Abstract
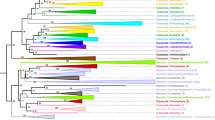
In C. elegans, four C2H2 zinc-finger proteins (ZIM-1, ZIM-2, ZIM-3, and HIM-8), which are arranged in tandem, mediate chromosome-specific pairing and synapsis during meiosis. The zim/him-8 genes from three Caenorhabditis species were contrasted in an effort to investigate the mechanisms driving their evolution. Here it is shown that the preservation of higher degree of sequence similarity in the N-terminal portion, particularly in several regions within the second exon between paralogous zim genes (especially between zim-1 and zim-3), is due to independent interparalogue gene conversions. However, the evolutionary force is not uniformly strong across species. The present data reveal that more frequent gene conversion events have occurred in C. elegans, whereas only gene conversions between zim-1 and zim-3 are detected in C. remanei. Although gene conversions are predicted to be present among zim-1, zim-2, and zim-3 in C. briggsae, the conversion tracts between zim-1/zim-2 and zim-2/zim-3 are very short. Moreover, positive selection analysis was performed on the basis of the significantly discordant phylogenies reconstructed using the N- and C-terminal sequences, respectively. Several codon sites located in the regions that are supposed not to have experienced gene conversions are predicted to be under the influence of positive selection. In comparison, stronger positive selection has acted on the C-terminal region relative to the N-terminal region. Thus, the zim/him-8 genes that evolve concertedly have also been shown to undergo adaptive diversifying selection.


Similar content being viewed by others
References
Anisimova M, Bielawski JP, Yang Z (2001) Accuracy and power of the likelihood ratio test in detecting adaptive molecular evolution. Mol Biol Evol 18:1585–1592
Beisswanger S, Stephan W (2008) Evidence that strong positive selection drives neofunctionalization in the tandemly duplicated polyhomeotic genes in Drosophila. Proc Natl Acad Sci USA 105:55447–55452
Blumenthal T, Gleason KS (2003) Caenorhabditis elegans operons: form and function. Nat Rev Genet 4:112–120
Blumenthal T, Evans D, Link CD, Guffanti A, Lawson D, Thierry-Mieg J, Thierry-Mieg D, Chiu WL, Duke K, Kiraly M, Kim SK (2002) A global analysis of Caenorhabditis elegans operons. Nature 417:851–854
Chen JM, Cooper DN, Chuzhanova N, Férec C, Patrinos GP (2007) Gene conversion: mechanism, evolution and human disease. Nat Genet 8:762–775
Cho S, Jin SW, Cohe A, Ellis RE (2004) A phylogeny of Caenorhabditis reveals frequent loss of introns during nematode evolution. Genome Res 14:1207–1220
Comeron JM (1995) A method for estimating the numbers of synonymous and nonsynonymous substitutions per site. J Mol Evol 41:1152–1159
Drouin G, Prat F, Ell M, Clarke GDP (1999) Detecting and characterizing gene conversions between multigene family members. Mol Biol Evol 16:1369–1390
Elliott B, Richardson C, Winderbaum J, Nickoloff JA, Jasin M (1998) Gene conversion tracts from double-strand break repair in mammalian cells. Mol Cell Biol 18:93–101
Englbrecht CC, Schoof H, Bohm S (2004) Conservation, diversification and expansion of C2H2 zinc finger proteins in the Arabidopsis thaliana genome. BMC Genomics 5:39
Fu YX (1997) Statistical tests of neutrality of mutations against population growth, hitchhiking, and background selection. Genetics 147:915–925
Garcia-Muse T, Boulton SJ (2007) Meiotic recombination in Caenorhabditis elegans. Chromosome Res 15:607–621
Goldstone HMH, Stegeman JJ (2006) A revised evolutionary history of the CYP1A subfamily: gene duplication, gene conversion, and positive selection. J Mol Evol 62:708–717
Holmes EC, Worobey M, Rambaut A (1999) Phylogenetic evidence for recombination in Dengue virus. Mol Biol Evol 16:405–409
Hurles ME, Willey D, Matthews L, Hussain SS (2004) Origins of chromosomal rearrangement hotspots in the human genome: evidence from the AZFa deletion hotspots. Genome Biol 5:R55
Innan H (2003) A two-locus gene conversion model with selection and its application to the human RHCE and RHD genes. Proc Natl Acad Sci USA 100:8793–8798
Katoh K, Kuma K, Miyata T, Toh H (2005) Improvement in the accuracy of multiple sequence alignment program MAFFT. Genome Inform 16:22–33
Kosakovsky Pond SL, Posada D, Gravenor MB, Woelk CH, Frost SDW (2006) Automated phylogenetic detection of recombination using a genetic algorithm. Mol Biol Evol 23:1891–1901
Kruithof EKO, Satta N, Liu JW, Dunoyer-Geindre S, Fish RJ (2007) Gene conversion limits divergence of mammalian TLR1 and TLR6. BMC Evol Biol 7:e148
Kumar S, Tamura K, Nei M (2004) MEGA3: integrated software for molecular evolutionary genetics analysis and sequence alignment. Brief Bioinform 5:150–163
McKim KS (2007) Meiotic pairing: a place to hook up. Curr Biol 17:R165–R168
Mita SD, Ronfort J, McKhann HI, Poncet C, Malki RE, Bataillon T (2007) Investigation of the demographic and selective forces shaping the nucleotide diversity of genes involved in nod factor signaling in Medicago truncatula. Genetics 177:2123–2133
Miyata T, Yasunaga T, Nishida T (1980) Nucleotide sequence divergence and functional constraint in mRNA evolution. Proc Natl Acad Sci USA 77:7328–7332
Moeller DA, Tiffin P (2005) Genetic diversity and the evolutionary history of plant immunity genes in two species of Zea. Mol Biol Evol 22:2480–2490
Mondragon-Palomino M, Gaut BS (2005) Gene conversion and the evolution of three Leucine-rich repeat gene families in Arabidopsis thaliana. Mol Biol Evol 22:2444–2456
Nelms BL, Hanna-Rose W (2006) C. elegans HIM-8 functions outside of meiosis to antagonize EGL-13 Sox protein function. Dev Biol 293:392–402
Noonan JP, Grimwood J, Schmutz J, Dickson M, Myers RM (2004) Gene conversion and the evolution of protocadherin gene cluster diversity. Genome Res 14:354–366
Ohta T (1997) Role of gene conversion in generating polymorphisms at major histocompatibility complex loci. Hereditas 127:97–103
Phillips CM, Dernburg AF (2006) A family of zinc-finger proteins is required for chromosome-specific pairing and synapsis during meiosis in C. elegans. Dev Cell 11:817–829
Phillips CM, Wong C, Bhalla N, Carlton PM, Weiser P, Meneely PM, Dernburg AF (2005) HIM-8 binds to the X chromosome pairing center and mediates chromosome-specific meiotic synapsis. Cell 123:1051–1063
Posada D (2002) Evaluation of methods for detecting recombination from DNA sequences: empirical data. Mol Biol Evol 19:708–717
Rozas J, Sánchez-DelBarrio JC, Messeguer X, Rozas R (2003) DnaSP, DNA polymorphism analyses by the coalescent and other methods. Bioinformatics 19:2496–2497
Sawyer S (1989) Statistical tests for detecting gene conversion. Mol Biol Evol 6:526–538
Sengupta S, Farheen S, Mukherjee N, Majumder PP (2007) Patterns of nucleotide sequence variation in ICAMI and TNF genes in twelve ethnic groups of India: roles of demographic history and natural selection. J Genet 86:225–239
Shannon M, Hamilton AT, Gordon L, Branscomb E, Stubbs L (2003) Differential expansion of zinc-finger transcription factor loci in homologous human and mouse gene clusters. Genome Res 13:1097–1110
Sun H, Nelms BL, Sleiman SF, Chamberlin HM, Hanna-Rose W (2007) Modulation of Caenorhabditis elegans transcription factor activity by HIM-8 and the related zinc-finger ZIM proteins. Genetics 177:1221–1226
Tajima F (1989) Statistical method for testing the neutral mutation hypothesis by DNA polymorphism. Genetics 123:585–595
Yang Z (2007) PAML4: A program package for phylogenetic analysis by maximum likelihood. Mol Biol Evol 24:1586–1591
Yang Z, Nielsen R, Goldman N, Pedersen AM (2000) Codon-substitution models for heterogeneous selection pressure at amino acid sites. Genetics 155:431–449
Zhang L (2008) Adaptive evolution and frequent gene conversion in the brain expressed X-linked gene family in mammals. Biochem Genet 46:293–311
Zhao Z, Hewett-Emmett D, Li WH (1998) Frequent gene conversion between human red and green opsin genes. J Mol Evol 46:494–496
Acknowledgments
I thank Dr. David Posada and the anonymous referees for their critical and constructive comments on an early draft of the manuscript. Likewise, great gratitude and appreciation are due to Professor Amitabh Joshi (chief editor of Journal of Genetics; Evolutionary and Organismal Biology Unit, Jawaharlal Nehru Centre for Advanced Scientific Research) and Dr. Xin’ai Zhao (Plant Molecular Biology Laboratory, International Rice Research Institute) for their help correcting the writing of the manuscript. This work was supported by an intramural fund from Zhejiang Forestry University (to Q. Liu).
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Liu, Q. Gene Conversion and Positive Selection Driving the Evolution of the Caenorhabditis ssp. ZIM/HIM-8 Protein Family. J Mol Evol 68, 217–226 (2009). https://doi.org/10.1007/s00239-009-9203-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00239-009-9203-z