Abstract
Water-filled sinkholes known locally as cenotes, found on the Yucatán Peninsula, have remarkable biodiversity. The primary objective of this study was to explore the biotechnological potential of Gram-positive cultivable bacteria obtained from sediment samples collected at the coastal cenote Pol-Ac in Yucatán, Mexico. Specifically, the investigation aimed to assess production of hydrolytic enzymes and antimicrobial compounds. 16 S rRNA gene sequencing led to the identification of 49 Gram-positive bacterial isolates belonging to the phyla Bacillota (n = 29) and Actinomycetota (n = 20) divided into the common genera Bacillus and Streptomyces, as well as the genera Virgibacillus, Halobacillus, Metabacillus, Solibacillus, Neobacillus, Rossellomorea, Nocardiopsis and Corynebacterium. With growth at 55ºC, 21 of the 49 strains were classified as moderately thermotolerant. All strains were classified as halotolerant and 24 were dependent on marine water for growth. Screening for six extracellular hydrolytic enzymes revealed gelatinase, amylase, lipase, cellulase, protease and chitinase activities in 93.9%, 67.3%, 63.3%, 59.2%, 59.2% and 38.8%, of isolated strains, respectively. The genes for polyketide synthases type I, were detected in 24 of the strains. Of 18 strains that achieved > 25% inhibition of growth in the bacterial pathogen Staphylococcus aureus ATCC 6538, 4 also inhibited growth in Escherichia coli ATCC 35,218. Isolates Streptomyces sp. NCA_378 and Bacillus sp. NCA_374 demonstrated 50–75% growth inhibition against at least one of the two pathogens tested, along with significant enzymatic activity across all six extracellular enzymes. This is the first comprehensive report on the biotechnological potential of Gram-positive bacteria isolated from sediments in the cenotes of the Yucatán Peninsula.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The Yucatán Peninsula in the Gulf of Mexico is a karst landscape with many natural flooded sinkholes, formed by the collapse of limestone bedrock and known locally as cenotes [1]. These cenotes are of profound cultural significance as they served as vital water sources for the ancient Maya civilization and were revered as sacred sites for their ritual activities [2]. Their unexploited biotechnological potential now requires urgent investigation [3], for example as source of novel halotolerant bacteria possessing antimicrobial properties [4].
Cenotes close to the coast show a saltwater intrusion beneath the freshwater surface, and this creates a marked shallow halocline [5]. Low concentrations of oxygen, high concentrations of sulfate and other nutrients, and stable water temperature, make them ideal habitats for the growth of distinctive and diverse species of bacteria [5, 6]. Their bacterial communities not only help in the decomposition of organic matter, but are also crucial in biogeochemical cycling, nutrient recycling, and the delicate ecosystem equilibrium [7, 8]. Therefore, they may produce a range of enzymes with biotechnological potential, such as extracellular carbohydrases, lipases and proteases [9, 10].
Since extracellular enzymes can break down complex molecules into simpler ones, they are of high commercial value in industrial biotechnology [11, 12], with applications such as food and beverage production, biofuel generation, and waste treatment. Carbohydrases, such as amylases, cellulases and chitinases, hold a significant share in the global industrial enzymes market, which was worth 6.0 billion US$ in 2021 [13]. For example, amylases are used to hydrolyze starch into polymers composed of glucose units, which can then be fermented to produce bioethanol [14]. Cellulases can break down plant biomass such as cellulose into simple sugars, which can be further processed into biofuels [15] Lipases are used in the production of detergents, cosmetics, and pharmaceuticals [16], and proteases are used for protein hydrolysis in food technology [17]. Chitinases have potential particularly in the agro-industry, but also in wastewater treatment, the food industry, cosmetics, and medicine [18]. The unexplored microbial biodiversity found in cenotes offers the potential discovery of novel enzymes with applications in these industries. Bacteria from coastal cenote sediments are influenced by marine-like conditions; they produce halotolerant extracellular enzymes, which are in high demand in the biotechnological industry [19].
Another application of bacteria from this ecological niche is the production of antibiotics. Bacteria from cenotes produce antimicrobial compounds with potential medical applications [3, 4]. Some members of the phyla Actinomycetota and Bacillota have emerged as prominent sources of antimicrobial compounds [20] accounting for > 50% of all antimicrobial activities reported in recent decades, and they merit further exploration in unusual habitats such as cenotes. Actinomycetota produce many substances relevant to biotechnology, agriculture and medicine, including the majority of currently employed antibiotics [21]. Indeed, > 75 antibiotics isolated from Actinomycetota are used in human therapy [22]. Due to the increasing resistance of clinically important microorganisms to therapeutic drugs [23], and the steady decline in the discovery of new antibiotics [24], the need for novel antibiotics rises steadily. Polyketides such as the glycopeptide vancomycin, the lipopeptide daptomycin and the macrolide erythromycin have important antibiotic effects [25], notably against Gram-positive bacteria, and can also have antitumor and immunosuppressive properties [26]. The biosynthesis of polyketides is catalyzed by complex multicomponent enzyme systems, the polyketide synthases (PKS), classed as type I, type II and type III [27]. From these, modular type I PKS complexes offer attractive prospects for producing novel metabolites through genetic engineering and combinatorial biosynthesis [28].
Despite exploration of bacterial communities from the underground river in Yucatán and its cenotes [29,30,31], there is little information on their biotechnological potential. By 2020, only 18 of the > 7000 mapped cenotes [1] had been assessed for their cultivable microbial diversity [3], and microorganisms from only six had been assessed for their biotechnological potential [3]; most studies have focused on inland cenotes, whereas the present study concerns a coastal cenote, Pol-Ac, with the aim of characterizing 49 Gram-positive bacteria isolated from sediment samples. This work provides insights into the diversity, enzymatic activity, and antimicrobial potential of bacteria inhabiting these unexplored habitats.
Materials and Methods
Materials
Chemicals, product standards, and solvents were obtained at the highest purity available from Sigma-Aldrich (St. Louis, US).
Sampling Site
Pol-Ac is an open coastal cenote within a mangrove environment in the El Palmar State Reserve in Yucatán, Mexico (21.0816118, -90.2029322), ~ 780 m east of the Gulf of Mexico (Fig. S1). It is cylindrical, with a cave opening of 4000 m2, and reaches a depth of 63 m at its lowest point. Water temperature, salinity, dissolved oxygen, and pH were determined continuously every 2 s to a depth of 53.5 m with a multiparameter EXO1 water probe (Xylem Analytics, Norway).
Microorganism Isolation
Marine sediment samples were collected in April 2021 under three different depths; a low depth at 14 m which is exposed to high solar radiation, a medium depth at 24 m which is exposed to low solar radiation and a deep depth at 54 m which is exposed to no solar radiation. Three replicates were collected from each sampled point in sterile hermetic seabags. Samples were cooled immediately on ice and stored at 4 °C until further processing. Additionally, 40 L of water from the surface was collected, stored at 4 °C, and later sterilized in an autoclave to be used in preparing the media.
All subsequent steps were performed under sterile conditions. Sediment samples were resuspended 1:10 in sterilized water from Pol-Ac. Thereafter, serial dilutions up to 10‑7 were prepared and streaked onto four different agar plate media. The first employed A1 media (10 g L−1 starch, 4 g L−1 yeast extract, 2 g L−1 peptone, 16 g L−1 agar) [32] with surface water from Pol-Ac (to ensure the exact composition of salts and nutrients), the second one A1 media with marine sea water from the coast of Sisal (A1m agar plates), the third one Brain Heart Infusion agar (8 g L−1 Brain and Heart Infusion, 5 g L−1 peptic digest of animal tissue, 16 g L−1 pancreatic digest of casein, 5 g L−1 sodium chloride, 2 g L−1 glucose, 2.4 g L−1 disodium hydrogen phosphate and 13.5 g L−1 agar) prepared with ddH2O to target more generalist bacteria, and the fourth one only agar (16 g L−1) with surface water from Pol-Ac. All plates were incubated at 27 °C for 1 to 2 weeks. Single colonies, selected on the basis of their morphology, pigmentation and the formation of a halo around the colony [33], were subcultured on A1m agar until axenic cultures were obtained.
Gram-positive strains, determined via the nonstaining KOH method [34], were inoculated in 50 mL of A1m liquid medium and incubated for four days at 27 °C. These liquid cultures were homogenized, mixed 1:1 with 70% glycerol and stored as glycerol stocks at -80 °C until further use. In total, 49 strains were preserved in glycerol stocks.
DNA Extraction and 16 S Sequencing
Genomic DNA extraction used the Quick-DNA Fungal/Bacterial Microprep Kit (Zymo Research, Irvine, US) and followed the manufacturer’s instructions. DNA concentrations were quantified with a Nanodrop One spectrophotometer (Thermo Scientific, Waltham, US), and DNA integrity was assessed on agarose gel (1%). Amplification of the 16 S rDNA gene used the universal primer pair 27 F (5′-AGAGTTTGATCCTGGCTCAG-3′) and 1492R (5′-TACGGYTACCTTGTTACGACTT-3′) [35]. The PCR was performed on a CFX96 Real-Time System thermal cycler (Bio-Rad, Hercules, US) using the following protocol: Initial denaturation for 5 min at 95 °C, followed by 30 cycles of 40 s at 95 °C for denaturation, 50 s at 60 °C for annealing, 40 s at 72 °C for extension, and 10 min at 72 °C for a final extension. The amplified products were visualized by electrophoresis on agarose gel (1%) using the Biorad Molecular Imager Gel Doc XR + System (Bio-Rad, Hercules, US) (Fig. S2). The amplicons were sequenced by the Sanger technique at the Institute of Biotechnology of the National Autonomous University of Mexico (UNAM, Mexico City).
Phylogenetic Analysis
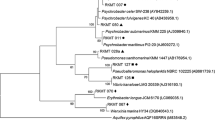
The partial 16 S DNA sequences obtained (503–970 bp) were trimmed with SnapGene v6.2.1 software (GSL Biotech, US) and were compared against the NCBI database via a BLAST search. Based on sequence identity the nearest neighbors and nearest type strains to those neighbors were selected to accurately classify the genera of the strains (Table S2). A phylogenetic tree was generated with the default mode of the online tool phylogeny.fr, which uses MUSCLE (v. 3.8.31) for multiple sequence alignment, Gblocks (v. 0.91b) for alignment refinement, PhyML (v. 3.1/3.0 aLRT) for phylogeny and finally TreeDyn (v. 198.3) for tree drawing [36].
Determination of Thermotolerant Strains
The influence of increasing temperatures on bacterial growth was assessed on strains growing on agar plates. Therefore, test tubes with liquid A1m media were inoculated with glycerol stocks of the strains and grown at 27 °C for five days. 2 µL of these cultures were drop spotted onto one square agar plate containing A1m agar (Fig. S3) in biological duplicates. After incubation for 5 days at 25 °C, 35 °C, 45 °C, 55–65 °C, the plates were examined for bacterial growth.
Marine Water Requirement for Growth
The reliance of the bacterial strains on marine water for growth was investigated in an agar plate assay. Test tubes with liquid A1m media were inoculated with glycerol stocks of the strains and grown at 27 °C for five days. 2 µL of these cultures were drop spotted in biological duplicates onto one square agar plate containing either A1m agar or A1 agar prepared with bidistilled water, ddH2O (A1ddH2O agar plates). After incubation for 5 days at 27 °C, the plates were examined for bacterial growth.
Agar Plate Screening of Extracellular Hydrolytic Enzymes
To evaluate extracellular enzymatic activity of the isolated strains, agar plate assays were conducted with the respective substrates, employing drop spots. The composition of each culture medium for the assays is described below. In contrast to the commonly used formulation with ddH2O, all agar plates were prepared with filtered marine water to support the halophilic nature of the bacterial strains. Media were autoclaved for 20 min at 121 °C and plates stored at 4 °C until further use. Test tubes with liquid A1m media were inoculated with glycerol stocks of the strains and grown at 27 °C for five days. On each agar plate, six different strains were spotted by adding 5 µL of the corresponding culture and incubating at 27 °C for five days, unless stated otherwise. All assays were performed in biological duplicates. The hydrolytic activities of the corresponding strains were expressed as level of enzymatic activity (LEA), as described previously [37, 38], by dividing the diameter of the clearance zone by the diameter of the colony in millimeters. For all agar plate assays, Escherichia coli XL1-blue was the negative control since E. coli is not only a poor secretor of enzymes [39], but also tests negative for extracellular amylase [40], cellulase [41], chitinase [42], lipase [43], gelatinase [44] and protease [45].
Amylase Activity
A1m agar plates were used to determine the level of amylase activity. After incubation for five days, the plates were flooded with Lugol’s iodine solution (Materiales y Abastos Especializados S.A. de C.V., Mexico City, MX) for 1 min. A clear halo around a colony would indicate starch hydrolysis, thereby confirming amylase activity [37].
Cellulase Activity
Cellulase activity was determined on carboxymethylcellulose (CMC)-based agar plates [38] containing CMC (5 g L−1), NaNO3 (2 g L−1), K2HPO4 (1 g L−1), MgSO4 (0.5 g L−1), KCl (0.5 g L−1), peptone (0.2 g L−1) and agar (18 g L−1). After autoclaving, CMC precipitates partially in marine water. Thus, the CMC degradation can be monitored without the need of a staining solution. After incubation for five days, the plates were examined. A clear halo around a colony would indicate CMC hydrolysis, thereby confirming cellulase activity.
Chitinase Activity
Chitinase activity was determined on chitin agar plates [46] with a simplified formula: colloidal chitin (100 g L−1, equal to 5 g dried chitin L−1), yeast extract (0.4 g L−1), peptone (0.2 g L−1) and agar (16 g L−1). Colloidal chitin was prepared according to Hsu and colleagues [47]. The degree of chemical modification of chitin is higher than that of cellulose and starch, so a ten-day incubation period was chosen for examination. A clear halo around a colony would indicate chitin hydrolysis, thereby confirming chitinase activity.
Lipase Activity
Olive oil agar plates [48] were used to determine lipase activity. Plates were prepared with olive oil (10 mL L−1), CaCl2 (1 g L−1), phenol red (0.1 g L−1) and agar (20 g L−1) and the pH was adjusted with NaOH to 7.3–7.4. Following a five-day incubation period, the plates were examined. The principle of the lipase agar plate assay hinges on a pH decrease from 7.3, representing the endpoint of the phenol red dye, to a slightly more acidic pH, inducing a distinctive color transition from red to yellow. This decrease in pH, attributed to the release of fatty acids formed by the lipase-catalyzed breakdown of olive oil, serves as the indicator of lipase activity. Thus, a discernible yellow halo encircling a colony would indicate olive oil hydrolysis, thereby confirming lipase activity.
Gelatinase Activity
Gelatinase activity was assessed on gelatin agar plates [44]: gelatin (12 g L−1), yeast extract (1 g L−1), peptone (4 g L−1), and agar (18 g L−1). After incubation for five days, the plates were treated with a saturated (NH4)2SO4 solution, causing the precipitation of unhydrolyzed gelatin and thereby obscuring the corresponding agar plate areas. Subsequently, the plates were checked after five minutes, with a clear halo around a colony indicating gelatin hydrolysis, thereby confirming gelatinase activity.
Protease Activity
Protease activity was determined on skimmed milk agar plates as prepared by Menasria and collaborators (Menasria et al. (2018): skimmed milk (10 g L−1), yeast extract (1 g L−1) and agar (20 g L−1). After incubation for five days, the plates were examined. A clear halo around a colony would indicate protein hydrolysis, confirming protease activity.
Determination of Polyketide Synthases
To show the presence of polyketide synthase genes encoding for type I PKS enzymes, genomic DNA from each of the 49 cultivated strains was amplified via PCR using the two degenerate oligonucleotide primer pairs 5LL/4UU and KPF/KPR: 5LL (5’-GGRTCNCCIARYTGIGTICCIGTICCRTGIGC-3’) with 4UU (5’-MGIGARGCIYTICARATGGAYCCICARCARMG-3’) [49]; and DKF (5’-GTGCCGGTNCCRTGNGYYTC-3’) with DKR (5’-GCGATGGAYCCNCARCARYG-3’) [50]. Following amplification, presence of products was determined by visualizing in a 1% agarose gel. The anticipated amplicon products for both primer pairs were expected to be approximately 700 base pairs in length.
Antimicrobial Activity Assessment
Extracts of all 49 strains were examined to assess their antimicrobial activity against the Gram-negative bacterium E. coli ATCC 35,218 and the Gram-positive bacterium Staphylococcus aureus (S. aureus) ATCC 6538. Antibiotic activities of all extracts and reference antibiotics (kanamycin and gentamycin) were tested with the broth dilution method based on the Clinical and Laboratory Standards Institute (CLSI) recommendations [51].
For the preparation of the extracts, the strains were reactivated from cryopreserved glycerol stocks on A1m agar plates and a single colony was transferred to a 250 mL Erlenmeyer flask containing 50 mL of A1m broth. Cultures were grown for 7 days, harvested and centrifuged at 4000 rpm. The supernatants were recovered, filtered through syringe filters of 0.8 μm, and lyophilized to generate dry powders, which were stored at 4ºC until further processing. Before performing the antimicrobial evaluation, 100 mg of each lyophilized powder were solubilized in ultrapur water by vigorous vortexing and then desalted by addition of ethanol, precipitating the solid salt. To remove salts, salted-out samples were centrifuged at maximum speed for 2 min. The supernatants were recovered in 2 mL Eppendorf tubes and dried under vacuo (Eppendorf® Concentrator Plus™ for microtubes). The obtained desalted samples were then employed to evaluate the antimicrobial activity. To perform the antibiotic experiments, 10 mg of each desalted sample were resuspended in 1 mL of 70% ethanol for use as a stock solution.
The activity assay used Mueller–Hinton broth medium with incubation at 150 rpm and 37ºC for the duration of the experiment. To perform the test, 5 µL of stock solution of an extract were added to 195 µL of pathogenic bacterial culture (final concentration 250 µg mL−1) at the beginning of the exponential growth phase (108 CFU mL−1). Reference antibiotics were solubilized in water and added at a final concentration of 25 µg mL−1. Optical density at 600 nm was determined at the initial and final times of the experiment. The growth inhibition (%) of the bacterial pathogen strains was assessed by use of negative controls grown with the addition of 5 µL ethanol (70%) instead of bacterial extract stock solution. All the experiments were performed in triplicate and at least three independent experiments were recorded.
Results
Physicochemical Characteristics of Water in Pol-Ac
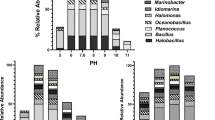
Salinity at the surface of the water column in Pol-Ac was 20.9 psu (Fig. 1a). As depth increased, salinity rose rapidly to 36.0 psu at a depth of 2.5 m, and then continued to increase gradually to 39.5 psu at a depth of 53.5 m. Thus, Pol-Ac can be considered a thalassic environment. pH values decreased with depth (Fig. 1b): 7.59 at the surface and 7.1 within the saltwater layer at 2.5 m. Beyond this depth, a further decrease gradually reached 6.95 at a depth of 53.5 m.
The water temperature is 27.4 °C at the surface, decreases to 26.0 °C at 2.5 m, and gradually increases to 26.8 °C at a depth of 30.0 m, after which it remains constant (Fig. 1c). Dissolved oxygen concentrations ranged from a minimum of 0.03 mg L−1 to a maximum of 0.06 mg L−1 with a mean value of 0.04 ± 0.01 mg L−1, indicative of a hypoxic environment (Fig. 1d).
Microbial Diversity of Pol-Ac
In total, 49 Gram-positive strains were isolated from the sediment of the cenote, whereby 44 strains were isolated from sediment collected at a depth of 14 m, four at a depth of 24 m and one from 54 m. Due to the limited number of strains isolated at medium and deep depths, a significant correlation between the depth of sample collection and strain affiliation, as well as enzymatic or antimicrobial activity was not possible. Further, of the 49 isolated strains, 39 were isolated from A1m agar plates and 10 from A1 agar plates prepared with surface water from Pol-Ac. No Gram-positive bacteria could be isolated from either Brain Heart Infusion agar or plates containing only agar and surface water of Pol-Ac. The BLAST search showed identities between the partial gene sequences and the nearest neighbor that ranged from 98.1 to 100%. Of the 49 strains, 29 represented seven genera of the Bacillota: Bacillus (17 strains), Virgibacillus (4), Halobacillus (3), Metabacillus (2), Solibacillus (1) Neobacillus (1) and Rossellomorea (1). The other 20 strains represented three genera of the Actinomycetota: Streptomyces (18 strains), Nocardiopsis (1) and Corynebacterium (1) (Fig. 2, Fig. S4 and Fig. S5). All sequences were deposited in the NCBI GenBank database with the accession numbers OR844320-OR844368 (Table S2).
Determination of thermo- and Halotolerant Strains
At the lower incubation temperatures (25 °C and 35 °C) all 49 strains grew robustly, in accordance with the measured water temperature range in Pol-Ac (26.0–27.4 °C); at 45 °C, the lower threshold for thermotolerance [52], 45 (91.8%) had substantial growth; at 55 °C, growth of 21 strains (42.9%) was characterized by reduced colony diameters ranging from 0.3 mm to 2 mm; and at 65 °C none grew. Therefore 21 of the strains isolated from Pol-Ac can be considered as moderately thermotolerant and 28 as mesophilic. Of the 21 moderately thermotolerant strains, 18 belonged to the Bacillota and 3 to the Actinomycetota.
All strains grew on the A1m agar plates, with a salt concentration of 3.5%, classifying them as slightly halotolerant [53]. Of the 49 strains, 24 were not capable of growing on A1ddH2O agar plates, indicating that these have a specific requirement for marine water and are likely halophiles [19]; 19 belonged to the Bacillota (six genera) and 5 to the Actinomycetota (to one genus).
Extracellular Hydrolytic Enzymes
Screening of bacterial isolates for extracellular hydrolytic enzymes (Fig. 3) showed that each of the 49 strains could produce at least one of these enzymes (Fig. 4); 47 could produce at least two extracellular enzymes; 41 at least three; 33 at least four; 15 at least five; and 2 could produce all six extracellular enzymes. Gelatinase activity was the most prevalent, seen in 46 of the isolates; amylase activity in 33; lipase activity in 31; cellulase and protease activity each in 29 of the strains; and chitinase activity in only 19.
Representative agar plate assays for the detection of extracellular hydrolytic enzymes. All plates used an uninoculated empty control (1), E. coli XL1-blue as a negative control (2), and Streptomyces sp. NCA_378 (3), which exhibited activity for all six assays. Colonies 4–9 represent strains with very high activities. a Amylase assay with Solibacillus sp. NCA_420 (4); b Cellulase assay with Halobacillus sp. NCA_370 (5); c Chitinase assay with Bacillus sp. NCA_375 (6); d Lipase assay with Bacillus sp. NCA_353 (7); e Gelatinase assay with Bacillus sp. NCA_358 (8); f Protease assay with Bacillus sp. NCA_419 (9)
Activity profile of the 49 bacteria isolated from the cenote Pol-Ac. From left to right: T: thermotolerance (light green to dark green); M: Marine water dependent for growth (grey), LEA: level of enzyme activity (light blue to dark blue); PKS: assessment of type I PKS genes (purple); AA: antimicrobial activity (light yellow to brown); taxonomic designation and internal strain number (NCA)
Since marine water was used for all six agar plate assays, all detected enzymes and their corresponding activities are halotolerant. Actinomycetota, on average, showed a higher cumulative enzyme activity rate (70.0%) than that observed in Bacillota (59.2%). In addition to being the most frequently isolated genus (n = 18), all Streptomyces strains exhibited enzymatic activity in the forms of amylase, lipase, and gelatinase; 11 of the Streptomyces strains showed chitinase activity; 10 showed cellulase activity; and 4 showed protease activity. From Pol-Ac, Streptomyces NCA_378 showed activities of all six extracellular enzymes: high for amylase and gelatinase; medium for lipase; and low for cellulase, chitinase and protease. Hence, this strain merits further study.
In contrast, Bacillus, the second most commonly isolated genus (n = 17), displayed a more varied distribution of enzymatic capabilities: 16 of the Bacillus strains exhibited gelatinase activity; 15 showed protease activity; 11 showed cellulose; 7 amylase; 6 chitinase; and 4 lipase. From Pol-Ac, Bacillus NCA_374 displayed high biotechnological potential across all six extracellular enzymes, with high cellulase activity, medium gelatinase activity, and low activity levels for amylase, chitinase, lipase, and protease. In addition, strain Halobacillus NCA_393 showed high activity of cellulase, gelatinase, and protease activities, and Bacillus NCA_375 showed high activity in cellulase, chitinase, and gelatinase. Hence, these two Bacillus strains, too, merit further biotechnological study.
Detection of Type I PKS Genes
In the current study, putative type I polyketide synthase (PKS) genes were identified in 24 of the isolated strains; 22 of these showed a PCR product with the KPF/KPR primer pair, and 10 with the 5LL/4UU primer pair. Of the 20 Actinomycetota strains, 14 (70.0%) showed a PCR product with the PKS primers, whereas of the 29 Bacillota strains only 10 (34.5%) showed a product.
Determination of Antimicrobial Activity
Four strains (8.2%) slightly inhibited the growth of E. coli ATCC 35,218 (25–50% inhibition in comparison to negative control) (Fig. 4). All four strains that slightly inhibited E. coli ATCC 35,218 were also active against S. aureus ATCC 6538. In contrast to the low number of strains inhibiting E. coli, 18 (36.7%) inhibited growth of S. aureus ATCC 6538, 10 (20.4%) of these showing medium inhibition (50–75%). Two strains, Streptomyces NCA_366 and Streptomyces NCA_387, achieved high levels of inhibition (75–100%; Fig. 4) and merit further study. Higher percentages of Actinomycetota (50%) than Bacillota (27.6%) showed inhibition against one of the two tested pathogens. These findings are promising and suggest that bacteria isolated from cenotes such as Pol-Ac have the potential to produce antimicrobial compounds with activity against Gram-positive bacteria such as S. aureus and should therefore be further investigated for secondary metabolites.
Discussion
This study focused on the bioprospection of cultivable Gram-positive bacteria isolated from sediment samples of the cenote Pol-Ac. In contrast to other coastal cenotes, Pol-Ac lacks a freshwater layer on its surface due to a surface salinity of 20.9 psu and a halocline at a shallow depth of only 0.4 m. For instance, the coastal cenotes Tábano [54] and Na’ach Wennen Ha [55], both within 1 km of the Caribbean Sea, have surface salinities of 2 psu and 0.5 psu, and much deeper haloclines at 11 m and 5 m, respectively. With an average water temperature of 26.4 °C, Pol-Ac follows the trend of other coastal cenotes like Tabano and Odyssey [54] during this time of the year [5, 56].
However, the dissolved oxygen concentrations along the whole water column were remarkably low, with a maximum of only 0.06 mg L−1. A study involving 30 coastal cenotes of the Yucatán Peninsula reported minimum oxygen concentrations to be more than 10 times higher than Pol-Acs [5]. Thus, the low oxygen levels correlated with the high salinity levels and with the eutrophication conditions [57]. The scarcity of macroscopic organisms in this cenote can be attributed to these low oxygen concentrations, as has been shown for hypoxic costal ecosystems [58]. These parameters suggest that Pol-Ac may host a unique microbial community that has adapted to these extreme conditions.
Previous studies of bacterial diversity in cenotes have reported a majority of Gram-negative isolates (80–90%) [4, 29]. However, among Gram-positive bacteria, the two most abundant phyla found by Escobar-Zepeda [29] were Bacillota and Actinomycetota, in the water and plant roots of a coastal cenote, similar than in our findings from Pol-Ac sediments. Indeed, Bacillota and Actinomycetota, found ubiquitously in soil and water ecosystems, actively contribute to the maintenance of ecological balance by efficiently degrading both organic and inorganic matter in their surroundings [59,60,61]. Further, the presence of Virgibacillus and Halobacillus, frequently found in saline environments [62, 63], is consistent with the marine-like nature of the sampling site.
The unexplored microbial biodiversity found in cenotes offers the potential for the discovery of novel enzymes that could have various applications in industries. Thus, one of the aims of this study was the exploration of extracellular hydrolytic enzymes secreted by the 49 isolated Gram-positive bacteria. Indeed, all the isolated strains were capable to produce at least one out of six of the screened enzymes. The present findings align with previous studies that reported the production of extracellular hydrolytic enzymes, including amylase, cellulase, chitinase, protease, gelatinase, and lipase, by marine coastal Streptomyces [38, 64] and marine Bacillus strains [65,66,67], which collectively accounted for 71.4% of the isolated strains in this study.
Another aim of the current work was the detection of PKS I genes of the isolated strains as well as the determination of their antimicrobial activity against the two model pathogen strains E. coli ATCC 35,218 and S. aureus ATCC 6538. PKS I amplicons were more abundant in the Actinomycetota group (70.0%) in comparison to the Bacillota one (34.5%). These results agree with reports of Actinomycetota, particularly Streptomyces species, as important producers of secondary metabolites, including pharmaceutically relevant polyketides [26, 68]. Nevertheless, PKS genes and their associated metabolites have also been found in Bacillota, such as species of Bacillus [69]. Further, 8.2% of the strains inhibited the growth of the pathogen E. coli ATCC 35,218 and 36.7% inhibited the growth of the Gram-positive pathogen S. aureus ATCC 6538. Low numbers of strains with activity against E. coli are expected, given the inherent resistance of Gram-negative bacteria to many antibiotics due to the presence of an outer membrane that restricts the penetration of many molecules into the cell [70]. Considering the remarkable capacity of Actinomycetota, especially Streptomyces, to produce abundant antimicrobial compounds, it is unsurprising to observe a higher percentage of antimicrobial activity in Actinomycetota (50.0%) compared to Bacillota (27.6%) [71,72,73,74]. Of the 49 strains, 24 showed a PCR product with the PKS I primers used for amplification, and 18 strains showed antimicrobial activity. However, only 11 strains showed both PKS I gene product amplification and antimicrobial activity. There was no significant correlation (chi-square 1.676; p = 0.196) between type I PKS gene product amplification and antimicrobial activity. This suggests that factors other than type I PKS genes, such as type II or type III PKS, non-ribosomal peptide synthetase genes [75] or other secondary-metabolite biosynthetic pathways, may contribute to the observed inhibition of these pathogens. Moreover, strains displaying amplicons for type I PKS genes, yet lacking antimicrobial activity against the two tested pathogens, may potentially yield polyketides that are active against pathogens not examined in the course of this study.
The present study emphasizes the potential of the microbial communities in cenote sediments of the Yucatán Peninsula to be a novel source of biotechnologically important bacteria. It demonstrates that the characteristics of the coastal cenote Pol-Ac form an environment that is ideal for further studies regarding the cultivation of halotolerant bacteria with hydrolytic enzymes of industrial value and potent antimicrobial compounds against known bacterial pathogens. Additionally, this study is the first report on the bioprospection of Gram-positive bacteria isolated from sediments in a cenote of the Yucatán Peninsula.
References
Bauer-Gottwein P, Gondwe BRN, Charvet G et al (2011) Review: the Yucatán Peninsula Karst aquifer, Mexico. Hydrogeol J 19:507–524. https://doi.org/10.1007/s10040-010-0699-5
Lopez-Maldonado Y, Berkes F (2017) Restoring the environment, revitalizing the culture: Cenote conservation in Yucatan, Mexico. Ecol Soc 22. https://doi.org/10.5751/ES-09648-220407
Moreno-Pérez PA, Hernández‐Téllez M, Ramirez‐Durán N et al (2020) Microorganisms and spatial distribution of the sinkholes of the Yucatan Peninsula, underestimated biotechnological potential? Water Environ J 34:41–49. https://doi.org/10.1111/wej.12502
De La Rosa-García SC, Muñoz-García AA, Barahona-Pérez LF, Gamboa-Angulo MM (2007) Antimicrobial properties of moderately halotolerant bacteria from cenotes of the Yucatan Peninsula. Lett Appl Microbiol 45:289–294. https://doi.org/10.1111/j.1472-765X.2007.02185.x
Schmitter-Soto JJ, Comín FA, Escobar-Briones E et al (2002) Hydrogeochemical and biological characteristics of cenotes in the Yucatan Peninsula (SE Mexico). In: Alcocer J, Sarma SSS (eds) Advances in Mexican limnology: Basic and Applied aspects. Developments in Hydrobiology. Springer Netherlands, Dordrecht, NL, pp 215–228
Sahl JW, Gary MO, Harris JK, Spear JR (2011) A comparative molecular analysis of water-filled limestone sinkholes in north-eastern Mexico. Environ Microbiol 13:226–240. https://doi.org/10.1111/j.1462-2920.2010.02324.x
Brankovits D, Pohlman JW, Ganju NK et al (2018) Hydrologic Controls of Methane Dynamics in Karst Subterranean estuaries. Glob Biogeochem Cycles 32:1759–1775. https://doi.org/10.1029/2018GB006026
Ritter SM, Isenbeck-Schröter M, Scholz C et al (2019) Subaqueous speleothems (Hells Bells) formed by the interplay of pelagic redoxcline biogeochemistry and specific hydraulic conditions in the El Zapote sinkhole, Yucatán Peninsula, Mexico. Biogeosciences 16:2285–2305. https://doi.org/10.5194/bg-16-2285-2019
Dumorné K, Severe R (2018) Marine enzymes and their industrial and biotechnological applications. Minerva Biotecnol 30:113–119. https://doi.org/10.23736/S1120-4826.18.02442-4
Cheng TH, Ismail N, Kamaruding N et al (2020) Industrial enzymes-producing marine bacteria from marine resources. Biotechnol Rep 27:e00482. https://doi.org/10.1016/j.btre.2020.e00482
Ferrari E, Jarnagin AS, Schmidt BF (2014) Commercial production of extracellular enzymes. In: Sonenshein AL, Hoch JA, Losick R (eds) Bacillus subtilis and other Gram-positive Bacteria. ASM, Washington, DC, US, pp 917–937
Kumar M, Kumar P, Das P et al (2020) Potential applications of extracellular enzymes from Streptomyces spp. in various industries. Arch Microbiol 202:1597–1615. https://doi.org/10.1007/s00203-020-01898-9
BlueWeave Consulting and Research Pvt. &, Ltd (2021) Global industrial enzymes market projected to reach worth $9.2 bn by 2027. Focus Catal 2021:2. https://doi.org/10.1016/j.focat.2021.11.004
de Souza PM, de Magalhães P O (2010) Application of microbial α-amylase in industry - A review. Braz J Microbiol 41:850–861. https://doi.org/10.1590/S1517-83822010000400004
Ejaz U, Sohail M, Ghanemi A (2021) Cellulases: from bioactivity to a variety of industrial applications. Biomimetics 6:1–11. https://doi.org/10.3390/biomimetics6030044
Chandra P, Enespa, Singh R, Arora PK (2020) Microbial lipases and their industrial applications: a comprehensive review. Microb Cell Fact 19:169. https://doi.org/10.1186/s12934-020-01428-8
Tavano OL, Berenguer-Murcia A, Secundo F, Fernandez-Lafuente R (2018) Biotechnological Applications of proteases in Food Technology. Compr Rev Food Sci Food Saf 17:412–436. https://doi.org/10.1111/1541-4337.12326
Poria V, Rana A, Kumari A et al (2021) Current perspectives on chitinolytic enzymes and their agro-industrial applications. Biology (Basel) 10:1319. https://doi.org/10.3390/biology10121319
Sekar A, Kim K (2020) Halophilic Bacteria in the Food Industry. In: Kim S-K (ed) Encyclopedia of Marine Biotechnology. Wiley, Hoboken, New Jersey, US, pp 2061–2070
Srinivasan R, Kannappan A, Shi C, Lin X (2021) Marine bacterial secondary metabolites: a treasure house for structurally unique and effective antimicrobial compounds. Mar Drugs 19:530. https://doi.org/10.3390/md19100530
n der Meij A, Worsley SF, Hutchings MI, van Wezel GP (2017) Chemical ecology of antibiotic production by actinomycetes. FEMS Microbiol Rev 41:392–416. https://doi.org/10.1093/femsre/fux005
Bérdy J (2005) Bioactive microbial metabolites. J Antibiot 58:1–26. https://doi.org/10.1038/ja.2005.1
Plackett B (2020) No money for new drugs. Nature 586:S50–S52. https://doi.org/10.1038/d41586-020-02884-3
Aminov RI (2010) A brief history of the antibiotic era: lessons learned and challenges for the future. Front Microbiol 1:134. https://doi.org/10.3389/fmicb.2010.00134
Baltz RH (2006) Molecular engineering approaches to peptide, polyketide and other antibiotics. Nat Biotechnol 24:1533–1540. https://doi.org/10.1038/nbt1265
Risdian C, Mozef T, Wink J (2019) Biosynthesis of polyketides in Streptomyces. Microorganisms 7:124. https://doi.org/10.3390/microorganisms7050124
Moffitt MC, Neilan BA (2003) Evolutionary affiliations within the superfamily of ketosynthases reflect complex pathway associations. J Mol Evol 56:446–457. https://doi.org/10.1007/s00239-002-2415-0
Kittendorf JD, Sherman DH (2006) Developing tools for engineering hybrid polyketide synthetic pathways. Curr Opin Biotechnol 17:597–605. https://doi.org/10.1016/j.copbio.2006.09.005
Escobar-Zepeda A, Rosas-Escobar P, Marquez Valdelamar L et al (2021) Distinctive prokaryotic microbiomes in sympatric plant roots from a Yucatan cenote. BMC Res Notes 14:1–6. https://doi.org/10.1186/s13104-021-05746-x
Maldonado Desena F, De la Cruz Ceferino N, Gómez Cornelio S et al (2022) Bacteria Halotolerant from Karst sinkholes as a source of Biosurfactants and Bioemulsifiers. Microorganisms 10:1–18. https://doi.org/10.3390/microorganisms10071264
Suárez-Moo P, Remes-Rodríguez CA, Márquez-Velázquez NA et al (2022) Changes in the sediment microbial community structure of coastal and inland sinkholes of a karst ecosystem from the Yucatan Peninsula. Sci Rep 12:1–11. https://doi.org/10.1038/s41598-022-05135-9
Marfil-Santana MD, Martínez-Cárdenas A, Ruíz-Hernández A et al (2021) A Meta-omics analysis unveils the Shift in Microbial Community structures and metabolomics profiles in Mangrove Sediments treated with a selective actinobacterial isolation Procedure. Molecules 26:7332. https://doi.org/10.3390/molecules26237332
Goodfellow M (2012) Phylum XXVI. Actinobacteria phyl. Nov. In: Goodfellow M, Kämpfer P, Busse H-J et al (eds) Bergey’s Manual of systematic bacteriology, 2nd edn. Springer New York, New York, US, pp 33–2028
Buck JD (1982) Nonstaining (KOH) method for determination of gram reactions of marine bacteria. Appl Environ Microbiol 44:992–993. https://doi.org/10.1128/aem.44.4.992-993.1982
Heuer H, Krsek M, Baker P et al (1997) Analysis of actinomycete communities by specific amplification of genes encoding 16S rRNA and gel-electrophoretic separation in denaturing gradients. Appl Environ Microbiol 63:3233–3241. https://doi.org/10.1128/aem.63.8.3233-3241.1997
Dereeper A, Guignon V, Blanc G et al (2008) Phylogeny.fr: robust phylogenetic analysis for the non-specialist. Nucleic Acids Res 36:W465-9. https://doi.org/10.1093/nar/gkn180
Menasria T, Aguilera M, Hocine H et al (2018) Diversity and bioprospecting of extremely halophilic archaea isolated from Algerian arid and semi-arid wetland ecosystems for halophilic-active hydrolytic enzymes. Microbiol Res 207:289–298. https://doi.org/10.1016/j.micres.2017.12.011
González V, Vargas-Straube MJ, Beys-da-Silva WO et al (2020) Enzyme Bioprospection of Marine-Derived Actinobacteria from the Chilean Coast and New Insight in the mechanism of keratin degradation in Streptomyces sp. G11C. Mar Drugs 18:1–26. https://doi.org/10.3390/md18110537
Ni Y, Chen R (2009) Extracellular recombinant protein production from Escherichia coli. Biotechnol Lett 31:1661–1670. https://doi.org/10.1007/s10529-009-0077-3
Rosales-Colunga LM, Martínez-Antonio A (2014) Engineering Escherichia coli K12 MG1655 to use starch. Microb Cell Fact 13:1–8. https://doi.org/10.1186/1475-2859-13-74
Gao D, Luan Y, Wang Q et al (2015) Construction of cellulose-utilizing Escherichia coli based on a secretable cellulase. Microb Cell Fact 14:1–8. https://doi.org/10.1186/s12934-015-0349-7
Francetic̃ O, Badaut C, Rimsky S, Pugsley AP (2000) The ChiA (YheB) protein of Escherichia coli K-12 is an endochitinase whose gene is negatively controlled by the nucleoid-structuring protein H-NS. Mol Microbiol 35:1506–1517. https://doi.org/10.1046/j.1365-2958.2000.01817.x
Lee LP, Karbul HM, Citartan M et al (2015) Lipase-Secreting Bacillus Species in an Oil-Contaminated Habitat: Promising Strains to Alleviate Oil Pollution. Biomed Res Int 2015:820575. https://doi.org/10.1155/2015/820575
dela Cruz TEE, Torres JMO (2012) Gelatin Hydrolysis Test Protocol. ASM 1–10
Helianti I, Furgeva N, Mulyawati L et al (2018) Cloning of a Gene Encoding Protease from Bacillus halodurans CM1 into Escherichia coli DH5α and expression analyses of the gene product. Makara J Sci 22. https://doi.org/10.7454/mss.v22i3.9900
Kawase T, Saito A, Sato T et al (2004) Distribution and phylogenetic analysis of Family 19 chitinases in Actinobacteria. Appl Environ Microbiol 70:1135–1144. https://doi.org/10.1128/AEM.70.2.1135-1144.2004
Hsu SC, Lockwood JL (1975) Powdered chitin Agar as a Selective Medium for Enumeration of Actinomycetes in Water and Soil. Appl Microbiol 29:422–426. https://doi.org/10.1128/am.29.3.422-426.1975
Ramnath L, Sithole B, Govinden R (2017) Identification of lipolytic enzymes isolated from bacteria indigenous to Eucalyptus wood species for application in the pulping industry. Biotechnol Rep 15:114–124. https://doi.org/10.1016/j.btre.2017.07.004
Parsley LC, Linneman J, Goode AM et al (2011) Polyketide synthase pathways identified from a metagenomic library are derived from soil Acidobacteria. FEMS Microbiol Ecol 78:176–187. https://doi.org/10.1111/j.1574-6941.2011.01122.x
Moffitt MC, Neilan BA (2001) On the presence of peptide synthetase and polyketide synthase genes in the cyanobacterial genus Nodularia. FEMS Microbiol Lett 196:207–214. https://doi.org/10.1111/j.1574-6968.2001.tb10566.x
Weinstein MP, Patel JB, Campeau S et al (2018) CLSI. Performance standards for Antimicrobial susceptibility testing, 28th edn. Clinical and Laboratory Standards Institute, Wayne, Pennsylvania, US
Yamini C, Sharmila G, Muthukumaran C et al (2022) Proteomic perspectives on thermotolerant microbes: an updated review. Mol Biol Rep 49:629–646. https://doi.org/10.1007/s11033-021-06805-z
Larsen H (1986) Halophilic and halotolerant microorganisms-an overview and historical perspective. FEMS Microbiol Lett 39:3–7. https://doi.org/10.1111/j.1574-6968.1986.tb01835.x
Benítez S, Iliffe TM, Quiroz-Martínez B, Alvarez F (2019) How is the anchialine fauna distributed within a cave? A study of the ox Bel ha System, Yucatan Peninsula, Mexico. Subterr Biol 31:15–28. https://doi.org/10.3897/subtbiol.31.34347
Brankovits D, Pohlman JW, Lapham LL (2022) Oxygenation of a karst subterranean estuary during a tropical cyclone: mechanisms and implications for the carbon cycle. Limnol Oceanogr 67:2691–2705. https://doi.org/10.1002/lno.12231
Pérez-Ceballos R, Canul-Macario C, Pacheco-Castro R et al (2021) Regional Hydrogeochemical Evolution of Groundwater in the Ring of cenotes, Yucatán (Mexico): an inverse Modelling Approach. Water (Switz) 13:614. https://doi.org/10.3390/w13050614
Herrera-Silveira JA, Cirerol NA, Ghinaglia LT et al (2005) Eutrofización costera en la Península de Yucatán. In: Caso M, Pisanty I, Ezcurra E (eds) Diagnóstico ambiental del Golfo de México. Mexico City, MX, pp 823–850
Altieri AH, Diaz RJ (2018) Dead zones: oxygen depletion in coastal ecosystems. In: Sheppard C (ed) World seas: an environmental evaluation, 2nd edn. Academic, Cambridge, Massachusetts, US, pp 453–473
Haldar S, Nazareth SW (2018) Taxonomic diversity of bacteria from mangrove sediments of Goa: metagenomic and functional analysis. 3 Biotech 8:. https://doi.org/10.1007/s13205-018-1441-6
Jagannathan SV, Manemann EM, Rowe SE et al (2021) Marine actinomycetes, new sources of biotechnological products. Mar Drugs 19. https://doi.org/10.3390/md19070365
Juottonen H, Eiler A, Biasi C et al (2017) Distinct anaerobic bacterial consumers of cellobiose-derived carbon in boreal fens with different CO2/CH4 production ratios. Appl Environ Microbiol 83:e02533-16. https://doi.org/10.1128/AEM.02533-16
Heyrman J, Vos P, De, Logan N (2015) Virgibacillus. In: Trujillo ME, Dedysh S, DeVos P et al (eds) Bergey’s Manual of Systematics of Archaea and Bacteria. Wiley, Hoboken, New Jersey, US, pp 1–15
Spring S (2015) Halobacillus. In: Trujillo ME, Dedysh S, DeVos P et al (eds) Bergey’s Manual of Systematics of Archaea and Bacteria. Wiley, Hoboken, New Jersey, US, pp 1–10
Han Y, Yang B, Zhang F et al (2009) Characterization of antifungal chitinase from marine Streptomyces sp. DA11 associated with South China sea sponge Craniella australiensis. Mar Biotechnol 11:132–140. https://doi.org/10.1007/s10126-008-9126-5
Farha AK, TR T, Purushothaman A et al (2018) Phylogenetic diversity and biotechnological potentials of marine bacteria from continental slope of eastern Arabian Sea. J Genet Eng Biotechnol 16:253–258. https://doi.org/10.1016/j.jgeb.2018.06.002
Lu J, Zhao Y, Hu R et al (2022) Screening and characteristics of Marine Bacillus velezensis Z-1 protease and its application of enzymatic hydrolysis of mussels to prepare antioxidant active substances. Molecules 27:6570. https://doi.org/10.3390/molecules27196570
Akeed Y, Atrash F, Naffaa W (2020) Partial purification and characterization of chitinase produced by Bacillus licheniformis B307. Heliyon 6. https://doi.org/10.1016/j.heliyon.2020.e03858
Li S, Li Z, Pang S et al (2021) Coordinating precursor supply for pharmaceutical polyketide production in Streptomyces. Curr Opin Biotechnol 69:26–34. https://doi.org/10.1016/j.copbio.2020.11.006
Chen XH, Vater J, Piel J et al (2006) Structural and functional characterization of three polyketide synthase gene clusters in Bacillus amyloliquefaciens FZB 42. J Bacteriol 188:4024–4036. https://doi.org/10.1128/JB.00052-06
Nikaido H (2003) Molecular basis of bacterial outer membrane permeability revisited. Microbiol Mol Biol Rev 67:593–656. https://doi.org/10.1128/mmbr.67.4.593-656.2003
Subramani R, Aalbersberg W (2012) Marine actinomycetes: an ongoing source of novel bioactive metabolites. Microbiol Res 167:571–580. https://doi.org/10.1016/j.micres.2012.06.005
Genilloud O (2017) Actinomycetes: still a source of novel antibiotics. Nat Prod Rep 34:1203–1232. https://doi.org/10.1039/c7np00026j
Passari AK, Leo VV, Chandra P et al (2018) Bioprospection of actinobacteria derived from freshwater sediments for their potential to produce antimicrobial compounds. Microb Cell Fact 17:1-4. https://doi.org/10.1186/s12934-018-0912-0
Mast Y, Stegmann E (2019) Actinomycetes: the antibiotics producers. Antibiotics 8:10–13. https://doi.org/10.3390/antibiotics8030105
Singh M, Chaudhary S, Sareen D (2017) Non-ribosomal peptide synthetases: identifying the cryptic gene clusters and decoding the natural product. J Biosci 42:175–187. https://doi.org/10.1007/s12038-017-9663-z
Acknowledgements
We would like to thank CONAHCyT, FQ, UNAM-PAIP, and FCT-Fundação para a Ciência e a Tecnologia for their financial support. J.L.W. and J.C.P.-F. are grateful for financial aid provided by UNAM-DGAPA in the form of a postdoctoral fellowship. We are also grateful to the Brady Laboratory at the Rockefeller University N.Y. for their support with funding for cave sampling, and to Efraín Chávez Solís, Luis A. Liévano Beltrán and Kay Vilchis Zapata for their invaluable help reaching the remote depths and lengths of the anchialine cave system in the Yucatán. We would like to thank Sebastien Santini (CNRS/AMU IGS UMR7256) and the PACA Bioinfo platform for the availability and management of the phylogeny.fr website used to generate alignments and the phylogenetic trees.
Funding
Funding for this research was provided by multiple sources. J.L.W. and J.C.P.-F. were both supported in the form of postdoctoral fellowships provided by UNAM-DGAPA. W.E.-H. was supported by the UNAM-PAPIIT IA203722. S.P.G. was supported by national funds from FCT-Fundação para a Ciência e a Tecnologia, IP, in the scope of the project UIDP/04378/2020 of the Research Unit on Applied Molecular Biosciences–UCIBIO and the project LA/P/0140/2020 of the Associate Laboratory Institute for Health and Bioeconomy. A.P.-D. received support from the Consejo Nacional de Humanidades Ciencias y Tecnologías (CONAHCyT) through the Ciencia Básica grant number A1-S-10785, as well as a sabbatical grant. Additionally, A.P.-D. received support from the School of Chemistry, UNAM PAIP program grant 5000-9149.
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic Supplementary Material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Wissner, J.L., Parada-Fabián, J.C., Márquez-Velázquez, N.A. et al. Diversity and Bioprospection of Gram-positive Bacteria Derived from a Mayan Sinkhole. Microb Ecol 87, 77 (2024). https://doi.org/10.1007/s00248-024-02392-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00248-024-02392-1