Abstract
Type IB topoisomerases relax the torsional stress associated with DNA metabolism in the nucleus and mitochondria and constitute important molecular targets of anticancer drugs. Vertebrates stand out among eukaryotes by having two Type IB topoisomerases acting specifically in the nucleus (TOP1) and mitochondria (TOP1MT). Despite their major importance, the origin and evolution of these paralogues remain unknown. Here, we examine the molecular evolutionary processes acting on both TOP1 and TOP1MT in Chordata, taking advantage of the increasing number of available genome sequences. We found that both TOP1 and TOP1MT evolved under strong purifying selection, as expected considering their essential biological functions. Critical active sites, including those associated with resistance to anticancer agents, were found particularly conserved. However, TOP1MT presented a higher rate of molecular evolution than TOP1, possibly related with its specialized activity on the mitochondrial genome and a less critical role in cells. We could place the duplication event that originated the TOP1 and TOP1MT paralogues early in the radiation of vertebrates, most likely associated with the first round of vertebrate tetraploidization (1R). Moreover, our data suggest that cyclostomes present a specialized mitochondrial Type IB topoisomerase. Interestingly, we identified two missense mutations replacing amino acids in the Linker region of TOP1MT in Neanderthals, which appears as a rare event when comparing the genome of both species. In conclusion, TOP1 and TOP1MT differ in their rates of evolution, and their evolutionary histories allowed us to better understand the evolution of chordates.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
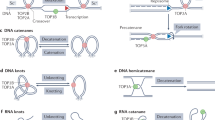
DNA topoisomerases introduce reversible breaks in the DNA phosphodiester backbone allowing for modifications in DNA topology during DNA replication, recombination, transcription and chromosome condensation (Pommier et al. 2022, 2016). Concerning Type I topoisomerases, they are monomeric and cleave one DNA strand at a time without requiring an energy cofactor. These topoisomerases are traditionally classified into two groups (Type IA and Type IB) without sequence or structural similarity. Indeed, while Type IA breaks the DNA by forming a covalent bond to the 5′ end, Type IB binds covalently to the 3′ end of the break (Capranico et al. 2017; Cheng et al. 1998; Redinbo et al. 1998).
Type IB topoisomerases were found in some bacteria and Poxviruses and in eukaryotes (Champoux 2001; Forterre et al. 2007). All eukaryotes have at least one topoisomerase I (TOP1) for relaxing both negative and positive supercoils in front of moving polymerases during replication and transcription. Studies in yeast suggest that a single TOP1 may act in both the nuclear and mitochondrial genomes (de la Loza and Wellinger 2009; Wang et al. 1995). However, a second Type IB topoisomerase (TOP1MT) was identified in vertebrates, encoded in the nuclear genome. The TOP1MT exclusively localizes to mitochondria via a mitochondrial targeting sequence (MTS) at its N-terminal domain (Zhang et al. 2001). Among model organisms, TOP1 is essential for mouse and fruit fly development (Lee et al. 1993; Morham et al. 1996). TOP1MT seems to be dispensable for mouse development, but its absence causes increased negative supercoiling of mitochondrial DNA (mtDNA) and affects cellular energy metabolism (Douarre et al. 2012; Zhang et al. 2014) by interfering with biological processes such as liver regeneration (Khiati et al. 2015). Despite the biological relevance of both genes, their origin and molecular evolutionary patterns are still unknown.
In humans, the TOP1 gene is located in the chromosome region 20q12 (Juan et al. 1988) and encodes a 91 kDa protein with 765 amino acids. Two TOP1 pseudogenes have been identified on chromosomes 1 (ψ1-hTOP1) and 22 (ψ2-hTOP1) resulting from truncated mRNA transcripts of the active gene (Fig. 1A) (Yang et al. 1990). The TOP1MT gene maps to chromosome region 8q24 resulting in a 70 kDa protein with 601 amino acids (Zhang et al. 2001). Although TOP1 has 21 exons and TOP1MT has 14 exons, the terminal 13 exons are conserved between both genes (Zhang et al. 2004).
Organization of human nuclear (TOP1) and mitochondrial (TOP1MT) DNA topoisomerases I. A Multiple sequence alignment of human TOP1 and TOP1MT mRNA sequences and the two TOP1 pseudogenes identified in chromosomes 1 (ψ1-hTOP1) and 22 (ψ2-hTOP1). B Pairwise alignment of TOP1 and TOP1MT protein sequences, annotated with the most relevant protein domains and sites. C Illustrative representation of the human TOP1 protein structure with major domains highlighted
Considering the molecular structure and sequence conservation, TOP1 and TOP1MT proteins are organized into four distinct domains: N-terminal, Core, Linker and C-terminal domains (Fig. 1B, C). The N-terminal domain is poorly conserved across species and varies considerably when comparing both proteins. In particular, the TOP1 N-terminal is highly charged and relatively unstructured, being dispensable for the enzyme activity, mediates protein–protein interactions and includes nuclear localization signals (NLSs) (Alsner et al. 1992; Mo et al. 2000; Palle et al. 2008). The TOP1MT N-terminal is much shorter than that from TOP1 and includes a MTS. The core domain is highly conserved and contains essential catalytic residues, being connected to the C-terminal domain by a poorly conserved Linker region formed by an extended pair of α-helices. TOP1 forms a toroidal fold with two modules entrapping the DNA molecule, a capping module matching the first half of the core domain (CAP domain or core sub-domains I and II) and a catalytic module comprising the second half of the core domain (CAT domain or core sub-domain III), the Linker and the C-terminal domain (Redinbo et al. 1998; Stewart et al. 1998; Takahashi et al. 2022). The catalytic module includes several active sites relevant for the protein activity (Champoux 2001). The Hinge is a five-residue loop connecting the capping and catalytic modules whose flexibility permits the opening/closing of the enzyme and the entry of DNA (Takahashi et al. 2022). The C-terminal domain is highly conserved and includes the Tyr723 active site which forms a transient phosphotyrosyl linkage to one DNA strand, catalysing changes in DNA topology (Stewart et al. 1996).
Importantly, TOP1 is the target of the camptothecin family of anticancer agents that binds to and reversibly stabilizes the covalent TOP1-DNA complex, resulting in double stranded DNA breaks and apoptosis, preferentially in cancer cells that often overexpress TOP1 (Pommier 2006; Pommier et al. 2010). TOP1MT is also sensitive to camptothecin agents, but it is not an in vivo target due to the alkaline mitochondria matrix that inactivates the drug (Tua et al. 1997; Zhang et al. 2001; Zhang and Pommier 2008). However, several mutations in TOP1 are known to impact the efficacy of camptothecin (Chrencik et al. 2004; Cretaio et al. 2007; Saleem et al. 2000).
Previous works have compared Type IB topoisomerases from different species, but often focused on a specific section of the protein or explored only a few animal species [e.g., (Champoux 1998; Takahashi et al. 2022; Zhang et al. 2004)]. Here, we present a detailed examination of the evolutionary history of Type IB topoisomerases using a variety of animals that represent the main taxonomic groups of Chordata. In particular, we evaluated the molecular evolution and adaptation processes and the origin of the TOP1 and TOP1MT paralogues in vertebrates.
Material and Methods
TOPIB Sequences
TOPIB protein sequences from the main Metazoa phyla were retrieved from the NCBI non-redundant protein sequences (nr) database via the protein–protein BLAST (blastp) suite, using as query sequences from species close to the target taxonomic group (Supplementary Fig. S1). Short sequences with less than half of the average of TOPIB length were ignored since they often represent partial protein sequences derived from gaps in assembled genomes in which the contigs do not cover the complete genomic region. Possibly by the same reason, we fail to detect one or both the paralogues in the sequenced genome of some species.
Denisovan and Neanderthal TOP1 and TOP1MT sequences were downloaded from the UCSC Genome Browser (http://genome.ucsc.edu/) (Kent et al. 2002). All BAM reads for tracks Denisova and Neanderthal Cntgs matching the Human Mar. 2006 (NCBI36/hg18) chr20:39,090,876–39,186,540 (TOP1) and chr8:144,462,903–144,488,425 (TOP1MT) were downloaded. The BAM reads from each track were then reassembled against the human TOP1 (NC_000020.11) and TOP1MT (NC_000008.11) reference sequences using Geneious v2022.1.1 (http://www.geneious.com). We only considered a variable position in Denisovan and Neanderthal genomes when: (1) at least two reads overlap in that position; (2) the variant represents more than 75% of all the reads and (3) the difference is not at the end of a read. The variations between modern humans and Neanderthals were also confirmed in the assembly available at The Neandertal Genome Project (http://neandertal.ensemblgenomes.org).
TOPIB Sequence Alignments
The TOPIB protein sequences were aligned with the Geneious alignment in three datasets: Metazoa (n = 161), Chordata TOP1 (n = 48) and Chordata TOP1MT (n = 48). The conservation across the alignments was measured with the percentage of pairwise identity (PI) that compares base pairs at every site. The same species were used in the Chordata alignments to avoid biases and facilitate the comparison of results. The coding domain sequences (CDS) of the orthologues of human TOP1 (ENSG00000198900) and TOP1MT (ENSG00000184428) were obtained from the Ensembl Genome Server (Hunt et al. 2018).
Phylogenetic Analyses
We analysed the TOPIB duplication events in chordates with a phylogenetic tree built with 37 protein sequences from Cephalochordata, Tunicata and Vertebrata species, and considering Acanthaster planci and Strongylocentrotus purpuratus (Echinodermata) as outgroups. We used Gblocks 0.91b server, running on Phylogeny.fr (Dereeper et al. 2008), to remove poorly aligned positions under the settings for a less stringent selection (Castresana 2000; Talavera and Castresana 2007). The best-fitting amino acid substitution model of evolution (LG + I + G4 + F) was determined with ModelTest-NG (Darriba et al. 2020; Flouri et al. 2015). Next, we build a Bayesian phylogenetic tree with MrBayes v3.2.7a (Huelsenbeck and Ronquist 2001; Ronquist and Huelsenbeck 2003) running on the CIPRES Science Gateway v3.3 (Miller et al. 2010). The Metropolis-coupled Markov chain Monte Carlo (MCMC) process was set with two independent runs, each with four independent chains that ran simultaneously during 4,000,000 iterations. The average standard deviation of split frequencies of the final tree was 0.002339, indicating convergence among the independent runs. A burn-in value of 0.25 was applied following the program recommendation. The resulting phylogenetic tree was edited with FigTree v1.4.3 (http://tree.bio.ed.ac.uk/software/figtree).
Evaluation of Selection
Molecular adaptation signatures in TOP1 and TOP1MT protein-coding sequence alignments were evaluated with the nonsynonymous/synonymous substitution rates ratio (dN/dS) (Del Amparo et al. 2021; Jeffares et al. 2015). First, we selected the best-fitting substitution model of DNA evolution and reconstructed a maximum likelihood (ML) phylogenetic tree. Next, we estimated dN/dS under a ML method, considering the reconstructed phylogenetic tree, implemented in the evolutionary framework Hyphy (Kosakovsky Pond and Frost 2005; Kosakovsky Pond et al. 2020). In particular, we applied the single-likelihood ancestor counting (SLAC) method for the dN/dS estimation, which has an accuracy similar to that from other likelihood-based methods and includes statistical evaluations (Kosakovsky Pond and Frost 2005).
Template of TOPIB Protein Structure
We considered as an illustrative template of the human TOPIB protein structure, the protein structure of the Protein Data Bank (PDB) (Berman et al. 2000) with code 1A36 (Stewart et al. 1998). The structure was analysed with Mol* (Sehnal et al. 2021) and RCSB PDB.
Results and Discussion
TOP1 and TOP1MT Paralogues Originated in the First Round of Vertebrate Tetraploidization (1R)
Previous works have shown that TOPIB topoisomerases are ubiquitous in eukaryotes, and that only vertebrates have two TOPIB paralogues, named TOP1 and TOP1MT (Forterre et al. 2007; Zhang et al. 2004). Our extensive search for TOPIB genes in the genome of all available chordates only retrieved paralogues in the cyclostomes (jawless vertebrates) and gnathostomes (jawed vertebrates), confirming the previous claiming that TOPIB paralogues only occur in vertebrates (Zhang et al. 2004). Our phylogeny placed cephalochordates at the root of Chordata (Fig. 2). The Tunicata (Urochordata) and Vertebrata form a clade known as Olfactores (Delsuc et al. 2006; Putnam et al. 2008; Satoh et al. 2014). The fast-evolving Oikopleura dioica forms a particularly long branch, as we previously found for TOPIIA (Moreira et al. 2022).
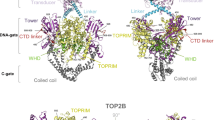
Phylogenetic analysis of TOPIB in chordates. Bayesian phylogenetic tree built with an alignment of 37 TOPIB protein sequences from chordates and considering two Echinodermata species as outgroup. The Bayesian posterior probabilities are shown on the internal nodes. The scale bar indicates substitutions per site. The putative occurrence of two rounds of tetraploidization (1R and 2R) is indicated
The timing of the duplication event that gave rise to both paralogues remains unclear, particularly considering that the origin of vertebrates is associated with several gene and genome duplication events. Two rounds of tetraploidization, known as 1R and 2R, are believed to have occurred early in vertebrate evolution (Ohno 2013; Smith and Keinath 2015; Van de Peer et al. 2009). The timing of the tetraploidization events is still a matter of debate, but it was recently proposed that 1R preceded the divergence between cyclostomes and gnathostomes and 2R only occurred in gnathostomes (Aase-Remedios and Ferrier 2021; Nakatani et al. 2021; Simakov et al. 2020). Previous works observed that vertebrata TOP1 and TOP1MT form two separate clusters (Forterre et al. 2007; Wang et al. 2009; Zhang et al. 2007), but were performed without sequences from cyclostomes. Our search for TOPIB genes in cyclostomes allowed us to retrieve two complete TOPIB sequences in two species, Petromyzon marinus and Eptatretus burgeri. We also noticed the presence of at least two paralogues in other cyclostomes (e.g., Lethenteron camtschaticum, Entosphenus tridentatus), but the genomic sequences were incomplete and thus were not used in the phylogenies. Therefore, it is likely that cyclostomes have at least two TOPIB paralogues, as observed in other vertebrates. In this concern, the TOPIB paralogues from P. marinus and E. burgeri did not cluster together in our phylogeny (Fig. 2). Instead, one pair clusters with TOP1MT sequences. Indeed, these two paralogues also display long branches, which are typical for the fast-evolving TOP1MT. Therefore, our analyses suggest that cyclostomes have a mitochondrial Type IB topoisomerase. The other pair of TOPIB paralogues from P. marinus and E. burgeri split from Gnathostomata TOP1 and TOP1MT at similar times. Our analysis is compatible with the idea that the duplication event that originated TOP1 and TOP1MT is related with the first round of tetraploidization (1R). In this situation, TOP1 and TOP1MT originated during the whole genome duplication in the early vertebrate evolution. The paralogues then diverged independently during the evolution of vertebrates, clustering in two separate branches (Fig. 2). The main difference between the phylogeny of the two genes is the placement of TOP1 from cyclostomes, which does not cluster with TOP1 from gnathostomes, as in the TOP1MT clade. Further analyses with additional sequences from Cyclostomata are necessary to better define the evolutionary history of these genes.
The specialization for acting on mtDNA may have occurred early in the radiation of vertebrates. In this concern, we previously identified that TOPIIA paralogues (TOP2A and TOP2B) present a different origin within chordates (Moreira et al. 2022). Here, we found that TOP2A and TOP2B paralogues from Cyclostomata cluster together in a separate branch from all Gnathostomata paralogues. Altogether, our findings suggest that the different classes of topoisomerases present different evolutionary histories in chordates.
Strong Purifying Selection Acting on TOP1 and TOP1MT
We estimated the dN/dS ratio to evaluate selection acting on TOP1 and TOP1MT paralogues of chordates (Table 1). We found that both genes present genetic signatures of negative (purifying) selection (dN/dS < 1), as noticed before in other topoisomerases (TOP3B, TOP2A, TOP2B) (Moreira et al. 2021, 2022). The paralogue pairs TOP1/TOP1MT and TOP2B (dN/dS = 0.156) / TOP2A (dN/dS = 0.238) (Moreira et al. 2022) presented higher dN/dS ratios than TOP3B (dN/dS = 0.076) (Moreira et al. 2022), which has no paralogue.
Paralogues can exhibit asymmetric rates of sequence evolution (Conant and Wagner 2003; Scannell and Wolfe 2008; Van de Peer et al. 2001). The strength of negative selection was higher in TOP1 (dN/dS = 0.154) than in TOP1MT (dN/dS = 0.307). Indeed, TOP1 also exhibits a lower diversity compared with TOP1MT (Table 1). The essential activity of TOP1 across species (Lee et al. 1993; Morham et al. 1996) in different biological processes can explain its relatively high conservation. On the other hand, TOP1MT presents the highest dN/dS ratio among all the topoisomerases studied by us (Moreira et al. 2021, 2022). Although it still evolved under negative selection, TOP1MT seems more permissive to accept amino acid changes than other topoisomerases. The higher diversity estimated in TOP1MT (in comparison to TOP1) can also be observed in the Chordata phylogeny, where TOP1MT branches are considerably longer than those for TOP1 (Fig. 2). The fast rate of change in TOP1MT can explain why finding orthologues for this gene is difficult. For example, the Ensembl genome browser only recognizes 77 orthologues for TOP1MT, in comparison with the 272 orthologues identified for TOP1 (accessed in April 2022). TOP1MT was also recognized as the only topoisomerase with highly frequent single nucleotide variants (SNVs) in the human population (Zhang et al. 2017). It was speculated that TOP1MT varies more than other topoisomerases due to several factors: (i) it is a nonessential gene under less constraints to mutate; (ii) it is in a subtelomeric end of a chromosome and/or (iii) it is a relatively recent gene under adaptation to its activity in mitochondria (Zhang et al. 2017). Thus, the observed pattern can be the result from a combination of those factors. Comparing with our previous results, TOP2B and TOP2A are more conserved than TOP1MT despite being also paralogues that originated early in vertebrate evolution (Moreira et al. 2022). Thus, we believe that these paralogues could be a good comparative model to study TOP1MT in future investigations.
Two Missense Mutations Identified in the Neandertals TOP1MT Linker Region
Neanderthals and Denisovans are extinct groups of hominins that inhabited Eurasia until around 40,000 years ago (Green et al. 2010; Reich et al. 2010). Previous works identified a few amino acid changes among modern humans and other hominins, some of which may have contributed to unique human traits (Green et al. 2010; Kuhlwilm and Boeckx 2019). Here, we searched for sequence differences in coding regions among modern human, Denisovan and Neanderthal TOP1 and TOP1MT genes. However, we did not identify polymorphic positions in TOP1 coding regions covered by Neanderthals or Denisovans sequence reads. On the contrary, we identified three nucleotide differences in the coding regions of TOP1MT (Table 2). A silent mutation in the CAP TOP1MT domain occurred in the human lineage. Next, two missense mutations were identified in the Neanderthal lineage. In particular, the mutations involved changes in two close amino acid positions (533 and 536) that belong to the Linker region (Fig. 3). Notice that the occurrence of missense mutations between modern humans and Neanderthals is rare (Green et al. 2010; Kuhlwilm and Boeckx 2019). When comparing present-day human and Neanderthals, Kuhlwilm and Boeckx (2019) identified 647 amino acid-changes in 571 genes. Among those genes, only 68 had two or more amino acid changes. Assuming that humans have 19,969 genes (Nurk et al. 2022), only 0.34% of those genes have more than one amino acid change, making it a rare event.
Structural conservation of TOPIB. The sequence identity plot was estimated from the 161 TOPIB protein sequences of the major chordate groups. The most conserved positions are indicated with brown bars, while the less conserved positions are shown using red bars. The sequence logo and an illustration of the protein structure for the highlighted regions are included (Color figure online)
Two mutations occurring in the same sequence read seems particularly improbable. However, we identified the mutations in several reads, including both our assembly and the assembly available at the Neandertal Genome Project (Supplementary Fig. S2). Moreover, we fail to align the Neandertal reads with any other available sequence in GenBank, including TOP1 gene and pseudogenes, which excludes a possible misplacement of reads from those regions in TOP1MT. The two mutations involved amino acids with different physicochemical properties. In particular, two glutamines (polar uncharged side chain) were replaced by an arginine and a lysine (positively charged, basic, side chain). These different properties could affect the protein function, but further experimental analyses are required to corroborate this possibility. We previously identified two missense mutations in TOP2A when comparing present-day humans and Neanderthals (Moreira et al. 2022). It is interesting to note that missense mutations were only identified in the two topoisomerases (TOP1MT and TOP2A) that are less conserved in chordates, which supports the credibility of the identified sequence differences. The sequencing of additional Neanderthal and Denisovan samples will allow us to confirm if these sequence variations were fixed among these species.
Relevant TOP1 and TOP1MT Sites for Catalytic Activities Tend to be Conserved Across Animals
The alignment of TOP1 and TOP1MT protein sequences from 48 representative chordate species confirms that TOP1MT (77% of pairwise sequence identity) is less conserved than TOP1 (sequence identity of 83.6%) (Fig. 3, Table 3). This result agrees with the long branches of TOP1MT in the Chordata phylogeny (Fig. 2) and its higher genetic diversity (Table 2). The N-terminal domain is the less conserved region in both proteins (sequence identities of 64.1% in TOP1 and 53.3% in TOP1MT), as noticed since the first studies on TOP1 (Champoux 1998, 2001; Stewart et al. 1996). The function of the N-terminal domain remains poorly understood, partially due to a lack of structural information. However, it is dispensable for the catalytic activity of the enzyme (Alsner et al. 1992), suggesting that it could accept mutations without compromising the protein activity. Moreover, the N-terminal domain mediates TOP1 interactions with other proteins (Czubaty et al. 2005). These protein–protein interactions might experience different co-evolution processes among species that could explain the poor sequence conservation of the domain. The protein–protein binding regions identified in N-terminal domains of TOP1 (NLSs) and TOP1MT (MTS) are also poorly conserved, possibly due to evolution driven by different species requirements. Only TOP1 NLS-II and NLS-IV are relatively conserved in chordates (Table 3).
The core domain (CAP, Hinge and CAT) is highly conserved due to its fundamental function on DNA binding during catalysis. We also found high conservation in the DNA-binding regions in other topoisomerases (Moreira et al. 2021, 2022), suggesting that these regions cannot accommodate changes due to maintaining the topoisomerase activity through a proper interaction with DNA. The CAT region is slightly more conserved than the CAP region, which agrees with the observation that only the CAT region is conserved in bacterial, viral and eukaryotic topoisomerases (Patel et al. 2010; Perry et al. 2006). The five-residue loop Hinge is conserved across metazoan (84.8% sequence identity), specifically the first two residues (TOP1 positions 428–429) that present the same amino acids in all the analysed species (Fig. 3). In addition, the tyrosine upstream of the Hinge (position 426) was also found conserved, in agreement with a previous work suggesting that this position interacts with the DNA duplex and guides the motion of the CAP domain upon DNA binding to enable the enzyme closing (Takahashi et al. 2022). Within the CAT region, we noticed that near the Linker there are two conserved stretches of around 20 amino acids that flank a poorly conserved region (Fig. 3). In particular, we identified a region with 6 amino acids AKVFRT (TOP1 reference positions 586–591) that is 100% conserved across all the 161 metazoan analysed species. This region includes several active sites. The CAP and CAT regions include sites conferring resistance to camptothecin and all of them are 100% conserved. The only variable sites conferring resistance to camptothecin were observed in the Linker (site 653, sequence identity of 77.2%) and C-terminal (site 729, sequence identity of 66.3%) regions.
We found that the Linker region is more variable than the surrounding core and C-terminal domains (Fig. 3, Table 3). The Linker consists of two long alpha helices connected by a short turn, forming an antiparallel coiled-coil configuration that protrudes away from the remainder of the enzyme (Stewart et al. 1998). We found that its conservation decreases with the increasing distance to the flanking domains and to the catalytic region of the enzyme (Fig. 3). The short turn at the end of the Linker (TOP1 positions 675–678) is extremely variable across species (21.4% of sequence identity), including some variation in length, suggesting that it can vary without affecting the protein function. The increase in conservation of the Linker in regions closer to the core of the enzyme indicates that amino acid replacements are less tolerated if they occur close to the catalytic region, possibly due to affecting the protein activity or the Linker connections to the DNA strand.
The C-terminal domain folds into a globular structure (Figs. 1C, 3) that includes the active-site nucleophile Tyr723 (Redinbo et al. 1998). This region also includes 8 residues near the Linker (718–722) with significant structural similarity with the bacteriophage family of DNA integrases (Redinbo et al. 1998). Our results confirmed previous observations about the high conservation of the C-terminal domain (Champoux 2001). In particular, we found that the 14 amino acids closer to the Linker (human TOP1 positions 715–728) are almost 100% conserved in all the analysed animal species (Fig. 3).
Conclusions
Type IB topoisomerases are widespread in the animal kingdom. Indeed, vertebrates present specialized topoisomerases to operate with the nuclear and mitochondrial genomes. However, little is known about its evolution and its genetic similarities among species. Here we analysed the molecular evolution of topoisomerases among a variety of animal species. Our phylogenetic investigation placed the event that originated the specialized TOP1 and TOP1MT proteins in the early evolution of vertebrates, possibly associated with whole-genome duplications. After the duplication event, the long-term evolution of both paralogues was primarily driven by strong purifying selection probably to maintain the protein function. However, we found that TOP1MT evolved much faster than TOP1 and other topoisomerases, perhaps related with its specific role within the mitochondria. The fast evolution of TOP1MT was also evident in the missense mutations detected in the Neanderthals, displaying a rare case of protein differences among hominids. Finally, comparison of topoisomerases among species showed that the relevant protein sites for catalytic activities are mainly conserved across animals, again probably caused by their relevant biological roles.
Code Availability
Not applicable.
Data Availability
Not applicable.
References
Aase-Remedios ME, Ferrier DEK (2021) Improved understanding of the role of gene and genome duplications in chordate evolution with new genome and transcriptome sequences. Front Ecol Evol 9:429
Alsner J, Svejstrup J, Kjeldsen E, Sørensen B, Westergaard O (1992) Identification of an N-terminal domain of eukaryotic DNA topoisomerase I dispensable for catalytic activity but essential for in vivo function. J Biol Chem 267:12408–12411
Berman HM et al (2000) The protein data bank. Nucleic Acids Res 28:235–242
Capranico G, Marinello J, Chillemi G (2017) Type i DNA topoisomerases. J Med Chem 60:2169–2192
Castresana J (2000) Selection of conserved blocks from multiple alignments for their use in phylogenetic analysis. Mol Biol Evol 17:540–552
Champoux JJ (1998) Domains of human topoisomerase I and associated functions. Prog Nucleic Acid Res Mol Biol 60:111–132
Champoux JJ (2001) DNA topoisomerases: structure, function, and mechanism. Annu Rev Biochem 70:369–413
Cheng C, Kussie P, Pavletich N, Shuman S (1998) Conservation of structure and mechanism between eukaryotic topoisomerase I and site-specific recombinases. Cell 92:841–850
Chrencik JE, Staker BL, Burgin AB, Pourquier P, Pommier Y, Stewart L, Redinbo MR (2004) Mechanisms of camptothecin resistance by human topoisomerase I mutations. J Mol Biol 339:773–784
Conant GC, Wagner A (2003) Asymmetric sequence divergence of duplicate genes. Genome Res 13:2052–2058
Cretaio E, Pattarello L, Fontebasso Y, Benedetti P, Losasso C (2007) Human DNA topoisomerase IB: structure and functions. Ital J Biochem 56:91–102
Czubaty A et al (2005) Proteomic analysis of complexes formed by human topoisomerase I. Biochim Et Biophys Acta (BBA)-Proteins Proteom 1749:133–141
Darriba D, Posada D, Kozlov AM, Stamatakis A, Morel B, Flouri T (2020) ModelTest-NG: a new and scalable tool for the selection of DNA and protein evolutionary models. Mol Biol Evol 37:291–294
de la Loza MD, Wellinger RE (2009) A novel approach for organelle-specific DNA damage targeting reveals different susceptibility of mitochondrial DNA to the anticancer drugs camptothecin and topotecan. Nucleic Acids Res 37:e26–e26
Del Amparo R, Branco C, Arenas J, Vicens A, Arenas M (2021) Analysis of selection in protein-coding sequences accounting for common biases. Brief Bioinform 22:bbaa431
Delsuc F, Brinkmann H, Chourrout D, Philippe H (2006) Tunicates and not cephalochordates are the closest living relatives of vertebrates. Nature 439:965–968
Dereeper A et al (2008) Phylogeny. fr: robust phylogenetic analysis for the non-specialist. Nucleic Acids Res 36:W465–W469
Douarre C et al (2012) Mitochondrial topoisomerase I is critical for mitochondrial integrity and cellular energy metabolism. PLoS ONE 7:e41094
Flouri T et al (2015) The phylogenetic likelihood library. Syst Biol 64:356–362
Forterre P, Gribaldo S, Gadelle D, Serre M-C (2007) Origin and evolution of DNA topoisomerases. Biochimie 89:427–446
Green RE et al (2010) A draft sequence of the Neandertal genome. Science 328:710–722
Huelsenbeck JP, Ronquist F (2001) MRBAYES: Bayesian inference of phylogenetic trees. Bioinformatics 17:754–755
Hunt SE et al (2018) Ensembl variation resources. Database 2018:bay 119
Jeffares DC, Tomiczek B, Sojo V, dos Reis M (2015) A beginners guide to estimating the non-synonymous to synonymous rate ratio of all protein-coding genes in a genome. Parasite genomics protocols. Springer, New York, pp 65–90
Juan C-C et al (1988) Human DNA topoisomerase I is encoded by a single-copy gene that maps to chromosome region 20q12-13.2. Proc Natl Acad Sci 85:8910–8913
Kent WJ, Sugnet CW, Furey TS, Roskin KM, Pringle TH, Zahler AM, Haussler D (2002) The human genome browser at UCSC. Genome Res 12:996–1006
Khiati S et al (2015) Lack of mitochondrial topoisomerase I (TOP1mt) impairs liver regeneration. Proc Natl Acad Sci 112:11282–11287
Kosakovsky Pond SL, Frost SD (2005) Not so different after all: a comparison of methods for detecting amino acid sites under selection. Mol Biol Evol 22:1208–1222
Kosakovsky Pond SL et al (2020) HyPhy 2.5—a customizable platform for evolutionary hypothesis testing using phylogenies. Mol Biol Evol 37:295–299
Kuhlwilm M, Boeckx C (2019) A catalog of single nucleotide changes distinguishing modern humans from archaic hominins. Sci Rep 9:1–14
Lee MP, Brown SD, Chen A, Hsieh T-s (1993) DNA topoisomerase I is essential in Drosophila melanogaster. Proc Natl Acad Sci 90:6656–6660
Miller MA, Pfeiffer W, Schwartz T (2010) Creating the CIPRES Science Gateway for inference of large phylogenetic trees. 2010 gateway computing environments workshop (GCE). Ieee, United States, pp 1–8
Mo Y-Y, Wang C, Beck WT (2000) A novel nuclear localization signal in human DNA topoisomerase I. J Biol Chem 275:41107–41113
Moreira F, Arenas M, Videira A, Pereira F (2021) Molecular evolution of DNA topoisomerase III beta (TOP3B) in metazoa. J Mol Evol 89:384–395
Moreira F, Arenas M, Videira A, Pereira F (2022) Evolutionary history of TOPIIA topoisomerases in animals. J Mol Evol 90:149–165
Morham SG, Kluckman KD, Voulomanos N, Smithies O (1996) Targeted disruption of the mouse topoisomerase I gene by camptothecin selection. Mol Cell Biol 16:6804–6809
Nakatani Y, Shingate P, Ravi V, Pillai NE, Prasad A, McLysaght A, Venkatesh B (2021) Reconstruction of proto-vertebrate, proto-cyclostome and proto-gnathostome genomes provides new insights into early vertebrate evolution. Nat Commun 12:1–14
Nurk S et al (2022) The complete sequence of a human genome. Science 376:44–53
Ohno S (2013) Evolution by gene duplication. Springer Science & Business Media, Germany
Palle K, Pattarello L, van der Merwe M, Losasso C, Benedetti P, Bjornsti M-A (2008) Disulfide cross-links reveal conserved features of DNA topoisomerase I architecture and a role for the N terminus in clamp closure. J Biol Chem 283:27767–27775
Patel A, Yakovleva L, Shuman S, Mondragón A (2010) Crystal structure of a bacterial topoisomerase IB in complex with DNA reveals a secondary DNA binding site. Structure 18:725–733
Perry K, Hwang Y, Bushman FD, Van Duyne GD (2006) Structural basis for specificity in the poxvirus topoisomerase. Mol Cell 23:343–354
Pommier Y (2006) Topoisomerase I inhibitors: camptothecins and beyond. Nat Rev Cancer 6:789–802
Pommier Y, Leo E, Zhang H, Marchand C (2010) DNA topoisomerases and their poisoning by anticancer and antibacterial drugs. Chem Biol 17:421–433
Pommier Y, Sun Y, Shar-yin NH, Nitiss JL (2016) Roles of eukaryotic topoisomerases in transcription, replication and genomic stability. Nat Rev Mol Cell Biol 17:703–721
Pommier Y, Nussenzweig A, Takeda S, Austin C (2022) Human topoisomerases and their roles in genome stability and organization. Nat Rev Mol Cell Biol. https://doi.org/10.1038/s41580-022-00452-3
Putnam NH et al (2008) The amphioxus genome and the evolution of the chordate karyotype. Nature 453:1064–1071
Redinbo MR, Stewart L, Kuhn P, Champoux JJ, Hol WG (1998) Crystal structures of human topoisomerase I in covalent and noncovalent complexes with DNA. Science 279:1504–1513
Reich D et al (2010) Genetic history of an archaic hominin group from Denisova Cave in Siberia. Nature 468:1053–1060
Ronquist F, Huelsenbeck JP (2003) MrBayes 3: Bayesian phylogenetic inference under mixed models. Bioinformatics 19:1572–1574
Saleem A, Edwards TK, Rasheed Z, Rubin EH (2000) Mechanisms of resistance to camptothecins. Ann N Y Acad Sci 922:46–55
Satoh N, Rokhsar D, Nishikawa T (2014) Chordate evolution and the three-phylum system. Proc R Soc b: Biol Sci 281:20141729
Scannell DR, Wolfe KH (2008) A burst of protein sequence evolution and a prolonged period of asymmetric evolution follow gene duplication in yeast. Genome Res 18:137–147
Sehnal D et al (2021) Mol* viewer: modern web app for 3D visualization and analysis of large biomolecular structures. Nucleic Acids Res 49:w431–w437
Simakov O et al (2020) Deeply conserved synteny resolves early events in vertebrate evolution. Nat Ecol Evol 4:820–830
Smith JJ, Keinath MC (2015) The sea lamprey meiotic map improves resolution of ancient vertebrate genome duplications. Genome Res 25:1081–1090
Stewart L, Ireton GC, Parker LH, Madden KR, Champoux JJ (1996) Biochemical and biophysical analyses of recombinant forms of human topoisomerase I (∗). J Biol Chem 271:7593–7601
Stewart L, Redinbo MR, Qiu X, Hol WG, Champoux JJ (1998) A model for the mechanism of human topoisomerase I. Science 279:1534–1541
Takahashi DT et al (2022) Topoisomerase I (TOP1) dynamics: conformational transition from open to closed states. Nat Commun 13:1–11
Talavera G, Castresana J (2007) Improvement of phylogenies after removing divergent and ambiguously aligned blocks from protein sequence alignments. Syst Biol 56:564–577
Tua A, Wang J, Kulpa V, Wernette C (1997) Mitochondrial DNA topoisomerase I of saccharomyces cerevisiae. Biochimie 79:341–350
Van de Peer Y, Taylor JS, Braasch I, Meyer A (2001) The ghost of selection past: rates of evolution and functional divergence of anciently duplicated genes. J Mol Evol 53:436–446
Van de Peer Y, Maere S, Meyer A (2009) The evolutionary significance of ancient genome duplications. Nat Rev Genet 10:725–732
Wang J, Kearney K, Derby M, Wernette CM (1995) On the relationship of the ATP-independent, mitochondrial-associated DNA topoisomerase of saccharomyces cerevisiae to the nuclear topoisomerase I. Biochem Biophys Res Commun 214:723–729
Wang X, Huang Y, Lavrov DV, Gu X (2009) Comparative study of human mitochondrial proteome reveals extensive protein subcellular relocalization after gene duplications. BMC Evol Biol 9:1–11
Yang G, Kunze N, Baumgärtner B, Jiang Z, Sapp M, Knippers R, Richter A (1990) Molecular structures of two human DNA topoisomerase I retrosequences. Gene 91:247–253
Zhang H, Pommier Y (2008) Mitochondrial topoisomerase I sites in the regulatory D-loop region of mitochondrial DNA. Biochemistry 47:11196–11203
Zhang H, Barceló JM, Lee B, Kohlhagen G, Zimonjic DB, Popescu NC, Pommier Y (2001) Human mitochondrial topoisomerase I. Proc Natl Acad Sci 98:10608–10613
Zhang H, Meng LH, Zimonjic DB, Popescu NC, Pommier Y (2004) Thirteen-exon-motif signature for vertebrate nuclear and mitochondrial type IB topoisomerases. Nucleic Acids Res 32:2087–2092
Zhang H, Meng L-H, Pommier Y (2007) Mitochondrial topoisomerases and alternative splicing of the human TOP1mt gene. Biochimie 89:474–481
Zhang H, Zhang Y-W, Yasukawa T, Dalla Rosa I, Khiati S, Pommier Y (2014) Increased negative supercoiling of mtDNA in TOP1mt knockout mice and presence of topoisomerases IIα and IIβ in vertebrate mitochondria. Nucleic Acids Res 42:7259–7267
Zhang H, Seol Y, Agama K, Neuman KC, Pommier Y (2017) Distribution bias and biochemical characterization of TOP1MT single nucleotide variants. Sci Rep 7:1–11
Funding
Open access funding provided by FCT|FCCN (b-on). This work was partially supported by a research grant to FM (SFRH/BD/131584/2017) from Fundação para a Ciência e a Tecnologia. MA was supported by the Spanish Ministerio de Ciencia e Innovación through the Grants [RYC-2015–18241] and [PID2019-107931GA-I00].
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
None to declare.
Ethical Approval
Not applicable.
Consent for Publication
Not applicable.
Consent to Participate
Not applicable.
Additional information
Handling editor: David Alvarez-Ponce.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Moreira, F., Arenas, M., Videira, A. et al. Evolution of TOP1 and TOP1MT Topoisomerases in Chordata. J Mol Evol 91, 192–203 (2023). https://doi.org/10.1007/s00239-022-10091-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00239-022-10091-z