Abstract
Thin films of conjugated polymer and enzyme can be used to unravel the interaction between components in a biosensor. Using artificial neural networks (ANNs) improves data interpretability and helps construct models with great capacity for classifying and processing information. The present work used kinetic data from the catalytic activity of urease immobilized in different conjugated polymers to create ANN models using time, substrate concentration, and absorbance as input variables since the models had absorbance in a posterior instant as output value to explore the predictivity of the ANNs. The performance of the models was evaluated by Pearson’s correlation coefficient (ρ) and mean squared error (MSE) values. After the learning process, a series of new experiments were performed to verify the generality of the models. As the main results, the best ANN model presented 0.9980 and 3.0736 × 10–5 for ρ and MSE, respectively. For the simulation step, intermediary values of substrate concentration were used. The mean absolute percentage error (MAPE) values were 3.34, 3.07, and 3.78 for 12 mM, 22 mM, and 32 mM concentrations, respectively. Overall, with the simulations, it was possible to ascertain the interpolatory capacity of the model, which has a learning mechanism based on absorbance and time as variables. Thus, the potential of ANNs would be in their use in pre-evaluations, helping to determine the substrate concentration at which there is higher catalytic activity or in determining the linear range of the sensor.
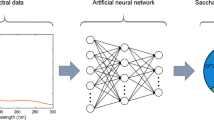
Graphical Abstract






Similar content being viewed by others
References
Arbter P, Widderich N, Utesch T, Hong Y, Zeng AP. Control of redox potential in a novel continuous bioelectrochemical system led to remarkable metabolic and energetic responses of Clostridium pasteurianum grown on glycerol. Microb Cell Fact. 2022;21(1):1–18. https://doi.org/10.1186/s12934-022-01902-5.
Ranganathan S, Mahesh S, Suresh S, Nagarajan A, Sen TZ, Yennamalli RM. Experimental and computational studies of cellulases as bioethanol enzymes. Bioengineered. 2022;13(5):14028–46. https://doi.org/10.1080/21655979.2022.2085541.
Fong M, Berrin JG, Paës G. Investigation of the binding properties of a multi-modular GH45 cellulase using bioinspired model assemblies. Biotechnol Biofuels. 2016;9(1):1–11. https://doi.org/10.1186/s13068-016-0428-y.
Tajik S, Ayoubi S, Nourbakhsh F. Prediction of soil enzymes activity by digital terrain analysis: comparing artificial neural network and multiple linear regression models. Environ Eng Sci. 2012;29(8):798–806. https://doi.org/10.1089/ees.2011.0313.
Li FF, Wang ZY, Zhao X, Xie E, Qiu J. Decomposition-ANN methods for long-term discharge prediction based on Fisher’s ordered clustering with MESA. Water Resour Manage. 2019;33(9):3095–110. https://doi.org/10.1007/s11269-019-02295-8.
Otchere DA, Ganat TOA, Gholami R, Ridha S. Application of supervised machine learning paradigms in the prediction of petroleum reservoir properties: comparative analysis of ANN and SVM models. J Petrol Sci Eng. 2021;200:108182. https://doi.org/10.1016/j.petrol.2020.108182.
Al-Sammarraie NA, Al-Mayali YMH, El-Ebiary YAB. Classification and diagnosis using back propagation artificial neural networks (ANN) algorithm. 2018 International Conference on Smart Computing and Electronic Enterprise, ICSCEE 2018 (2018) 1–5. https://doi.org/10.1109/ICSCEE.2018.8538383.
Ottaiano GY, da Cruz INS, da Cruz HS, Martins TD. Estimation of vaporization properties of pure substances using artificial neural networks. Chem Eng Sci. 2021;231:116324. https://doi.org/10.1016/j.ces.2020.116324.
Tijanić K, Car-Pušić D, Šperac M. Cost estimation in road construction using artificial neural network. Neural Comput Appl. 2020;32(13):9343–55. https://doi.org/10.1007/s00521-019-04443-y.
Shafi I, Ansari S, Din S, Jeon G, Paul A. Artificial neural networks as clinical decision support systems. Concurr Comput: Pract Exper. 2021;33(22):1–29. https://doi.org/10.1002/cpe.6342.
Tiryaki S, Aydin A. An artificial neural network model for predicting compression strength of heat treated woods and comparison with a multiple linear regression model. Constr Build Mater. 2014;62:102–8. https://doi.org/10.1016/j.conbuildmat.2014.03.041.
Barhmi S, Elfatni O, Belhaj I. Forecasting of wind speed using multiple linear regression and artificial neural networks. Energy Systems. 2020;11(4):935–46. https://doi.org/10.1007/s12667-019-00338-y.
Bekesiene S, Meidute-Kavaliauskiene I, Vasiliauskiene V. Accurate prediction of concentration changes in ozone as an air pollutant by multiple linear regression and artificial neural networks. Mathematics. 2021;9(4):1–21. https://doi.org/10.3390/math9040356.
Rodrigues RR, Aquino TP, Caseli L, Péres LO. Enzyme activity of thiophene-fluorene based-copolymer blended with urease in thin films. Colloids Surf A Physicochem Eng Asp. 2020;603:125139. https://doi.org/10.1016/j.colsurfa.2020.125139.
Barbosa CG, Caseli L, Péres LO. Conjugated polymers nanostructured as smart interfaces for controlling the catalytic properties of enzymes. J Colloid Interface Sci. 2016;476:206–13. https://doi.org/10.1016/j.jcis.2016.05.033.
Vidic J, Manzano M. Electrochemical biosensors for rapid pathogen detection. Curr Opin Electrochem. 2021;29:100750. https://doi.org/10.1016/j.coelec.2021.100750.
Janssen J, Lambeta M, White P, Byagowi A. Carbon nanotube-based electrochemical biosensor for label-free protein detection. Biosensors. 2019;9(4):144. https://doi.org/10.3390/bios9040144.
Tang L, Zeng G, Liu J, Xu X, Zhang Y, Shen G, Li Y, Liu C. Catechol determination in compost bioremediation using a laccase sensor and artificial neural networks. Anal Bioanal Chem. 2008;391(2):679–85. https://doi.org/10.1007/s00216-008-2049-1.
Torres-Gamez J, Rodriguez JA, Paez-Hernandez ME, Galan-Vidal CA. Application of multivariate statistical analysis to simultaneous spectrophotometric enzymatic determination of glucose and cholesterol in serum samples. International Journal of Analytical Chemistry. 2019;2019:4–9. https://doi.org/10.1155/2019/7532687.
Maleki N, Kashanian S, Maleki E, Nazari M. A novel enzyme based biosensor for catechol detection in water samples using artificial neural network. Biochem Eng J. 2017;128:1–11. https://doi.org/10.1016/j.bej.2017.09.005.
de Jesus CG, Rodrigues RR, Caseli L, Péres LO. Conducting polymers modulating the catalytic activity of urease in thin composite films. Colloids Surf, A. 2022;654:130–6. https://doi.org/10.1016/j.colsurfa.2022.130136.
Rodrigues RT, Nordi CFS, Junior JRS, Caseli L. Effect of interfering agents for urease immobilized in Langmuir-Blodgett films of controlled molecular architecture. Thin Solid Films. 2020;704:138043. https://doi.org/10.1016/j.tsf.2020.138043.
Menandro AS, Parolin GA, Barbosa CG, Faez R, Péres LO. New strategy to prepare luminescent blend by spin coating. Macromol Symp. 2019;383:1–5. https://doi.org/10.1002/masy.201800023.
Bento DC, Barbosa CG, Roncaselli LKM, Renzi W, Duarte JL, Olivati CA, Péres LO, Santana H. Thin films of poly[(9,9-dioctylfluorene)-co-thiophene] deposited on ITO by the Langmuir-Schaefer and Langmuir-Blodgett techniques. J Mater Sci Mater Electron. 2017;28:3875–83. https://doi.org/10.1007/s10854-016-6000-5.
Goto TE, Sakai A, Iost RM, Silva WC, Crespilho FN, Péres LO, Caseli L. Langmuir-Blodgett films based on poly(p-phenylene vinylene) and protein-stabilised palladium nanoparticles: implications in luminescent and conducting properties. Thin Solid Films. 2013;540:202–7. https://doi.org/10.1016/j.tsf.2013.05.106.
Péres LO, Fernandes MR, Garcia JR, Wang SH, Nart FC. Synthesis and characterization of chloro and bromo substituted p-phenylene vinylene homopolymers and alternating copolymers. Synth Met. 2006;156:529–36. https://doi.org/10.1016/j.synthmet.2006.01.014.
Yin Z, Tian B, Zhu Q, Duan C. Characterization and application of PVDF and its copolymer films prepared by spin-coating and Langmuir-Blodgett Method. Polymers. 2019;11(12):2033. https://doi.org/10.3390/polym11122033.
Günay ME, Nikerel IE, Oner ET, Kirdar B, Yildirim R. Simultaneous modeling of enzyme production and biomass growth in recombinant Escherichia coli using artificial neural networks. Biochem Eng J. 2008;42(3):329–35. https://doi.org/10.1016/j.bej.2008.08.002.
Akaike H. A new look at the statistical model identification. IEEE Trans Autom Control. 1974;19(6):716–23. https://doi.org/10.1109/TAC.1974.1100705.
Fogel DB. An information criterion for optimal neural network selection. Conf Rec - Asilomar Conf Circuits Syst Comput. 1991;2(5):998–1002. https://doi.org/10.1109/acssc.1990.523488.
Fregoso J, Gonzalez CI, Martinez GE. Optimization of convolutional neural networks architectures using PSO for sign language recognition. Axioms. 2021;10(3):139. https://doi.org/10.3390/axioms10030139.
Gevrey M, Dimopoulos I, Lek S. Review and comparison of methods to study the contribution of variables in artificial neural network models. Ecol Modell. 2003;160(3):249–64. https://doi.org/10.1016/S0304-3800(02)00257-0.
Bannoud MA, Gomes BP, Abdalla MC de SP, Freire MV, Anderola K, Martins TD, Silva CAM, Souza LFG, Braga MB. Mathematical modeling of drying kinetics of ground Açaí (Euterpe oleracea) kernel using artificial neural networks. Chem Pap. 2023. https://doi.org/10.1007/s11696-023-03142-2.
Tang L, Zeng G, Shen G, Zhang Y, Huang G, Li J. Simultaneous amperometric determination of lignin peroxidase and manganese peroxidase activities in compost bioremediation using artificial neural networks. Anal Chim Acta. 2006;579(1):109–16. https://doi.org/10.1016/j.aca.2006.07.021.
Sun Q, Zhang M, Mujumdar AS. Recent developments of artificial intelligence in drying of fresh food: a review. Crit Rev Food Sci Nutr. 2019;59(14):2258–75. https://doi.org/10.1080/10408398.2018.1446900.
Previdello BAF, de Carvalho FR, Tessaro AL, Souza VR, Hioka N. O pKa de indicadores ácido-base e os efeitos de sistemas coloidais. Quim Nova. 2006;29:600–6.
Walczak S, Cerpa N. Heuristic principles for the design of artificial neural networks. Inf Softw Technol. 1999;41(2):107–17. https://doi.org/10.1016/S0950-5849(98)00116-5.
Balas CE, Koç ML, Tür R. Artificial neural networks based on principal component analysis, fuzzy systems and fuzzy neural networks for preliminary design of rubble mound breakwaters. Appl Ocean Res. 2010;32(4):425–33. https://doi.org/10.1016/j.apor.2010.09.005.
Yogitha R, Mathivanan G. Performance analysis of transfer functions in an artificial neural network. 2018 International Conference on Communication and Signal Processing (ICCSP) 2018;393–397. https://doi.org/10.1109/ICCSP.2018.8524387.
Park ES, Shin JS. Free energy analysis of ω-transaminase reactions to dissect how the enzyme controls the substrate selectivity. Enzyme Microb Technol. 2011;49(4):380–7. https://doi.org/10.1016/j.enzmictec.2011.06.019.
Reetz MT, Carballeira JD, Peyralansv J, Höbenreich H, Maichele A, Vogel A. Expanding the substrate scope of enzymes: combining mutations obtained by CASTing. Chem Eur J. 2006;12(23):6031–8. https://doi.org/10.1002/chem.200600459.
Kargi F. Generalized rate equation for single-substrate enzyme catalyzed reactions. Biochem Biophys Res Commun. 2009;382(1):157–9. https://doi.org/10.1016/j.bbrc.2009.02.155.
Sigurdardóttir SB, Lehmann J, Ovtar S, Grivel JC, Negra MD, Kaiser A, Pinelo M. Enzyme immobilization on inorganic surfaces for membrane reactor applications: mass transfer challenges, enzyme leakage and reuse of materials. Adv Synth Catal. 2018;360(14):2578–607. https://doi.org/10.1002/adsc.201800307.
Rao SV, Anderson KW, Bachas LG. Oriented immobilization of proteins. Mikrochim Acta. 1998;128(3):127–43. https://doi.org/10.1007/bf01243043.
Acknowledgements
We gratefully acknowledge FAPESP (2020/04427-2 and 2014/50869-6), CNPq (310647/2020-7) and CAPES (Education Ministry) (23038.000776/201754) via the projects of the National Institute for Science and Technology on Organic Electronics (INEO) for the financial support and 88887.608793/2021-00. GAP receives a CAPES fellowship (Finance code 001). FTA receives a CAPES fellowship (Finance code 001).
Funding
This work was supported by CAPES (88887.608793/2021–00, 23038.000776/201754 and 001), FAPESP (grant number 2020/04427–2 and 2014/50869–6), and CNPq (310647/2020–7).
Author information
Authors and Affiliations
Contributions
Conceptualization; investigation; writing—original draft preparation: Cléber G. de Jesus. Writing—review and editing: Rebeca da R. Rodrigues, Carlos A. M. da Silva, Laura O. Péres. Supervision: Carlos A. M. da Silva and Laura O. Péres.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
de Jesus, C.G., da Rocha Rodrigues, R., da Silva, C.A.M. et al. Artificial neural networks in the modeling of the catalytic activity of a biosensor composed of conjugated polymers and urease. Anal Bioanal Chem 416, 1217–1227 (2024). https://doi.org/10.1007/s00216-023-05114-7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00216-023-05114-7