Abstract
Artemisinin is a component part of current frontline medicines for the treatment of malaria. The aim of this study is to make analogues of artemisinin using microbial transformation and evaluate their in vitro antimalarial activity. A panel of microorganisms were screened for biotransformation of artemisinin (1). The biotransformation products were extracted, purified and isolated using silica gel column chromatography and semi-preparative HPLC. Spectroscopic methods including LC-HRMS, GC–MS, FT-IR, 1D and 2D NMR were used to elucidate the structure of the artemisinin metabolites.1H NMR spectroscopy was further used to study the time-course biotransformation. The antiplasmodial activity (IC50) of the biotransformation products of 1 against intraerythrocytic cultures of Plasmodium falciparum were determined using bioluminescence assays. A filamentous fungus Aspergillus niger CICC 2487 was found to possess the best efficiency to convert artemisinin (1) to a novel derivative, 4-methoxy-9,10-dimethyloctahydrofuro-(3,2-i)-isochromen-11(4H)-one (2) via ring rearrangement and further degradation, along with three known derivatives, compound (3), deoxyartemisinin (4) and 3-hydroxy-deoxyartemisinin (5). Kinetic study of the biotransformation of artemisinin indicated the formation of artemisinin G as a key intermediate which could be hydrolyzed and methylated to form the new compound 2. Our study shows that the anti-plasmodial potency of compounds 2, 3, 4 and 5 were ablated compared to 1, which attributed to the loss of the unique peroxide bridge in artemisinin (1). This is the first report of microbial degradation and ring rearrangement of artemisinin with subsequent hydrolysis and methoxylation by A.niger.
Key points
• Aspergillus niger CICC 2487 was found to be efficient for biotransformation of artemisinin
• A novel and unusual artemisinin derivative was isolated and elucidated
• The peroxide bridge in artemisinin is crucial for its high antimalarial potency
• The pathway of biotransformation involves the formation of artemisinin G as a key intermediate
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
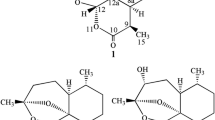
Artemisinin (1), qinghaosu, is a sesquiterpene lactone with an unique endoperoxide moiety first isolated from a Chinese medicinal plant in 1970s and showed extreme potency to malaria (Tu 2016). Artemisinin combination therapies are front-line medicines for the treatment of malaria. Artemisinin has also been intensively investigated for the treatment of cancer (Efferth 2017). Recent reports show the development and global spread of artemisinin resistance (Ashley et al. 2014; Balikagala et al. 2021; Miotto et al. 2020). This threat highlights the need to continue to search for novel antimalarial therapies, for example, novel analogues of artemisinin.
Many derivatives of artemisinin have been made chemically or biologically and tested for antimalarial (Kumari et al. 2019; Patel et al. 2021) and anticancer activity (Crespo-Ortiz and Wei 2012; Zhang et al. 2016). In particular, various artemisinin analogues including dihydroartemisinin, arteether, artemether, artesunate, artelinic acid, sodium artelinate and artesunic acid have been developed through chemical modification and approved for use (Kumari et al. 2019). Microbial transformation is the structural modification of complex substrates by microbial cells, that is, the catalytic reaction of specific groups of substrates by one or a series of enzymes produced in the process of microbial metabolism. It has become an effective tool widely used, especially for structurally complex natural products (Abourashed et al. 1999; de Carvalho and da Fonseca 2006). Microbial transformations can be considered as ‘‘green chemistry’’ for their ecological technology using mild conditions with less pollution. Additional benefits of biotransformation include its low cost, efficiency, strong chiral and stereoselectivity (de Carvalho and da Fonseca 2006), which are superior to conventional chemical synthesis methods. Microbial transformation is also an efficient way to mimic the mammalian metabolism (Abourashed et al. 1999). Previously, artemisinin (1) was transformed to various metabolites via deoxygenation (reduction) (Lee et al. 1989; Yu et al. 2017; Zhan et al. 2002b), hydroxylation (Gaur et al. 2014; Parshikov et al. 2006, 2004; Zhan et al. 2002a), oxidation (Liu et al. 2006), C-acetoxylation (Goswami et al. 2010), deoxygenation and hydroxylation (Lee et al. 1989; Ponnapalli et al. 2018), dihydroxylation (Zhan et al. 2017) and epoxylation (Zhan et al. 2015).
In this study, we report the biotransformation of artemisinin using a panel of microorganisms, further extraction, isolation, structural elucidation, kinetic monitoring and evaluation of in vitro anti-plasmodial activity of the transformation products.
Materials and methods
Chemicals
Artemisinin (98%) was purchased from Macklin Co., Ltd (China) which was used for biotransformation and bioactivity studies. HPLC grade formic acid, acetonitrile, hexane and ethyl acetate were purchased from Merck (Darmstadt, Germany). The other chemicals were purchased from Sinopharm Chemical Reagent Co., Ltd. The thin layer chromatography (TLC) plate was obtained from Qingdao Haiyang Chemical Group Co., People’s Republic of China.
Microbial strains
Aspergillus niger CICC 2487, Bacillus subtilis CICC 10157, Streptococcus thermophiles CICC 6038 and Phanerochaete chrysosporium CICC 40719 were purchased from the China General Microbiological Culture Collection Center. Trichoderma reesei ATCC 56765 and Lactococcus lactis ATCC 11454 were purchased from the American Type Culture Collection. Saccharomyces cerevisiae is from Mauripan Co.
Cultivation of microbial strains
The fungal strain A. niger was maintained on potato-dextrose agar plates at 4 ℃ and freshly sub-cultured before the transformation experiment. Fungal mycelia from agar slope cultures were transferred into 2-L Erlenmeyer flasks containing 1 L of potato dextrose liquid (PDL) medium, and incubated for 3 days at 180 rpm and 28 ℃ in a rotary shaker.
The screening procedure for small scale of biotransformation of artemisinin with microbial strains
Fungal mycelia were moved into 100-mL shaker flasks containing 40 mL of different types of culture media depending on the microbial strains (Supplementary Materials). After 2 days of incubation at 28 ℃ and 180 rpm on a rotary shaker, 10 mL of cultures containing mycelia were transferred to inoculate 100 mL of medium in 250-mL Erlenmeyer flasks. The inoculated flasks were incubated for 2 days on rotary shakers at the same speed and temperature as before. One flask was kept as control without the addition of artemisinin. 2 mL of artemisinin in acetone at a concentration of 100 mg/mL was added into each flask. A total of 200 mg of substrate was used in the biotransformation, and the final concentration of artemisinin in the flask was 2 mg/mL. The cultures were incubated under the same conditions for an additional 14 days. The mycelia were separated by filtration or centrifugation and discarded. The filtrate or supernatant was extracted three times with ethyl acetate. The combined extract was evaporated under vacuum and under 40 °C using a rotary evaporator.
Large scale of biotransformation of artemisinin with A.niger
A. niger mycelia were transferred into 100-mL Erlenmeyer flasks containing 40 mL of PDL medium for 3 days of incubation. The fermentation broth from the above Erlenmeyer flask was transferred into 2-L Erlenmeyer flasks containing 1 L of PDL medium with the same culture conditions. 20 mL of artemisinin (100 mg/mL in acetone) was added into each of the above flask with 1L-fermentation broth to achieve a final concentration of 2 mg/mL. The control was kept without addition of artemisinin. Substrate controls consisted of sterile medium containing the same amount of artemisinin and incubated under the same conditions. A total of 4 g of artemisinin was transformed with 2 L of fermentation broth. The cultures were maintained under the same conditions for 14 days. Later on, all fermentation broth was combined and the sediment was separated by centrifugation at 8500 g for 20 min. The supernatant (about 2 L) was extracted three times with ethyl acetate. The extract was evaporated under reduced pressure to afford a residue for further analysis and purification.
LC-HRMS analysis of biotransformed products of artemisinin by A.niger
Liquid chromatography and high-resolution mass spectrometry (LC-HRMS) were applied for detecting the biotransformation products of artemisinin by the culture supernatant of A. niger. The re-dissolved transformation product was analyzed using LC-HRMS (Agilent 1290—Bruker micrOTOF QII, Bruker). The resolving power was 17,500 for full-scan and mass accuracy for 5 ppm. Chromatographic separation was achieved on an Agilent zorbax SB18 column (50 mm × 2.1 mm i.d., 1.8 µm). The LC–MS experiments were performed using a mobile phase A of 0.1% formic acid and a mobile phase B of acetonitrile containing 0.1% formic acid at a flow rate of 0.7 ml/min. The high-performance liquid chromatography (HPLC) gradient system for metabolites identification started with 30% A and linearly increased to 80% A in 5 min.
Purification and isolation of biotransformation products by A.niger
The ethyl acetate extract (2.0 g) consisting of artemisinin transformation products was added to 4 mL of methanol, the precipitated solid was removed and identified as the untransformed artemisinin. The solution phase was evaporated and submitted for silica gel chromatography (50 g silica gel, 70 A size) eluting with hexane with increasing ethyl acetate (from 5–100%). 15 fractions were collected and the solvents were evaporated. The sub-fractions were further purified by semi-preparative high-performance liquid chromatography (HPLC) using Agilent 1220 LC (USA) system. The mobile phase A consisted of water and solvent B was methanol. The mobile phase calibration rose from 20% by 80% (A + B) over a period of 25 min to 100% B and kept at 100% for 10 min at a flow rate of 4 mL/min at 215 nm on semi-preparative HPLC column (Phenomenex, UK; 5 µm particle size: 9.4 × 250 mm). Four compounds: compound 2 (3 mg, 98% purity), compound 3 (25 mg, 95% purity), a mixture of compound 1 and 4 (1:1, 15 mg), and compound 5 (55 mg, 98% purity) were isolated from the extract of A. niger.
Time-course biotransformation of artemisinin (1) monitored by 1H NMR spectroscopy
Each 100 mg of artemisinin dissolved in 2 mL acetone was transferred into ten Erlenmeyer flask containing 200 mL of medium with inoculation of A.niger. 100 mg of artemisinin in 2 mL acetone was also transferred to a flask containing only medium and incubated for 336 h. Cell culture of A. niger without addition of 1 was used as control. The transformation flasks were incubated and the extraction of transformation products at 48, 120, 192, 264 and 336 h in two replicates was performed as described above. The transformation products were identified by 1H NMR spectroscopy (400 MHz Bruker AVANCE NEO instrument) in CDCl3 in comparison with the 1H NMR data of 1–5 obtained before and that of 7 as reported (Zhan et al. 2015). The change of the percentages of the substrate 1 and the biotransformation products were plotted according to the integration values of the distinct peaks for each compound.
NMR spectroscopy
1D and 2D NMR spectra of compounds 2, 3, 4 and 5 were obtained with a Bruker Ascend 400 NMR spectrometer (MA, USA). Bruker TopSpin 4.1.3 software or ACD/Labs 10 Freeware (Advanced Chemistry Development Inc., Ontario, Canada) was used to analyze the NMR spectra.
Gas chromatography mass spectrometry
1–2 μL of the isolated compounds dissolved in ethyl acetate was injected into gas chromatography mass spectrometry (GC–MS) system consisting of an Agilent 7890 coupled to Agilent MS model 5975C MSD (Agilent Technologies, US). The GC started from150 °C increasing to 300 °C at the rate of 10 °C/min, which was then kept at 300 °C for 4 min with a constant helium pressure (10 psi). The mass spectra data were acquired in the scan mode in the m/z range 40–1000.
High resolution electrospray mass spectrometry
The purified compounds 2, 3, 5, and a mixture of 1 and 4 were dissolved in methanol and injected to a LTQ Orbitrap mass spectrometer system (Thermo Scientific, MA, USA) to determine their high-resolution molecular masses and their fragment ions (MS/MS) using a positive or negative electrospray ion source.
Fourier transform infrared spectroscopy (FT-IR)
FT-IR instrument (Nicolet AVATAR360, Thermal Fisher, USA) was used to determine the functional groups of isolated compounds. FT-IR spectra were recorded from 4000 to 800 cm−1 using an attenuated total reflection (ATR) mode. The powder of isolated artemisinin derivatives were pressed against the ATR wafer.
Compound 2: white powder. GC-EI-MS, m/z (%): 223.1 (1), 195.0 (1), 166.1 (80), 151.1 (92), 137.0 (100), 98.0 (40), 69 (35), 41.1 (50) (Figure S1). HRMS (positive ESI) m/z calcd for 2 ( [M + Na]+ 277.1416, found 277.1416 (Figure S2). HR MS/MS, 245.1152 [M -CH3OH]+. ART-FT-IR (νmax, cm−1): 3081, 2940, 2860, 1786, 1732, 1458, 1382, 1129, 992, 987. 1H, 13C NMR, and 2D NMR data (see Table 1 and Figure S4-9).
Compound 3: white powder. GC-EI-MS, m/z (%): 211.1 (50), 165.1 (100), 137.1 (55), 109.1 (10), 89.0 (20), 41.1 (30) (Figure S10). HRMS (negative ESI) m/z calcd for [M-H]+ 239.1283, found 239.1182. 1H and 13C NMR data (see Table 1).
3-Hydroxy-deoxyartemisinin (5): white powder. HRMS (positive ESI) m/z calcd for [M + Na]+, 305.1365, found 305.1366. GC-EI-MS, m/z (%): 222.1 (70), 179.1 (30), 150.1 (55), 93.1 (40), 43.1 (100) (Figure S11). ART-FT-IR (νmax, cm−1): 3243, 2988, 2860, 1734, 1459, 1391, 1198, 1137, 985. 1H and 13C NMR data are consisted with those reported (Lee et al. 1989).
Determination of antiplasmodial activity
Parasite strain Dd2luc was maintained at 2% haematocrit, 0.5–5% parasitaemia and incubated at 37 °C in a gas chamber with 1% oxygen, 3% carbon dioxide and 96% nitrogen as described (Wong et al. 2011). Growth inhibition assays were carried out using a two-fold dilution of test compounds on 96-well plates, starting from 4 µM. A supralethal concentration (10 µM) of chloroquine was added to the negative (0% growth) control wells. The positive control (100% growth) wells were left untreated. P. falciparum culture (1–2% parasitaemia, 2% hematocrit) was added to all wells then incubated for 48 h at 37 °C in a gas chamber with 1% oxygen, 3% carbon dioxide and 96% nitrogen. A single-step lysis and bioluminescence assay to determine growth relative to control was carried out as previously described (Hasenkamp et al. 2012). IC50 values were estimated from nonlinear regression (sigmoidal concentration–response/variable slope equation) of log10-transformed concentration versus normalized growth (%) using GraphPad Prism v5.0 (GraphPad Software, Inc., San Diego, CA, USA).
Results
Structural elucidation of biotransformation products of artemisinin (1)
A panel of seven microorganisms was screened for their capability to biotransform artemisinin by thin layer chromatography (TLC) and/or analytical high-performance liquid chromatography (HPLC) of the ethyl acetate extract of microbial cells after incubation with artemisinin (1). Aspergillus niger was found to be most efficient to produce different products and selected for preparative scale biotransformation with artemisinin, while others did not yield detectable products. TLC analysis (Figure S1) and HPLC demonstrated the formation of new products compared to the artemisinin and the control supernatant of A. niger. Further liquid chromatography (LC) high-resolution (HR) mass spectrometry (MS) analysis of the extract of A. niger after incubation with artemisinin (1) confirmed the formation of a number of transformation products (2–5) (Fig. 1).
LC-HR-MS analysis of biotransformation products of artemisinin (1) by A. niger. A) Total ion chromatogram of high-resolution positive electrospray ionization mass spectrometry of the biotransformation products. B) Full scan mass spectrum of peaks from 0.5 to 4 min. HR-( +) ESI MS of peaks of 2, 3, 4 and 1/5/7 showed the masses of 241.144 [M + H]+, 277.215 [M + Na]+, 267.172 [M + H]+ (289.154 [M + Na]+) and 283.155 [M + H]+ (305.138 [M + Na]+), respectively. A peak labelled by * showing a major mass of 223.136 is an unknown compound
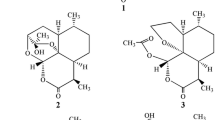
A larger scale of transformation of artemisinin at gram scale with A. niger was performed to obtain more products. Using a combination of silica gel chromatography and semi-preparative HPLC, compounds 2, 3 and 5 were isolated in high purity, while compound 4 was isolated as a mixture with recovered artemisinin (1:1). These compounds were identified using HR-ESI–MS, GC–MS, FT-IR, 1D and 2D NMR. Among them, compound 2 is a novel metabolite of artemisinin (Fig. 2).
Compound 2 was isolated as white powder with high purity (98%) as evidenced by GC–MS (Figure S2). High-resolution ESI–MS (Figure S3) determines the formula of 2 as C14H22O4 (found 277.1416, calculated for C14H22O4Na [M + Na]+, 277.1416 as sodium adduct) indicating the presence of four unsaturation units. This is consistent with the mass for compound 2 found in LC-HRMS (Fig. 1). FT-IR spectrum of 2 indicates the presence of an ester bond (1732 cm−1). 1H NMR (Figure S4), 13C NMR, DEPT (Figure S5) and HSQC (Figure S6) analysis indicates the presence of eight of a sum of CH and CH3, four methylene (CH2) and two quaternary carbons. There is an ester carbon (δc, 172.0 ppm), three oxygenated carbons (δc, 69.4, 79.6, and 105.1 ppm) and a methoxy group (δc, 55.1 ppm). 1H NMR, HSQC and 1H-1H COSY (Figure S7) support the presence of the two methyl groups (δH, 1.13 (3H, d, 8.0 Hz); 0.94 (3H, d, 8.0 Hz)) and a OCH3 group (δH, 3.52 (3H, s)). HR ESI MS/MS showed a strong fragmentation ion at m/z 245.1152 [MCH3OH]+ after collision of ion at m/z 277.1512 [M + Na]+, which indicates a loss of CH3OH group (a mass difference of 32.0360).
Comparing the 1H and 13C NMR of 2 with those of compound 6, 4-hydroxy-9,10-dimethyloctahydrofuro-(3,2-i)-isochromen-11(4H)-one (Table S1), which was reported as a biotransformation product of artemisinin in hairy root cell culture (Pandey et al. 2015), compound 2 has an extra OCH3 group, which suggests that 2 is a methyl ether of compound 6 at C4-OH position. The change of chemical shifts of neighboring carbons such as C-4, 5, 6, 10 and 11 also supports such structure. HSQC and 1H-1H COSY allow to complete the assignment of 1H and 13C NMR peaks (Table 1). Furthermore, the observation of the crossing spots between the peaks at 5.17 (H-4, s) and 3.52 (OCH3, s) on the NOESY spectrum (Figure S8) and the spot between the peak at δH 5.17 (H-4, s) and at 105.1 (C-4) on the HMBC spectrum (Figure S9) both support that the OCH3 group is attached to C-4 position. The 1H and 13C NMR data of compound 2 are also similar to those of artemisinin G (7) (Zhan et al. 2015), an acetylated compound 6 at position C-4 (Table S1). Therefore, compound 2 is identified as 4-methoxy-9,10-dimethyloctahydrofuro-(3,2-i)-isochromen-11(4H)-one, a new compound (Fig. 2).
Compound 3 was isolated as white powder. Both positive and negative ESI-HRMS confirmed its molecular formula as C13H20O4, suggesting to be a degradation product of artemisinin by loss of two carbons and an oxygen atom. 1H and 13C NMR analysis indicates the presence of a carboxylic acid group, an aldehyde group and a furan ring. GC-EI-MS analysis of 3 (Figure S10) indicates the presence of a strong ion m/z at 211.1 due to loss of a mass of 29 (CHO) from its molecular ion m/z at 240.1. After comparison of 1H and 13C NMR data of 3 with those of compound 2 (Table 1), most of peaks associated with 6-membered and 5-membered tetrahydrofuran rings are similar except those for the lactone moiety in 2; therefore, compound 3 is suggested to be the hydrolyzed form of 4-hydroxy-9,10-dimethyloctahydrofuro-(3,2-i)-isochromen-11(4H)-one (compound 6) (Fig. 2).
Compound 4 was not separated from compound 1 by HPLC, but is identified as deoxyartemisinin by GC–MS and NMR data. Compound 5 was identified as 3-hydroxy-deoxyartemisinin by GC–MS (Figure S11), HR-ESI–MS, 1D and 2D NMR and comparison with literature data (Lee et al. 1989).
Time-course of the biotransformation of artemisinin (1)
To better understand the kinetics and pathways of the biotransformation of artemisinin (1) by A niger, the transformation products were monitored using 1H NMR spectroscopy (Fig. 3). In the control experiment, artemisinin could be slowly and chemically transformed to three compounds 4, 5 and 7 (yields less than 10%) in the medium at 30 ºC after 14 days without addition of A. niger (Fig. 3). When incubating 1 with A niger for 2 days, significant increase of 7, 5 and 4 (in this order) were observed as well as the formation of compounds 2 and 3. This indicated that certain enzymes from the fungi could catalyze these transformation processes. The levels of compounds 2, 3, 4 and 5 hardly changed from day 2 to day 14. Compound 7 decrease from day 2 to day 8, and artemisinin (1) was almost fully consumed on day 8. Interestingly and surprisingly, on day 11 and 14, artemisinin was found to be reformed and increased, while compound 7 was significantly decreased (Fig. 3B, Figure S12).
Kinetics of biotransformation of artemisinin (1) over 14 days (336 h) by 1H NMR spectroscopy. A) pure artemisinin. B) artemisinin 1 in the culture medium without A. niger over 336 h; most of the artemisinin remained except for the appearance of compounds 4 (3%), 5 (6%) and 7 (10%). C) metabolites of A. niger without the addition of 1; no significant peaks between 5-7 ppm are present. D) incubation of 1 with A. niger for 48 h; E) incubation of 1 with A. niger for 120 h; F) incubation of 1 with A. niger for 192 h; G) incubation of 1 with A. niger for 264 h; H) incubation of 1 with A. niger for 336 h; I) The change of percentage of starting compound 1 and its biotransformation products (2–5, and 7) over 336 h. The percentage of each compound at each time interval is calculated based on the integration value of the distinct singlet peaks at 5.88, 5.18, 9.95, 5.70, 5.64 and 6.66 ppm for the single proton in the respective compounds 1, 2, 3, 4, 5 and 7 (% of each compound represents as mean ± standard deviation (SD), n = 2)
Determining antiplasmodial activity of biotransformed artemisinin products
The in vitro antiplasmodial activity of biotransformed artemisinin products was tested against intraerythrocytic stages of the Plasmodium falciparum chloroquine-resistant strain Dd2luc (Ullah et al. 2020; Wong et al. 2011). Growth inhibition assays were carried out after 48-h incubation with each compound, with IC50 values estimated from data developed over three independent assays (Fig. 4). Compounds 2, 3 and 7 were unfortunately not active (Table 2, Figure S13).
IC50 determination of artemisinin and transformed fractions. Log-transformed concentration versus a normalized % parasite growth (compared to untreated control) of P. falciparum Dd2luc parasites after 48-h incubation with the indicated fraction. Data are presented as mean ± standard deviation with n = ≤ 4 with a minimum of 2 biological repeats
Discussion
Over the last three decades, extensive work has been carried out to make analogues of artemisinin using biotransformation with a range of microorganisms and plant cell cultures. The biotransformation of artemisinin (1) by Nocardia corallina (ATCC 19,070) and Penicillium chrysogenum (ATCC 9480) to deoxyartemisinin and 3- α-hydroxydeoxyartemisinin, respectively, was first reported in 1989 (Lee et al. 1989). Artemisinin biotransformation by Cunninghamella echinulata afforded 10-β-hydroxyartemisinin (Zhan et al. 2002a), and by C. elegans yielded 7-β-hydroxyartemisinin as the major product along with other minor bioconversion products: 7-β-hydroxy-9-α-hydroxy-artemisinin, 4-α-hydroxy-1-deoxoartemisinin, 6-β-hydroxyartemisinin (Parshikov et al. 2004), 7-α-hydroxyartemisinin and 6-β-7-α-dihydroxyartemisinin (Zhan et al. 2017). Microbial metabolism of artemisinin by Mucor polymorphosporus resulted in the production of 9-β-hydroxyartemisinin, 3-β-hydroxyartemisinin and deoxyartemisinin (Zhan et al. 2002b). Artemisinin (1) was converted to 5-β-hydroxyartemisinin and 7-β-hydroxyartemisinin by Eurotium amstelodami (Parshikov et al. 2006). Biotransformation of artemisinin by fungi Rhizopus stolonifer afforded a new compound, 1-α-hydroxyartemisinin and 10-β-hydroxyartemisinin and deoxyartemisinin (Gaur et al. 2014). Artemisitone-9, a ketone derivative of artemisinin, was produced by cultured Streptomyces griseus ATCC 13,273 with artemisinin (Liu et al. 2006). Artemisinin was biotransformed to C-9 acetoxy artemisinin using Penicillium simplissimum along with C-9 hydroxy derivative (Goswami et al. 2010).
The fungi like Aspergillus showed to be an efficient machinery to metabolize artemisinin. Artemisinin was converted to deoxyartemisinin, 3-α-hydroxydeoxyartemisinin and 1-α-hydroxydeoxyartemisinin by Aspergillus niger AS 3.1858 (Zhan et al. 2002b). Further biotransformation of artemisinin (1) by A. niger resulted in 5-β-hydroxyartemisinin and 7-β-hydroxyartemisinin (Parshikov et al. 2006) similar as by E. amstelodami, and two novel compounds, 3-β-hydroxy-4,12-epoxy-1-deoxyartemisinin and 3,13-epoxyartemisinin together with known artemisinin G (7) and 3-α-hydroxydeoxyartemisinin by A. niger VKM F-1119 (Zhan et al. 2015). A. terreus transformed artemisinin to deoxyartemisinin and 4-α-hydroxy-1-deoxyartemisinin (= 3-α-hydroxydeoxyartemisinin) (Yu et al. 2017). The filamentous fungus A. flavus MTCC-9167 converted artemisinin to a new hydroxy derivative, 14-hydroxydeoxyartemisinin along with known deoxyartemisinin, artemisinin G (7) and 3-α-hydroxydeoxyartemisinin (Ponnapalli et al. 2018).
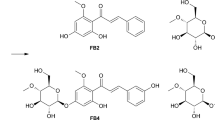
In this study, a novel metabolite (2) of artemisinin was identified, which is structurally similar to artemisinin G (7), previously found from a biotransformation product of artemisinin by A. niger VKM F-1119 (Zhan et al. 2015). The strain of A. niger CICC 2487 used in our study may produce different enzymes compared to other A.niger strains, thus leading to the formation of the novel product 2. The possible biotransformation pathway of artemisinin by A. niger CICC 2487 is proposed in Fig. 5. Artemisinin was found to be unstable in the culture PBL media and could be transformed to compound 7, 5 and 4 (Fig. 5A). The change of artemisinin (1) to compound 4 in the culture medium was reported before (Zhan et al. 2002b). The thermal degradation of artemisinin to compounds 5 and 7 at a high temperature of 190 ºC for 10 min was also reported (Lin et al. 1985). The reducing agents such as Fe (II) could also lead to the formation of compounds 4, 5 and 7 (Kapetanaki and Varotsis 2001; Wu et al. 1998). Our results further indicated the sensitivity and instability of artemisinin in agreement with these previous reports. However, this process was hugely accelerated by A. niger with further generation of compounds 2 and 3 (Fig. 5B). Artemisinin G (7) was initially formed via ring rearrangement of artemisinin (1) as also found previously by A. niger (Zhan et al. 2015), whose acetyl group could be subsequently hydrolyzed by an esterase to form compound 6 with a free hydroxy group, which was then methoxylated by a methyltransferase to form compound 2. However, compound 6 was not detected by 1H NMR spectroscopy, probably due to a very fast conversion step, which suggested the presence of an active esterase and methyltransferase allowing the full conversion of the intermediate 6 to the final and stable product 2. Our kinetic study by 1H NMR spectroscopy (Fig. 3B) supported this proposed pathway. An interesting and surprising observation is that artemisinin was reformed after 11 days with a significant loss of artemisinin G (7), suggesting that on day 8 the fungus A niger might produce an unique enzyme allowing the convertion of the degradation product (7) back to the original artemisinin via unususal ring closure and rearrangement reactions. This is somehow consistent with our large scale of biotransformation of artemisinin, where a portion of artemisinin was recovered over the purification process while compound 7 was not isolated at all. The possible enzymes involved and associated mechanisms are under further investigation.
Proposed transformation pathways for the generation of compounds (2–5) from artemisinin (1) by the cell culture medium (A) and A. niger CICC 2487 (B), respectively. Compounds 4, 5 and 7 were slowly formed due to inherent instability of artemisinin in the aqueous medium. The formation of these compounds were accelerated by A. niger, while also generating a new compound 2 via a possibly short-lived intermediate 6 from the initial product 7 together with the formation of compound 3
Compound 3 is also a degradation product of artemisinin with the loss of two carbons, which has not been found through biotransformation by either A.niger or other microbes but through thermal degradation at a high temperature of 180 °C (Xuan-De Luo et al. 1985). This is the first report of the production of 3 via mild biotransformation by a fungus. Compounds 4 and 5 were found in this study, in agreement with previous findings in the biotransformation of artemisinin by A. niger (Yu et al. 2017; Zhan et al. 2002b, 2015). Compounds 4 and 5 were suggested to be formed simultaneously and independently as happened at a high temperature (Lin et al. 1985) by reductive cleavage of the peroxide bridge of artemisinin (Fig. 3 and 5). This process of forming 4 and 5 seems to be different from the plausible biotransformation pathway of artemisinin by Aspergillus flavus (Ponnapalli et al. 2018) where compound 5 was suggested to be formed from compound 4 by hydroxylation reaction.
The IC50 of the artemisinin (1) was 28.1 ng/mL with the antiplasmodial activity almost completely ablated following biotransformation for compounds 2, 3 and 5. Whilst one fraction has antiplasmodial activity at a reduced potency (IC50 value of 46.2 ng/mL) compared to the artemsininin, this fraction contains both compound 1 (artemisinin) and compound 4 as a 1:1 mix. This suggests that the antiplasmodial activity of this fraction likely results from the artemsisinin content, and that compound 4 has no or almost none antiplasmodial activity. The antiplasmodial activity data is in agreement with repored data for compound 4 and 5 (Ponnapalli et al. 2018) and are not unexpected as the antiplasmodial activity of artemisinin and its active derivatives is attributable to the peroxide-containing moiety. In the presence of ferous ions (FeII), this moiety is cleaved to produce carbon and oxygen centred radicles (O'Neill et al. 2010). Following the biotransformation of artemisinin (1), this peroxide-containing moiety was removed and thus antiplasmodial activity lost. Nevertheless, the biotransformaton produced new molecular entities which are not easily accessible by chemical synthesis. Further derivation of these derivatives as precursors using chemical or biological means and additional biological activities including cytotoxicty against cancer are worth investigating. Our studies along with others indicated that A. niger could produce mainly the deoxyartemisinin derivatives, which can cause the significant loss of antiplasmodial activity. However, fortunately other microorganisms such as Mucor polymorphosporus (Zhan et al. 2002b), Cunninghamella echinulata (Zhan et al. 2002a), C. elegans (Parshikov et al. 2004), Eurotium amstelodami (Parshikov et al. 2006) and Rhizopus stolonifer (Gaur et al. 2014) could yield (di)hydroxylation derivatives of artemisinin while retaining the crucial and sensitive peroxide bridge group for their antiplasmodial activity. In the future, those derivatives and novel hydroxy derivatives will be made using the same or related microorganism species by further targeted screening and tested for their likely improved activity. The hydroxylation artemisinin derivatives generated from microbial transformation may provide advantages over artemisinin itself by increasing water solubility and binding affinity to its targets. Conjugation of artemisinin with (iso)quinoline and other compounds into hybrid compounds has been demonstrated to be an effective way to tackle multidrug-resistant malaria (Capci et al. 2019; Peter et al. 2021). Therefore, making novel hybrid molecules based on the hydroxylation artemisinin derivative with a reactive hydroxy group and other antimalarial compounds such as (iso)quinonline (Capci et al. 2019), gallic acid (Aldulaimi et al. 2017), thymoquinone (Johnson-Ajinwo et al. 2018) and cycleanine derivatives (Uche et al. 2021, 2018) with different mechanisms of action could be a promising approach to attack artemisinin resistance.
In conclusion, a novel atemisinin derivative (2) along with three known compounds has been generated using mild microbial transformation of atemisinin with a strain of A. niger CICC 2487, whose structure was elucidated using extensive spectroscopic analysis. The antiplasmodial activity of all the transfomed products signficantly decreased due to the loss of the peroxide moiety in artemisinin. Hydroxylation artemisinin derivatives retaining a peroxide bridge through microbial transformation should be the future focus, which will allow facile chemical conjugation of artemisinin with other antimalarial compounds to tackle artemisinin-sensitive or artemisinin-resistant malaria through dual or multiple mechanisms.
Data availability
All data generated or analyzed during this study are included in this published article (and its supplementary information files).
References
Abourashed EA, Clark AM, Hufford CD (1999) Microbial models of mammalian metabolism of xenobiotics: An updated review. Curr Med Chem 6(5):359–374
Aldulaimi O, Uche FI, Hameed H, Mbye H, Ullah I, Drijfhout F, Claridge TDW, Horrocks P, Li WW (2017) A characterization of the antimalarial activity of the bark of Cylicodiscus gabunensis Harms. J Ethnopharmacol 198:221–225. https://doi.org/10.1016/j.jep.2017.01.014
Ashley EA, Dhorda M, Fairhurst RM, Amaratunga C, Lim P, Suon S, Sreng S, Anderson JM, Mao S, Sam B, Sopha C, Chuor CM, Nguon C, Sovannaroth S, Pukrittayakamee S, Jittamala P, Chotivanich K, Chutasmit K, Suchatsoonthorn C, Runcharoen R, Hien TT, Thuy-Nhien NT, Thanh NV, Phu NH, Htut Y, Han KT, Aye KH, Mokuolu OA, Olaosebikan RR, Folaranmi OO, Mayxay M, Khanthavong M, Hongvanthong B, Newton PN, Onyamboko MA, Fanello CI, Tshefu AK, Mishra N, Valecha N, Phyo AP, Nosten F, Yi P, Tripura R, Borrmann S, Bashraheil M, Peshu J, Faiz MA, Ghose A, Hossain MA, Samad R, Rahman MR, Hasan MM, Islam A, Miotto O, Amato R, MacInnis B, Stalker J, Kwiatkowski DP, Bozdech Z, Jeeyapant A, Cheah PY, Sakulthaew T, Chalk J, Intharabut B, Silamut K, Lee SJ, Vihokhern B, Kunasol C, Imwong M, Tarning J, Taylor WJ, Yeung S, Woodrow CJ, Flegg JA, Das D, Smith J, Venkatesan M, Plowe CV, Stepniewska K, Guerin PJ, Dondorp AM, Day NP, White NJ, Tracking Resistance to Artemisinin C (2014) Spread of artemisinin resistance in Plasmodium falciparum malaria. N Engl J Med 371(5):411–423. https://doi.org/10.1056/NEJMoa1314981
Balikagala B, Fukuda N, Ikeda M, Katuro OT, Tachibana SI, Yamauchi M, Opio W, Emoto S, Anywar DA, Kimura E, Palacpac NMQ, Odongo-Aginya EI, Ogwang M, Horii T, Mita T (2021) Evidence of Artemisinin-Resistant Malaria in Africa. N Engl J Med 385(13):1163–1171. https://doi.org/10.1056/NEJMoa2101746
Capci A, Lorion MM, Wang H, Simon N, Leidenberger M, Borges Silva MC, Moreira DRM, Zhu Y, Meng Y, Chen JY, Lee YM, Friedrich O, Kappes B, Wang J, Ackermann L, Tsogoeva SB (2019) Artemisinin-(Iso)quinoline Hybrids by C-H Activation and Click Chemistry: Combating Multidrug-Resistant Malaria. Angew Chem Int Ed Engl 58(37):13066–13079. https://doi.org/10.1002/anie.201907224
Crespo-Ortiz MP, Wei MQ (2012) Antitumor activity of artemisinin and its derivatives: from a well-known antimalarial agent to a potential anticancer drug. J Biomed Biotechnol 2012:247597. https://doi.org/10.1155/2012/247597
de Carvalho CC, da Fonseca MM (2006) Biotransformation of terpenes. Biotechnol Adv 24(2):134–142. https://doi.org/10.1016/j.biotechadv.2005.08.004
Efferth T (2017) From ancient herb to modern drug: Artemisia annua and artemisinin for cancer therapy. Semin Cancer Biol 46:65–83. https://doi.org/10.1016/j.semcancer.2017.02.009
Gaur R, Darokar MP, Ajayakumar PV, Shukla RS, Bhakuni RS (2014) In vitro antimalarial studies of novel artemisinin biotransformed products and its derivatives. Phytochemistry 107:135–140. https://doi.org/10.1016/j.phytochem.2014.08.004
Goswami A, Saikia PP, Barua NC, Bordoloi M, Yadav A, Bora TC, Gogoi BK, Saxena AK, Suri N, Sharma M (2010) Bio-transformation of artemisinin using soil microbe: Direct C-acetoxylation of artemisinin at C-9 by Penicillium simplissimum. Bioorg Med Chem Lett 20(1):359–361. https://doi.org/10.1016/j.bmcl.2009.10.097
Hasenkamp S, Wong EH, Horrocks P (2012) An improved single-step lysis protocol to measure luciferase bioluminescence in Plasmodium falciparum. Malar J 11:42. https://doi.org/10.1186/1475-2875-11-42
Johnson-Ajinwo OR, Ullah I, Mbye H, Richardson A, Horrocks P, Li WW (2018) The synthesis and evaluation of thymoquinone analogues as anti-ovarian cancer and antimalarial agents. Bioorg Med Chem Lett 28(7):1219–1222. https://doi.org/10.1016/j.bmcl.2018.02.051
Kapetanaki S, Varotsis C (2001) Fourier transform infrared investigation of non-heme Fe(III) and Fe(II) decomposition of artemisinin and of a simplified trioxane alcohol. J Med Chem 44(19):3150–3156. https://doi.org/10.1021/jm010848d
Kumari A, Karnatak M, Singh D, Shankar R, Jat JL, Sharma S, Yadav D, Shrivastava R, Verma VP (2019) Current scenario of artemisinin and its analogues for antimalarial activity. Eur J Med Chem 163:804–829. https://doi.org/10.1016/j.ejmech.2018.12.007
Lee IS, elSohly HN, Croom EM, Hufford CD (1989) Microbial metabolism studies of the antimalarial sesquiterpene artemisinin. J Nat Prod 52(2):337–341. https://doi.org/10.1021/np50062a020
Lin AJ, Klayman DL, Hoch JM, Silverton JV, George CF (1985) Thermal Rearrangement and Decomposition Products of Artemisinin (Qinghaosu). J Org Chem 50(23):4504–4508
Liu JH, Chen YG, Yu BY, Chen YJ (2006) A novel ketone derivative of artemisinin biotransformed by Streptomyces griseus ATCC 13273. Bioorg Med Chem Lett 16(7):1909–1912. https://doi.org/10.1016/j.bmcl.2005.12.076
Luo X-D, Yeh HJC, Brossi A, Flippen-Anderson JL, Gilardi R (1985) The Chemistry of Drugs. VI. Thermal Decomposition of Qinghaosu. Heterocycles 23:881–887
Miotto O, Sekihara M, Tachibana SI, Yamauchi M, Pearson RD, Amato R, Goncalves S, Mehra S, Noviyanti R, Marfurt J, Auburn S, Price RN, Mueller I, Ikeda M, Mori T, Hirai M, Tavul L, Hetzel MW, Laman M, Barry AE, Ringwald P, Ohashi J, Hombhanje F, Kwiatkowski DP, Mita T (2020) Emergence of artemisinin-resistant Plasmodium falciparum with kelch13 C580Y mutations on the island of New Guinea. PLoS Pathog 16(12):e1009133. https://doi.org/10.1371/journal.ppat.1009133
O’Neill PM, Barton VE, Ward SA (2010) The molecular mechanism of action of artemisinin–the debate continues. Molecules 15(3):1705–1721. https://doi.org/10.3390/molecules15031705
Pandey P, Singh S, Tewari N, Srinivas KVNS, Shukla A, Gupta N, Vasudev PG, Khan F, Pal A, Bhakuni RS, Tandon S, Kumar JK, Banerjee S (2015) Hairy root mediated functional derivatization of artemisinin and their bioactivity analysis. J Mol Catal B Enzym 113:95–103
Parshikov IA, Muraleedharan KM, Avery MA, Williamson JS (2004) Transformation of artemisinin by Cunninghamella elegans. Appl Microbiol Biotechnol 64(6):782–786. https://doi.org/10.1007/s00253-003-1524-z
Parshikov IA, Miriyala B, Muraleedharan KM, Avery MA, Williamson JS (2006) Microbial transformation of artemisinin to 5-hydroxyartemisinin by Eurotium amstelodami and Aspergillus niger. J Ind Microbiol Biotechnol 33(5):349–352. https://doi.org/10.1007/s10295-005-0071-2
Patel OPS, Beteck RM, Legoabe LJ (2021) Exploration of artemisinin derivatives and synthetic peroxides in antimalarial drug discovery research. Eur J Med Chem 213:113193. https://doi.org/10.1016/j.ejmech.2021.113193
Peter S, Jama S, Alven S, Aderibigbe BA (2021) Artemisinin and Derivatives-Based Hybrid Compounds: Promising Therapeutics for the Treatment of Cancer and Malaria. Molecules 26(24) https://doi.org/10.3390/molecules26247521
Ponnapalli MG, Sura MB, Sudhakar R, Govindarajalu G, Sijwali PS (2018) Biotransformation of Artemisinin to 14-Hydroxydeoxyartemisinin: C-14 Hydroxylation by Aspergillus flavus. J Agric Food Chem 66(40):10490–10495. https://doi.org/10.1021/acs.jafc.8b03573
Tu Y (2016) Artemisinin-A Gift from Traditional Chinese Medicine to the World (Nobel Lecture). Angew Chem Int Ed Engl 55(35):10210–10226. https://doi.org/10.1002/anie.201601967
Uche FI, McCullagh J, Claridge TWD, Richardson A, Li WW (2018) Synthesis of (aminoalkyl)cycleanine analogues: cytotoxicity, cellular uptake, and apoptosis induction in ovarian cancer cells. Bioorg Med Chem Lett 28(9):1652–1656. https://doi.org/10.1016/j.bmcl.2018.03.038
Uche FI, Guo X, Okokon J, Ullah I, Horrocks P, Boateng J, Huang C, Li WW (2021) In Vivo Efficacy and Metabolism of the Antimalarial Cycleanine and Improved In Vitro Antiplasmodial Activity of Semisynthetic Analogues. Antimicrob Agents Chemother 65(2) https://doi.org/10.1128/AAC.01995-20
Ullah I, Sharma R, Mete A, Biagini GA, Wetzel DM, Horrocks PD (2020) The relative rate of kill of the MMV Malaria Box compounds provides links to the mode of antimalarial action and highlights scaffolds of medicinal chemistry interest. J Antimicrob Chemother 75(2):362–370. https://doi.org/10.1093/jac/dkz443
Wong EH, Hasenkamp S, Horrocks P (2011) Analysis of the molecular mechanisms governing the stage-specific expression of a prototypical housekeeping gene during intraerythrocytic development of P. falciparum. J Mol Biol 408(2):205–21
Wu WM, Wu YK, Wu YL, Yao ZJ, Zhou CM, Li Y, Shan F (1998) Unified mechanistic framework for the Fe(II)-induced cleavage of Qinghaosu and derivatives/analogues. The first spin-trapping evidence for the previously postulated secondary C-4 radical (vol 120, pg 3316, 1998). J Am Chem Soc 120(49):13002–13002
Yu H, Zhu B, Zhan Y (2017) Microbial transformation of artemisinin by Aspergillus terreus. Bioresour Bioprocess 4(1):33. https://doi.org/10.1186/s40643-017-0164-6
Zhan JX, Guo HZ, Dai JG, Zhang YX, Guo DA (2002) Microbial metabolism of artemisinin by Cunningghamella echinulata and Aspergillus niger. Tetrahedron Lett 43:4519–4523
Zhan JX, Zhang YX, Guo HZ, Han J, Ning LL, Guo DA (2002) Microbial metabolism of artemisinin by Mucor polymorphosporus and Aspergillus niger. J Nat Prod 65(11):1693–1695. https://doi.org/10.1021/np020113r
Zhan Y, Liu H, Wu Y, Wei P, Chen Z, Williamson JS (2015) Biotransformation of artemisinin by Aspergillus niger. Appl Microbiol Biotechnol 99(8):3443–3446. https://doi.org/10.1007/s00253-015-6464-x
Zhan Y, Wu Y, Xu F, Bai Y, Guan Y, Williamson JS, Liu B (2017) A novel dihydroxylated derivative of artemisinin from microbial transformation. Fitoterapia 120:93–97. https://doi.org/10.1016/j.fitote.2017.05.015
Zhang CJ, Wang J, Zhang J, Lee YM, Feng G, Lim TK, Shen HM, Lin Q, Liu B (2016) Mechanism-Guided Design and Synthesis of a Mitochondria-Targeting Artemisinin Analogue with Enhanced Anticancer Activity. Angew Chem Int Ed Engl 55(44):13770–13774. https://doi.org/10.1002/anie.201607303
Acknowledgements
We thank Keele University for ILAS fellowship to XDR and WWL.
Author information
Authors and Affiliations
Contributions
JL and XDR performed the microbial transformation, TLC, HPLC and LCHRMS analysis of transformation products; WWL performed the isolation, GC–MS measurement and structural elucidation of products. FD performed ESI–MS analysis; SW performed NMR measurement; RM and PH performed antiplasmodial activity; WWL and XDR. designed and conceived the studies; JL, XDR and WWL drafted, and all authors contributed to the article. All authors have read and agreed to the published version of the manuscript.
Corresponding authors
Ethics declarations
Conflicts of Interest
All authors do not have conflict of interest to disclose.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
253_2022_11888_MOESM1_ESM.pdf
Supplementary file1 TLC analysis, GC–MS, NMR data and IC50 curves against P. falciparum Dd2luc parasites of artemisinin transformation products are available (PDF 522 KB)
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Luo, J., Mobley, R., Woodfine, S. et al. Biotransformation of artemisinin to a novel derivative via ring rearrangement by Aspergillus niger. Appl Microbiol Biotechnol 106, 2433–2444 (2022). https://doi.org/10.1007/s00253-022-11888-0
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-022-11888-0