Abstract
Heavily pigmented glacier ice algae Ancylonema nordenskiöldii and Ancylonema alaskanum (Zygnematophyceae, Streptophyta) reduce the bare ice albedo of the Greenland Ice Sheet, amplifying melt from the largest cryospheric contributor to eustatic sea-level rise. Little information is available about glacier ice algae interactions with other microbial communities within the surface ice environment, including fungi, which may be important for sustaining algal bloom development. To address this substantial knowledge gap and investigate the nature of algal-fungal interactions, an ex situ co-cultivation experiment with two species of fungi, recently isolated from the surface of the Greenland Ice Sheet (here proposed new species Penicillium anthracinoglaciei Perini, Frisvad and Zalar, Mycobank (MB 835602), and Articulospora sp.), and the mixed microbial community dominated by glacier ice algae was performed. The utilization of the dark pigment purpurogallin carboxylic acid-6-O-β-D-glucopyranoside (C18H18O12) by the two fungi was also evaluated in a separate experiment. P. anthracinoglaciei was capable of utilizing and converting the pigment to purpurogallin carboxylic acid, possibly using the sugar moiety as a nutrient source. Furthermore, after 3 weeks of incubation in the presence of P. anthracinoglaciei, a significantly slower decline in the maximum quantum efficiency (Fv/Fm, inverse proxy of algal stress) in glacier ice algae, compared to other treatments, was evident, suggesting a positive relationship between these species. Articulospora sp. did uptake the glycosylated purpurogallin, but did not seem to be involved in its conversion to aglycone derivative. At the end of the incubation experiments and, in conjunction with increased algal mortality, we detected a substantially increasing presence of the zoosporic fungi Chytridiomycota suggesting an important role for them as decomposers or parasites of glacier ice algae.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Two species belonging to the Zygnematophyceae, Ancylonema alaskanum [1] and Ancylonema nordenskioeldii, dominate the surface of the ablation zone of the Greenland Ice Sheet (GrIS) [2,3,4,5]. These so-called glacier ice algae represent the most important biological light-absorbing particle in the GrIS ‘Dark Zone’ (western margin) [3, 6,7,8,9,10,11,12], contributing an additional 10% of runoff generation during high-melt years [13]. Their impact on surface ice albedo is driven by high production of secondary phenolic pigmentation [8]. High-performance liquid chromatography (HPLC) analysis of the aqueous raw extract of Ancylonema alaskanum identified the main pigment as purpurogallin carboxylic acid-6-O-β-D-glucopyranoside (C18H18O12) [14], which is present at 11 times the cellular content of chlorophyll a within glacier algal cells [8]. The structure of the compound includes a benzotropolone backbone, hydroxyl and carboxyl groups and a sugar moiety (glucopyranoside) as described by Remias et al. [14]. Primarily, purpurogallin has a photoprotective activity, shielding the chloroplasts from the high irradiance (~ 1700 μmol photos m−2 s−1) characteristic of the GrIS surface ice environment during summer melt seasons [8], but this phenolic pigment may also play an antimicrobial function [14, 15], or potentially be a carbon source for the heterotrophic community inhabiting the ice surface [16]. However, bacterial biomass and production measured on the surface ice associated with high glacier ice algae abundance are far lower than expected, considering the amount of algal biomass available [2, 16].
Fungi have only recently been identified from the GrIS, with surface ice of the Dark Zone found to harbour a diverse and abundant fungal community [17]. Culture-dependent methods, verified by ergosterol quantification, detected up to 105 CFU/100 ml (colony-forming unit/100 ml) of surface ice [17]. In comparison with other ice habitats, ice with high algal abundance contained a high number of species otherwise known as endophytes and plant pathogens, suggesting that glacier ice algae might act as an environmental filter, shaping the associated fungal diversity and abundance. Overall, two filamentous fungi were noteworthy from a previous study [17]; Penicillium anthracinoglaciei was the only Penicillium that was also recovered from high algal content ice, and Articulospora sp. was observed in close association with glacier algal clumps [17]. Penicillium is a cosmopolitan genus, present in Arctic glaciers with high abundance and species diversity [18, 19]. The species P. anthracinoglaciei dominated samples from all the environments (surface ice, cryoconite, supraglacial water) collected in 2016 (July–August), and was recovered in 2017 as well, albeit in lower numbers (June–July) [17]. However, its occurrence was not only limited to glacial environments, but the species were also recovered from other habitats, such as refrigerators and museums (Fig. S1). Articulospora sp. is an aquatic Ingoldian fungus (from Ingold, the mycologist who discovered the typical habitat of these fungi which grow and sporulate underwater) belonging to the order Helotiales, with a recognized role in carbon dynamics of cryoconite holes in the Arctic (Svalbard) and the potential to influence plant colonisation in glacier forefields [20]. Members of the same genus are recognized as important freshwater decomposers of leaf detritus in Mediterranean regions [21, 22].
In many ecosystems, in addition to playing a role in nutrient cycling, fungi can also act as predators, pathogens and parasites and form symbiotic associations with plants, algae, animals and other organisms [23]. Chytridiomycota, often referred to as chytrids, are a group of saprotrophic and necrotrophic fungi widespread in both aquatic and terrestrial environments [24]. In marine habitats, they have been recognized as important microalgae parasites, able to influence algal bloom dynamics [25]. The presence of chytridiomycetous fungi in a variety of cold habitats has been proven by culture-independent methods [19, 26,27,28,29].
Experimental evidence of the interaction between glacier ice algae and other members of the microbial community is currently lacking since glacier ice algae have never been successfully grown in pure culture, despite several attempts [30,31,32]. In the present study, we focussed on two species of fungi from the surface ice of the GrIS, Articulospora sp. and the novel P. anthracinoglaciei, and their potential interactions with glacier ice algae. Mixed samples of glacier ice algae and other associated microbes were obtained directly from the GrIS and used during ex situ incubation experiments to investigate potential positive or negative impact of fungi on glacier algal assemblages. The potential for fungal utilization of glacier algal phenolic pigmentation, fungal impacts to algal photophysiology and establishment of putative lichenoid-like relationships was also examined. Finally, to test the possible growth modulation of third microbial partners of the mixed community, antimicrobial compound production was tested for all fungal species incubated during microcosm experiments. The fungal-algal interactions explored here are of particular importance due to the scale of changes occurring in the extremely cold environment of the GrIS, contributing to our understanding of the fungal role in the propagation of glacier algal blooms, which accelerate ice melt.
Materials and Methods
Description of Sampling Site and Samples
Sampling for fungal isolation was performed in the southwestern ablation zone of the GrIS in the 2016 (July–August) and 2017 (June–July) summer melt seasons. Glacier ice algae for the co-cultivating experiment were recovered from ice collected ∼60 km east of Kangerlussuaq (67°04′43′′ N 49°20′29′′ W), within the so-called Dark Zone, characterized by extensive glacier algal blooms and consequently, by low albedo [2, 3, 5, 17]. The surface ice of the Dark Zone is dominated by glacier ice algae (Ancylonema nordenskiöldii and Ancylonema alaskanum), but also contains a mixed microbial community of bacteria, fungi and viruses (e.g. 16, 17, 32). Since in this study we were not able to grow pure algal cultures, when we mention glacier ice algae in the experimental setups, that encompasses the whole microbial community, albeit with a high abundance of glacier ice algae. Surface ice samples of 2 cm depth (5 replicates, approximately 1830 ± 37 ml of ice) and containing high glacier algal abundance (104 cells/ml) were removed using a clean ice saw and transferred into sterile Whirl-Pak® plastic bags. Samples were stored in a dark fridge until transportation to the University of Bristol, UK.
Isolation of Fungi
For isolation of cultivable fungi, ice was collected and melted aseptically at in situ air temperature (approx. 4 °C). The samples were filtered, and filters were placed onto two enumerations (dichloran rose bengal chloramphenicol agar – DRBC [33], and Reasoner’s 2A agar – R2A [34]) and four different selective agar media (dichloran 18% glycerol – DG-18 [35], Malt Yeast Extract 10% sodium chloride and 12% glucose – MY10-12 [35], synthetic nutrient-poor agar – SNA [36], minimal medium – MM [37]) and incubated at 10 °C as described in Perini et al. [17]. Additionally, samples of melted ice containing a high abundance of glacier ice algae communities were directly inoculated (100 μl) in triplicates on MM, SNA, DRBC, R2A and DG18, and spread with a Drigalski spatula. To prevent bacterial growth, media contained chloramphenicol (50 mg/l) except for R2A. Pure cultures of the identified fungal isolates were deposited in the Ex Culture Collection of the Infrastructural Centre Mycosmo (MRIC UL) at the Department of Biology, Biotechnical Faculty, University of Ljubljana, Slovenia.
Identification of Fungi
DNA from pure fungal cultures was extracted as described by Perini et al. [17]. For filamentous fungi, a portion of rDNA including the Internal Transcribed Spacer region 1 (ITS1), 5.8S rDNA and ITS region 2 (ITS2) was amplified using ITS5 and ITS4 primers [38]. The ITS nucleotide sequences were determined by Sanger sequencing, performed by Microsynth AG, Switzerland. MUSCLE software [39] implemented in the MEGA5 package [40] was used to align the sequences. The maximum likelihood method implemented in MEGA 5 [40] was used to build phylogenetic trees, which were used to identify the unknown sequences by its proximity to the closest type and reference sequences previously retrieved with the BLAST software from the National Centre for Biotechnology Information (NCBI) GenBank database [41].
A total of 50 different fungal species belonging to 36 genera were isolated and identified from cryoconite, snow, dark and clean supraglacial ice and supraglacial water [17]. Based on cultivable diversity (CFU/100 ml), two of the most abundant filamentous fungi recovered from both seasons were P. anthracinoglaciei (42 isolates) and Articulospora sp. (59 isolates). Phylogenetic analysis of the combined partial β-tubulin and RNA polymerase 2 (RPB2) genes supported the recognition of a novel species: P. anthracinoglaciei with a well-supported clade containing all the isolates from Greenland (Table S1, Fig. S1). Detailed methods and results regarding gene amplification and phylogenetic analysis are provided in the supplementary material. The abundance and repeated isolation of these two fungal species identified them as the best candidates for a co-cultivation experiment to explore the nature of interactions with glacier ice algal communities on the Greenland Ice Sheet. The description of the proposed new species P. anthracinoglaciei is provided at the beginning of the “Results” section of the manuscript (Fig. 1). Methods and results for screening of extracellular enzymes and antimicrobial compound production for both fungal species used in the experiment and phylogeny with secondary metabolites profile of P. anthracinoglaciei are provided in the supplementary material (Fig. S2, Tables S2–S5).
Co-cultivation Experiment Setup
Surface ice samples, characterized by a mixed microbial community of bacteria, fungi, viruses and glacier ice algae, were homogenized, transferred into vented sterile cell culture flasks (Corning Inc, USA) and kept at +4 °C in the dark for 3 days prior to the start of the incubation experiment to limit the light stress. Glacier algal maximum quantum efficiency in the dark-adapted state (Fv/Fm, inverse proxy of stress in microalgae) was monitored throughout using pulse-amplitude modulation (PAM) fluorometry as in Williamson et al. [8]. The incubation comprised 5 treatments in triplicates containing varying combinations of glacier ice algae (Ancylonema nordenskiöldii and Ancylonema alaskanum), and fungi P. anthracinoglaciei EXF-11445, and Articulospora sp. EXF-13072, isolated from GrIS surface ice (Table 1).
Glacier ice algae suspensions were transferred to a 10 L Duran bottle, previously cleaned with 15% hydrogen peroxide and washed with sterile Milli-Q water, wrapped in ice packs to ensure that the suspension remained cold. The Duran bottle was continuously stirred to prevent algal cells from sinking and to ensure a homogenous distribution of glacier ice algae was achieved in subsequent treatments. A sterile glass cylinder was used to divide the suspension randomly between sterile culture flasks (Corning Inc, USA) – final volume of 250 ml. A sterile filter unit containing a 0.45-μm polycarbonate filter (Whatman, England) was used to remove glacier ice algae for fungal control treatments. The filtrate from each unit was divided between cell culture flasks for the respective treatment.
Glacier ice algae Ancylonema alaskanum and Ancylonema nordenskioeldii were incubated without any fungal culture (treatment 1), or incubated with 5% v/v inoculum of Articulospora sp. mycelium (treatment 2), since the fungus does not sporulate in culture, or P. anthracinoglaciei spores (treatment 3). Fungal controls in filtered melted ice without the presence of glacier ice algae were also included (Articulospora sp. (treatment 4); P. anthracinoglaciei (treatment 5)).
Epifluorescence microscopy (Leica DM 2000 epifluorescence microscope) with a Fuchs-Rosenthal haemocytometer (0.2 depth, 1/16 mm2, Lancing, UK) was used to quantify initial algal concentrations of 5 × 103 cells/ml across treatments, consistent with their ambient concentration in GrIS surface ice (103–104 cells/ml) [5]. Data available from the GrIS showed that relative ambient frequencies of Penicillium and Articulospora genera from ITS2 amplicon sequencing results in surface ice are up to 0.3 and 0.06% of all reads, respectively [17]. To verify the interaction of these fungi with glacier algal assemblages, cultures were inoculated with an initial concentration of 3 × 106 spores/ml P. anthracinoglaciei across treatments. Articulospora sp. biomass inoculum was measured as 1.74 mg dry weight per ml (8.7 mg in 5 ml). Sterile spore suspension solution was also added to treatment 1 to account for any additional nutrients that may be available from this substrate.
The flasks were incubated at 4 °C under a 14/10 h cycle of light-darkness at 181.45 ± 24.5 μmol m−2 s−1 photosynthetically active radiation (PAR) for 5 months. Light/dark cycles reflected the ambient irradiance regime apparent at our sampling location on the southwestern GrIS. A parallel experiment with continuous dark conditions was performed for all treatments by wrapping flasks with aluminium foil. Culture flasks were mixed every other day and samples randomly redistributed to ensure an even distribution of light across all samples.
Samples for light microscopy, scanning electron microscopy and pulse-amplitude-modulated (PAM) fluorometry were collected in triplicates at five time points; day one of incubations and after 1, 2, 3 weeks, and after 2 and 5 months of incubation.
Photophysiology
Pulse-amplitude modulation (PAM) fluorometry was used to assess algal photophysiology throughout incubations at each sampling time point for those treatments containing glacier ice algae (up to 5-month incubation). Rapid light response curves (RLCs) were performed following Perkins et al. [42] using a Walz Water-PAM fluorometer (Walz GmBH) consisting of a PAM-control unit with attached cuvette emitter-detector WATER-ED. Samples were dark-adapted for a minimum of 5 min prior to all RLCs, which were performed as 9 incremental light steps of 20 s duration ranging from 0 to 3480 μmol photons m−2 s−1 PAR, calibrated using a cosine-corrected sensor LI-COR Quantum Sensor (LI-COR Instruments, Lincoln, NE, USA). The maximum quantum efficiency (Fv/Fm) was calculated from minimum (Fo) and maximum (Fm) fluorescence yields in the dark-adapted state as Fv/Fm = (Fm – Fo)/Fm. Electron transport through photosystem II (PSII) was calculated from PSII quantum efficiencies (Y[PSII]) in relative units as rETR = Y(PSII) × PAR × 0.5, assuming an equal division of PAR between photosystems I and II. Analysis of RLCs (rETR ~ PAR) followed Perkins et al. (2006), with iterative curve fitting (R, v.3.6.0) and calculation of the relative maximum electron transport rate (rETRmax), maximum light utilization coefficient (α) and light saturation coefficient (Ek) following Eilers and Peeters [43].
Light Microscopy
For the determination of the morphological characteristics and monitoring of interactions between fungi and algae, microscope slides were prepared from incubations and mixed (1:1) with a water solution of 0.1% (w/v) poly-L-lysine. The slides were examined under oil immersion with a BX51 microscope (Olympus, Japan) by differential interference contrast (DIC), at up to ×100 magnification. Digital micrographs were taken with a DP73 digital camera (Olympus, Japan), and analyzed using the cellSens software (Olympus, Japan). Pigmented glacier ice algae were classified as active/alive, whereas unpigmented algae were classified as unactive/dead by manual counting.
Scanning Electron Microscopy (SEM)
For SEM observation, the samples of co-cultivated algae and fungi were fixed in an equal volume of 0.5% glutaraldehyde + 0.2% formaldehyde at room temperature for up to 2 h. The samples were then washed 3 times with 500 μl of phosphate buffer (7.2 – 7.4 pH) for 7 min, transferred on a polycarbonate membrane (pore size 200 nm) in a filtrating column system and post-fixed in 0.25% OsO4 for 30 min. After washing in Milli-Q water, the samples were dehydrated in an EtOH series of ascending concentrations (50%, 70%, 90% and 100%) and acetone for 3 min in each. Acetone was gradually replaced by hexamethyldisilazane and allowed to air-dry overnight. Filters with dried samples were attached to metal holders, covered with platinum and observed with a JEOL JSM-7500F field-emission scanning electron microscope.
Quantification and Utilization of the Glycosylated Purpurogallin
To investigate the potential for glacier algal phenolic pigmentation to act as a nutrient source for our two fungal isolates, a complementary incubation experiment was established. Five hundred microliters of glacier algal water-extractable phenolic pigmentation (extracted after Williamson et al. [5] and confirmed here by HPLC to be dominated by purpurogallin carboxylic acid-6-O-β-D-glycopyranoside) was incubated with either P. anthracinoglaciei or Articulospora mycelium in sterile 5-ml Eppendorf tubes containing 500 μl of sterile minimal medium (MM) 0.5 M without carbon source [37]. Eppendorf tubes were incubated at 15°C for 6 weeks. Fungi were previously grown in malt extract broth (MEB) for 7 days at 15°C. Mycelium were washed with sterile Milli-Q water prior to inoculation.
Pigment concentration was calculated based on the chromatographic peak height (mAU, milli-absorption unit) monitored at 350 nm. Even though peak areas for the detected compounds were also calculated, we preferred to use the peak height of the compounds.
Confirmation of Identity of Purpurogallin Carboxylic Acid Aglycone by HPLC-MS-MS-DAD
HPLC-MS-MS-DAD analysis was run on an Agilent Q-TOF 6545 using an Agilent Poroshell 120 Phenylhexyl column (2.1mm diameter × 150 mm length, with 1.9 μm particle size) in eluent A consisting of 100% water with 20 mM formic acid and eluent B 100% acetonitrile with 20 mM formic acid, in a gradient running from 10% eluent B to 100% B in 10 min, held at 100% B at 2 min, back to 10% B over 0.1. min, and held at 10% B for 1.9 min. The column temperature was 40°C, and the eluent flow rate was 350 μl/min. The MS range measured was from 100 to 1700 Dalton, with a scan rate of 10 spectra/s, using positive and negative ionization separately. Fragmentation (for MS-MS) was done at 10, 20 and 40 eV [44]. The UV spectrum was compared to the published spectrum [14] and to a standard of purpurogallin (purpurogallin standard purchased from Sigma-Aldrich, as nr. P7380 MSDS), the latter having a slightly different UV spectrum (and retention time) than purpurogallin carboxylic acid. The UV spectra of purpurogallin carboxylic acid and purpurogallin carboxylic acid glycopyranoside were identical (Fig. S3).
Data Analysis
All statistical analyses of data were performed using R v.3.4.1 [45]. Levene’s test for homogeneity of variance and Shapiro-Wilk's normality test were performed on all data prior to the application of parametric analyses. Consequently, a two-way analysis of variance (ANOVA) was conducted in relation to algal photophysiology for each treatment. Post-hoc Tukey HSD was performed on all significant ANOVA analyses.
Results
Description of Penicillium anthracinoglaciei Perini, Frisvad and Zalar sp. nov.
Penicillium anthracinoglaciei Perini, Frisvad and Zalar sp. nov. Mycobank (MB 835602) (Fig. 1).
In: Penicillium subgenus Penicillium section Brevicompacta [46].
Etymology: anthrăcĭnus: “of black colour” (coal black), glăcĭēs, glăcĭēi: “ice”; referring to the origin of this fungus - coal black–coloured ice due to algal bloom causing blackening of Greenland Ice Sheet.
Morphological description: colony diameter after 1 week at 25 °C, in mm: CYA: 13–20 (pale yellow reverse); CYAS: 14–20; MEA: 12–15.5; YES: 17–18.5 (green with yellow reverse); CREA: 7–9, moderate growth, acid production. CYA, 5 °C: microcolonies 0–2; CYA, 10 °C: 3–9.5; CYA, 15 °C:13–16.5; CYA, 30 °C, 37 °C: no growth. CYA 15 °C/CYA 25 °C: 0.9 [0.8–1.1], psychrotolerant. Optimum growth temperature: 15–24 °C. CYA/CYAS: 1.1 [0.8–1.4], halotolerant. Conidia en masse dark green.
Synnemata or fasciculation: none; sclerotia and ascomata: none; colony texture on CYA: velutinous, radially furrowed; conidium colour on CYA: green to dark green; exudate droplets on CYA: none; reverse colour on CYA: pale yellow; reverse colour on YES: cream yellow; diffusible colour: none; Ehrlich reaction: no or weak violet reaction; conidiophores arising from agar surface; stipes (100–)150–300(–500) × 3–4 (6.5) μm, smooth-walled; penicilli predominantly terverticillate; rami 2–3 per stipe, mostly appressed, (7–)8.5–20 × 2.5–5.5 (–8) μm; metulae 3–4 per ramus, 7–13 (–17) × 3–6 μm, clavate to apically inflated up to 10 μm diam; phialides 4–6 per metula, ampulliform with distinguishable necks, 5.5–11 × 2–4 μm; conidia subglobose to ellipsoidal, 2.5–3.5 × 2–3 μm, walls finely roughened, born in chains that form loose masses (Fig. 1).
Distribution: Greenland, Slovenia
Ecology and habitats: Greenland, various glacial environments, dust (deposition on museum items), refrigerators.
Chemotaxonomy (Table S4): Raistrick phenols, breviones, asperphenamate, quinolactacin 1 and 2, quinolactacin A1, xanthoepocin, mycophenolic acids, orthosporine 1, Alk-747 (an indole alkaloid), Alk-788 (an indole alkaloid), linoleic acid.
Holotype: EXF-11443H
Cultures ex type: EXF-11443 = IBT 34739, dark ice, collected 27.10.2016, Greenland, 67°04′43′′ N 49°20′29′′ W, adjacent to the S6 PROMICE weather station, Laura Perini.
The holotype, EXF-11443H, originated from dark ice in Greenland Ice Sheet, approximately 35 km from the southwestern Ice Sheet margin, 67° 04′43″ N 49° 20′29″ W, adjacent to the S6 PROMICE weather station, in July 2017 by Laura Perini. It is permanently preserved in a metabolically inactive state at the Ex Culture Collection of the Infrastructural Centre Mycosmo (MRIC UL), Slovenia (www.ex-genebank.com) at the Department of Biology, Biotechnical Faculty, University of Ljubljana, Slovenia. Accession numbers of DNA sequences derived from ex-type strain EXF-11443, deposited into the above mentioned Ex Culture Collection: MT080493 (BenA), MT080519 (RPB2), MT080552 (CaM).
Cultures ex type: EXF-11443 = IBT 34739.
The results on the phylogenetic placement of P. anthracinoglaciei are provided in the supplementary material (Fig. S1) and detailed results of the comparison of produced secondary metabolites in related species in Table S4.
Higher Algal Fv/Fm Ratios in P. anthracinoglaciei Treatments After 3 Weeks of Incubation
RLCs confirmed active and healthy glacier algal assemblages at the onset of incubation (day0), with Fv/Fm averaging 0.67 ± 0.01 and rETRmax 102.7 ± 2.5 across treatments (Fig. 2A, B). To give context to these values, in situ experiments assessing the glacier ice algae photophysiology highlighted that Fv/Fm ranged around 0.5 and rETRmax ranged between 120 and 160 when incubated under 50% or 0% of irradiance [8]. Values were lower when glacier ice algae were incubated at natural irradiance [8]. After 5 months of laboratory incubation, some degree of variable chlorophyll fluorescence was apparent across treatments with an average Fv/Fm of 0.51 ± 0.01; however, no measurable electron cycling through photosystem II (PSII) was detectable over RLCs performed using PAM fluorescence (Fig. S5, Table S6). Considering that no active photophysiology was detected after 5 months of incubation, we only present and discuss here the PAM data from up to 3-week incubation, where the glacier ice algae showed signs of active photosynthesis. Across the first 3 weeks of incubation, glacier algal assemblages remained active, though a decrease in their photosynthesis rate (rETRmax) was observed across all treatments (Fig. 2B), likely reflecting the detrimental impacts of laboratory incubations on glacier algal health [30, 31]. Consistent with this, a notable decline was apparent in Fv/Fm across all incubations under light conditions (Fig. 2A) despite increasing algal abundance (day0 media in light = 4530 ± 1416 cell ml−1, 3week-incubation media in light = 8123 ± 2252 cell ml−1) (Fig. S4). Fv/Fm under all dark conditions remained consistent or increased. After 3 weeks of incubation, algae incubated under dark conditions exhibited significantly higher Fv/Fm as compared to those in the light (algae only: F= 59.5, p< 0.001; Alg+Art: F= 58.6, p< 0.001; Alg + Pen: F= 272, p< 0.001), possibly due to the lack of light stress compared to the light treatment. Furthermore, Fv/Fm was significantly higher in glacier ice algae + Penicillium treatment in the light as compared to the algal control (F= 22.8, p< 0.05). On the contrary, Fv/Fm was not significant for the glacier ice algae + Articulospora treatment as compared to the algal control in the light (F= 22.8, p>0.05) or the dark (F= 0.8, p>0.05). No significant differences in α or Ek were apparent between treatments of light/dark incubations (Fig. S6 and S7).
A Glacier algae maximum quantum efficiency Fv/Fm (mean ± standard error, n = 3) during the incubation period across both light (orange) and dark (black) treatments. B rETRmax (maximum relative electron transport rate) (mean ± standard error, n = 3) over the incubation period in each treatment for light (orange) and dark (black) conditions
Penicillium anthracinoglaciei Had a Beneficial Impact on Glacier Ice Algae Pigmentation After 5-Month Incubation in the Dark
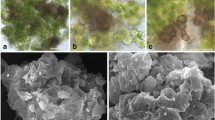
Microscopy showed that after 3 weeks of incubation, glacier ice algae remained pigmented and formed clumps containing also mineral particles (Fig. 3A). Articulospora sp. had densely septated brownish hyphae with many vacuolar inclusions (not shown). No sporulation of Articulospora sp. during the experimental incubation was observed. During its growth in co-culture with algae, it formed a net-like hyphal structure that incorporated individual algal cells (Fig. 3B). P. anthracinoglaciei spore germination occurred after 1 week of incubation, and after 3 weeks of incubation, its hyphae were found to be in association with algal clumps (Fig. 3C). After 2 months of incubation, P. anthracinoglaciei mycelium appeared full of vacuolar inclusions in the fungal control (not shown). Both fungi grew in an extremely low nutrient medium (melted ice) and at a low temperature (+4 °C).
A–C Light microscopy, light condition, 3-week incubation. A algal control; B glacier ice algae + Articulospora, in this treatment algae are entrapped in the mycelial network; C glacier ice algae + Penicillium anthracinoglaciei. D–F Light microscopy, dark condition, 5-month incubation. D algal control; E glacier ice algae + Articulospora; F glacier ice algae + Penicillium anthracinoglaciei. In the latter treatment, the algae retained their characteristic dark pigmentation. Algal cells: black arrows (Aa, Ancylonema alaskanum; An, Ancylonema nordenskioeldii); fungal hyphae: white arrows
Light microscopy observations of the co-cultivation treatments showed that the phenolic pigment was retained in glacier ice algae under both light and dark conditions after 2 months of incubation. After 5 months, glacier ice algae showed loss of their dominant pigmentation, (Fig. 3D and E) presumably as a consequence of cell mortality except in the P. anthracinoglaciei treatment under dark conditions, where glacier ice algae retained their characteristic pigmentation (Fig. 3F). Around 73% of the algal cells were pigmented in the Penicillium treatment, compared to 34% of the algal control (Table S7).
After 5 months of incubation, glacier ice algae in all treatments deteriorated considerably, resulting in the growth and release of a non-targeted group in our experiments, Chytridiomycota fungi and their zoospores (Fig. 4A and B). Eucarpic-monocentric thalli had a spherical shape and were on long pedicel-like rhizoids with extensive branching. Morphological characteristics of the Chytridiomycota are shown in Fig. 4C. All attempts to grow chytridiomycetous fungi thereafter in pure culture were unsuccessful, but since these findings became an interesting feature of the experiments at the end of the incubations, a discussion regarding them will be held.
Light microscopy, light condition, 5-month incubation. Penicillium treatment. Chytrids releasing zoospores (A, black arrow) and interacting with the algae through rhizoids (white arrow) (B). Eucarpic-monocentric thalli containing zoospores (white arrowheads) on long pedicel-like rhizoids (black arrowheads) are clearly visible (C)
Articulospora Embedded Glacier Ice Algae on its Hyphal Network
Scanning electron microscopy revealed preserved outer morphology of both algal species Ancylonema alaskanum and Ancylonema nordenskioeldii under light and dark conditions after 2 months of incubation (Fig. 5), whilst numerous deflated algae appeared after 5-month incubation time (Fig. 5D), indicating algal mortality. The combination of Articulospora sp. and glacier ice algae showed entanglement of the latter in the fibre-like network of Articulospora sp. hyphae, with many contact points between the partners after 2 months of incubation (Fig. 5B). After 5 months of incubation, co-cultivated samples were densely packed together (Fig. 5E, F), revealing the presence of chytridiomycetous fungi (Fig. 6A) and their rhizoidal system branching outside the algal substrate (Fig. 6B).
Scanning electron micrographs of algal-fungal co-cultivations. A, D Algal control (Aa, Ancylonema alaskanum; An, Ancylonema nordenskioeldii; asterisk - dead cell). Algae (black arrows) incubated with Articulospora sp. (B, E), and P. anthracinoglaciei (C, F) hyphae (white arrows) after 2 months and 5 months of incubation in light condition. B Articulospora sp. started developing a putative lichenoid structure with embedded glacier ice algae (photobiont) inside the hyphal network. D–F After 5 months of incubation, cultures appeared more compact after the preparation for SEM (p, mineral particles)
Penicillium anthracinoglaciei Was Able to Interconvert Purpurogallin Glucopyranoside into an Aglycone Derivative
The uptake of purpurogallin glucopyranoside by P. anthracinoglaciei and Articulospora sp. was tested. After 6 weeks of incubation, the negative control (containing no fungal culture) showed in solution (extracellularly) the presence of 337.5 ±162.5 mAU of purpurogallin glucopyranoside and a lower amount of the aglycone derivative (1.1 ±0.9 mAU) (Fig. 7, Table S8). Fungi are preferentially cleaving the sugar moiety for utilization in metabolism and thus driving the interconversion from purpurogallin glucopyranoside to purpurogallin aglycone. Both compounds were identified intracellularly, in the mycelium of both fungi. The aglycone derivative peak in P. anthracinoglaciei mycelium was higher (49 ±0 mAU) than purpurogallin glucopyranoside peak (31.25 ±1.75 mAU), which represented a 157 ±9% interconversion within the Penicillium treatment. Articulospora sp. mycelium contained 46 ±2 mAU of purpurogallin glucopyranoside and 1 ±0 mAU of purpurogallin aglycone, with a lower percentage of interconversion (2.2 ±0.1%).
HPLC results as monitored at 350 nm indicated the relative amounts of purpurogallin carboxylic acid-6-O-β-D-glycopyranoside and purpurogallin carboxylic acid aglycone across fungal and pigment controls and treatments after 6 weeks of incubation, both total (extracellular + intracellular) and intracellular. Values were calculated based on the chromatographic peak height and expressed in mAU (mean ± standard error, n = 3). The relative percentage on the top of the bars indicates the amount of purpurogallin glycopyranoside transformed into purpurogallin aglycone
Discussion
In this study, we have attempted to map interactions between glacier ice algae (in a mixed microbial community from the ice) and fungal communities isolated from the surface of the GrIS. Whilst fungal concentrations were most likely elevated as compared to ambient concentrations apparent within GrIS surface ice, the purpose of our experiments was not to recreate ambient concentration, but to promote a response between the partners. Fungal concentrations were therefore applied as a treatment to identify the fungal impact on glacier algal assemblages. The microbial community in glacial environments is complex and the interactions between microorganisms may well be more elaborate than implied here. Particularly, besides filamentous fungi that were tested here, abundant species of yeast might have a role in modulating the growth and survival of ice algae.
Our in vitro co-cultivation experiments showed potential associations, with both fungi uptaking the purpurogallin glucopyranoside and with P. anthracinoglaciei improving the photosynthetic capabilities of the ice algae, compared to the other treatments. The two fungal species used in the experiments also produced several active compounds, emphasising their ability to regulate the growth of other microbial partners in these associations (Fig. S2, Tables S2–S5). Based on the overall results, we hypothesize different types of fungal-algal interactions occurring in GrIS surface ice.
Penicillium—Positive Association Between Fungi and Glacier Ice Algae
After 3 weeks of incubation, PAM data showed a significantly slower decline in Fv/Fm (inverse proxy of photosynthetic algal stress) in co-cultures that contained P. anthracinoglaciei in comparison with other treatments, indicating that Penicillium improved the survivability of glacier ice algae. Whilst this might simply be a result of a shading effect conferred by the fungi, no such positive effect on Fv/Fm was conferred by Articulospora sp. that entrapped algal cells in its hyphal network and would presumably have the same shading impact on glacier ice algae. The beneficial effects of P. anthracinoglaciei on algae in vitro might be driven additionally by the production of beneficial secondary metabolites, or mobilization of otherwise unavailable organic or inorganic compounds. For example, several species of penicillia are able to mobilize phosphorus, which has been demonstrated to be a limiting nutrient for glacier ice algae communities [47], making it available to plants, promoting and increasing their growth [48,49,50,51,52,53,54,55]. P. anthracinoglaciei showed its ability to produce several secondary metabolites in standard conditions (pure culture, 25 °C) (Table S4). These might affect glacier ice algae directly or indirectly, by modulating the growth or metabolism of a third partner in this mixed community, with a potentially beneficial effect on algal growth. HPLC analysis revealed the presence of mycophenolic acid and its derivatives, xanthoepocin, Raistrick phenols, asperphenamate, quinolactacins and breviones. Mycophenolic acid reportedly proved to inhibit Candida albicans and Staphylococcus aureus, and several other microorganisms [56]. Xanthoepocin is an anthraquinone dimer with antifungal and antibiotic properties, isolated for the first time from Penicillium simplicissimum [57]. Breviones, isolated from Penicillium sp. by Tikikawa et al. [58], are bioactive diterpenoid derivatives with potential use in agriculture as herbicides. We hypothesize that P. anthracinoglaciei might exert antibacterial activity during active algal growth and photosynthesis. The screening of antimicrobial compounds revealed that P. anthracinoglaciei organic extract had a wide antimicrobial spectrum, ranging from fungi to bacteria (Table S5, Fig. S2). It was active both against pathogenic bacteria Bacillus subtilis subsp. spizizenii, Staphylococcus aureus subsp. aureus and opportunistic yeast Candida albicans, and to environmental isolates of Micrococcus lactis, Sphingomonas sp., Cryobacterium psychrotolerans and Bacillus sp.
Light microscopy also highlighted the potential beneficial impact of P. anthracinoglaciei on glacier ice algae pigmentation, as evidenced by the prevalence of fully pigmented ("overall" cellular pigmentation) glacier algal cells (73%) following 5 months of incubation in the dark (Fig. 3). In contrast, the proportion of pigmented cells were conspicuously lower (34%) after 5 months of the algal control treatment. Taken together with measurable variable fluorescence at the end of the experiment (average Fv/Fm = 0.49 ± 0.04; Table S6), these data potentially suggest the entrance on some form of less active stage within our algal assemblages co-incubated with P. anthracinoglaciei under dark conditions for a prolonged period. Glacier ice algae are known to over-winter in a vegetative state, presumably as an adaptation to allow rapid onset of metabolism at the start of their relatively short growth season [59]. For example, studies on Alpine glaciers have demonstrated the high freezing tolerance of Ancylonema alaskanum and recovery to an active state after ~ 1 week following the end of the winter period and onset of spring ablation [31]. Similarly, active glacier algal assemblages have been observed within shallow sub-surface ice cores at the beginning of the ablation season on the GrIS [16]. In this study, sustained pigmentation of the glacier ice algae during a prolonged (5 months) incubation under complete darkness in the presence of P. anthracinoglaciei indicated a direct or indirect beneficial role of this fungus in algal survival. Further research is required to clarify whether glacier ice algae could recover their active physiology after such a prolonged period of darkness.
Incubation of P. anthracinoglaciei in a medium with purpurogallin glucopyranoside as the only carbon source resulted in the detection of purpurogallin carboxylic acid aglycone, a compound that exhibited an identical UV spectrum to purpurogallin, but a longer retention time (Fig. S3). These data indicated the ability of P. anthracinoglaciei of uptaking the molecule and removing the sugar moiety, possibly utilizing the latter as a nutrient source. This interconversion could occur in the fungus either extracellularly or intracellularly, since both forms of purpurogallin were detected in the medium and mycelium. Since a variety of factors (algal cell burst, death, and release through exudation) can involve the presence of purpurogallin glucopyranoside in the extracellular environment, these data do not contrast with the ability of Penicillium to sustain the algal pigmentation after a long period of incubation in the dark. The fungal ability to utilize the glucopyranoside sugar as a nutrient source will be confirmed in a future experiment.
Articulospora—Primitive Lichenoid Association Between Fungi and Glacier Ice Algae
Articulospora sp. exhibited a potential lichenoid-like relationship with glacier ice algae. Both light and scanning electron microscopies showed a progressive accumulation of the glacier ice algae, embedded in extracellular polymeric substances (EPS), in Articulospora hyphal network, that became particularly distinct after 2 months of incubation. Articulospora sp. showed the ability to uptake purpurogallin glucopyranoside in the mycelium; however, its interconversion into purpurogallin aglycone was negligible compared to Penicillium in pure culture with purpurogallin glucopyranoside in the medium.
According to the literature, in borderline lichens, two participating species do not form structures resembling a lichen thallus. In these associations, fungal and algal cells grow intertwined with loose contacts observed in both native samples and culture experiments [60, 61]. Because these fungi share strongly oligotrophic environments with green algae, they might take advantage from the presence of the primary producers as a first selective advantage to promote lichenoid-like associations. These loose associations can better proliferate under fluctuating hygroscopic conditions of the subaerial/ice-water native habitat, even when algal cells become more embedded in a thick layer of exopolysaccharides, hampering the formation of contacts. Fungal association with microscopic algae potentially improves their meagre carbon supplies in oligotrophic conditions [62]. We therefore speculate this putative lichenoid-like structure is formed with direct alignment of hyphae and algal cells and many contact points between the partners, possibly with additional contributions of additional symbionts (photosynthetic algae, bacteria and yeasts) [63]. However, after 3 weeks of incubation, the presence of Articulospora sp. did not highlight any significant increase in the glacier ice algae Fv/Fm (inverse proxy of algal stress), either in light or dark treatments, suggesting a neutral effect (neither beneficial nor detrimental) on the algal photophysiology, and/or implying that the incubation conditions may not have been optimal to test the interactions between glacier ice algae and Articulospora.
Chytridiomycota—Parasitic and/or Saprotrophic Fungi
Although this study was not initially designed to investigate the role of Chytridiomycota, a considerable growth of this fungal group was observed by light microscopy in conjunction with increased algal mortality at the termination of our microcosm experiments (after 5 months). Chytridiomycota were only observed sporadically and in smaller numbers during previous sampling time points of the experiment. However, the presence of unidentified Chytridiomycota in dark ice samples from the GrIS was also supported by NGS biodiversity analysis [17], confirming their ubiquity in the GrIS surface ice environments, although none were recovered by cultivation. Moreover, cells of glacier ice algae A. nordenskioeldii were found to be colonized by chytridiomycetous fungi in ice samples from Svalbard glaciers [64], hinting at a possible host-specific infection of such glacier ice algae. We intend to address this interesting topic in future research. The length and branching of chytrid rhizoids outside of the algal substrate suggested that a proportion of their nourishment was derived from water dissolved algal or other compounds. Since phenolic pigments might play an antimicrobial function [14], the loss of dark pigmentation apparent during the loss of cell viability may stimulate an overgrowth of the first parasitic and later saprotrophic Chytridiomycota. Chytrids are known for their negative impact on algal biomass in freshwater systems, where they are able to influence algal bloom dynamics, their host gene pool, and phytoplankton size distribution and succession [25]. No information exists to date on the potential top-down controls of glacier algal blooms on the GrIS or indeed across the wider cryosphere [59].
Conclusions
A study of interactions between glacier ice algae communities and two selected fungal species found in the supraglacial ice of the Greenland Ice Sheet indicated that fungi and algae have different types of interactions.
The interconversion of the pigment purpurogallin glucopyranoside into its aglycone derivative by P. anthracinoglaciei suggests its ability of metabolizing the sugar moiety of the pigment, possibly using it as a carbon source. Conversely, Articulospora sp. embedded algae into its hyphal network, but failed to provide the algae with any benefits based on the analyses conducted in the experiments. Given the complexity of the systems used in this study, investigation of the potential involvement of additional partners in the fungal-algal interactions would also shed more light on the impact of non-photosynthetic organisms on the GrIS algal blooms.
On the one hand, the increasing presence of Chytridiomycota at the end of the incubation experiment in parallel with higher algal mortality indicates that these zoosporic fungi could act as decomposers or parasites of algal blooms. On the other hand, other fungal species (e.g., Penicillium anthracinoglaciei and Articulospora sp.) could have a role in sustaining glacier algal blooms during the ablation season or during the over-wintering period, providing various benefits from the mobilization of organic or inorganic compounds, to the secretion of secondary metabolites, physical shielding from light or water currents, induction of dormancy or even modulation of the growth of other microorganisms in putative lichenoid-like structures.
Data availability
The dataset generated for this study can be found in GenBank with the following accession numbers: MK460372, MK460374, MK460412, MK460414, MK460417, MK460418, MK460422, MT080468- MT080567.
References
Procházková L, Řezanka T, Nedbalová L, Remias D (2021) Unicellular versus filamentous: the glacial alga Ancylonema alaskana comb. et stat. nov. and its ecophysiological relatedness to Ancylonema nordenskioeldii (zygnematophyceae, streptophyta). Microorganisms 9:1–15. https://doi.org/10.3390/microorganisms9051103
Yallop ML, Anesio AM, Perkins RG et al (2012) Photophysiology and albedo-changing potential of the ice algal community on the surface of the Greenland ice sheet. ISME J 6:2302–2313. https://doi.org/10.1038/ismej.2012.107
Stibal M, Box JE, Cameron KA et al (2017) Algae drive enhanced darkening of bare ice on the Greenland ice sheet. Geophys Res Lett 44:11,463–11,471. https://doi.org/10.1002/2017GL075958
Lutz S, McCutcheon J, McQuaid JB, Benning LG (2018) The diversity of ice algal communities on the Greenland ice sheet as revealed by oligotyping. Microb Genomics 4:1–10. https://doi.org/10.1099/mgen.0.000159
Williamson CJ, Anesio AM, Cook J et al (2018) Ice algal bloom development on the surface of the Greenland ice sheet. FEMS Microbiol Ecol 94:fiy025. https://doi.org/10.1093/femsec/fiy025
Shimada R, Takeuchi N, Aoki T (2016) Inter-annual and geographical variations in the extent of bare ice and dark ice on the Greenland ice sheet derived from MODIS satellite images. Front Earth Sci 4:1–10. https://doi.org/10.3389/feart.2016.00043
Tedstone AJ, Bamber JL, Cook JM et al (2017) Dark ice dynamics of the south-west Greenland ice sheet. Cryosphere Discuss 2011:1–22. https://doi.org/10.5194/tc-2017-79
Williamson CJ, Cook J, Tedstone A et al (2020) Algal photophysiology drives darkening and melt of the Greenland ice sheet. Proc Natl Acad Sci 201918412. https://doi.org/10.1073/pnas.1918412117
Gokul JK, Cameron KA, Irvine-Fynn TDL et al (2019) Illuminating the dynamic rare biosphere of the Greenland ice sheet’s Dark Zone. FEMS Microbiol Ecol 95:1–46. https://doi.org/10.1093/femsec/fiz177
Wientjes IGM, Van De Wal RSW, Reichart GJ et al (2011) Dust from the dark region in the western ablation zone of the Greenland ice sheet. Cryosphere 5:589–601. https://doi.org/10.5194/tc-5-589-2011
Stibal M, Gözdereliler E, Cameron KA et al (2015) Microbial abundance in surface ice on the Greenland ice sheet. Front Microbiol 6:225. https://doi.org/10.3389/fmicb.2015.00225
Ryan JC, Hubbard A, Stibal M et al (2018) Dark zone of the Greenland ice sheet controlled by distributed biologically-active impurities. Nat Commun 9:1065. https://doi.org/10.1038/s41467-018-03353-2
Cook JM, Tedstone AJ, Williamson C et al (2020) Glacier algae accelerate melt rates on the south-western Greenland ice sheet. Cryosphere 14:309–330. https://doi.org/10.5194/tc-14-309-2020
Remias D, Schwaiger S, Aigner S et al (2012) Characterization of an UV- and VIS-absorbing, purpurogallin-derived secondary pigment new to algae and highly abundant in Mesotaenium berggrenii (Zygnematophyceae, Chlorophyta), an extremophyte living on glaciers. FEMS Microbiol Ecol 79:638–648. https://doi.org/10.1111/j.1574-6941.2011.01245.x
Inamori Y, Muro C, Sajima E et al (1997) Biological activity of purpurogallin. Biosci Biotechnol Biochem 61:890–892
Nicholes MJ, Williamson CJ, Tranter M et al (2019) Bacterial dynamics in supraglacial habitats of the Greenland ice sheet. Front Microbiol 10:1366. https://doi.org/10.3389/fmicb.2019.01366
Perini L, Gostinčar C, Anesio AM et al (2019) Darkening of the Greenland ice sheet: fungal abundance and diversity are associated with algal bloom. Front Microbiol 10:557. https://doi.org/10.3389/fmicb.2019.00557
Sonjak S, Frisvad JC, Gunde-Cimerman N (2006) Penicillium mycobiota in Arctic subglacial ice. Microb Ecol 52:207–216. https://doi.org/10.1007/s00248-006-9086-0
Perini L, Gostinčar C, Gunde-Cimerman N (2019) Fungal and bacterial diversity of Svalbard subglacial ice. Sci Rep 9:20230. https://doi.org/10.1038/s41598-019-56290-5
Edwards A, Douglas B, Anesio AM et al (2013) A distinctive fungal community inhabiting cryoconite holes on glaciers in Svalbard. Fungal Ecol 6:168–176. https://doi.org/10.1016/j.funeco.2012.11.001
Seena S, Duarte S, Pascoal C, Cássio F (2012) Intraspecific variation of the aquatic fungus Articulospora tetracladia: an ubiquitous perspective. PLoS One 7:e35884. https://doi.org/10.1371/journal.pone.0035884
Seena S, Marvanová L, Letourneau A, Bärlocher F (2018) Articulospora – phylogeny vs morphology. Fungal Biol 122:965–976. https://doi.org/10.1016/j.funbio.2018.06.001
Naranjo-Ortiz MA, Gabaldón T (2019) Fungal evolution: major ecological adaptations and evolutionary transitions. Biol Rev 94:1443–1476. https://doi.org/10.1111/brv.12510
Powell MJ (2017) Chytridiomycota. In: Handbook of the Protists: Second Edition. Springer International Publishing, pp 1523–1558
Frenken T, Alacid E, Berger SA et al (2017) Integrating chytrid fungal parasites into plankton ecology: research gaps and needs. Environ Microbiol 19:3802–3822. https://doi.org/10.1111/1462-2920.13827
Brown SP, Olson BJSC, Jumpponen A (2015) Fungi and algae co-occur in snow: an issue of shared habitat or algal facilitation of heterotrophs? Arct Antarct Alp Res 47:729–749. https://doi.org/10.1657/aaar0014-071
Hassett BT, Gradinger R (2016) Chytrids dominate arctic marine fungal communities. Environ Microbiol 18:2001–2009. https://doi.org/10.1111/1462-2920.13216
Yakimovich KM, Engstrom CB, Quarmby LM (2020) Alpine snow algae microbiome diversity in the Coast Range of British Columbia. Front Microbiol 11:1721. https://doi.org/10.3389/fmicb.2020.01721
Rojas-Jimenez K, Wurzbacher C, Bourne EC et al (2017) Early diverging lineages within Cryptomycota and Chytridiomycota dominate the fungal communities in ice-covered lakes of the McMurdo Dry Valleys. Antarctica Sci Rep 7:15348. https://doi.org/10.1038/s41598-017-15598-w
Remias D, Holzinger A, Aigner S, Lütz C (2012) Ecophysiology and ultrastructure of Ancylonema nordenskiöldii (Zygnematales, Streptophyta), causing brown ice on glaciers in Svalbard (high arctic). Polar Biol 35:899–908. https://doi.org/10.1007/s00300-011-1135-6
Remias D, Holzinger A, Lütz C (2009) Physiology, ultrastructure and habitat of the ice alga Mesotaenium berggrenii (Zygnemaphyceae, Chlorophyta) from glaciers in the European Alps. Phycologia 48:302–312. https://doi.org/10.2216/08-13.1
Procházková L, Remias D, Řezanka T, Nedbalová L (2018) Chloromonas nivalis subsp. tatrae, subsp. nov. (Chlamydomonadales, Chlorophyta): re–examination of a snow alga from the High Tatra Mountains (Slovakia). Fottea 18:1–18. https://doi.org/10.5507/fot.2017.010
King AD, Hocking AD, Pitt JI (1979) Dichloran-rose bengal medium for enumeration and isolation of molds from foods. Appl Environ Microbiol 37:959–964
Reasoner DJ, Geldreich EE (1985) A new medium for the enumeration and subculture of bacteria from potable water. Appl Environ Microbiol 49:1–7
Pitt JI, Hocking AD (2011) Methods for isolation, enumeration and identification. Fungi and Food Spoilage. Springer, US, pp 19–52
Nirenberg HI (1981) A simplified method for identifying Fusarium spp. occurring on wheat. Can J Bot 59:1599–1609. https://doi.org/10.1139/b81-217
De VRP, Burgers K, van de Vondervoort PJI, Frisvad JC, Samson RA, Visser J (2004) A new black Aspergillus Species, A. vadensis, is a promising host for homologous and heterologous protein production. Appl Environ Microbiol 70:3954–3959. https://doi.org/10.1128/AEM.70.7.3954
White TJ, Bruns T, Lee S, Taylor J (1990) Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. PCR Protoc 315–322. https://doi.org/10.1016/b978-0-12-372180-8.50042-1
Edgar RC (2004) MUSCLE: Multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res 32:1792–1797. https://doi.org/10.1093/nar/gkh340
Tamura K, Peterson D, Peterson N et al (2011) MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Biol Evol 28:2731–2739. https://doi.org/10.1093/molbev/msr121
Altschul SF, Gish W, Miller W et al (1990) BLAST_article.pdf. J Mol Biol 215:403–410
Perkins RG, Mouget JL, Lefebvre S, Lavaud J (2006) Light response curve methodology and possible implications in the application of chlorophyll fluorescence to benthic diatoms. Mar Biol 149:703–712. https://doi.org/10.1007/s00227-005-0222-z
Eilers PHC, Peeters JCH (1988) A model for the relationship between light intensity and the rate of photosynthesis in phytoplankton. Ecol Model 42:199–215. https://doi.org/10.1016/0304-3800(88)90057-9
Theobald S, Vesth TC, Rendsvig JK et al (2018) Uncovering secondary metabolite evolution and biosynthesis using gene cluster networks and genetic dereplication. Sci Rep 8:17957. https://doi.org/10.1038/s41598-018-36561-3
R Core Team (2017) R: a language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria https://www.R-project.org/
Houbraken J, Samson RA (2011) Phylogeny of Penicillium and the segregation of Trichocomaceae into three families. Stud Mycol 70:1–51. https://doi.org/10.3114/sim.2011.70.01
McCutcheon J, Lutz S, Williamson C et al (2021) Mineral phosphorus drives glacier algal blooms on the Greenland ice sheet. Nat Commun 12:1–11. https://doi.org/10.1038/s41467-020-20627-w
Qiao H, Sun XR, Wu XQ et al (2019) The phosphate-solubilizing ability of Penicillium guanacastense and its effects on the growth of Pinus massoniana in phosphate-limiting conditions. Biol Open 8:bio046797. https://doi.org/10.1242/bio.046797
Yin Z, Shi F, Jiang H et al (2015) Phosphate solubilization and promotion of maize growth by Penicillium oxalicum P4 and Aspergillus niger P85 in a calcareous soil. Can J Microbiol 61:913–923. https://doi.org/10.1139/cjm-2015-0358
Wakelin SA, Warren RA, Harvey PR, Ryder MH (2004) Phosphate solubilization by Penicillium spp. closely associated with wheat roots. Biol Fertil Soils 40:36–43. https://doi.org/10.1007/s00374-004-0750-6
Scervino JM, Papinutti VL, Godoy MS et al (2011) Medium pH, carbon and nitrogen concentrations modulate the phosphate solubilization efficiency of Penicillium purpurogenum through organic acid production. J Appl Microbiol 110:1215–1223. https://doi.org/10.1111/j.1365-2672.2011.04972.x
Asea PEA, Kucey RMN, Stewart JWB (1988) Inorganic phosphate solubilization by two Penicillium species in solution culture and soil. Soil Biol Biochem 20:459–464. https://doi.org/10.1016/0038-0717(88)90058-2
Pandey A, Das N, Kumar B et al (2008) Phosphate solubilization by Penicillium spp. isolated from soil samples of Indian Himalayan region. World J Microbiol Biotechnol 24:97–102. https://doi.org/10.1007/s11274-007-9444-1
Raymond NS, Stöver DM, Peltre C et al (2018) Use of Penicillium bilaiae to improve phosphorus bioavailability of thermally treated sewage sludge – a potential novel type biofertiliser. Process Biochem 69:169–177. https://doi.org/10.1016/j.procbio.2018.03.021
Gómez-Muñoz B, Pittroff SM, de Neergaard A et al (2017) Penicillium bilaii effects on maize growth and P uptake from soil and localized sewage sludge in a rhizobox experiment. Biol Fertil Soils 53:23–35. https://doi.org/10.1007/s00374-016-1149-x
Hedstrom L, Liechti G, Goldberg JB, Gollapalli DR (2011) The antibiotic potential of prokaryotic IMP dehydrogenase inhibitors. Curr Med Chem 18:1909–1918. https://doi.org/10.2174/092986711795590129
Igarashi Y, Kuwamori Y, Takagi K et al (2000) Xanthoepocin, a new antibiotic from Penicillium simplicissimum IF05762. J Antibiot (Tokyo) 53:928–933. https://doi.org/10.7164/antibiotics.53.928
Takikawa H, Hirooka M, Sasaki M (2003) The first synthesis of (±)-brevione B, an allelopathic agent isolated from Penicillium sp. Tetrahedron Lett 44:5235–5238. https://doi.org/10.1016/S0040-4039(03)01275-9
Williamson CJ, Cameron KA, Cook JM et al (2019) Glacier algae: a dark past and a darker future. Front Microbiol 10:524. https://doi.org/10.3389/fmicb.2019.00524
Ametrano CG, Selbmann L, Muggia L (2017) A standardized approach for co-culturing dothidealean rock-inhabiting fungi and lichen photobionts in vitro. Symbiosis 73:35–44. https://doi.org/10.1007/s13199-017-0479-2
Muggia L, Kraker S, Gößler T, Grube M (2018) Enforced fungal-algal symbioses in alginate spheres. FEMS Microbiol Lett 365:1–7. https://doi.org/10.1093/femsle/fny115
Gostinĉar C, Muggia L, Grube M (2012) Polyextremotolerant black fungi: oligotrophism, adaptive potential, and a link to lichen symbioses. Front Microbiol 3:1–6. https://doi.org/10.3389/fmicb.2012.00390
Spribille T, Tagirdzhanova G, Goyette S et al (2020) 3D biofilms: in search of the polysaccharides holding together lichen symbioses. FEMS Microbiol Lett 367:1–17. https://doi.org/10.1093/femsle/fnaa023
Fiołka MJ, Takeuchi N, Sofińska-Chmiel W et al (2021) Morphological and spectroscopic analysis of snow and glacier algae and their parasitic fungi on different glaciers of Svalbard. Sci Rep 11:1–18. https://doi.org/10.1038/s41598-021-01211-8
Bellas CM, Anesio AM, Barker G (2015) Analysis of virus genomes from glacial environments reveals novel virus groups with unusual host interactions. Front Microbiol 6:1–14. https://doi.org/10.3389/fmicb.2015.00656
Acknowledgements
We sincerely thank Lucia Muggia and Martin Grube for their help in the interpretation of the light microscopy and scanning electron microscopy pictures.
Funding
This project has received funding from the European Union’s Horizon 2020 research and innovation programme under the Marie Skłodowska-Curie grant MicroArctic agreement No. 675546 and the European Research Council (ERC) under the European Union’s Horizon 2020 Research and Innovation Programme (ERC Synergy Grant ‘Deep Purple’ under grant agreement No. 856416). We further acknowledge support from the UK Natural Environment Research Council Consortium Grant ‘Black and Bloom’ (NE/M021025/1). We received financial support from the Slovenian Research Agency to the Infrastructural Centres Mycosmo and Microscopy of Biological Samples (both MRIC UL), as well as to the programs P1-0170 and P1-0198 and the project J4-2549. JCF thanks the Danish National Research Foundation (DNRF137) for support to Center for Microbial Secondary Metabolites and Agilent for a Thought Leader Award (#2871).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Context
The significant role that photosynthetic microorganisms, mainly glacier ice algae Ancylonema nordenskiöldii and Ancylonema alaskanum, play on the Greenland Ice Sheet darkening has been recognized only recently. With their dark pigments that shield them from the high UV intensities present at those latitudes, glacier ice algae significantly contribute to ice albedo reduction resulting in an acceleration of ice melt. Made of circa 1.7 million square km of ice, the Greenland Ice Sheet would add around 7.2 m of water to the global sea level if fully melted. Currently, very little is known about the interactions between glacier ice algae with other members of the ice microbial community. The significance of our research is in increasing the understanding of glacier algal bloom dynamics by providing, for the first time, experimental evidence of the fungal physiology and their interaction with glacier ice algae.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Perini, L., Gostinčar, C., Likar, M. et al. Interactions of Fungi and Algae from the Greenland Ice Sheet. Microb Ecol 86, 282–296 (2023). https://doi.org/10.1007/s00248-022-02033-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00248-022-02033-5