Abstract
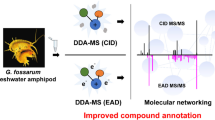
Mass spectrometry-based plant metabolomics allow large-scale analysis of a wide range of compounds and the discovery of potential new active metabolites with minimal sample preparation. Despite recent tools for molecular networking, many metabolites remain unknown. Our objective is to show the complementarity of collision cross section (CCS) measurements and calculations for metabolite annotation in a real case study. Thus, a systematic and high-throughput investigation of root, bark, branch, and leaf of the Gabonese plant Zhanthoxylum heitzii was performed through ultra-high performance liquid chromatography high-resolution tandem mass spectrometry (UHPLC-QTOF/MS). A feature-based molecular network (FBMN) was employed to study the distribution of metabolites in the organs of the plants and discover potential new components. In total, 143 metabolites belonging to the family of alkaloids, lignans, polyphenols, fatty acids, and amino acids were detected and a semi-quantitative analysis in the different organs was performed. A large proportion of medical plant phytochemicals is often characterized by isomerism and, in the absence of reference compounds, an additional dimension of gas phase separation can result in improvements to both quantitation and compound annotation. The inclusion of ion mobility in the ultra-high performance liquid chromatography mass spectrometry workflow (UHPLC-IMS-MS) has been used to collect experimental CCS values in nitrogen and helium (CCSN2 and CCSHe) of Zhanthoxylum heitzii features. Due to a lack of reference data, the investigation of predicted collision cross section has enabled comparison with the experimental values, helping in dereplication and isomer identification. Moreover, in combination with mass spectra interpretation, the comparison of experimental and theoretical CCS values allowed annotation of unknown features. The study represents a practical example of the potential of modern mass spectrometry strategies in the identification of medicinal plant phytochemical components.
Graphical abstract





Similar content being viewed by others
Data availability
All the data are described within the manuscript. The raw data and metadata analyzed during the current study are available from the corresponding author on request. The molecular networking jobs can be publicly accessed at https://gnps.ucsd.edu/ProteoSAFe/status.jsp?task=813b6121f4a840dbaa2c5d9a969f7056 for positive mode and https://gnps.ucsd.edu/ProteoSAFe/status.jsp?task=022432bae4e94699aa73b973c8214451 for negative mode.
References
Kokoska L, Kloucek P, Leuner O, Novy P. Plant-derived products as antibacterial and antifungal agents in human health care. Curr Med Chem. 2019. https://doi.org/10.2174/0929867325666180831144344.
Barbieri R, Coppo E, Marchese A, Daglia M, Sobarzo-Sánchez E, Nabavi SF, Nabavi SM. Phytochemicals for human disease: an update on plant-derived compounds antibacterial activity. Microbiol Res. 2017. https://doi.org/10.1016/j.micres.2016.12.003.
Fabiani R. Antitumoral properties of natural products. Volume 2, molecules, MDPI edition. 2020. https://doi.org/10.3390/molecules25030650.
Majolo F, Delwing LK, Marmitt DJ, Bustamante-Filho IC, Goettert MI. Medicinal plants and bioactive natural compounds for cancer treatment: important advances for drug discovery. Phytochem Lett. 2019. https://doi.org/10.1016/j.phytol.2019.04.003.
Wolfender JL, Nuzillard JM, Van Der Hooft JJ, Renault JH, Bertrand S. Accelerating metabolite identification in natural product research: toward an ideal combination of liquid chromatography–high-resolution tandem mass spectrometry and NMR profiling, in silico databases, and chemometrics. Anal Chem. 2018. https://doi.org/10.1021/acs.analchem.8b05112.
Luo MD, Zhou ZW, Zhu ZJ. The application of ion mobility-mass spectrometry in untargeted metabolomics: from separation to identification. J Anal Test. 2020. https://doi.org/10.1007/s41664-020-00133-0.
Schauer N, Fernie AR. Plant metabolomics: towards biological function and mechanism. Trends Plant Sci. 2006. https://doi.org/10.1016/j.tplants.2006.08.007.
Hall RD. Plant metabolomics: from holistic hope, to hype, to hot topic. New Phytol. 2006. https://doi.org/10.1111/j.1469-8137.2005.01632.x.
Šimura J, Antoniadi I, Široká J, Tarkowská DE, Strnad M, Ljung K, Novák O. Plant hormonomics: multiple phytohormone profiling by targeted metabolomics. Plant Phys. 2018. https://doi.org/10.1104/pp.18.00293.
Gachet MS, Schubert A, Calarco S, Boccard J, Gertsch J. Targeted metabolomics shows plasticity in the evolution of signaling lipids and uncovers old and new endocannabinoids in the plant kingdom. Sci Reports. 2017. https://doi.org/10.1038/srep41177.
Vanderplanck M, Glauser G. Integration of non-targeted metabolomics and automated determination of elemental compositions for comprehensive alkaloid profiling in plants. Phytochemistry. 2018. https://doi.org/10.1016/j.phytochem.2018.06.011.
Kårlund A, Hanhineva K, Lehtonen M, McDougall GJ, Stewart D, Karjalainen RO. Non-targeted metabolite profiling highlights the potential of strawberry leaves as a resource for specific bioactive compounds. J Sci Food Agric. 2017. https://doi.org/10.1002/jsfa.8027.
Perez de Souza L, Alseekh S, Naake T, Fernie A. Mass spectrometry-based untargeted plant metabolomics. Curr Protoc Plant Biol. 2019. https://doi.org/10.1002/cppb.20100.
Lisec J, Schauer N, Kopka J, Willmitzer L, Fernie AR. Gas chromatography mass spectrometry–based metabolite profiling in plants. Nat Protoc. 2006. https://doi.org/10.1038/nprot.2006.59.
Rivera-Pérez A, Romero-González R, Garrido Frenich A. Feasibility of applying untargeted metabolomics with GC-Orbitrap-HRMS and chemometrics for authentication of black pepper (Piper nigrum L.) and identification of geographical and processing markers. J Agric Food Chem. 2021. https://doi.org/10.1021/acs.jafc.1c01515.
Aydoğan C. Recent advances and applications in LC-HRMS for food and plant natural products: a critical review. Anal Bioanal Chem. 2020. https://doi.org/10.1007/s00216-019-02328-6.
Wolfender JL, Litaudon M, Touboul D, Ferreira QE. Innovative omics-bases approaches for prioritisation and targeted isolation of natural products—new strategies for drug discovery. Nat Prod Reports. 2019;36:855. https://doi.org/10.1039/c9np00004f.
Vinaixa M, Schymanski EL, Neumann S, Navarro M, Salek RM, Yanes O. Mass spectral databases for LC/MS-and GC/MS-based metabolomics: state of the field and future prospects. TrAC Trends Anal Chem. 2016. https://doi.org/10.1016/j.trac.2015.09.005.
Smith CA, O’Maille G, Want EJ, Qin C, Trauger SA, Brandon TR, Custodio DE, Abagyan R, Siuzdak G. METLIN: a metabolite mass spectral database. Ther Drug Monit. 2005. https://doi.org/10.1097/01.ftd.0000179845.53213.39.
Gerasimoska T, Ljoncheva M, Simjanoska M. MSL-ST: development of mass spectral library search tool to enhance compound identification. Proceedings of the 14th International Joint Conference on Biomedical Engineering Systems and Technologies. Volume 3: BIOINFORMATICS. 2021. https://doi.org/10.5220/0010424101950203
Nothias LF, Petras D, Schmid R, Dührkop K, Rainer J, Sarvepalli A, Protsyuk I, Ernst M, Tsugawa H, Fleischauer M, Aicheler F. Feature-based molecular networking in the GNPS analysis environment. Nat Methods. 2020. https://doi.org/10.1038/s41592-020-0933-6.
Olivon F, Elie N, Grelier G, Roussi F, Litaudon M, Touboul D. MetGem software for the generation of molecular networks based on the t-SNE algorithm. Anal Chem. 2018. https://doi.org/10.1021/acs.analchem.8b03099.
Hoffmann MA, Nothias LF, Ludwig M, Fleischauer M, Gentry EC, Witting M, Dorrestein PC, Dührkop K, Böcker S. High-confidence structural annotation of metabolites absent from spectral libraries. Nat Biotechnol. 2021. https://doi.org/10.1038/s41587-021-01045-9.
Baker ES, Livesay EA, Orton DJ, Moore RJ, Danielson WF III, Prior DC, Ibrahim YM, LaMarche BL, Mayampurath AM, Schepmoes AA, Hopkins DF. An LC-IMS-MS platform providing increased dynamic range for high-throughput proteomic studies. J Proteome Res. 2010. https://doi.org/10.1021/pr900888b.
Chalet C, Hollebrands B, Janssen HG, Augustijns P, Duchateau G. Identification of phase-II metabolites of flavonoids by liquid chromatography–ion-mobility spectrometry–mass spectrometry. Anal Bioanal Chem. 2018. https://doi.org/10.1007/s00216-017-0737-4.
Kanu AB, Dwivedi P, Tam M, Matz L, Hill HH Jr. Ion mobility–mass spectrometry. J Mass Spectrom. 2008. https://doi.org/10.1002/jms.1383.
Mason EA, Schamp HW Jr. Mobility of gaseous ions in weak electric fields. Ann Phys. 1958. https://doi.org/10.1016/0003-4916(58)90049-6.
Giles K, Pringle SD, Worthington KR, Little D, Wildgoose JL, Bateman RH. Applications of a travelling wave-based radio-frequency-only stacked ring ion guide. Rapid Commun Mass Spectrom. 2004. https://doi.org/10.1002/rcm.1641.
Stephan S, Jakob C, Hippler J, Schmitz OJ. A novel four-dimensional analytical approach for analysis of complex samples. Anal Bioanal Chem. 2016. https://doi.org/10.1007/s00216-016-9460-9.
Neumann EK, Migas LG, Allen JL, Caprioli RM, Van de Plas R, Spraggins JM. Spatial metabolomics of the human kidney using MALDI trapped ion mobility imaging mass spectrometry. Anal Chem. 2020. https://doi.org/10.1021/acs.analchem.0c02051.
Zhang X, Kew K, Reisdorph R, Sartain M, Powell R, Armstrong M, Quinn K, Cruickshank-Quinn C, Walmsley S, Bokatzian S, Darland E. Performance of a high-pressure liquid chromatography-ion mobility-mass spectrometry system for metabolic profiling. Anal Chem. 2017. https://doi.org/10.1021/acs.analchem.6b04628.
Dwivedi P, Wu P, Klopsch SJ, Puzon GJ, Xun L, Hill HH. Metabolic profiling by ion mobility mass spectrometry (IMMS). Metabolomics. 2008. https://doi.org/10.1007/s11306-007-0093-z.
Schroeder M, Meyer SW, Heyman HM, Barsch A, Sumner LW. Generation of a collision cross section library for multi-dimensional plant metabolomics using UHPLC-trapped ion mobility-MS/MS. Metabolites. 2020. https://doi.org/10.3390/metabo10010013.
Zhou Z, Luo M, Chen X, Yin Y, Xiong X, Wang R, Zhu ZJ. Ion mobility collision cross-section atlas for known and unknown metabolite annotation in untargeted metabolomics. Nat Commun. 2020. https://doi.org/10.1038/s41467-020-18171-8.
Colby SM, Thomas DG, Nuñez JR, Baxter DJ, Glaesemann KR, Brown JM, Pirrung MA, Govind N, Teeguarden JG, Metz TO, Renslow RS. ISiCLE: a quantum chemistry pipeline for establishing in silico collision cross section libraries. Anal Chem. 2019. https://doi.org/10.1021/acs.analchem.8b04567.
Little JL, Cleven CD, Brown SD. Identification of “known unknowns” utilizing accurate mass data and chemical abstracts service databases. J Am Soc Mass Spectrom. 2011. https://doi.org/10.1007/s13361-010-0034-3.
Little JL, Williams AJ, Pshenichnov A, Tkachenko V. Identification of “known unknowns” utilizing accurate mass data and ChemSpider. J Am Soc Mass Spectrom. 2012. https://doi.org/10.1007/s13361-011-0265-y.
McCullagh M, Goshawk J, Eatough D, Mortishire-Smith RJ, Pereira CA, Yariwake JH, Vissers JP. Profiling of the known-unknown Passiflora variant complement by liquid chromatography-ion mobility-mass spectrometry. Talanta. 2021. https://doi.org/10.1016/j.talanta.2020.121311.
Celma A, Sancho JV, Schymanski EL, Fabregat-Safont D, Ibáñez M, Goshawk J, Barknowitz G, Hernández F, Bijlsma L. Improving target and suspect screening high-resolution mass spectrometry workflows in environmental analysis by ion mobility separation. Environ Sci Technol. 2020. https://doi.org/10.1021/acs.est.0c05713.
Ollivier S, Fanuel M, Rogniaux H, Ropartz D. Molecular networking of high-resolution tandem ion mobility spectra: a structurally relevant way of organizing data in glycomics? Anal Chem. 2021. https://doi.org/10.1021/acs.analchem.1c01244.
Goodman CD, Austarheim I, Mollard V, Mikolo B, Malterud KE, McFadden GI, Wangensteen H. Natural products from Zanthoxylum heitzii with potent activity against the malaria parasite. Malar J. 2016;15(1):1–8. https://doi.org/10.1186/s12936-016-1533-x.
Ntchapda F, Maguirgue K, Adjia H, Etet PF, Dimo T. Hypolipidemic, antioxidant and anti—atherosclerogenic effects of aqueous extract of Zanthoxylum heitzii stem bark in diet—induced hypercholesterolemic rats. Asian Pac J Trop. 2015. https://doi.org/10.1016/S1995-7645(14)60344-8.
Mokondjimobe E, Miantezila Basilua J, Barkha S, Dzeufiet PD, Chenal H, Otsudi’andjeka JB, Bipolo S, Besse M, Mamadou G, Limas-Nzouzi N, Kamtchouing P. Fagaricine, a new immunorestorative phytomedicine from Zanthoxylum heitzii: preclinical and multicenter cohort clinical studies based on HIV-infected patients in six countries. Phytopharmacology. 2012;2(1):26–45.
Pieme CA, Santosh GK, Tekwu EM, Askun T, Aydeniz H, Ngogang JY, Bhushan S, Saxena AK. Fruits and barks extracts of Zanthozyllum heitzii a spice from Cameroon induce mitochondrial dependent apoptosis and Go/G1 phase arrest in human leukemia HL-60 cells. Biol Res. 2014. https://doi.org/10.1186/0717-6287-47-54.
Ntchapda F, Kakesse M, Fokam MA, Pancha OM, Abakar D, Dimo T. Evaluation of the diuretic effects of crude stem bark extraction of Zanthoxylum heitzii (Rutaceae) in Wistar rats. J Integr Med. 2015. https://doi.org/10.1016/S2095-4964(15)60188-1.
Tian Y, Zhang C, Guo M. Comparative study on alkaloids and their anti-proliferative activities from three Zanthoxylum species. BMC Complement Altern Med. 2017. https://doi.org/10.1186/s12906-017-1966-y.
Bongui JB, Blanckaert A, Elomri A, Seguin E. Constituents of Zanthoxylum heitzii (Rutaceae). Biochem Syst Ecol. 2005;33(8):845–7. https://doi.org/10.1016/j.bse.2004.12.019.
Ngouela S, Tsamo E, Connolly JD. Lignans and other constituents of Zanthoxylum heitzii. Phytochemistry. 1994. https://doi.org/10.1016/S0031-9422(00)90373-X.
Wangensteen H, Ho GT, Tadesse M, Miles CO, Moussavi N, Mikolo B, Malterud KE. A new benzophenanthridine alkaloid and other bioactive constituents from the stem bark of Zanthoxylum heitzii. Fitoterapia. 2016. https://doi.org/10.1016/j.fitote.2016.01.012.
Giles K, Williams JP, Campuzano I. Enhancements in travelling wave ion mobility resolution. Rapid Commun Mass Spectrom. 2011. https://doi.org/10.1002/rcm.5013.
Yu JS, Nothias LF, Wang M, Kim DH, Dorrestein PC, Kang KB, Yoo HH. Tandem mass spectrometry molecular networking as a powerful and efficient tool for drug metabolism studies. Anal Chem. 2022. https://doi.org/10.1021/acs.analchem.1c04925.
Wang M, Carver JJ, Phelan VV, Sanchez LM, Garg N, Peng Y, Nguyen DD, Watrous J, Kapono CA, Luzzatto-Knaan T, Porto C. Sharing and community curation of mass spectrometry data with Global Natural Products Social Molecular Networking. Nat Biotechnol. 2016. https://doi.org/10.1038/nbt.3597.
Katajamaa M, Miettinen J, Orešič M. MZmine: toolbox for processing and visualization of mass spectrometry based molecular profile data. Bioinformatics. 2006. https://doi.org/10.1093/bioinformatics/btk039.
Pluskal T, Castillo S, Villar-Briones A, Orešič M. MZmine 2: modular framework for processing, visualizing, and analyzing mass spectrometry-based molecular profile data. BMC Bioinforma. 2010. https://doi.org/10.1186/1471-2105-11-395.
Myers OD, Sumner SJ, Li S, Barnes S, Du X. One step forward for reducing false positive and false negative compound identifications from mass spectrometry metabolomics data: new algorithms for constructing extracted ion chromatograms and detecting chromatographic peaks. Anal Chem. 2017. https://doi.org/10.1021/acs.analchem.7b00947.
Mohimani H, Gurevich A, Shlemov A, Mikheenko A, Korobeynikov A, Cao L, Shcherbin E, Nothias LF, Dorrestein PC, Pevzner PA. Dereplication of microbial metabolites through database search of mass spectra. Nature Comm. 2018. https://doi.org/10.1038/s41467-018-06082-8.
Shannon P, Markiel A, Ozier O, Baliga NS, Wang JT, Ramage D, Amin N, Schwikowski B, Ideker T. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res. 2003. https://doi.org/10.1101/gr.1239303.
Bush MF, Campuzano ID, Robinson CV. Ion mobility mass spectrometry of peptide ions: effects of drift gas and calibration strategies. Anal Chem. 2012. https://doi.org/10.1021/ac3014498.
Gabelica V, Shvartsburg AA, Afonso C, Barran P, Benesch JLP, Bleiholder C, Bowers MT, Bilbao A, Bush MF, Campbell JL, Campuzano IDG, Causon T, Clowers BH, Creaser CS, De Pauw E, Far J, Fernandez-Lima F, Fjeldsted JC, Giles K, Groessl M, Hogan CJ Jr, Hann S, Kim HI, Kurulugama RT, May JC, McLean JA, Pagel K, Richardson K, Ridgeway ME, Rosu F, Sobott F, Thalassinos K, Valentine SJ, Wyttenbach T. Recommendations for reporting ion mobility mass spectrometry measurements. Mass Spectrom Rev. 2019. https://doi.org/10.1002/mas.21585.
Smith DP, Knapman TW, Campuzano I, Malham RW, Berryman JT, Radford SE, Ashcroft AE. Deciphering drift time measurements from travelling wave ion mobility spectrometry-mass spectrometry studies. Eur J Mass Spectrom. 2009. https://doi.org/10.1255/ejms.947.
Vainio MJ, Johnson MS. Generating conformer ensembles using a multiobjective genetic algorithm. J Chem Inf Model. 2007. https://doi.org/10.1021/ci6005646.
Puranen JS, Vainio MJ, Johnson MS. Accurate conformation-dependent molecular electrostatic potentials for high-throughput in silico drug discovery. J Comp Chem. 2010. https://doi.org/10.1002/jcc.21460.
Halgren TA. Merck molecular force field I Basis, form, scope, parameterization, and performance of MMFF94. J Comp Chem. 1996. https://doi.org/10.1002/(SICI)1096-987X(199604)17:5/6%3c490::AID-JCC1%3e3.0.CO;2-P.
Zeng X, Wang Z, Liu X, Chen M, Fahr A, Zhang K. Predicting the skin-permeating components of externally-applied medicinal herbs: application of a newly constructed linear free-energy relationship equation for human skin permeation. New J Chem. 2018. https://doi.org/10.1039/C8NJ00929E.
Liang M, Zhang W, Hu J, Liu R, Zhang C. Simultaneous analysis of alkaloids from Zanthoxylum nitidum by high performance liquid chromatography–diode array detector–electrospray tandem mass spectrometry. J Pharm Biomed Anal. 2006. https://doi.org/10.1016/j.jpba.2006.03.031.
Sumner LW, Amberg A, Barrett D, Beale MH, Beger R, Daykin CA, Fan TW, Fiehn O, Goodacre R, Griffin JL, Hankemeier T. Proposed minimum reporting standards for chemical analysis. Metabolomics. 2007. https://doi.org/10.1007/s11306-007-0082-2.
Bessonova IA, Faizutdinova ZS, Rashkes YV, Yunusov SY. Mass spectrometry of dubinidine and dubinine. Chem Nat Compd. 1970. https://doi.org/10.1007/BF00564249.
Mbaze LM, Lado JA, Wansi JD, Shiao TC, Chiozem DD, Mesaik MA, Choudhary MI, Lacaille-Dubois MA, Wandji J, Roy R, Sewald N. Oxidative burst inhibitory and cytotoxic amides and lignans from the stem bark of Fagara heitzii (Rutaceae). Phytochemistry. 2009. https://doi.org/10.1016/j.phytochem.2009.08.007.
Hanhineva K, Rogachev I, Aura AM, Aharoni A, Poutanen K, Mykkänen H. Identification of novel lignans in the whole grain rye bran by non-targeted LC–MS metabolite profiling. Metabolomics. 2012. https://doi.org/10.1007/s11306-011-0325-0.
Ross DH, Seguin RP, Krinsky AM, Xu L. High-throughput measurement and machine learning-based prediction of collision cross sections for drugs and drug metabolites. bioRxiv. 2021. https://doi.org/10.1101/2021.05.13.443945.
Hines KM, Ross DH, Davidson KL, Bush MF, Xu L. Large-scale structural characterization of drug and drug-like compounds by high-throughput ion mobility-mass spectrometry. Anal Chem. 2017. https://doi.org/10.1021/acs.analchem.7b01709.
Qing Z, Xu Y, Yu L, Liu J, Huang X, Tang Z, Cheng P, Zeng J. Investigation of fragmentation behaviours of isoquinoline alkaloids by mass spectrometry combined with computational chemistry. Sci Rep. 2020. https://doi.org/10.1038/s41598-019-57406-7.
Liu YL, Gao LL, Song TT, Guo T, Chang J. Two new sesquiterpenoid glycosides from the stems of Zanthoxylum armatum DC. Nat Prod Res. 2020. https://doi.org/10.1080/14786419.2019.1607332.
Winkler A, Puhl M, Weber H, Kutchan TM, Gruber K, Macheroux P. Berberine bridge enzyme catalyzes the six electron oxidation of (S)-reticuline to dehydroscoulerine. Phytochemistry. 2009. https://doi.org/10.1016/j.phytochem.2009.06.005.
Sheen WS, Tsai IL, Teng CM, Ko FN, Chen IS. Indolopyridoquinazoline alkaloids with antiplatelet aggregation activity from Zanthoxylum integrifoliolum. Planta Med. 1996. https://doi.org/10.1055/s-2006-957846.
Li W, Sun YN, Yan XT, Yang SY, Kim EJ, Kang HK, Kim YH. Coumarins and lignans from Zanthoxylum schinifolium and their anticancer activities. J Agric Food Chem. 2013. https://doi.org/10.1021/jf403479c.
Acknowledgements
The authors gratefully acknowledge Waters Corporation for the kind provision of the UNIFI software, Kevin Giles, PhD, for his advices, and the COBRA team for the support. The authors gratefully acknowledge Dr. Jean-Bernard Bongui for Z. heitzii providing.
Funding
This work has been partially supported by University of Rouen Normandy, INSA Rouen Normandy, the Centre National de la Recherche Scientifique (CNRS), European Regional Development Fund (ERDF), Labex SynOrg (ANR-11-LABX-0029), Carnot Institut I2C, the graduate school for research Xl-Chem (ANR-18-EURE-0020 XL CHEM), and by Region Normandie.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Calabrese, V., Schmitz-Afonso, I., Prevost, C. et al. Molecular networking and collision cross section prediction for structural isomer and unknown compound identification in plant metabolomics: a case study applied to Zhanthoxylum heitzii extracts. Anal Bioanal Chem 414, 4103–4118 (2022). https://doi.org/10.1007/s00216-022-04059-7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00216-022-04059-7