Abstract
Heavy metal (HM) accumulation in soil affects plants and soil fauna, yet the effect on microbial alpha-diversity remains unclear, mainly due to the absence of dedicated research synthesis (e.g. meta-analysis). Here, we report the first meta-analysis of the response of soil microbial alpha-diversity to the experimental addition of cadmium (Cd) and copper (Cu). We considered studies conducted between 2013 and 2022 using DNA metabarcoding of bacterial and fungal communities to overcome limitations of other cultivation- and electrophoresis-based techniques. Fungi were discarded due to the limited study number (i.e. 6 studies). Bacterial studies resulted in 66 independent experiments reported in 32 primary papers from four continents. We found a negative dose-dependent response for Cu but not for Cd for bacterial alpha-diversity in the environments, only for Cu additions exceeding 29.6 mg kg−1 (first loss of − 0.06% at 30 mg kg−1). The maximal loss of bacterial alpha-diversity registered was 13.89% at 3837 mg kg−1. Our results first highlight that bacterial communities behave differently to soil pollution depending on the metal. Secondly, our study suggests that even extreme doses of Cu do not cause a dramatic loss in alpha-diversity, highlighting how the behaviour of bacterial communities diverges from soil macro-organisms.
Graphical abstract

Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
The biodiversity measurement is based on three fundamental components: alpha- (local or within-sample), beta- (among-samples), and gamma- (biome-scale) diversity [1,2,3,4,5]. Alpha-diversity considers the number of species and/or their abundance distribution, and it is commonly used to characterize species community structure [6,7,8,9]. Alpha-diversity variability is usually associated with environmental perturbation and is widely used in biomonitoring to assess environmental health [10,11,12,13]. Specifically, ecosystems influenced by anthropic activities are generally characterized by lower levels of alpha-diversity due to habitat alteration and loss [14, 15]. Intensive agriculture and urbanization, pesticides, heavy metals (HMs) and other pollutants are among the major drivers of worldwide biodiversity decline [16,17,18]. Of the different biomes, soil and sediments are the final repository of these potentially toxic compounds, including HM, which are severely accumulated, posing a potential threat to fauna, plants and humans [19,20,21]. HM can be transferred to plants and animals throughout ecosystems in different ways, such as food web contamination (e.g. soil–plant-animal) or direct ingestion by the soil mesofauna [21,22,23]. When ingested by soil animals, HM can bioaccumulate in the body causing body size alteration and population size declines (e.g. earthworms) [24, 25]. Generally, the biological organization of soil invertebrates at the community level is also negatively impacted by HM [26, 27]. For example, community structure and species richness alterations strongly related to soil HM content have been observed in Oribatida and Mesostigmata mites [28]. The community structure of nematodes is also strongly altered by HM pollution, and the diversity of major nematode trophic groups decreases along a pollution gradient [29, 30].
While the role of HM in declining soil fauna species richness is generally clear [28, 31], their effects on soil bacterial alpha-diversity diverge [32, 33]. One gram of soil is typically inhabited by more than 4 × 103 bacterial species [34,35,36,37,38], and this considerable diversity plays a pivotal role in ecosystem services involved in, to name a few, the biogeochemical cycling of elements, soil formation and interaction with macro-organisms (i.e. plants, soil fauna) [39,40,41,42,43]. Furthermore, freshwater and marine sediments are considered to be hotspots of bacterial diversity [44,45,46]; the top 50 cm of marine sediments can contain ~ 1 × 1029 bacterial species [47,48,49,50].
Previous investigations of polluted soils and sediments suggested that bacterial and fungal communities can exhibit differences in their response to HM [51,52,53,54,55]. This is partly based on general distinctions in biochemical pathways activated by bacteria and fungi in response to HM [56,57,58,59]. In addition, there are cases where experimental conditions could inherently contribute to the estimation of HM ecotoxicity, limiting the possibility of extending conclusions to a broader ecological scale [59]. Indeed, recent investigations have reported negative, neutral and positive estimated responses of the richness of bacterial taxa to HM exposure [60,61,62,63]. There are various, debated, hypotheses to explain these contrasting results [59]. Many studies are not comparable due to differences in soil and experimental conditions, background pollution levels and other experimental covariates that influence the response of microorganisms to HM [64]. Furthermore, some studies consider only a small number of replicates, limiting robust ecological conclusions [65,66,67], or only specific soil ecosystems (i.e. vineyards, orchards, arable lands or grasslands) [68].
After an accurate selection of studies and the rejection of fungal entries due to insufficient existing research from 2013 to 2022, we carried out a large-scale meta-analysis investigating bacterial alpha-diversity responses to HM (Cd and Cu) contamination in soils and sediments along a gradient ranging from 0.3 to 3837 mg kg−1. The study aims to clarify (a) the relationship between the dose of HM addition and the response of microbial alpha-diversity, (b) if microorganisms respond differently in different environments (i.e. soil, rhizosphere, and sediments) and (c) if the chemical properties of the specific environment (pH and Organic Matter (OM) content) influence the direction and magnitude of the alpha-diversity response to HM.
Methods
Data Selection and Collection
We conducted a meta-analysis collecting data from primary papers reporting experimental results on the response of bacterial diversity to lab-scaled and controlled Cu and Cd additions in soils and sediments. A preliminary search was conducted to understand the quantity of studies for each HM. The literature on HM is highly heterogeneous, focusing not only on different environments (e.g. soil, sediment, freshwater and marine environments) but also on different approaches, from descriptive biomonitoring experiments to pollution experiments and those focusing on remediation of historically polluted sites.
Based on our search, we found that the only metals eligible for contrast-based meta-analysis (i.e. experiments using replicated treatments and control samples) were Cd and Cu (e.g. most studies of arsenic focused on single-species toxicological experiments rather than an extensive and exhaustive bacterial community analysis). At first, we focused on Cu and Cd due to the environmental issues related to their intensive usage (e.g. mining, agriculture, industrial wastewaters) [69,70,71,72]. Moreover, from the preliminary search for other HM eligible for being included in the meta-analysis, we found that only Cu and Cd have been sufficiently studied in compliance with the selection criteria we adopted. First, we searched Scopus and Web of Science databases (last access in January 2022). The search strings contained Boolean statements, e.g. ‘OR’ and ‘AND’, combining key search terms such as ‘bacterial diversity’ and ‘copper’ or ‘cadmium’ and other additional terms (Supplementary Table S1). The search was restricted to peer-reviewed articles published between 2013 and 2021, as primary papers published after 2013 were more likely to adopt metabarcoding and next-generation sequencing (NGS) techniques for bacterial ecology analysis. This approach resulted in a selection process starting from an initial 10,687 unique entries for which the title and abstract were screened to remove non-relevant studies. Of this initial number, after this first screening, 374 relevant primary papers were identified.
We further evaluated the eligibility of these primary papers using the following criteria: (1) the study was based on the addition of Cu or Cd and replicated treated samples were compared to replicated control samples; (2) the bacterial communities were inferred through amplicon sequencing of the 16S rRNA gene (bacteria) and ITS region (fungi) as taxonomic barcodes since comprehensiveness and homogeneity of databases are also required for reliable comparison; (3) both gradual or abrupt HM addition was considered, but the primary papers had a metal-free control to calculate contrast-based effect size (i.e. the log-response ratio) [73]. After applying the selection criteria, 54 primary papers were eligible for the meta-analysis that reported direct relationships between HM addition and variation in soil bacterial diversity indices. From these, only those with proper alpha-diversity data were retained (some papers did not report sufficient information to calculate diversity indices).
The remaining 32 primary papers selected for our meta-analysis were mainly conducted in the People’s Republic of China (25) and a small number in Europe (2), North America (1) and Australia (2) (Fig. 1; PRISMA workflow shown in Fig. S1).
We extracted metal doses and alpha-diversity data directly from manuscripts, both from tables and figures. We further extracted additional information regarding the soil environment, the type of metal, the bacterial communities investigated, the environmental conditions possibly modifying the outcome of addition (soil pH and OM), the sample size and the time after treatment waited for the collection of samples.
Sample sizes in the selected papers were variable, ranging from 2 to 8 while most adopted an experimental design conducting metabarcoding analyses on 3–4 replicates per sample. HM additions spanned from 0.3 to 3837 mg kg−1. The maximal doses reached by Cd and Cu were respectively 1770 to 3837 mg kg−1. The response of bacterial communities to these addition ranges was measured over time, with sample collection occurring from 2 h to over 5 years after addition.
Calculation of the Effect Size
The effect of HM addition on bacterial diversity metrics was calculated, for each case study, as the natural logarithm-transformed (ln) response ratio (Eq. 1) [73]:
where \(\overline{X }\) t and \(\overline{X }\) c are the average bacterial diversity values for treatment and control (metal-free) samples, respectively. Accordingly, we calculated multiple response ratios in cases where an experiment reported multiple treatments sharing the same control. Since this violates the assumption of independence among effect sizes, namely when multiple treated samples are compared to a single control group [74], the variance–covariance (VCV) matrix was computed to account for correlation of effect sizes, as proposed by [75]. In case the experiment was biomonitoring bacterial diversity over time, the metal-free control at the time zero was taken. The inverse of the sampling variance for each effect size in the VCV matrix was used to weight the meta-analysis. Sampling variance was calculated as follows, based on Hedges et al. [73] (Eq. 2):
where SD is the standard deviation of the mean X (see Eq. 1), N is the sample size (i.e. the number of soil samples from which DNA was extracted, amplified and sequenced). When sample size was not reported, we imputed it from the median across all the primary papers included in the meta-analysis. The median of the sample size imputed this way was equal to 3 for both the treatment and control, for a total of 41 out of 502 observations that were missing the sample size. Similarly, in cases where the standard deviation of the mean was not reported, these were imputed by considering the coefficient of variation of the mean from all complete observations, following Hedges et al. [73].
Data Analysis
Data were first split according to the microbial domain investigated, and then split according to the metal. Data were then analysed using a multilevel meta-analytic linear mixed-effect model using the rma.mv function within the metafor R package [76]. We first fitted a null model to estimate the weighted mean pooled effect size, namely the overall amount of alpha bacterial diversity changes across all experiments, independently from the dose of metal added.
Seven moderators were considered as influencing factors affecting bacterial community response in a meta-regression analysis: (a) diversity metrics considered in the primary papers (i.e. Shannon–Wiener, Simpson, observed OTU, Chao and ACE), (b) the ecosystem type where the experiment was conducted (sediment, soil or rhizosphere; categorical), (c) taxonomic kingdom (bacteria or fungi; categorical), (d) chemical form of the metal added (either in salt or nanoparticle form; categorical), (e) type of HM considered (either Cu or Cd; categorical), (f) soil pH and (g) soil OM content (% w/w of soil; continuous). Where no information on soil pH and OM content was retrievable from the primary papers (only 7 papers in which at least one of the two parameters was not available), geographic locations of the studies were used to access the SoilGrids database (at 250 m resolution) and extract estimates of pH (H2O) and OM data from 5 to 15 cm depth [77, 78].
Models were fitted with a crossed and nested random effect structure, as follows: (1|study/experiment/id), (time | experiment). The first component structure considers the possibility that the individual estimates within each observation (id) could be nested within the experiment and study grouping level [79]. The second component takes into account that the experiments could be nested within the time of sampling, considering an autoregressive correlation structure of true effects over time (this was achieved by setting struct = ‘CAR’ in the rma.mv function), since the results could have been collected at multiple time points within the same experiment [80, 81].
After computing the null model, we then fitted single meta‐regression models using the addition and the type of metal as the sole moderators, to compare changes among the metrics for a given amount of metal added to the matrices considered (i.e. rhizosphere, soil and sediment). Then, we fitted multiple meta‐regression models including other moderators and relevant interactions between the dose added and any other moderator since, in all the studies included, the dose was the sole discriminant factor between control and treated samples. We performed stepwise backward selection based on the Bayesian information criterion (BIC), selecting the model displaying the lowest BIC, excluding a moderator only if it was also dropped from the interaction term. Potential overparameterization of the model was checked by plotting the profile of the (restricted) log‐likelihood over the variance and the correlation components of the models [76].
Finally, publication bias was tested through visual examination of funnel plots and performing the Egger’s test for asymmetry by fitting null model residuals with the precision of the observations (1/SE) [82, 83]. Results of null model and publication bias are reported in Fig. S2 and Supplementary Table S2.
All analyses were performed in the R environment (version 4.2; R Core Team, 2022).
Results
The meta-analysis was conducted on observations obtained by investigating bacterial communities since for fungi, the body of research that complied with our selection criteria was not sufficiently large to be included for quantitative analysis (see “Lack of Eligible Studies on Fungal Alpha-Diversity and HM”; Supplementary Table S2). Specifically, we ensured the robustness of obtained results by discarding all papers that did not focus on controlled additions of HM or did not rely on high-throughput sequencing to measure the response of soil microbial communities. This brought 32 final studies.
We found that Cu addition caused a 9.81% mean pooled bacterial diversity loss (− 6.69 to − 12.82% with 95% CI) compared to untreated samples, conversely to Cd, that did not provoke any change in bacterial alpha-diversity (Fig. 2). The other experimental covariates did not influence the response of soil bacterial alpha-diversity to metal addition, except for the dose of Cu and pH (Final equation: LnRR ~ dose + pH) (Supplementary Table S4). For Cd, the model selection did not find any model explaining the effect size distribution better than the null model (Fig. 2, Supplementary Table S4). The multilinear mixed-effect meta-regression model finally obtained (i.e. the lowest BIC model obtained) accounted for a Cu-dependent dose–effect influencing the variation of bacterial alpha-diversity effect sizes across the studies (Fig. 3). The final model estimated a minimum inhibitory concentration of 29.6 mg kg−1 to register a significant bacterial alpha-diversity decrease (Fig. 3). From that minimum concentration upwards, the model fit indicated a linear decrease in alpha-diversity. For instance, at 100 mg kg−1, bacterial communities lost 5%, at 1000 mg kg−1 lost 11.1% and at 3837 mg kg−1, the bacterial communities investigated lost 13.89% of alpha-diversity (Fig. 3). The final model obtained for Cu contained pH as moderator of the effect size distribution as influencing increases in effect sizes moving from acidic to alkaline environmental conditions (lnRR = − 0.0628 + 0.073 × pH).
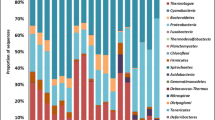
Distribution of the effect sizes of microbial alpha-diversity variation. Violet and green bars indicate the distribution of significant LnRRs (p‐value < 0.05) after respectively Cd and Cu addition. Blue vertical lines indicate the mean pooled estimate (solid line) with its 95% CI (shade) from a null mixed-effect model. Effect sizes reported consider alpha-diversity indices jointly
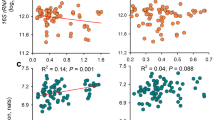
a Scatterplot showing dose–response relationship between alpha-diversity change (lnRR) and metal dose (on a log scale; back-transformed to mg kg−1 on the x axis, back-transformed to relative change (%) on the y axis). Cd on the left, Cu on the right. The horizontal black line indicates bacterial alpha-diversity LnRR effect size equal to zero. The meta-analysis revealed a dose–effect on bacterial alpha-diversity only for Cu. The vertical black line indicates the minimum alpha-diversity inhibitory dose corresponding to no overlap with zero by the 95% CI of the slope estimate (29.6 mg kg−1). b Scatterplot showing relationship between alpha-diversity change (lnRR) and the environment pH (on a log scale; back-transformed on the x axis, back-transformed to relative change (%) on the y axis). Cd on the left, Cu on the right. Effect sizes reported consider alpha-diversity indices jointly
Among the alpha-diversity metrics, we found no difference in their relative dose–response, except for richness that differed from the others (Simpson, Shannon–Wiener, Chao1 and ACE) (p = 0.0084) (Fig. 3). However, the same result was not recorded at the end of the BIC selection process; thus, this did not influence the dose–response.
Discussion
Bacterial Communities’ Metal-Dependent Response
Heavy metals, including Cu and Cd, are antimicrobial agents, and their activity has been tested on different bacterial species [84, 85]. Nevertheless, addressing the differential toxicity effect of Cu and Cd on bacteria is not straightforward. This is mainly due to the different direct and side effects that the two metals can cause to bacterial species [86]. Direct evidence pinpoints Cu as more detrimental for bacteria than Cd [84, 87]. For instance, iron-sulphur clusters of dehydratases (e.g., fumarase) are known as the main target of Cu in bacteria (Macomber and Imlay, 2009). On the other hand, for Cd, evidence showed that the toxicity of iron-sulphur clusters is less pronounced than that of Cu [88, 89]. Nevertheless, bacteria live in highly diverse and complex communities [90] in soils and sediments [91, 92]; thus, it is crucial to address HM toxicity at the community level rather than the effect on the single species. Our meta-analysis found that the response of soils and sediments bacterial alpha-diversity to HM is metal-dependent. Cu, differently from Cd, was correlated with a decrease in bacterial alpha-diversity (Fig. 2). While Cd-related response exploits mainly detoxification mechanisms (e.g., efflux, chelation etc.), Cu also causes significant oxidative stress to bacterial cells, forcing them to manage reactive oxygen species (ROS) and uncontrolled redox reactions in addition to metal detoxification [93, 94]. However, the bacteria alpha-diversity response to Cu was moderate (Fig. 3). Indeed, the dose must exceed 29.6 mg kg−1 to register an initial significant decrease in bacterial alpha-diversity (Fig. 3). The maximum loss of alpha-diversity (− 13.89%) was registered at super-excessive HM additions peaking to 3837 mg kg−1 (Fig. 3). The dose we registered are far beyond the levels of HM loads registered worldwide [95,96,97,98]. European topsoils display on average 60–100 mg kg−1 of Cu and 0.5–1 mg kg−1 Cd, similar to soils in other continents [72, 95, 99,100,101]. Exceptional concentrations coherent with our results can be found in specific intensive agricultural areas and mining sites where these values reach approximately thousands of mg kg−1 [102,103,104,105,106].
The negative Cu-related response was generally moderate compared to the extreme doses added to soils and sediments, which contrasts with the response of macro-organism biodiversity to HM. Soil macro-organisms (e.g. plants, nematodes and wild bees) are severely affected by HM at doses even lower than those reported in our work. For instance, nematode alpha-diversity decreased from 0.7 to 10 mg kg−1 of Cd and 42.7 to 1000 mg kg−1 of Cu [30], with a loss of more than 13.89% of total biodiversity (the maximum decrease we registered). Likewise, the diversity of wild bee species is known to be severely reduced in locations with soils polluted with HM (20–350 mg Cd kg−1 soil; [107]). Nevertheless, microorganisms live in complex communities and have already been demonstrated more resistant than single species subjected to HM in toxicological studies (i.e. higher minimum inhibitory concentrations) [108]. For microorganisms, the abrupt exposure to HM may not necessarily severely compromise alpha-diversity levels, though beta-diversity may rapidly change [109,110,111]. In fact, the use of alpha-diversity measures has been debated [112], highlighting that alpha-diversity is meaningless without additional bacterial analysis investigating both metabolic and ecological responses (i.e. biomass, respiration, enzymatic activities, beta-diversity analysis, bacterial functional ecology and inter-kingdom relationships through ecological network analysis).
Although alpha-diversity represents a specific component of biodiversity, it can be measured using different indices. In our work, both qualitative measures based on presence/absence (e.g. Chao 1, ACE, observed OTUs number) [113, 114] and quantitative measures based on taxon relative abundance (e.g. Shannon and Simpson) [115, 116] were recorded across the studies we considered. The synergic variation of diversity indices (i.e. Shannon–Wiener and Simpson) (Fig. 4) is explained by their mathematical relatedness in estimating diversity in bacterial communities. Contrarily, the discordancy between the Chao estimator and bacterial richness has different potential explanations. Indeed, there is evidence that the Chao estimator could fail in precisely estimating the correct level of richness of bacterial communities, being rather an accurate estimate of the lower bound of bacterial richness [117, 118]. Further, the decrease in richness may indicate a specialization of bacterial communities towards living in a metal-rich environment through metal-resistant species selection.
Distribution of the effect sizes according to the alpha-diversity metric considered. Cd-related effect sizes on the left, Cu-related ones on the right. The horizontal dashed line indicates bacterial alpha-diversity LnRR effect size equal to zero. No significant difference was detected across different diversity indices. Red square points indicate mean pooled effect (and 95% CI) obtained from mixed‐effect models using diversity metric type as predictor
Environment Conditions Might Influence the Bacterial Alpha-Diversity Response to HM
The toxicity of HM to soil organisms is partially dependent on the chemical composition of the surrounding matrix (e.g. pH, OM), which modulates HM bioavailability [19, 119]. By considering studies of different soil environments contaminated with HM in our analysis, we aimed to collect the largest possible soils and sediment matrix physical–chemical variability (i.e. pH 4–9, OM 1–8%). pH is one of the most important drivers of bacterial diversity [10, 120] and a crucial moderator of element availability in soil [121, 122]. Most complexed Cu is adsorbed to negative binding sites of dissolved organic matter. In acidic conditions, the protonation of these sites might cause a mobilization of adsorbed Cu and release to the soil solution [123, 124] where it can exert strong pressure over bacterial biomass [125, 126]. Indeed, pH has been already described as more important than total Cu content for its effect on bacterial communities [127]. Our study demonstrates that the response of bacterial alpha-diversity to Cu toxicity is not only dependent on the dose of Cu applied, but also on the pH of the soil/sediment (Fig. 2). The decrease in alpha-diversity of the bacterial communities subjected to Cu contamination is enhanced moving from alkaline to sub-acid, acid environmental conditions.
The recorded response of bacterial communities to HM did not vary across the soil environments investigated (i.e. bulk soil, rhizosphere, marine and freshwater sediments) (Supplementary Table S4). For instance, the response of bacterial community alpha-diversity to HM addition was not altered by the close proximity of plants. Although plants can modulate the local availability of HM in the rhizosphere, for example due to root exudation [128,129,130], their effect was likely masked by the extreme loads of Cu and Cd oversaturating the soil. Indeed, the modulation of HM availability by soluble organic ligands has been shown at low concentrations [131, 132].
Lack of Eligible Studies on Fungal Alpha-Diversity and HM
Although, the crucial ecological significance of fungal communities (e.g. interaction with plants, construction of soil food-web, and carbon sequestration) [133,134,135,136], the number of works and consequently the available datasets regarding the effect of HM on their diversity is considerably lower than those for bacteria. Thus, we could not conduct a large-scale meta-analysis on fungi due to the limited number (n 6) of eligible works (Supplementary Table S2). Some authors have already postulated that bacterial and fungal communities could display differences in their response to HM [55]. There is evidence that fungal communities should be more resistant to HM [137], probably due to the different genetics and physiology characterizing the two kingdoms [55]. In addition, some authors hypothesized that, at least in short-term experiments of HM addition, fungi could benefit from carbon released by bacterial death cells, thus balancing the adverse effect of HM [55].
Conclusions
We show that the response of bacterial alpha-diversity to Cd is absent. On the other hand, bacterial alpha-diversity decreases up to 14%, although extreme doses of Cu have been applied to soils and sediments (up to 3837 mg kg−1). We also demonstrate that bacterial communities respond differently to Cu addition in relation to soil and sediment pH, showing decreases in alpha-diversity in acidic conditions. In contrast to bacteria, we remark that more research should be conducted to explore the response of fungal communities to HM pollution in soils and sediments.
Our findings shed new light on the role of HM on microbial communities, unravelling a general overview of the response of microbial alpha-diversity to HM.
Data and Code Availability
The dataset and R code are freely available at Figshare data repository (https://figshare.com) following this link: https://doi.org/10.6084/m9.figshare.19188917.v2.
References
Dantas de Miranda M, Pereira HM, Corley MFV, Merckx T (2019) Beta diversity patterns reveal positive effects of farmland abandonment on moth communities. Sci Rep 9(1):1–9. https://doi.org/10.1038/s41598-018-38200-3
Grilli J (2020) Macroecological laws describe variation and diversity in microbial communities. Nat Commun 11(1):1–11. https://doi.org/10.1038/s41467-020-18529-y
Jost L (2007) Partitioning diversity into independent alpha beta concepts. Ecology 88(10):2427–2439. https://doi.org/10.1890/06-1736.1
Prober SM et al (2015) Plant diversity predicts beta but not alpha diversity of soil microbes across grasslands worldwide. Ecol Lett 18(1):85–95. https://doi.org/10.1111/ele.12381
Walters KE, Martiny JBH (2020) Alpha-, beta-, and gamma-diversity of bacteria varies across habitats. PLoS One 15(9 September):1–17. https://doi.org/10.1371/journal.pone.0233872
Edward O (1992) Wilson, The diversity of life. Belknap Press of Harvard University Press, Massachusetts
Hagerty SL, Hutchison KE, Lowry CA, Bryan AD (2020) An empirically derived method for measuring human gut microbiome alpha diversity: demonstrated utility in predicting healthrelated outcomes among a human clinical sample. PLoS One 15(3):1–21. https://doi.org/10.1371/journal.pone.0229204
T. H. M. P. Consortium (2012) Structure, function and diversity of the healthy human microbiome. Nature 486(7402):207–214. https://doi.org/10.1038/nature11234
Whittaker RH (1972) Evolution and measurement of species diversity. Taxon 21(2/3):213–251. https://doi.org/10.2307/1218190
Fierer N, Jackson RB (2006) The diversity and biogeography of soil bacterial communities. Proc Natl Acad Sci USA 103(3):626–631. https://doi.org/10.1073/pnas.0507535103
Kaarlejärvi E, Salemaa M, Tonteri T, Merilä P, Laine AL (2021) Temporal biodiversity change following disturbance varies along an environmental gradient. Glob Ecol Biogeogr 30(2):476–489. https://doi.org/10.1111/geb.13233
Kondratyeva A et al (2020) Urbanization effects on biodiversity revealed by a two-scale analysis of species functional uniqueness vs. redundancy. Front Ecol Evol 8(March):1–16. https://doi.org/10.3389/fevo.2020.00073
Willig MR, Presley SJ (2017) Biodiversity and disturbance, vol. 1–5. Elsevier Inc. https://doi.org/10.1016/B978-0-12-809665-9.09813-X
Johnson CN et al (2017) Biodiversity losses and conservation responses in the Anthropocene. Science (80) 356(6335):270–275. https://doi.org/10.1126/science.aam9317
Seddon N et al (2016) Biodiversity in the anthropocene: prospects and policy. Proc R Soc B Biol Sci 283(1844):1–9. https://doi.org/10.1098/rspb.2016.2094
Krinner G et al (2013) Long-term climate change: projections, commitments and irreversibility. Clim Chang 2013 Phys Sci Basis Work Gr I Contrib Fifth Assess Rep Intergov Panel Clim Chang 9781107057:1029–1136. https://doi.org/10.1017/CBO9781107415324.024
Monchanin C, Devaud JM, Barron AB, Lihoreau M (2021) Current permissible levels of metal pollutants harm terrestrial invertebrates. Sci Total Environ 779:146398. https://doi.org/10.1016/j.scitotenv.2021.146398
Sánchez-Bayo F, Wyckhuys KAG (2019) Worldwide decline of the entomofauna: a review of its drivers. Biol Conserv 232(September 2018):8–27. https://doi.org/10.1016/j.biocon.2019.01.020
Alloway BJ (2013). Heavy Met Soils. https://doi.org/10.1007/978-94-007-4470-7
Genova G et al (2021) Copper and zinc as a window to past agricultural land-use. J Hazard Mater (March): 126631. https://doi.org/10.1016/j.jhazmat.2021.126631
Tovar-Sánchez E, Hernández-Plata I, Martínez MS, Valencia-Cuevas L, Galante PM (2018) Heavy metal pollution as a biodiversity threat. Heavy Met. https://doi.org/10.5772/intechopen.74052
Hu C, Shui B, Yang X, Wang L, Dong J, Zhang X (2021) Trophic transfer of heavy metals through aquatic food web in a seagrass ecosystem of Swan Lagoon, China. Sci Total Environ 762:143139. https://doi.org/10.1016/j.scitotenv.2020.143139
Michelutti N et al (2010) Trophic position influences the efficacy of seabirds as metal biovectors. Proc Natl Acad Sci USA 107(23):10543–10548. https://doi.org/10.1073/pnas.1001333107
Heikens A, Peijnenburg WJGM, Hendriks AJ (2001) Bioaccumulation of heavy metals in terrestrial invertebrates. Environ Pollut 113(3):385–393. https://doi.org/10.1016/S0269-7491(00)00179-2
Moyson S, Town RM, Vissenberg K, Blust R (2018) The effect of metal mixture composition on toxicity to C. elegans at individual and population levels. PLoS One 14(6):1–23. https://doi.org/10.1371/journal.pone.0218929
Nahmani J, Lavelle P (2002) Effects of heavy metal pollution on soil macrofauna in a grassland of Northern France. Eur J Soil Biol 38(3–4):297–300. https://doi.org/10.1016/S1164-5563(02)01169-X
Spurgeon DJ, Hopkin SP (1999) Seasonal variation in the abundance, biomass and biodiversity of earthworms in soils contaminated with metal emissions from a primary smelting works. J Appl Ecol 36(1):173–183. https://doi.org/10.1046/j.1365-2664.1999.00389.x
Manu M, Honciuc V, Neagoe A, Băncilă RI, Iordache V, Onete M (2019) Soil mite communities (Acari: Mesostigmata, Oribatida) as bioindicators for environmental conditions from polluted soils. Sci Rep 9(1):1–13. https://doi.org/10.1038/s41598-019-56700-8
Gutiérrez C et al (2016) Effect of soil properties, heavy metals and emerging contaminants in the soil nematodes diversity. Environ Pollut 213:184–194. https://doi.org/10.1016/j.envpol.2016.02.012
Chauvin C et al (2020) Soil nematodes as indicators of heavy metal pollution: a meta-analysis. Open J Soil Sci 10(12):579–601. https://doi.org/10.4236/ojss.2020.1012028
Pouyat RV et al (2015) Multi-scale assessment of metal contamination in residential soil and soil fauna: a case study in the Baltimore-Washington metropolitan region, USA. Landsc Urban Plan 142:7–17. https://doi.org/10.1016/j.landurbplan.2015.05.001
Dance A (2020) The search for microbial dark matter. Nature 582(7811):301–303. https://doi.org/10.1038/d41586-020-01684-z
Thaler DS (2021) Is global microbial biodiversity increasing, decreasing, or staying the same? Front Ecol Evol 9(April). https://doi.org/10.3389/fevo.2021.565649
FAO (2020) State of knowledge of soil biodiversity - status, challenges and potentialities. https://doi.org/10.4060/cb1928en
Raynaud X, Nunan N (2014) Spatial ecology of bacteria at the microscale in soil. PLoS One 9(1). https://doi.org/10.1371/journal.pone.0087217.
Roesch LFW et al (2007) Pyrosequencing enumerates and contrasts soil microbial diversity. ISME J 1(4):283–290. https://doi.org/10.1038/ismej.2007.53
Torsvik V, Goksoyr J, Daae FL (1990) High diversity in DNA of soil bacteria. Appl Environ Microbiol 56(3):782–787. https://doi.org/10.1128/aem.56.3.782-787.1990
Trevors JT (2010) One gram of soil: a microbial biochemical gene library, Antonie van Leeuwenhoek. Int J Gen Mol Microbiol 97(2):99–106. https://doi.org/10.1007/s10482-009-9397-5
Almela P, Velázquez D, Rico E, Justel A, Quesada A (2019) Carbon pathways through the food web of a microbial mat from byers peninsula, antarctica. Front Microbiol 10(MAR):1–11. https://doi.org/10.3389/fmicb.2019.00628
Antunes PM, Koyama A (2017) Mycorrhizas as nutrient and energy pumps of soil food webs: multitrophic interactions and feedbacks, no. November. Elsevier Inc. https://doi.org/10.1016/B978-0-12-804312-7.00009-7
Heijboer A, de Ruiter PC, Bodelier PLE, Kowalchuk GA (2018) Modulation of litter decomposition by the soil microbial food web under influence of land use change. Front Microbiol 9(NOV):1–11. https://doi.org/10.3389/fmicb.2018.02860
Holtkamp R, Kardol P, van der Wal A, Dekker SC, van der Putten WH, de Ruiter PC (2008) Soil food web structure during ecosystem development after land abandonment. Appl Soil Ecol 39(1):23–34. https://doi.org/10.1016/j.apsoil.2007.11.002
Potapov AM et al (2021) Size compartmentalization of energy channeling in terrestrial belowground food webs. Ecology 102(8):1–14. https://doi.org/10.1002/ecy.3421
Borruso L, Zerbe S, Brusetti L (2015) Bacterial community structures as a diagnostic tool for watershed quality assessment. Res Microbiol 166(1):38–44. https://doi.org/10.1016/j.resmic.2014.11.004
Hoshino T et al (2020) Global diversity of microbial communities in marine sediment. Proc Natl Acad Sci USA 117(44):27587–27597. https://doi.org/10.1073/pnas.1919139117
Varliero G, Bienhold C, Schmid F, Boetius A, Molari M (2019) Microbial diversity and connectivity in deep-sea sediments of the South Atlantic Polar Front. Front Microbiol 10(APR):1–18. https://doi.org/10.3389/fmicb.2019.00665
Cavicchioli R et al (2019) Scientists’ warning to humanity: microorganisms and climate change. Nat Rev Microbiol 17(9):569–586. https://doi.org/10.1038/s41579-019-0222-5
Danovaro R, Corinaldesi C, Rastelli E, Dell’Anno A (2015) Towards a better quantitative assessment of the relevance of deep-sea viruses, bacteria and archaea in the functioning of the ocean seafloor. Aquat Microb Ecol 75(1):81–90. https://doi.org/10.3354/ame01747
Flemming HC, Wuertz S (2019) Bacteria and archaea on Earth and their abundance in biofilms. Nat Rev Microbiol 17(4):247–260. https://doi.org/10.1038/s41579-019-0158-9
Wang Y et al (2012) Comparison of the levels of bacterial diversity in freshwater, intertidal wetland, and marine sediments by using millions of illumina tags. Appl Environ Microbiol 78(23):8264–8271. https://doi.org/10.1128/AEM.01821-12
Haferburg G, Kothe E (2007) Microbes and metals: interactions in the environment. J Basic Microbiol 47(6):453–467. https://doi.org/10.1002/jobm.200700275
Gadd GM, Griffiths AJ (1978) Microorganisms and heavy metal toxicity. Microb Ecol 4(4):303–317. https://doi.org/10.1007/BF02013274
Yi J, Lo LSH, Liu H, Qian P-Y, Cheng J (2021) Study of heavy metals and microbial communities in contaminated sediments along an urban estuary. Front Mar Sci 8(November):1–17. https://doi.org/10.3389/fmars.2021.741912
Frossard A, Hartmann M, Frey B (2017) Tolerance of the forest soil microbiome to increasing mercury concentrations. Soil Biol Biochem 105:162–176. https://doi.org/10.1016/j.soilbio.2016.11.016
Rajapaksha RMCP, Tobor-Kapłon MA, Bååth E (2004) Metal toxicity affects fungal and bacterial activities in soil differently. Appl Environ Microbiol 70(5):2966–2973. https://doi.org/10.1128/AEM.70.5.2966-2973.2004
Gerwien F, Skrahina V, Kasper L, Hube B, Brunke S (2018) Metals in fungal virulence. FEMS Microbiol Rev 42(1):1–21. https://doi.org/10.1093/femsre/fux050
Solioz M (2019) Copper disposition in bacteria. Elsevier Inc. https://doi.org/10.1016/b978-0-12-810532-0.00011-2
Vest KE, Zhu X, Cobine PA (2019) Copper disposition in yeast, in Clinical and Translational Perspectives on WILSON DISEASE, Elsevier Inc, pp. 115–126. https://doi.org/10.1016/b978-0-12-810532-0.00012-4
Hao X et al (2021) Recent advances in exploring the heavy metal(loid) resistant microbiome. Comput Struct Biotechnol J 19:94–109. https://doi.org/10.1016/j.csbj.2020.12.006
Xie Y et al (2016) Effect of heavy metals pollution on soil microbial diversity and bermudagrass genetic variation. Front Plant Sci 7(MAY2016):1–12. https://doi.org/10.3389/fpls.2016.00755
Chen Y et al (2018) Long-term and high-concentration heavy-metal contamination strongly influences the microbiome and functional genes in Yellow River sediments. Sci Total Environ 637–638:1400–1412. https://doi.org/10.1016/j.scitotenv.2018.05.109
Ding Z, Wu J, You A, Huang B, Cao C (2017) Effects of heavy metals on soil microbial community structure and diversity in the rice (Oryza sativa L. subsp. Japonica, Food Crops Institute of Jiangsu Academy of Agricultural Sciences) rhizosphere. Soil Sci Plant Nutr 63(1):75–83. https://doi.org/10.1080/00380768.2016.1247385
Tipayno SC et al (2018) The bacterial community structure and functional profile in the heavy metal contaminated paddy soils, surrounding a nonferrous smelter in South Korea. Ecol Evol 8(12):6157–6168. https://doi.org/10.1002/ece3.4170
Neaman A, Selles I, Martínez CE, Dovletyarova EA (2020) Analyzing soil metal toxicity: spiked or field-contaminated soils? Environ Toxicol Chem 39(3):513–514. https://doi.org/10.1002/etc.4654
Keiblinger KM et al (2018) Assessment of Cu applications in two contrasting soils—effects on soil microbial activity and the fungal community structure. Ecotoxicology 27(2):217–233. https://doi.org/10.1007/s10646-017-1888-y
Liu Y et al (2021) Application of low dosage of copper oxide and zinc oxide nanoparticles boosts bacterial and fungal communities in soil. Sci Total Environ 757:143807. https://doi.org/10.1016/j.scitotenv.2020.143807
Naveed M et al (2014) Simultaneous loss of soil biodiversity and functions along a copper contamination gradient: when soil goes to sleep. Soil Sci Soc Am J 78(4):1239–1250. https://doi.org/10.2136/sssaj2014.02.0052
Karimi B et al (2021) Ecotoxicity of copper input and accumulation for soil biodiversity in vineyards. Environ Chem Lett 0123456789. https://doi.org/10.1007/s10311-020-01155-x
Briffa J, Sinagra E, Blundell R (2020) Heavy metal pollution in the environment and their toxicological effects on humans. Heliyon 6(9):e04691. https://doi.org/10.1016/j.heliyon.2020.e04691
Cesco S et al (2021) A smart and sustainable future for viticulture is rooted in soil: How to face cu toxicity. Appl Sci 11(3):1–21. https://doi.org/10.3390/app11030907
Daehn KE, Cabrera Serrenho A, Allwood JM (2017) How will copper contamination constrain future global steel recycling? Environ Sci Technol 51(11):6599–6606. https://doi.org/10.1021/acs.est.7b00997
Lamichhane JR, Osdaghi E, Behlau F, Köhl J, Jones JB, Aubertot JN (2018) Thirteen decades of antimicrobial copper compounds applied in agriculture. A review. Agron Sustain Dev 38(3). https://doi.org/10.1007/s13593-018-0503-9
Hedges LV, Gurevitch J, Curtis PS (1999) The meta-analysis of response ratios in experimental ecology. Ecology 80(4):1150. https://doi.org/10.2307/177062
Gleser L, Olkin I (2009) Stochastically dependent effect sizes. In Cooper H, Hedges LV, Valentine JC (eds) The handbook of research synthesis and meta-analysis. Russell Sage Foundation, pp 357–376
Lajeunesse MJ (2011) On the meta-analysis of response ratios for studies with correlated and multi-group designs. Ecology 92(11):2049–2055. https://doi.org/10.1890/11-0423.1
Viechtbauer W (2010) Conducting meta-analyses in R with the metafor Package. J Stat Softw 36(3). https://doi.org/10.18637/jss.v036.i03
Hengl T et al (2017) SoilGrids250m: global gridded soil information based on machine learning. 12(2). https://doi.org/10.1371/journal.pone.0169748
Guerra CA et al (2020) Blind spots in global soil biodiversity and ecosystem function research. Nat Commun 11(1):1–13. https://doi.org/10.1038/s41467-020-17688-2
Konstantopoulos S (2011) Fixed effects and variance components estimation in three-level meta-analysis. Res Synth Methods 2(1):61–76. https://doi.org/10.1002/jrsm.35
Ishak KJ, Platt RW, Joseph L, Hanley JA, Caro JJ (2007) Meta-analysis of longitudinal studies. Clin Trials 4(5):525–539. https://doi.org/10.1177/1740774507083567
Trikalinos TA, Olkin I (2012) Meta-analysis of effect sizes reported at multiple time points: a multivariate approach. Clin Trials 9(5):610–620. https://doi.org/10.1177/1740774512453218
Jennions MD, Møller AP (2002) Relationships fade with time: a meta-analysis of temporal trends in publication in ecology and evolution. Proc R Soc B Biol Sci 269(1486):43–48. https://doi.org/10.1098/rspb.2001.1832
Nakagawa S, Santos ESA (2012) Methodological issues and advances in biological meta-analysis. Evol Ecol 26(5):1253–1274. https://doi.org/10.1007/s10682-012-9555-5
Luo J, Hein C, Mücklich F, Solioz M (2017) Killing of bacteria by copper, cadmium, and silver surfaces reveals relevant physicochemical parameters. Biointerphases 12(2):020301. https://doi.org/10.1116/1.4980127
Moyson S, Vissenberg K, Fransen E, Blust R, Husson SJ (2018) Mixture effects of copper, cadmium, and zinc on mortality and behavior of Caenorhabditis elegans. Environ Toxicol Chem 37(1):145–159. https://doi.org/10.1002/etc.3937
Giachino A, Waldron KJ (2020) Copper tolerance in bacteria requires the activation of multiple accessory pathways. Mol Microbiol 114(3):377–390. https://doi.org/10.1111/mmi.14522
Chen J, Zhang H, Li J, Liu Y, Shi W, Hu H (2020) The toxic factor of copper should be adjusted during the ecological risk assessment for soil bacterial community. Ecol Indic 111(October 2019):106072. https://doi.org/10.1016/j.ecolind.2020.106072
Fang F, Imlay JA (2012) Silver ( I ), mercury ( II ), cadmium ( II ), and zinc ( II ) target exposed enzymic iron-sulfur clusters when they toxify Escherichia coli. Appl Environ Microbiol 78(10):3614–3621. https://doi.org/10.1128/AEM.07368-11
Macomber L, Imlay JA (2009) The iron-sulfur clusters of dehydratases are primary intracellular targets of copper toxicity. Proc Natl Acad Sci USA 106(20):8344–8349. https://doi.org/10.1073/pnas.0812808106
Stubbendieck RM, Vargas-bautista C, Straight PD, Romero DF, Beauregard P (2016) Bacterial communities : interactions to scale. 7(August): 1–19. https://doi.org/10.3389/fmicb.2016.01234
Lozupone CA, Knight R (2007) Global patterns in bacterial diversity. 104(27). https://doi.org/10.1073/pnas.0611525104
Hoshino T, Doi H, Uramoto G, Wörmer L, Adhikari RR, Xiao N (2020) Global diversity of microbial communities in marine sediment. 117(44): 27587–27597. https://doi.org/10.1073/pnas.1919139117
Salah I, Parkin IP, Allan E (2021) Copper as an antimicrobial agent: recent advances. RSC Adv 11(30):18179–18186. https://doi.org/10.1039/d1ra02149d
Morey JR, Begg SL, Eijkelkamp BA, Luo Z, Couñago RM, Omara ML, Maher MJ, Ong C-LY, McEwan AG, Kobe B, Paton JC, McDevitt CA (2014) Disregulation of transition metal ion homeostasis is the molecular basis for cadmium toxicity in Streptococcus pneumoniae. Nat Commun. https://doi.org/10.1038/ncomms7418
Ballabio C et al (2018) Copper distribution in European topsoils: an assessment based on LUCAS soil survey. Sci Total Environ 636:282–298. https://doi.org/10.1016/j.scitotenv.2018.04.268
Jiang HH, Cai LM, Wen HH, Luo J (2020) Characterizing pollution and source identification of heavy metals in soils using geochemical baseline and PMF approach. Sci Rep 10(1):1–11. https://doi.org/10.1038/s41598-020-63604-5
Mihaileanu RG et al (2019) Assessment of heavy metals (total chromium, lead, and manganese) contamination of residential soil and homegrown vegetables near a former chemical manufacturing facility in Tarnaveni, Romania. Environ Monit Assess 191(1). https://doi.org/10.1007/s10661-018-7142-0
Tóth G, Hermann T, Szatmári G, Pásztor L (2016) Maps of heavy metals in the soils of the European Union and proposed priority areas for detailed assessment. Sci Total Environ 565:1054–1062. https://doi.org/10.1016/j.scitotenv.2016.05.115
Chen H, An J, Wei S, Gu J (2015) Spatial patterns and risk assessment of heavy metals in soils in a resource-exhausted city, Northeast China. PLoS One 10(9):1–12. https://doi.org/10.1371/journal.pone.0137694
Lado LR, Hengl T, Reuter HI (2008) Heavy metals in European soils: a geostatistical analysis of the FOREGS Geochemical database. Geoderma 148(2):189–199. https://doi.org/10.1016/j.geoderma.2008.09.020
Zhang W, Liu M, Li C (2020) Soil heavy metal contamination assessment in the Hun-Taizi River watershed, China. Sci Rep 10(1):1–10. https://doi.org/10.1038/s41598-020-65809-0
Dumestre A, Sauve S, Mcbride M, Baveye P, Berthelin J (1999) Environmental contamination and toxicology copper speciation and microbial activity in long-term contaminated soils. 131: 124–131. https://doi.org/10.1007/s002449900451
Mirlean N, Roisenberg A, Chies JO (2007) Metal contamination of vineyard soils in wet subtropics (southern Brazil). Environ Pollut 149(1):10–17. https://doi.org/10.1016/j.envpol.2006.12.024
Schaider LA, Senn DB, Estes ER, Brabander DJ, Shine JP (2014) Sources and fates of heavy metals in a mining-impacted stream: temporal variability and the role of iron oxides. Sci Total Environ 490:456–466. https://doi.org/10.1016/j.scitotenv.2014.04.126
Senkondo YH, Semu E, Tack FMG (2015) Vertical distribution of copper in copper-contaminated coffee fields in Kilimanjaro, Tanzania. Commun Soil Sci Plant Anal 46(10):1187–1199. https://doi.org/10.1080/00103624.2015.1019085
Wei B, Yang L (2010) A review of heavy metal contaminations in urban soils, urban road dusts and agricultural soils from China. Microchem J 94(2):99–107. https://doi.org/10.1016/j.microc.2009.09.014
Moroń D et al (2012) Abundance and diversity of wild bees along gradients of heavy metal pollution. J Appl Ecol 49(1):118–125. https://doi.org/10.1111/j.1365-2664.2011.02079.x
Mejias Carpio IE, Ansari A, Rodrigues DF (2018) Relationship of biodiversity with heavy metal tolerance and sorption capacity: a meta-analysis approach. Environ Sci Technol 52(1):184–194. https://doi.org/10.1021/acs.est.7b04131
Rillig MC et al (2019) The role of multiple global change factors in driving soil functions and microbial biodiversity. Science (80) 366(6467):886–890. https://doi.org/10.1126/science.aay2832
Rocca JD et al (2019) The Microbiome Stress Project: toward a global meta-analysis of environmental stressors and their effects on microbial communities. Front Microbiol 10(JAN). https://doi.org/10.3389/fmicb.2018.03272.
Zhou Z, Wang C, Luo Y (2020) Meta-analysis of the impacts of global change factors on soil microbial diversity and functionality. Nat Commun 11(1). https://doi.org/10.1038/s41467-020-16881-7
Shade A (2017) Diversity is the question, not the answer. ISME J 11(1):1–6. https://doi.org/10.1038/ismej.2016.118
Chao A (1984) Nonparametric estimation of the number of classes in a population. Scand J Stat 11(4):265–270
Chao A, Lee SM (1992) Estimating the number of classes via sample coverage. J Am Stat Assoc 87(417):210–217. https://doi.org/10.1080/01621459.1992.10475194
Shannon CE, Weaver W (1964) The mathematical theory of communications. Int Bus 131
Simpson EH (1949) Measurment of diversity. Nature 688(1943): 688 [Online]. https://doi.org/10.1038/163688a0
Kurm V, Geisen S, Gera Hol WH (2019) A low proportion of rare bacterial taxa responds to abiotic changes compared with dominant taxa. Environ Microbiol 21(2):750–758. https://doi.org/10.1111/1462-2920.14492
Shade A et al (2014) Conditionally rare taxa disproportionately contribute to temporal changes in microbial diversity. MBio 5(4):1–9. https://doi.org/10.1128/mBio.01371-14
Kabata-Pendias A (2011) Trace Elements in Soils and Plants - Fourth Edition 50 Suppl 1
Fierer N (2017) Embracing the unknown: disentangling the complexities of the soil microbiome. Nat Rev Microbiol 15(10):579–590. https://doi.org/10.1038/nrmicro.2017.87
Kicińska A, Pomykała R, Izquierdo-Diaz M (2022) Changes in soil pH and mobility of heavy metals in contaminated soils. Eur J Soil Sci 73(1):1–14. https://doi.org/10.1111/ejss.13203
Rieuwerts JS, Thornton I, Farago ME, Ashmore MR (1998) Factors influencing metal bioavailability in soils: preliminary investigations for the development of a critical loads approach for metals. Chem Speciat Bioavailab 10(2):61–75. https://doi.org/10.3184/095422998782775835
Oorts K (2013) Copper. In: Alloway B (eds) Heavy metals in soils. Environmental Pollution, vol 22. Springer, Dordrecht. https://doi.org/10.1007/978-94-007-4470-7_13
Bravin MN, Tentscher P, Rose J, Hinsinger P (2009) Rhizosphere pH gradient controls copper availability in a strongly acidic soil. Environ Sci Technol 43(15):5686–5691. https://doi.org/10.1021/es900055k
Broos K et al (2007) Soil factors controlling the toxicity of copper and zinc to microbial processes in Australian soils. Environ Toxicol Chem 26(4):583–590. https://doi.org/10.1897/06-302R.1
Fernández-Calviño D, Bååth E (2016) Interaction between pH and Cu toxicity on fungal and bacterial performance in soil. Soil Biol Biochem 96:20–29. https://doi.org/10.1016/j.soilbio.2016.01.010
Fernández-Calviño D, Arias-Estévez M, Díaz-Raviña M, Bååth E (2011) Bacterial pollution induced community tolerance (PICT) to Cu and interactions with pH in long-term polluted vineyard soils. Soil Biol Biochem 43(11):2324–2331. https://doi.org/10.1016/j.soilbio.2011.08.001
Chen YT, Wang Y, Yeh KC (2017) Role of root exudates in metal acquisition and tolerance. Curr Opin Plant Biol 39(Iii):66–72. https://doi.org/10.1016/j.pbi.2017.06.004
De Conti L et al (2020) Iron fertilization to enhance tolerance mechanisms to copper toxicity of ryegrass plants used as cover crop in vineyards. Chemosphere 243:125298. https://doi.org/10.1016/j.chemosphere.2019.125298
Montiel-Rozas MM, Madejón E, Madejón P (2016) Effect of heavy metals and organic matter on root exudates (low molecular weight organic acids) of herbaceous species: an assessment in sand and soil conditions under different levels of contamination. Environ Pollut 216:273–281. https://doi.org/10.1016/j.envpol.2016.05.080
Degryse F, Smolders E, Parker DR (2009) Partitioning of metals (Cd Co, Cu, Ni, Pb, Zn) in soils: concepts, methodologies, prediction and applications - a review. Eur J Soil Sci 60(4):590–612. https://doi.org/10.1111/j.1365-2389.2009.01142.x
Liu Y, Xu Z, Hu X, Zhang N, Chen T, Ding Z (2019) Sorption of Pb(II) and Cu(II) on the colloid of black soil, red soil and fine powder kaolinite: effects of pH, ionic strength and organic matter. Environ Pollut Bioavailab 31(1):85–93. https://doi.org/10.1080/26395940.2019.1578186
De Boer W, Folman LB, Summerbell RC, Boddy L (2005) Living in a fungal world: Impact of fungi on soil bacterial niche development. FEMS Microbiol Rev 29(4):795–811. https://doi.org/10.1016/j.femsre.2004.11.005
Emilia Hannula S, Morriën E (2022) Will fungi solve the carbon dilemma? Geoderma 413(February). https://doi.org/10.1016/j.geoderma.2022.115767
FracM, Hannula SE, Belka M, Jȩdryczka M (2018) Fungal biodiversity and their role in soil health. Front Microbiol 9(APR). https://doi.org/10.3389/fmicb.2018.00707
Egidi E et al (2019) A few Ascomycota taxa dominate soil fungal communities worldwide. Nat Commun 10(1). https://doi.org/10.1038/s41467-019-10373-z
Lin Y, Xiao W, Ye Y, Wu C, Hu Y, Shi H (2020) Adaptation of soil fungi to heavy metal contamination in paddy fields—a case study in eastern China. Environ Sci Pollut Res 27(22):27819–27830. https://doi.org/10.1007/s11356-020-09049-9
Acknowledgements
We thank all authors of the primary studies included in the meta‐analysis who indirectly contributed to this work. Specifically, we thank A. Avellan, M. Simonin and X. Guan for kindly providing the additional data on their studies. We extend our gratitude to W. Viechtbauer for his help while discussing methodological issues in the meta-analysis. The authors also thank Dr. Jenny Marie Booth for the English editing.
Funding
Open access funding provided by Libera Università di Bolzano within the CRUI-CARE Agreement. The research was supported by a grant from the Free University of Bolzano: TN200V (Pestiphere).
Author information
Authors and Affiliations
Contributions
L.B. planned and designed the work. M.S. screened the literature and collected the data. M.S. and G.M. conducted the meta-analysis. M.S. drafted the paper. G.M., L.B., T.M. and S.C. supervised the work. L.B. mainly contributed to the discussion of results.
Corresponding authors
Ethics declarations
Competing Interests
The authors declare no competing interests.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Signorini, M., Midolo, G., Cesco, S. et al. A Matter of Metals: Copper but Not Cadmium Affects the Microbial Alpha-Diversity of Soils and Sediments — a Meta-analysis. Microb Ecol 86, 1071–1081 (2023). https://doi.org/10.1007/s00248-022-02115-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00248-022-02115-4