Abstract
The effect of salinity on the activity of nitrifying bacteria, floc characteristics, and microbial community structure accessed by fluorescent in situ hybridization and polymerase chain reaction–denaturing gradient gel electrophoresis techniques was investigated. Two sequencing batch reactors (SRB1 and SBR2) treating synthetic wastewater were subjected to increasing salt concentrations. In SBR1, four salt concentrations (5, 10, 15, and 20 g NaCl/L) were tested, while in SBR2, only two salt concentrations (10 and 20 g NaCl/L) were applied in a more shock-wise manner. The two different salt adaptation strategies caused different changes in microbial community structure, but did not change the nitrification performance, suggesting that regardless of the different nitrifying bacterial community present in the reactor, the nitrification process can be maintained stable within the salt range tested. Specific ammonium oxidation rates were more affected when salt increase was performed more rapidly and dropped 50% and 60% at 20 g NaCl/L for SBR1 and SBR2, respectively. A gradual increase in NaCl concentration had a positive effect on the settling properties (i.e., reduction of sludge volume index), although it caused a higher amount of suspended solids in the effluent. Higher organisms (e.g., protozoa, nematodes, and rotifers) as well as filamentous bacteria could not withstand the high salt concentrations.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Nitrification is a key process in biological nitrogen removal. Commonly, it is considered as the rate-limiting step of the overall biological wastewater treatment process due to the low growth rate of the organisms involved. Moreover, in general, the autotrophic nitrifying bacteria have been reported to be more sensitive for the presence of inhibitory organic compounds (Juliastuti et al. 2003), heavy metals (Hu et al. 2002; Juliastuti et al. 2003; You et al. 2009), and high substrate concentrations (Anthonisen et al. 1976). Nitrification is also susceptible to inhibition by salt, which has been considered an instability factor in many wastewater treatment plants, especially in industrial settings (Moussa et al. 2006). High salt concentrations can be found in wastewaters generated by chemical, pharmaceutical and petroleum industries, fish caning, seafood processing, meatpacking, tannery, and cheese factories (Dahl et al. 1997). The use of seawater for toilet flushing can also contribute to increasing salt levels in sewage (Panswad and Anan 1999). Salinity is known to affect the metabolic activity of nitrifying bacteria, reducing microbial growth and ammonium oxidation rates (Moussa et al. 2006). Besides that, the structure and settling properties of sludge flocs, as well as the maximum oxygen solubility and its transfer to the liquid phase, are affected by salt (Van’t Riet and Tramper 1991). In this context, the detrimental effect of salt on nitrification has been the subject of several investigations (Panswad and Anan 1999; Dahl et al. 1997; Moussa et al. 2006; Campos et al. 2002). In many cases, the results presented in the literature are difficult to compare and show contradictory results. The configuration of the system and the instability of the experimental conditions (in relation to pH, temperature, and presence of inhibitory compounds) are among the factors that directly influence the extent of salt effect on the nitrification process. In addition, the way how the microbial community is adapted to increasing salt concentrations (either by gradual or pulse dosing) may considerably change the results obtained.
One operational strategy that can be applied to minimize the effect caused by salt on the nitrification process is to gradually adapt the microorganisms to high salinity levels. This procedure was successfully applied in a previous research project (Bassin et al. 2011), during operation of a moving bed biofilm reactor, when the salt concentration was progressively increased from 0 to 13 g NaCl/L over a period of 6 months. Panswad and Anan (1999) observed that the effect of a gradual increase in the salt concentration (from 0 to 30 g NaCl/L) on the chemical oxygen demand (COD) and N removal was less severe for the anaerobic/anoxic/aerobic system inoculated with salt-adapted biomass in comparison to one containing non-adapted biomass. The sludge acclimated to salt also showed a shorter recovery period after a salt shock loading of 70 g NaCl/L. Other studies regarding gradual microbial adaptation to salt showed that, depending on the salt concentration, the long-term effect of salt can be more pronounced than the short-term effect (Moussa et al. 2006). These studies do not report on the role of changes in the microbial community structure during adaptation. So, the goal of this research was to evaluate how different salt adaptation strategies will influence the microbial diversity, activity, and settling of the activated sludge. For this purpose, a combination of denaturing gradient gel electrophoresis (DGGE) and sequence analysis of polymerase chain reaction (PCR)-amplified 16S ribosomal RNA (rRNA) gene fragments and fluorescent in situ hybridization (FISH) was used to validate the PCR-based results and to observe the dominant bacterial populations, relating all these results to the nitrification process performance.
Materials and methods
Experimental setup and operational conditions
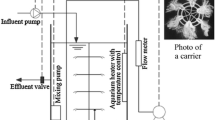
Two identical lab-scale sequencing batch reactors (SBR1 and SBR2), with a volume of 4 L, were operated in parallel in a 24-h cycle, which consisted of the following phases: (1) feeding for 3 min, (2) aeration for 23 h and 7 min, (3) settling for 45 min, and (4) effluent withdrawal for 5 min. The exchange volume was fixed at 75%. Both reactors were inoculated with activated sludge from a municipal wastewater treatment plant (ETIG-Rio de Janeiro) and fed with synthetic wastewater consisting of: 305 mg/L NH4Cl, 222 mg/L KH2PO4, 53 mg/L MgSO4, 889 mg/L NaCl, and 900 mg/L NaHCO3. A trace element solution (Vishniac and Santer 1957) was added to the culture medium in a proportion of 0.5 mL/L medium. The medium was also supplemented with different salt (NaCl) concentrations. The salt concentration was increased from 0 to 20 g/L in both reactors, although in a different manner in each system, as shown in Table 1. The operational phases of SBR1 and SBR2 are indicated in Table 1. As can be seen, the operational phases of both reactors were similar, although phases II and IV were skipped in SBR2. The experimental conditions were chosen in such a way to allow a more gradual adaptation to salt in SBR1 (four-step salt increase) in comparison with SBR2 (two-step salt increase). Air was supplied by diffusers placed on the bottom of each reactor. Dissolved oxygen (DO) level was kept above 6 mg/L and the pH was maintained between 6.8 and 7.5 by using either 1 M NaOH or 1 M HCl. All experiments were conducted at room temperature (24 ± 2°C).
Analytical methods and calculation procedures
Ammonium, biomass concentrations determined as total suspended solids (TSS), the fraction corresponding to the biomass as volatile suspended solids (VSS), and sludge volumetric index (SVI) were determined by standard methods (American Public Health Association 1995). Quantification of nitrate and nitrite was performed by ion chromatography (Dionex ICS 90) and an analytical kit provided by Hach Co., respectively. Dissolved oxygen was monitored with a DO meter (WTW Oxi-538). pH and temperature were measured with a specific electrode (Oakton pH 110 meter).
Ammonium removal efficiency was calculated on influent–effluent basis, and 100% efficiency was hence obtained when no ammonium was detectable in the effluent. The specific ammonium oxidation rate (q NH4) was determined by linear regression of the concentration of ammonium over time divided by the VSS concentration and was expressed as a fraction of the maximum specific ammonia oxidation rate (q max) observed at each salt concentration.
DNA extraction and PCR amplification
DNA was extracted from SBR1 and SBR2 samples at the end of phases I, III, and V. Samples were taken in duplicate from different parts of each reactor to confirm reproducibility of the results. No samples were taken at the end of phases II and IV of SBR1, since there were no corresponding phases in SBR2. Extraction was performed using the FastDNA® SPIN Kit (Qbiogene, Carlsbad, CA, USA), following the manufacturer’s recommendations. The extracted DNA was evaluated on 1% (wt/vol) agarose gel and stored at −20°C until further use. Around 10 ng of genomic DNA was used as template for PCR amplification of the 16S rRNA gene, which was performed with universal primers for the domain bacteria: BAC341F (containing a 40-bp GC clamp) and BAC907RM (M=A/C) (Schafer and Muyzer 2001). The standard thermal profile used included a pre-cooling phase of the thermocycler at 4°C for 1.5 min and initial denaturation at 95°C for 5 min, followed by 32 cycles of denaturing at 95°C for 30 s, primer annealing at 57°C for 40 s and elongation at 72°C for 40 s. After the last cycle, a final elongation at 72°C for 30 min took place and the amplification ended at 12°C. The primer set amoA-1F-GC and amoA-2R (Hornek et al. 2006) was used for amplification of the gene encoding ammonia monooxygenase of ammonia-oxidizing bacteria (amoA gene). The following conditions were chosen for the amplification of amoA gene: 4 min at 94°C (initial denaturation), then 35 cycles consisting of 30 s at 94°C (denaturation), 40 s at 60°C (annealing), and 40 s at 72°C (elongation) and 30 min at 72°C (final extension). PCR products of 16S rRNA and amoA genes were quantified in a 1% (wt/vol) agarose gel.
Denaturing gradient gel electrophoresis
DGGE was performed using the Bio-Rad DCode System (Bio-Rad, Richmond, USA). Electrophoresis was run in 1-mm-thick gels, containing either 6% or 8% polyacrylamide for 16S RNA and amoA gene PCR products, respectively. The denaturing gradients of the gels varied from 20% to 70% for 16S RNA fragments and from 10% to 50% for amoA fragments (100% denaturants is defined as 7 M urea and 40% (v/v) deionized formamide). Gels were submerged in 1× TAE buffer (40 mM Tris, 40 mM acetic acid, 1 mM EDTA, pH 7.4) for 16S RNA fragments, or 0.5× TAE buffer for the amoA gene fragments. Around 250 ng of the GC-clamped PCR products was added to each gel lane. Electrophoresis of 16S rRNA PCR products lasted for 16 h at a constant voltage of 100 V and a temperature of 60°C. For the amoA gene PCR products, the electrophoresis was run for 5 h at 200 V and a temperature of 55°C. After electrophoresis, the gels were stained for 30 min with a 5-mL 1× TAE solution containing SYBR green nucleic acid stain (Molecular Probes, Eugene, OR, USA) in the dark and visualized in a Safe Imager Blue-Light Transilluminator (Invitrogen, Carlsbad, CA). The gel images were captured with the GeneSnap system (Syngene, Cambridge, UK).
Individual bands from DGGE gels of both 16S RNA and amoA genes were excised using sterile razor blades, eluted in 1× Tris–HCl buffer, and stored overnight at 4°C. The same PCR programs as described previously were followed for the DNA re-amplification with non-GC-clamped primers, in which a volume of 1 μL of the DNA eluted from the DGGE band was used as template. DNA sequencing analysis was carried out by the commercial company Macrogen (Korea). The obtained 16S rRNA and amoA gene sequences were compared to sequences stored in GenBank using the Basic Local Alignment Search Tool algorithm (http://www.ncbi.nlm.nih.gov/blast). In a further step, the sequences were imported into the ARB software (http://www.arb-home.de), aligned by using the ARB automatic aligner. The alignment was further verified and corrected manually. Phylogenetic trees were generated by performing neighbor-joining algorithm. The DGGE banding pattern of different samples was analyzed and compared using the GelCompar II software package (Applied Maths, Belgium). Cluster analysis, displayed as a dendrogram, was done using the unweighted pair group method with mathematical averages, and calculation of the pair-wise similarities was based on the Dice correlation coefficient. The sequences were deposited in GenBank under the following accession numbers: JF710450–JF710456 and JF710458–JF710469 (for 16S rRNA sequences) and JF710470–JF710480 (for amoA sequences).
Fluorescent in situ hybridization
Sludge samples were taken from both the sequencing batch reactors at the end of phases I, III, and V. The samples were washed twice with 1× phosphate-buffered saline (PBS) (0.01 M K2HPO4 + KPO4, pH 7.0, 0.85% (wt/vol) NaCl) and were fixed with freshly prepared 4% (wt/vol) paraformaldehyde for 3 h at 4°C. Subsequently, the samples were centrifuged at 13,000×g for 1 min, washed twice in 1× PBS, and re-suspended in an ethanol/PBS solution (1:1) before storage at −20°C. During the hybridization step, samples previously fixed were spread on gelatin-coated microscope slides and were placed in the oven at 46°C for drying. The cover slips containing the dried cells were dehydrated in three steps (3 min each step) with 50%, 80%, and 96% (v/v) ethanol. After dehydration, 10 μL of a hybridization buffer solution containing 0.9 M NaCl, 0.02 M Tris–HCl, 35% (v/v) formamide for all probes, and 0.02% (w/v) sodium dodecyl sulfate (SDS), including fluorescently labeled oligonucleotide probes (0.5 pmol for Cy3/Cy5 and 0.83 pmol for fluorescein-labeled probes), was added to the cells. The hybridization was carried out in a humid chamber for at least 1.5-h incubation period at 46°C. A subsequent washing step to remove unbound oligonucleotides was performed by immersing the gelatin-coated slides in a buffer containing 20 mM Tris–HCl (pH 8), 0.01% (w/v) sodium dodecyl sulfate, 0.08 mM NaCl, and 0.005 mM EDTA for 10 min at 48°C. The wells of the slides were rinsed with Milli-Q water, dried by compressed air, and embedded in 2 μL of Vectashield H-1000 mounting oil for fluorescence (Vector Laboratories, Burlingame, CA, USA). Slides were observed with an epifluorescence microscope (Axioplan 2, Zeiss), and image acquisition was performed with a Leica D350F camera. The hybridization experiments were performed using different fluorochromes for each probe to validate the results. The images were exported as jpg format from the Zeiss microscopy imaging software (AxioVision version 4.7). The rRNA-targeted oligonucleotide probes, labeled with three different fluorescent dyes (Cy3, Fluos, and Cy5), are listed in Table 2.
Results
Nitrification
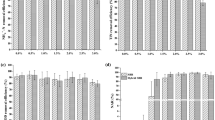
The influent ammonium concentration was kept around 80 mg N/L, and the effluent concentration was usually lower than 1 mg N/L during all experimental phases for both reactors (Fig. 1). Some small disturbances occurred due to problems related to pH adjustment or to an accidental high ammonium load. The average NH4–N removal efficiency was 95% and 96% for SBR1 and SBR2, respectively. The nitrite concentration was always below 0.5 mg N/L. According to the amount of ammonium oxidized and nitrate/nitrite produced, a nitrogen balance was performed. The soluble nitrogen recovery amounted approximately 88% for SBR1 and 86% for SBR2, independent of the actual salt concentration. Stripping of ammonia can be neglected, since the pH values were always lower than 7.5 and a low ammonium concentration was present in the reactors.
Cycle measurements were performed at the end of each operational phase for both sequencing batch systems. Ammonium profiles are shown in Fig. 2. The specific ammonia oxidation rate (q NH4) obtained during operation at different salt concentrations was calculated and expressed as a fraction of the maximum specific ammonia oxidation rate observed at non-elevated salt concentrations (q max). The actual biomass loading rate in the system was approximately 2 mg mN/g VSS/h. The results are shown in Fig. 3. In SBR1, in which salt adaptation was performed more slowly, the maximum specific ammonia oxidation rate was approximately 14 mg N/g VSS/h, value obtained at 5 g/L (phase II). The q NH4 obtained in phase I (no salt addition) and phase III was a bit lower and amounted around 13 mg N/g VSS/h. When the NaCl content was increased from 10 to 15 g/L (transition from phase III to phase IV), the q NH4 considerably decreased and reached only 70% of the q max. The lowest maximum specific ammonia activity was observed in phase V (20 g NaCl/L) and was equivalent to 50% of q max. The salt acclimation procedure in SBR2 caused a stronger impact on the specific ammonia oxidation rate compared to the results obtained during operation of SBR1. For SBR2, the maximum specific ammonia oxidation rate was observed in phase I (13.5 mg N/g VSS/h), when no salt was present in the synthetic media fed to the reactor. When the NaCl concentration was increased directly from 0 (phase I) to 10 g/L (phase III), q NH4 decreased to 20%. This activity was reduced by 60% at the end of phase V. Results from cycle measurements also showed that no accumulation of nitrite occurred during all the experimental phases of both reactors. This implies that the specific nitrite oxidation rate (qNO2) was roughly equal to the specific ammonia oxidation rate (q NH4).
Specific ammonia oxidation rate (q NH4) as a fraction of the maximum ammonia oxidation rate (q max) obtained during the operation of SBR1 (filled square) and SBR2 (empty square) reactor under different salt concentration. Data from Moussa et al. (2006) are shown in dotted lines
Biomass concentration and sludge properties
TSS and VSS of SBR1 and SBR2 along the whole experimental period are shown in Table 3. During phase I, biomass concentration tended to be constant in both reactors. When the salt concentration was increased from 0 to 5 g/L in SBR1, a decrease in the biomass concentration was observed, which can be attributed to a higher washout of cells as indicated by the increased turbidity of the supernatant. When phase III was started, a considerable washout of solids was observed, particularly in SBR2, where the NaCl content was increased directly from 10 to 20 g/L. In SBR1, the increase of salt from 5 (phase II) to 10 g/L (phase III) did not cause a big difference in VSS concentration. Conversely, with the transition from phase III (10 g/L) to phase IV (15 g/L), the concentration of biomass in the SBR1 supernatant considerably increased, which led to more suspended solids in the effluent and, consequently, to a decrease in VSS in the reactor. In phase V, just a slight decrease in biomass concentration was observed in both reactors, tending to reach a constant value. In general, the decrease in solids concentration was more pronounced in SBR2 in comparison with SBR1, especially when considering phase III of that reactor. Consequently, the sludge retention time (SRT) decreased more in SBR2, which reached values of around 20 days at the end of the operation. The percentage of volatile suspended solids among the total suspended solids was also reduced with the increment of salt in both reactors, indicating the accumulation of inorganic matter in the activated sludge. Biomass yield did not change very much with increasing salinity and amounted around 0.02 g VSS/g NH4–N removed.
The SVI was monitored during phases I–V (Table 3). A decrease in the SVI was observed with an increase in salinity levels in both reactors. In SBR1, the SVI decreased gradually from 110 mL/g (phase I) to 70 mL/g (phase IV). From phase IV to phase V, no variation was observed. In SBR2, the reduction of the SVI was slightly higher as compared to that observed in SBR1. The SVI dropped from 110 mL/g (phase I) to 60 mL/g (phase V).
Higher organisms, such as protozoa, nematodes, and rotifers (shown in the Supplementary Material) were found in the sludge at the beginning of the operation of both SBRs. When the salt concentration was increased from 5 to 10 g/L during the transition from phase II to phase III in SBR1, these organisms started to disappear. At the end of phase III, only a small number of them could be found. In SBR2, these organisms completely disappeared during the passage from phase I to phase III. The osmotic pressure seems to play a big role in the higher organism survival (Moussa et al. 2006), and the slower adaptation to salt in SBR1 enhanced their tolerance to increased salinity levels. Filamentous bacteria were also found in phases I and II of SBR1 and in phase I of SBR2. When the salt concentration was equal or above 10 g/L, these microorganisms were not detected anymore.
Microbial population dynamics revealed by PCR–DGGE
Dynamics of general bacterial community
The composition of the microbial community in the sequencing batch reactors was investigated through PCR–DGGE analysis. Figure 4 shows the typical DGGE fingerprints of both reactors at the end of phases I, III, and V. A dendogram representing the percent similarity of the different DGGE fingerprints is also presented. SBR1 and SBR2 initially contained a very similar community, but there was a strong divergence upon increasing the salt concentration. The more shock-wise salt adaptation strategy performed in SBR2 caused a more pronounced change in the microbial community compared to the change observed in SBR1. Duplicates of each sample were assayed to check the reproducibility of the results. The similarity between all the duplicates was above 97%. Since each duplicate sample was taken from a different place in the reactors, no microbial population gradient was present inside the reactors due to the perfect mixture in the liquid phase. The DGGE profiles indicate the presence of a diverse microbial community. A total of 27 bands were excised from the DGGE gel and sequenced. From those, six bands (i.e., bands B13, B16, B20, B22, B24, and B25) gave ambiguous sequencing results and were neglected in the further phylogenetic analysis. Figure 5 illustrates a phylogenetic tree based on partial sequences of the 16S rRNA gene. The microorganisms found belong to several phyla, such as Proteobacteria (α-, β-, and γ-subclass), Bacteroidetes, Chloroflexi, Firmicutes, and Actinobacteria. At the end of phase I, the community fingerprints of SBR1 and SBR2 were very similar, as indicated by the cluster analysis (similarity of 93%). This seems logical, since both reactors were operated at the same conditions in phase I (no salt addition). The majority of the excised bands were already found in the sample taken at the end of the first experimental phase, with the exception of bands B1, B4, B7, B17, B18, and B21, which were retrieved in further phases. Among the bands of phase I, ammonia-oxidizing bacteria (AOB) were detected in bands B9–B12, all of them belonging to the Nitrosomonas group. Particularly, bands B9 and B11 gave identical sequencing results. Nitrite-oxidizing bacteria were found in bands B14, B15, B21, and B26. Band B14 showed high sequence similarity with Nitrobacter vulgaris, and bands B15 and B26 were closely related to Nitrobacter hamburgensis. Even though the reactors were fed only with autotrophic media without any organic carbon source, some heterotrophic bacteria were present in the microbial community in phase I. These microorganisms were found in bands B3, B5, B6, B8, B19, B23, and B27. From those, bands B3, B5, and B6 clustered well with members of the Bacteroidetes phylum. Comparative sequence analysis showed that band B8 belongs to α-Proteobacteria. Bands B19 and B23 represented bacteria belonging to the Firmicutes and Chloroflexi, respectively. Band B27 was closely related to an uncultured Gemmatimonadetes bacterium, isolated from an enhanced biological phosphate removal process.
The banding pattern shown in phase III revealed clear differences between the microbial composition of SBR1 and SBR2, which is likely due to the different salt adaptation strategy adopted in each reactor. In SBR1, it was observed that bands B2, B3, and B6 have disappeared from phase I to phase II. Conversely, the intensity of band B12 and bands B23–B27 became higher when the salt concentration was increased from 0 (phase I) to 5 g/L (phase II) and then to 10 g/L (phase III). The banding pattern similarity between phases I and III of SBR1 was around 85%. With the increase of salt concentration directly from 0 (phase I) to 10 g/L (phase III), the microbial population in SBR2 shifted in a different manner than observed in SBR1. A significant variation in the microbial community was observed from phase I to phase III, as indicated by the relatively low percentage of similarity (50%) between the fingerprints of these two experimental phases. It was observed that new bands (B1, B4, B7, and B21) appeared. With the exception of band 21, in all other bands, microorganisms belonging to the Bacteroidetes group were retrieved. Bands B1 and B4, in particular, showed high sequence similarity to members of the family Chitinophagaceae. The closest relative to band B21 among the culture bacteria was Nitrospira moscoviensis. The ammonia oxidizers related to the Nitrosomonas group retrieved in bands B9–B12 were not detected anymore, the intensity of bands B3 (Bacteroidetes) and B23 (Chloroflexi) increased, and the intensity of the band B19 decreased.
According to the SBR1 banding profile, the differences between the microbial community structure in phase III and phase V are minimal. According to cluster analysis, more than 90% of similarity between the microbial fingerprints of these two phases was observed. The only differences were the appearance of bands B17 (closely related to Denitromonas sp.) and B18 (closely related to Arthrobacter soli) and the decrease in the intensity of band B19. The transition from phase III to phase V in SBR2 was also accompanied by small variations in the bacterial fingerprint, particularly referred to the appearance of band B17 and the increase in the intensity of bands B8, B15, and B16. According to the cluster analysis, the similarity between phases III and V for SBR2 was approximately 85%.
Dynamics of ammonia-oxidizing bacteria
In order to get a better insight into the AOB dynamics in the course of the experiment and to obtain sufficient sequence information from DGGE bands for phylogenetic analysis of AOB community, a study was carried out using DGGE analysis of PCR-amplified amoA gene fragments (Fig. 6). According to the 16S rRNA gene DGGE gel, the duplicate samples were practically identical. Therefore, only one sample representative of each experimental phase was taken into consideration in the DGGE analysis of amoA gene fragments.
Eleven dominant bands were selected, excised, and sequenced in order to reveal the identity of the microorganisms involved. The phylogenetic affiliation of the dominant bands was analyzed and depicted in a phylogenetic tree, shown in Fig. 7. As observed for the 16S rRNA gene DGGE, the microbial community fingerprint in phase I (no salt addition) was the same in SBR1 and SBR2. Furthermore, DGGE analysis showed that all the main bands representing the diverse community of AOB were already detected in the steady-state operation of phase I in both reactors. With the increasing salt concentration from 0 (phase I) to 5 g/L (phase II) and subsequently to 10 g/L (phase III) in SBR1, bands B3, B6, B8, B9, and B11 have completely disappeared. For SBR2, a similar behavior was observed with the transition from phase I directly to phase III, although some differences were observed: band B4 was not detected anymore, band B8 did not disappear and its intensity was even increased, and bands B9 and B11 were still present at the end of phase III.
The same bands of phase III were detected in phase V in SBR1. For SBR2, the only modification in the banding pattern from phase III to phase V refers to the disappearance of bands B8, B9, and B11. For both reactors, it was observed that the intensity of band B5 increased along the different phases, and bands B1, B2, B7, and B10 remained stable over all experimental phases.
The phylogenetic analysis showed that all AOB belonged to the Nitrosomonas group. Sequencing results showed that bands B1–B4, B6, B7, and B9 were closely related to Nitrosomonas europaea. The intensity of band B5 (closely related to Nitrosomonas sp.) was strongly enhanced at the highest salt concentration. Bands B8, B10, and B11 showed high sequence similarity with Nitrosomonas eutropha. Although the banding profile changed differently from phase I to phase III in SBR1 and SBR2, the AOB community in both systems was very similar at the end of the experiment (phase V). It is clear that the changes in microbial diversity observed with the amoA gene are less than with the 16S RNA gene, indicating that the impact of salt on the ammonium-oxidizing bacteria was small.
AOB and NOB populations accessed by FISH analysis
FISH analysis was performed using a combination of specific oligonucleotide probes to detect AOB and nitrite-oxidizing bacteria (NOB). Figure 8 shows representative FISH pictures obtained from both SBR samples taken at the end of phases I and V. It was observed that the proportion of nitrifying bacteria among the whole bacterial community was similar for both SBRs at the end of phase I, when both systems were operated at the same conditions (no salt addition). FISH results showed that the proportion of AOB and NOB in the whole bacterial community increased when salt concentration was increased from 0 to 10 g NaCl/L in SBR1, remaining relatively constant when the salt concentration was further increased to 20 g/L. In SBR2, the number of AOB cells decreased as the salt concentration was increased, although no significant variations were observed for the NOB population. In SBR1, the number of AOB and NOB cells was comparable, while in SBR2, the NOB cells were detected in a higher number in comparison with AOB.
AOB (in red) and NOB (in green) populations within the whole bacterial community (in blue): a SBR1, phase I; b SBR1, phase V; c SBR2, phase I; and d SBR2, phase V. AOB appear violet due to superposition of the red-labeled AOBmix and the blue-labeled EUBmix probes, while NOB appear turquoise due to the superposition of green-labeled NOBmix and the blue-labeled EUBmix probes. Scale bar indicates 5 μm
Discussion
Effect of salt on biomass concentration and sludge properties
The results have shown that TSS and VSS decreased gradually as the salinity was increased from phases I to V in both reactors, and the more rapid salt adaption procedure in SBR2 caused a higher washout of cells as compared to SBR1. Moreover, the sludge was enriched in inorganic material, which was evidenced by a decrease in the VSS/TSS ratio, especially at the highest salt concentration (20 g NaCl/L).
An increase of the supernatant turbidity, which resulted in higher washout of biomass, was also noticed. The fact that biomass yield remained practically constant over all the experiment runs in both reactors suggested that elevated biomass washout due to increasing salt concentrations was the main cause of lower sludge content in the reactor. Elevated salt concentrations are known to increase water density, which can potentially cause the washout of small and poor settling sludge flocs. According to Woolard and Irvine (1995), biological systems subjected to salt shock loads respond with increased effluent suspended solids and loss of volatile suspended solids, similar to that observed in this work. Panswad and Anan (1999) also observed a decrease in TSS when the salt concentration was increased. These authors observed a reduction of 60% of TSS with an increasing in salinity from 0 to 30 g/L of salt. In this study, the decrease in TSS was a bit lower (40% and 45% for SBR1 and SBR2, respectively), but the maximum salt concentration tested was only 20 g NaCl/L.
The effect of salt on the physical properties of activated sludge is a major concern in order to maintain good biomass retention in the reactor. Sludge volumetric index is an important parameter in activated sludge to evaluate the settling ability of the flocs. In this study, the presence of increasing salt concentrations had no negative effect on the sludge volume index. A gradual decrease of SVI was observed when the salt concentration was increased. This contrasts with an increased turbidity of the supernatant, mainly caused by the poor settling properties of smaller aggregates or dispersed organisms that were washed out from the reactor. According to Moussa et al. (2006), due to increased salinity levels and consequently increase in water density, lighter flocs will be washed out (increasing effluent turbidity), while dense flocs will remain in the reactor (decreasing SVI). The quicker salt adaptation in SBR2 caused a slightly higher decrease in the SVI than the one observed in SBR1. Moreover, the decrease in solid concentration was more pronounced in SBR2 in comparison with its counterpart, especially when considering phase III of that reactor, when salt was increased directly from 0 to 10 g/L. This result can be explained by the fact that gradual increase in salinity will stimulate the selection of dense flocs with minimum washout (as observed in SBR1), while sudden salt increase (as performed in SBR2) enhances water density, which can cause excessive washout of biomass (Moussa et al. 2006). Several investigations have been done considering the effect of salinity on SVI. In accordance to our experiments, Campos et al. (2002) and Moussa et al. (2006) also showed a decrease in SVI with increasing salt concentrations. Moon et al. (2002) pointed out that increasing salt concentrations incites physical modifications in aggregates and also observed an increase in the SVI immediately after the salt concentration was increased.
Microscopic observations (shown in the Supplementary Material) revealed that higher organisms, such as protozoa, nematodes, and rotifers, gradually disappear as salinity levels were increased. In addition, the more gradual four-step salt adaptation in SBR1 enhanced their tolerance to increased salinity levels when compared to the two-step salt adaptation in SBR2.
Moussa et al. (2006) also observed that rotifers, protozoa, and nematodes present in a nitrifying sludge could not survive when the salinity (in terms of chloride) was above 5 g Cl−/L (around 8 g NaCl/L). Salvadó et al. (2001) reported than an increase in salt concentration from 3 to 10 g NaCl/L gradually affected the protozoa and metazoan communities. Higher organisms, such as protozoa, are considered important predators which feed on particularly on suspended bacteria and other organic suspended particles in wastewater treatment systems (Curds et al. 1968; Wheale and Williamson 1980). As a consequence, they indirectly influence the formation of sludge flocs, favoring the biosolid/liquid separation. Due to their important clarification ability, the gradual disappearance of higher organisms when salt concentration was increased could contribute to the higher turbidity of the supernatant and consequent higher biomass washout observed at high salinity levels. The results of this study also indicated that filamentous bacteria, which are commonly related to sludge-bulking problems (Pipes 1978), could not withstand salt concentrations above 10 g NaCl/L in both SBRs. This can explain the fact that higher SVI values, directly reflecting poorer sludge settling characteristics, were observed when the salt concentration was below 10 g/L. On the other hand, filamentous bacteria are also important, providing a superstructure for activated sludge flocs that could enhance mechanical integrity beyond the cell interactions required for flocculation (Urbain et al. 1993). Therefore, a lack of filamentous organisms could also have some implications on the supernatant turbidity. The complete absence of these organisms from 10 g NaCl/L in both SBR1 and SBR2 possibly increased effluent turbidity, leading to higher biomass washout and, consequently, higher loss of biomass. Reduction of protozoa and filamentous organisms required for proper flocculation due to high salt levels was also reported by Woolard (1993) and Ludzack and Noran (1965).
Different salt adaptation strategies: implications for nitrification process and microbial community structure
The gradual salt adaptation process performed in both sequencing batch reactors allowed to obtain stable ammonia removal efficiency close to 100% in all experimental phases of both reactors. However, although the functionality of both systems remained the same, DGGE analysis of 16S rRNA fragments (Fig. 4) showed a significant variation in the ammonia-oxidizing bacteria community, especially in SBR2. Most of the bands whose sequences fell within the Nitrosomonas group (9–12) were practically not detected in phase III of that reactor, although ammonia removal efficiency was not affected along phases I–V. Moreover, FISH analysis (Fig. 8) using specific probes targeting Nitrosomonas genus showed that these organisms were still detected in SBR2 even at the highest salt concentration tested (20 g NaCl/L), although in a reduced number compared to the operational phase when no salt was added to the reactor influent media. The reason why AOB were almost not detected anymore in SBR2, particularly at the end of phases III and V, is that their relative abundance was below the detection limit of DGGE using universal 16S rRNA gene-targeted primers.
The fact that AOB were hardly detected in the 16S rRNA DGGE analysis is of course an unexpected result, since the reactor was fed with inorganic media during the whole operational period. Moreover, this fact was not observed in SBR1, submitted to the same process conditions, with the exception of the different salt increase procedure adopted. Therefore, the microbiological results suggest that the two-step salt adaptation which occurred in SBR2 was more severe to the AOB compared to the four-step salt adaptation performed in SBR1, a fact supported by FISH analysis results and in agreement with the lower ammonia oxidation activities obtained in SBR2 when compared to those obtained in SBR1, especially when salinity was increased from 10 to 20 g NaCl/L.
It is interesting to compare these results with the ones obtained during operation of an aerobic granular sludge reactor, subjected to salt increase up to 30 g NaCl/L (Bassin et al., submitted). For example, the shift in ammonia-oxidizing bacteria community within the granules due to salt increase was considerably lower compared to that observed in SBR1 and SBR2. The granular sludge reactor, operated for simultaneous nitrogen and phosphorus removal, was also fed with synthetic media containing ammonium as nitrogen source, although it also contained acetate as carbon source.
The low concentrations of salt had a positive effect on the ammonia oxidation activity, particularly when considering the increase of salt from 0 to 5 g/L in SBR1. At concentrations higher than 10 g NaCl/L, the specific ammonia-oxidizing activity significantly decreased. This result, combined with the decrease in SRT when the salinity was gradually increased, resulted in the substantial decrease in volumetric conversion rate.
Uygur and Kargi (2004) also observed that ammonia removal rate was not significantly affected by salt content when its concentration was lower than 10 g/L. These authors observed a reduction of around 75% of ammonia oxidation rate when the salt concentration was increased from 0 to 5 and then 10 g/L, which is roughly the same decrease observed in this study for SBR1 (70%), where salinity was increased in the same proportion. A considerable decrease in the ammonia oxidation rate with salt increase was also observed by Moussa et al. (2006), whose results are also shown in Fig. 3.
To have a better overview of the AOB community as salt concentration was increased, a functional gene (amoA) playing a key role in the ammonia-oxidizing process was used. It was demonstrated that the AOB community was quite diverse in both reactors. Moreover, the different acclimation to salt performed in SBR1 and SBR2 had only a slight influence on the AOB community.
Closely related to Nitrosomonas sp., the microorganism whose sequence was retrieved in band B5 seemed to have high affinity to salt, while others closely related to N. eutropha (bands B8 and B11) and to N. europaea (band B9) seemed to be more affected by the longer exposition time to salt which occurred in SBR1 than by more rapid changes in salinity levels in SBR2. It should be remarked that the amoA gene analysis does not reflect a quantitative evaluation, but a qualitative one. In this sense, although the AOB diversity was higher in SBR2 at 10 g/L (phase III), nothing can be said about their quantity.
It should be pointed out that the microbial population structure was changing faster than can be expected based on SRT. This can be observed in the DGGE analysis of both 16S rRNA and amoA fragments. For instance, the disappearance of several organisms in SBR1 was observed from the end of phase I to the end of phase III (33 days of operation), despite an SRT of 85 days. Often reported in the literature, accumulation of nitrite due to the salt inhibition of NOB (Ye et al. 2009; Jin et al. 2007) was not observed in the cycle measurements performed in this study, even at the highest salt concentration tested (20 g NaCl/L). Indeed, when considering the 16S rRNA DGGE profile, the bands representing NOB (B14, B15, and B26) were present along the whole experiment. Besides that, the intensity of bands B15 and B26 was even enhanced when salinity levels were increased. Moreover, a microorganism closely related to N. moscoviensis (band B21) showed high affinity for salt, since it was detected only in phases III and V of SBR2. Results obtained from FISH analysis also showed that NOB were not affected by increasing salt concentration and the NOB proportion of the whole bacterial community was even increased in the reactors.
Although the nitrifying sludge in both SBRs was cultured in inorganic media, not only autotrophic AOB and NOB were detected through PCR–DGGE analysis, but also several heterotrophic bacteria. Heterotrophic bacteria could possibly have grown on soluble products from nitrifying bacteria. Chemolithoautotrophic nitrifiers fix and reduce inorganic carbon (e.g., CO2) for cell synthesis (Brock and Madigan 1991) and produce and release soluble organic products into solution from substrate metabolism and decaying biomass (Rittman et al. 1994). Therefore, these microorganisms also interact with the exchange of organic materials. Okabe et al. (1999) also verified the coexistence of nitrifying and heterotrophic bacteria in a nitrifying biofilm fed only with inorganic synthetic media. Rittmann and Brunner (1984) observed that an active heterotrophic population was maintained in a biofilm reactor when the organic compounds were removed from the feed. These authors postulated that the active nitrifying bacteria produced a continuous flow of organic substrates for the heterotrophs. Rittmann et al. (1994) have done some experiments with Nitrobacter sp. and N. europaea and demonstrated their capability to produce soluble organic products that can be used by heterotrophic bacteria. Nevertheless, they also mentioned that just a small heterotrophic population was maintained in the autotrophic system. It remains unknown whether the diversity of heterotrophic populations growing in pure autotrophic systems can possibly be related to different usage of various organic compounds produced by autotrophs. A lack of information regarding the ecophysiological interaction between nitrifiers and heterotrophs limits further analysis. A mathematical model to describe the interaction between nitrifiers and heterotrophs in biological treatment systems was proposed by Moussa et al. (2005). Their results showed that the influent COD is responsible for around 40% of the total formed heterotrophic biomass and 60% result from biomass decay. Moreover, even at a low input COD (5–10 mg/L), heterotrophic biomass could be formed in an autotrophic reactor. Moussa et al. (2005) mentioned that such low values of COD can be indirectly introduced to the system by organic impurities in the media and in the air used for aeration. The long starvation period (up to 20 h in some cycles at 0 and 5 g NaCl/L) due to lack of substrate (totally consumed) could also favor the presence of a significant diversity of heterotrophic bacteria in both systems, enhancing the turnover of microbial population. In the absence of nitrogenous substrate (ammonium), nitrifying biomass decay could have happened, which in turn favored the heterotrophic growth at the expense of decaying nitrifying bacteria. Moreover, the considerable starvation period could also influence the low growth yield obtained (around 0.02 g VSS/g N), which is much lower compared to that obtained in autotrophic activated sludge systems reported by Blackburne et al. (2007) and by Brauer and Annachhatre (1992), who measured 0.14 and 0.4 g VSS/g N, respectively.
Nitrospira-like organisms play a big role in other ecosystems, such as soil and drinking water distribution systems, and seem to outcompete Nitrobacter under substrate limitation (Wagner and Loy 2002; Dionisi et al. 2002). Schramm et al. (2000) hypothesized that species of Nitrospira are K-strategists, thriving at low nitrite concentrations, while Nitrobacter is considered as an r-strategist, which means that they can compete only in environments containing significant amounts of nitrite. This hypothesis can explain that Nitrospira and Nitrobacter will coexist at higher nitrite concentrations (Daims et al. 2001). This hypothesis is not valid in our case. Even at really low nitrite concentrations, both Nitrobacter sp. (band B14) and N. vulgaris (band B26) coexisted with Nitrospira sp. (band B21), particularly in SBR2. The results of this work are different from those obtained by Moussa et al. (2006), who observed that Nitrospira sp. was the dominant nitrite oxidizer up to 10 g Cl−/L (16.5 g NaCl/L), whereas they could not be detected above that concentration.
Among the heterotrophs found in the bacterial community of the sludge, a microorganism closely related A. soli (band B18) was detected. Verstraete and Alexander (1972) found out that Arthrobacter sp. is an organism involved in heterotrophic nitrification, capable of oxidizing ammonium to hydroxylamine, hydroxamic acid, and nitrite. Several organic and inorganic substances can be used as nitrogen source for heterotrophic nitrification, and organic carbon source is necessary for growth of the microorganisms and for nitrification of inorganic N sources (Focht and Verstraete 1977). In our study, the occurrence of heterotrophic nitrification cannot be neglected due to the presence of organic C source derived from the soluble organic products released by active nitrifying bacteria and possibly from organic impurities within the culture medium or the air used for aeration.
As described in the “Results” section, soluble nitrogen recovery amounted approximately 88% for SBR1 and 86% for SBR2, which generates a nitrogen loss from solution of 12% and 14% in SBR1 and SBR2, respectively. The nitrogen loss from solution observed during the course of the operation can be attributed to other parallel processes, such as N2O or NO formation by nitrifying bacteria (Kampschreur et al. 2007). Moreover, the sludge produced will also contain nitrogen. Taking into account the amount of sludge (VSS) produced per day and considering a standard nitrogen content of biomass of 10%, it was estimated that the amount of nitrogen used for growth was around 5–8% of the influent ammonium nitrogen.
The results of this work reinforced the fact that activated sludge from a conventional domestic wastewater treatment can be used as a source of nitrifying bacteria which can be adapted to significant salt concentrations through gradual acclimation. The two different salt adaptation methods applied in this study allowed to reach good nitrification performance and caused different shifts in microbial community structure. The findings of this work suggest that independent of the different nitrifying bacterial community present in each reactor, the nitrification process can be maintained stable within the salt range tested. Ammonia-oxidizing activity was more affected when salt adaptation was performed more rapidly. Settling properties of the sludge improved at higher salt content, although higher organisms (protozoa, nematodes, and rotifers) and filamentous bacteria could not withstand high salinity.
References
APHA American Public Health Association (1995) Standard methods for the examination of water and wastewater, 21st, centennial edition. APHA, Washington
Amann RI, Binder BJ, Olson RJ, Chisholm SW, Devereux R, Stahl DA (1990) Combination of 16S rRNA-targeted oligonucleotide probes with flow cytometry for analyzing mixed microbial populations. Appl Environ Microbiol 56:1919–609
Anthonisen AC, Loher RC, Prakasam TBS, Srinath E (1976) Inhibition of nitrification by ammonia and nitrous acid. J Water Pollut Control Fed 48:835–852
Bassin JP, Dezotti M, Sant’Anna GL Jr (2011) Nitrification of industrial and domestic saline wastewaters in moving bed biofilm reactor and sequencing batch reactor. J Haz Mat 185:242–248
Blackburne R, Vadivelu VM, Yuan Z, Keller J (2007) Determination of growth rate and yield of nitrifying bacteria by measuring carbon dioxide uptake rate. Water Environ Res 79:2437–2445
Brauer H, Annachhatre AP (1992) Wastewater nitrification kinetics using reciprocating jet bioreactor. Bioprocess Eng 7:277–286
Brock TD, Madigan MT (1991) Biology of microorganisms, 6th edn. Prentice-Hall, Englewood Cliff
Campos JL, Mosquera-Corral A, Méndez R, Lema JM (2002) Nitrification in saline wastewater with high ammonia concentration in an activated sludge unit. Water Res 36:2555–2560
Curds CR, Cockburn A, Vandike JM (1968) An Experimental Study of the Role of the Ciliated Protozoa in the Activated-Sludge Process. Wat Pollut Control 67:312–329
Dahl C, Sund C, Kristensen GH, Vredenbregt L (1997) Combined biological nitrification and denitrification of high-salinity wastewater. Water Sci Technol 36:345–352
Daims H, Brühl A, Amann R, Schleifer KH, Wagner M (1999) The domain-specific probe EUB338 is insufficient for the detection of all Bacteria: Development and evaluation of a more comprehensive probe set. Syst Appl Microbiol 22:434–444
Daims H, Nielsen P, Nielsen JL, Juretschko S, Wagner M (2000) Novel Nitrospira-like bacteria as dominant nitriteoxidizers in biofilms from wastewater treatment plants: diversity and in situ-physiology. Water Sci Technol 41:85–90
Daims H, Nielsen JL, Nielsen PH, Schleifer KH, Wagner M (2001) In situ characterization of Nitrospira-like nitrite-oxidizing bacteria active in wastewater treatment plants. Appl Environ Microbiol 67:5273–5284
Dionisi HM, Layton AC, Harms G, Gregory IR, Robinson KG, Sayler GS (2002) Quantification of Nitrosomonas oligotropha-like ammonia-oxidizing bacteria and Nitrospira spp. from full-scale wastewater treatment plants by competitive PCR. Appl Environ Microbiol 68:245–253
Focht DD, Verstraete W (1977) Biochemical ecology of nitrification and denitrification. Adv Microb Ecol 1:135–214
Hornek R, Pommerening-Roser A, Koops HP, Farnleitner AH, Kreuzinger N, Kirschner A, Mach RL (2006) Primers containing universal bases reduce multiple amoA gene specific DGGE band patterns when analysing the diversity of beta-ammonia oxidizers in the environment. J Microbiol Method 66:147–155
Hu Z, Chandran K, Grasso D, Smets BF (2002) Effect of nickel and cadmium speciation on nitrification inhibition. Environ Sci Technol 36:3074–3078
Jin R-C, Zheng P, Mahmood Q, Hu B-L (2007) Osmotic stress on nitrification in an airlift bioreactor. J Haz Mat 146:148–154
Juliastuti SR, Baeyens J, Creemers C (2003) Inhibition of nitrification by heavy metals and organic compounds: the ISO 9509 test. Environ Eng Sci 20:79–90
Juretschko S, Timmermann G, Schmid M, Schleifer K-H, Pommerening-Roser A, Koops H-P, Wagner M (1998) Combined molecular and convetional analyses of nitrifying bacterium in activated sludge: Nitrosococcus mobilis and Nitrospira-like bacteria as dominant populations. Appl Environ Microbiol 64:3042–3051
Kampschreur MJ, Picioreanu C, Tan N, Kleerebezem R, Jetten MSM, van Loosdrecht MCM (2007) Unraveling the source of nitric oxide emission during nitrification. Water Environ Res 79:2499–2509
Ludzack FJ, Noran PK (1965) Tolerance of high salinities by conventional wastewater treatment process. J Wat Pollut Control Fed 37:1404–1416
Mobarry BK, Wagner M, Urbain V, Rittmann BE, Stahl DA (1996) Phylogenetic probes for analyzing abundance and spatial organization of nitrifying bacteria. Appl. Environ Microbiol 62:2156–2162
Moon BH, Seo GT, Lee TS, Kim SS, Yoon CH (2002) Effects of salt concentration on floc characteristics and pollutants removal efficiencies in treatment of seafood wastewater by SBR. Water Sci Technol 47:65–70
Moussa MS, Hooijmans CM, Lubberding HJ, GIjzen HJ, van Loosdrecht MCM (2005) Modelling nitrification, heterotrophic growth and predation in activated sludge. Water Res 39:5080–5098
Moussa MS, Sumanasekera DU, Irahim SH, Lubberding HJ, Hooijmans CM, Gijzen HJ, van Loosdrecht MCM (2006) Long term effects of salt on activity, population structure and floc characteristics in enriched bacterial cultures of nitrifiers. Water Res 40:1377–1388
Okabe S, Satoh H, Watanabe Y (1999) In situ analysis of nitrifying biofilms as determined by in situ hybridization and the use of microelectrodes. Appl Environ Microbiol 65:3182–3191
Panswad T, Anan C (1999) Impact of high chloride wastewater on an anaerobic/anoxic/aerobic process with and without inoculation of chloride acclimated seeds. Water Res 33:1165–1172
Pipes WO (1978) Microbiology of activated sludge bulking. Adv Appl Microbiol 24:85–127
Rittmann BE, Regan JM, Stahl DA (1994) Nitrification as source of soluble organic substrate in biological treatment. Water Sci Technol 30:1–8
Rittmann BE, Brunner CW (1984) The nonsteady-state process for advanced organics removal. J Water Pollut Control Fed 56:874–880
Salvadó H, Mas M, Menéndez S, Gracia MP (2001) Effects of shock loads of salt on protozoan communites of activated sludge. Acta Protozool 40:177–185
Schafer H, Muyzer G (2001) Denaturing gel electrophoresis in marine microbial ecology. Method Microbiol 30:425–468
Schramm A, De Beer D, Gieseke A, Amann R (2000) Microenvironments and distribution of nitrifying bacteria in a membrane-bound biofilm. Environ Microbiol 2:680–686
Urbain V, Block JC, Manen J (1993) Bioflocculation of activated sludge: an analytical approach. Water Res 27:829–838
Uygur A, Kargi F (2004) Salt inhibition on biological nutrient removal from saline wastewater in a sequencing batch reactor. Enz Microb Technol 34:313–318
Van’t Riet K, Tramper J (1991) Basic bioreactor design. Marcel Dekker, New York
Verstraete W, Alexander M (1972) Heterotrophic nitrification by Arthrobacter sp. J Bact 110:955–961
Vishniac W, Santer M (1957) The thiobacilli. Bact Rev 21:195–213
Wagner M, Rath G, Amann R, Koops H-P, Schleifer K-H (1995) In-situ identification of ammonia-oxidizing bacteria. System Appl Microbiol 18:251–264
Wagner M, Rath G, Koops HP, Schleifer HK (1996) In situ identification of nitrifying bacteria in sewage treatment plants. Wat Sci Tech 34:237–244
Wagner M, Loy A (2002) Bacterial community composition and function in sewage treatment system. Environ Biotechnol 13:218–227
Wheale G, Williamson, DJ (1980) Unusual behaviour of ciliated protozoa in a secondary settlement tank. Wat Pollut Control 80:496–500
Woolard CR (1993) Biological treatment of hypersaline wastewaters. In: Ph.D. dissertation, Univ. of Notre Dame, Notre Dame, IN
Woolard CR, Irvine RL (1995) Treatment of hypersaline wastewater in the sequencing batch reactor. Water Res 29:1159–1168
Ye L, Peng C-Y, Tang B, Wang S-Y, Zhao K-F, Peng Y-Z (2009) Determination effect of influent salinity and inhibition time on partial nitrification in a sequencing batch reactor treating saline sewage. Desalination 246:556–566
You S-J, Tsai Y-P, Huang R-Y (2009) Effect of heavy metals on nitrification performance in different activated sludge processes. J Haz Mat 165:987–994
Open Access
This article is distributed under the terms of the Creative Commons Attribution Noncommercial License which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and source are credited.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This is an open access article distributed under the terms of the Creative Commons Attribution Noncommercial License (https://creativecommons.org/licenses/by-nc/2.0), which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and source are credited.
About this article
Cite this article
Bassin, J.P., Kleerebezem, R., Muyzer, G. et al. Effect of different salt adaptation strategies on the microbial diversity, activity, and settling of nitrifying sludge in sequencing batch reactors. Appl Microbiol Biotechnol 93, 1281–1294 (2012). https://doi.org/10.1007/s00253-011-3428-7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-011-3428-7