochea M, Boland M, Burgin J, Cochrane G, Crusoe MR, Kale V, Potter SC, Richardson LJ, Sakharova E, Scheremetjew M, Korobeynikov A, Shlemov A, Kunyavskaya O, Lapidus A, Finn RD (2019) MGnify: the microbiome analysis resource in 2020. Nucleic Acids Res. https://doi.org/10.1093/nar/gkz1035
Mónico A, Martínez-Senra E, Cañada FJ, Zorrilla S, Pérez-Sala D (2017) Drawbacks of dialysis procedures for removal of EDTA. PLoS ONE 12:e0169843. https://doi.org/10.1371/journal.pone.0169843
Robert X, Gouet P (2014) Deciphering key features in protein structures with the new ENDscript server. Nucleic Acids Res 42:W320–W324. https://doi.org/10.1093/nar/gku316
Schnaar RL, Gerardy-Schahn R, Hildebrandt H (2014) Sialic acids in the brain: gangliosides and polysialic acid in nervous system development, stability, disease, and regeneration. Physiol Rev 94:461–518. https://doi.org/10.1152/physrev.00033.2013
Schnellbächer M (2017) Kinetische und präparative Studien mit neuen Sialinsäuresynthasen aus Neisseria meningitidis und Campylobacter jejuni. Ph.D. Thesis, Technische Universität Darmstadt
Schrödinger LLC (2015) The PyMOL Molecular Graphics System. Version 1:8
Sievers F, Wilm A, Dineen D, Gibson TJ, Karplus K, Li W, Lopez R, McWilliam H, Remmert M, Söding J, Thompson JD, Higgins DG (2011) Fast, scalable generation of high-quality protein multiple sequence alignments using Clustal Omega. Mol Syst Biol 7:539–539. https://doi.org/10.1038/msb.2011.75
Steinegger M, Meier M, Mirdita M, Vöhringer H, Haunsberger SJ, Söding J (2019) HH-suite3 for fast remote homology detection and deep protein annotation. BMC Bioinform 20:473. https://doi.org/10.1186/s12859-019-3019-7
Stetefeld J, McKenna SA, Patel TR (2016) Dynamic light scattering: a practical guide and applications in biomedical sciences. Biophys Rev 8:409–427. https://doi.org/10.1007/s12551-016-0218-6
Sudar M, Česnik M, Clapés P, Pohl M, Vasić-Rački Đ, Findrik Blažević Z (2021) A cascade reaction for the synthesis of D-fagomine precursor revisited: kinetic insight and understanding of the system. New Biotechnol 63:19–28. https://doi.org/10.1016/j.nbt.2021.02.004
Sudar M, Findrik Z, Vasić-Rački Đ, Clapés P, Lozano C (2013) Mathematical model for aldol addition catalyzed by two D-fructose-6-phosphate aldolases variants overexpressed in E. coli. J Biotechnol 167:191–200. https://doi.org/10.1016/j.jbiotec.2013.07.008
Sundaram AK, Pitts L, Muhammad K, Wu J, Betenbaugh M, Woodard RW, Vann WF (2004) Characterization of N-acetylneuraminic acid synthase isoenzyme 1 from Campylobacter jejuni. Biochem J 383:83–89. https://doi.org/10.1042/BJ20040218
Suryanti V, Nelson A, Berry A (2003) Cloning, over-expression, purification, and characterisation of N-acetylneuraminate synthase from Streptococcus agalactiae. Prot Expr Purif 27:346–356. https://doi.org/10.1016/S1046-5928(02)00633-2
Traving C, Schauer R (1998) Structure, function and metabolism of sialic acids. Cell Mol Life Sci 54:1330–1349. https://doi.org/10.1007/s000180050258
Valinger D, Vrsalović Presečki A, Kurtanjek Ž, Pohl M, Findrik Blažević Z, Vasić-Rački Đ (2014) Continuous enzymatic carboligation of benzaldehyde and acetaldehyde in an enzyme ultrafiltration membrane reactor and laminar flow microreactors. J Mol Catal B Enzym 102:132–137. https://doi.org/10.1016/j.molcatb.2014.02.003
van Kempen M, Kim SS, Tumescheit C, Mirdita M, Lee J, Gilchrist CLM, Söding J, Steinegger M (2023) Fast and accurate protein structure search with Foldseek. Nat Biotechnol. https://doi.org/10.1038/s41587-023-01773-0
Vann WF, Tavarez JJ, Crowley J, Vimr E, Silver RP (1997) Purification and characterization of the Escherichia coli K1 neuB gene product N-acetylneuraminic acid synthetase. Glycobiology 7:697–701. https://doi.org/10.1093/glycob/7.5.697
Varki A (1993) Biological roles of oligosaccharides: all of the theories are correct. Glycobiology 3:97–130. https://doi.org/10.1093/glycob/3.2.97
Varki A (1997) Sialic acids as ligands in recognition phenomena. FASEB J 11:248–255. https://doi.org/10.1096/fasebj.11.4.9068613
von ltzstein M, Thomson RJ (1997) Sialic acids and sialic acid-recognising proteins: drug discovery targets and potential glycopharmaceuticals. Curr Med Chem 4:185–210. https://doi.org/10.2174/0929867304666220313111728
Warwel M, Fessner W-D (2000) Indium-mediated chain-extension: an improved protocol for the concise, diastereospecific synthesis of KDN and other sialic acids. Synlett 2000:865–867. https://doi.org/10.1055/s-2000-6693
Yi D, He N, Kickstein M, Metzner J, Weiß M, Berry A, Fessner W (2013) Engineering of a cytidine 5′-monophosphate-sialic acid synthetase for improved tolerance to functional sialic acids. Adv Synth Catal 355:3597–3612. https://doi.org/10.1002/adsc.201300568
Zhang Y, Tao F, Du M, Ma C, Qiu J, Gu L, He X, Xu P (2010) An efficient method for N-acetyl-D-neuraminic acid production using coupled bacterial cells with a safe temperature-induced system. Appl Microbiol Biotechnol 86:481–489. https://doi.org/10.1007/s00253-009-2302-3
Funding
This project has received funding from the European Union’s Horizon 2020 research and innovation programme under the Marie Sklodowska-Curie grant agreement no. 956631 (CC-TOP).
Author information
Authors and Affiliations
Contributions
MMÇ, NM, and TA conducted experiments. MMÇ, NM, and ZFB designed the research. NM, ZFB, and SC supervised the research. ZFB, SC, and WDF acquired resources. ZFB, SC, and WDF acquired funding. All authors wrote, read, and approved the manuscript.
Corresponding author
Ethics declarations
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Çakar, M.M., Milčić, N., Andreadaki, T. et al. Kinetic characterization of two neuraminic acid synthases and evaluation of their application potential. Appl Microbiol Biotechnol 108, 446 (2024). https://doi.org/10.1007/s00253-024-13277-1
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00253-024-13277-1