Abstract
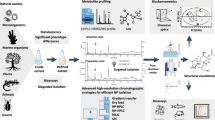
The evaluation of binding affinities between large biomolecules and small ligands is challenging and requires highly sensitive techniques. Microscale thermophoresis (MST) is an emerging biophysical technique used to overcome this limitation. This work describes the first MST binding method to evaluate binding affinities of small ligands to lipases from crude porcine pancreatic extracts. The conditions of the MST assay were thoroughly optimized to successfully evaluate the dissociation constant (Kd) between pancreatic lipases (PL) and triterpenoid compounds purified from oakwood. More precisely, the fluorescent labeling of PL (PL*) using RED-NHS dye was achieved via a buffer exchange procedure. The MST buffer was composed of 20 mM NaH2PO4 + 77 mM NaCl (pH 6.6) with 0.05% Triton-X added to efficiently prevent protein aggregation and adsorption, even when using only standard, uncoated MST capillaries. Storage at −20 °C ensured stability of PL* and its fluorescent signal. MST results showed that crude pancreatic extracts were suitable as a source of PL for the evaluation of binding affinities of small ligands. Quercotriterpenoside-I (QTT-I) demonstrated high PL* binding affinity (31 nM) followed by 3-O-galloylbarrinic acid (3-GBA) (500 nM) and bartogenic acid (BA) (1327 nM). To enrich the 50 kDa lipase responsible for the majority of hydrolysis activity in the crude pancreatic extracts, ammonium sulfate precipitation was attempted and its efficiency confirmed using capillary electrophoresis (CE)-based activity assays and HRMS. Moreover, to accurately explain enzyme modulation mechanism, it is imperative to complement binding assays with catalytic activity ones.
Graphical abstract






Similar content being viewed by others
References
Speer TW. Dissociation constant (Kd). In: Brady LW, Yaeger TE, editors. Encyclopedia of radiation oncology. Berlin, Heidelberg: Springer; 2013. https://doi.org/10.1007/978-3-540-85516-3_675.
Kairys V, Baranauskiene L, Kazlauskiene M, Matulis D, Kazlauskas E. Binding affinity in drug design: experimental and computational techniques. Expert Opin Drug Discovery. 2019;14:755–68. https://doi.org/10.1080/17460441.2019.1623202.
Mallik R, Yoo M, Briscoe C, Hage D. Analysis of drug-protein binding by ultrafast affinity chromatography using immobilized human serum albumin. J Chromatogr A. 2010;1217:2796–803. https://doi.org/10.1016/j.chroma.2010.02.026.
Zhang C, Rodriguez E, Bi C, Zheng X, Suresh D, Suh K, et al. High performance affinity chromatography and related separation methods for the analysis of biological and pharmaceutical agents. Analyst. 2018;143:374–91. https://doi.org/10.1039/C7AN01469D.
Hage DS. Analysis of biological interactions by affinity chromatography: clinical and pharmaceutical applications. Clin Chem. 2017;63:1083–93. https://doi.org/10.1373/clinchem.2016.262253.
Whately H. Basic principles and modes of capillary electrophoresis. In: Petersen J, Mohammad AA, editors. Clinical and forensic applications of capillary electrophoresis. Totowa: Humana Press; 2001. p. 21–58. https://doi.org/10.1007/978-1-59259-120-6.
Lipponen K, Tähkä S, Samuelsson J, Jauhiainen M, Metso J, Karhu GC, et al. Partial-filling affinity capillary electrophoresis and quartz crystal microbalance with adsorption energy distribution calculations in the study of biomolecular interactions with apolipoprotein E as interaction partner. Anal Bioanal Chem. 2014;406:4137–46. https://doi.org/10.1007/s00216-014-7821-9.
Ascoli GA, Domenici E, Bertucci C. Drug binding to human serum albumin: abridged review of results obtained with high-performance liquid chromatography and circular dichroism. Chirality. 2006;18:667–79. https://doi.org/10.1002/chir.20301.
Dawod M, Arvin NE, Kennedy RT. Recent advances in protein analysis by capillary and microchip electrophoresis. Analyst. 2017;142:1847–66. https://doi.org/10.1039/c7an00198c.
Sun L, Gidley MJ, Warren FJ. The mechanism of interactions between tea polyphenols and porcine pancreatic alpha-amylase: analysis by inhibition kinetics, fluorescence quenching, differential scanning calorimetry and isothermal titration calorimetry. Mol Nutr Food Res. 2017;61:1–13. https://doi.org/10.1002/mnfr.201700324.
Wu X, He W, Yao L, Zhang H, Liu Z, Wang W, et al. Characterization of binding interactions of (−)-epigallocatechin-3-gallate from green tea and lipase. J Agric Food Chem. 2013;61:8829–35. https://doi.org/10.1021/jf401779z.
Lin K, Wu G. Isothermal titration calorimetry assays to measure binding affinities in vitro. In: Hergovich A, editor. The Hippo pathway. Methods in molecular biology. New York: Humana Press; 2019. p. 257–72. https://doi.org/10.31826/9781463230777-019.
Bjelić S, Jelesarov I. A survey of the year 2007 literature on applications of isothermal titration calorimetry. J Mol Recognit. 2008;21:289–311. https://doi.org/10.1002/jmr.909.
Bhardwaj SK, Basu T. Study on binding phenomenon of lipase enzyme with tributyrin on the surface of graphene oxide array using surface plasmon resonance. Thin Solid Films. 2018;645:10–8. https://doi.org/10.1016/j.tsf.2017.10.021.
Sparks RP, Jenkins JL, Fratti R. Use of surface plasmon resonance (SPR) to determine binding affinities and kinetic parameters between components important in fusion machinery. Methods Mol Biol. 2019;1860:199–210. https://doi.org/10.1007/978-1-4939-8760-3_12.
Helmerhorst E, Chandler DJ, Nussio M, Mamotte CD. Real-time and label-free bio-sensing of molecular interactions by surface plasmon resonance: a laboratory medicine perspective. Clin Biochem Rev. 2012;33:161–73.
Sugiki T, Furuita K, Fujiwara T, Kojima C. Current NMR techniques for structure-based drug discovery. Molecules. 2018;23:1–27. https://doi.org/10.3390/molecules23010148.
Silva MS, Pietrobom D. Simplification of lipase design in the enzymatic kinetic resolution of amines by saturation transfer difference NMR. J Braz Chem Soc. 2016;27:1918–23. https://doi.org/10.5935/0103-5053.20160077.
Dias DM, Ciulli A. NMR approaches in structure-based lead discovery: recent developments and new frontiers for targeting multi-protein complexes. Prog Biophys Mol Biol. 2014;116:101–12. https://doi.org/10.1016/j.pbiomolbio.2014.08.012.
Vuignier K, Schappler J, Veuthey JL, Carrupt PA, Martel S. Drug-protein binding: a critical review of analytical tools. Anal Bioanal Chem. 2010;398:53–66. https://doi.org/10.1007/s00216-010-3737-1.
Baaske P, Wienken CJ, Reineck P, Duhr S, Braun D. Optical thermophoresis for quantifying the buffer dependence of aptamer binding. Angew Chem Int Ed. 2010;49:2238–41. https://doi.org/10.1002/anie.200903998.
Wienken CJ, Baaske P, Rothbauer U, Braun D, Duhr S. Protein-binding assays in biological liquids using microscale thermophoresis. Nat Commun. 2010;1:1–7. https://doi.org/10.1038/ncomms1093.
Schubert T, Längst G. Studying epigenetic interactions using microscale thermophoresis (MST). AIMS Biophys. 2015;2:370–80. https://doi.org/10.3934/biophy.2015.3.370.
Berleth M, Berleth N, Minges A, Hänsch S, Burkart RC, Stork B, et al. Molecular analysis of protein-protein interactions in the ethylene pathway in the different ethylene receptor subfamilies. Front Plant Sci. 2019;10:1–15. https://doi.org/10.3389/fpls.2019.00726.
Torres OB, Duval AJ, Sulima A, Antoline JFG, Jacobson AE, Rice KC, et al. A rapid solution-based method for determining the affinity of heroin hapten-induced antibodies to heroin, its metabolites, and other opioids. Anal Bioanal Chem. 2018;410:3885–903. https://doi.org/10.1007/s00216-018-1060-4.
Sloan J, Hakenjos JP, Gebert M, Ermakova O, Gumiero A, Stier G, et al. Structural basis for the complex DNA binding behavior of the plant stem cell regulator WUSCHEL. Nat Commun. 2020;11:1–16. https://doi.org/10.1038/s41467-020-16024-y.
Moon MH, Hilimire TA, Sanders AM, Schneekloth JS. Measuring RNA-ligand interactions with microscale thermophoresis. Biochemistry. 2018;57:4638–43. https://doi.org/10.1021/acs.biochem.7b01141.
Rubio MJ, Svobodová M, Mairal T, Schubert T, Künne S, Mayer G, et al. β -Conglutin dual aptamers binding distinct aptatopes. Anal Bioanal Chem. 2016;408:875–84. https://doi.org/10.1007/s00216-015-9179-z.
Syntia F, Nehmé R, Claude B, Morin P. Human neutrophil elastase inhibition studied by capillary electrophoresis with laser induced fluorescence detection and microscale thermophoresis. J Chromatogr A. 2016;1431:215–23. https://doi.org/10.1016/j.chroma.2015.12.079.
Lee N, Wiegand S. Thermophoretic micron-scale devices : practical approach and review. Entropy. 2020;22:1–24. https://doi.org/10.3390/e22090950.
Fayad S, Morin P, Nehmé R. Use of chromatographic and electrophoretic tools for assaying elastase, collagenase, hyaluronidase, and tyrosinase activity. J Chromatogr A. 2017;1529:1–28. https://doi.org/10.1016/j.chroma.2017.11.003.
Rainard JM, Pandarakalam GC, McElroy SP. Using microscale thermophoresis to characterize hits from high-throughput screening: a European lead factory perspective. SLAS Discov. 2018;23:225–41. https://doi.org/10.1177/2472555217744728.
Jerabek-Willemsen M, André T, Wanner R, Roth HM, Duhr S, Baaske P, et al. Microscale thermophoresis: interaction analysis and beyond. J Mol Struct. 2014;1077:101–13. https://doi.org/10.1016/j.molstruc.2014.03.009.
Jerabek-Willemsen M, Wienken CJ, Braun D, Baaske P, Duhr S. Molecular interaction studies using microscale thermophoresis. Assay Drug Dev Technol. 2011;9:342–53. https://doi.org/10.1089/adt.2011.0380.
Muntholib, Sulistyaningrum D, Subandi, Siti M. Identification of flavonoid isolates of papaya (Carica papaya L.) seed and their activity as pancreatic lipase inhibitors. AIP Conf Proc. 2020;2231:1–11. https://doi.org/10.1063/5.0003456.
Du X, Bai M, Huang Y, Jiang Z, Chen F, Ni H, et al. Inhibitory effect of astaxanthin on pancreatic lipase with inhibition kinetics integrating molecular docking simulation. J Funct Foods. 2018;48:551–7. https://doi.org/10.1016/j.jff.2018.07.045.
Xu-Dong H, Li-Lin S, Yun-Feng C, Yi-Nan W, Qi Z, Sheng-Quan F, et al. Pancreatic lipase inhibitory constituents from Fructus Psoraleae. Chin J Nat Med. 2020;18:369–78. https://doi.org/10.3724/SP.J.1009.2019.000000.
Hao G, Yang L, Mazsaroff I, Lin M. Quantitative determination of lipase activity by liquid chromatography-mass spectrometry. J Am Soc Mass Spectrom. 2007;18:1579–81. https://doi.org/10.1016/j.jasms.2007.05.019.
Al Hamoui Dit Banni G, Nasreddine R, Fayad S, Ngoc PC, Rossi JC, Leclercq L, et al. Screening for pancreatic lipase natural modulators by capillary electrophoresis hyphenated to spectrophotometric and conductometric dual detection. Analyst. 2021;146:1386–401. https://doi.org/10.1039/D0AN02234A.
Liu J, Ma R-T, Shi Y-P. An immobilization enzyme for screening lipase inhibitors from Tibetan medicines. J Chromatogr A. 1615;2020:1–8. https://doi.org/10.1016/j.chroma.2019.460711.
Tang Y, Li W, Wang Y, Zhang Y, Ji Y. Rapid on-line system for preliminary screening of lipase inhibitors from natural products by integrating capillary electrophoresis with immobilized enzyme microreactor. J Sep Sci. 2019;43:1003–10. https://doi.org/10.1002/jssc.201900523.
Fayad S, Nehmé R, Langmajerová M, Ayela B, Colas C, Maunit B, et al. Hyaluronidase reaction kinetics evaluated by capillary electrophoresis with UV and high-resolution mass spectrometry (HRMS) detection. Anal Chim Acta. 2017;951:140–50. https://doi.org/10.1016/j.aca.2016.11.036.
Nehmé H, Nehmé R, Lafite P, Routier S, Morin P. Human protein kinase inhibitor screening by capillary electrophoresis using transverse diffusion of laminar flow profiles for reactant mixing. J Chromatogr A. 2013;1314:298–305. https://doi.org/10.1016/j.chroma.2013.08.046 Accessed 9 Jan 2019.
Panuganti K, Nguyen M, Kshirsagar R. Obesity. In: StatPearls. Treasure Island (FL): StatPearls Publishing; 2020. https://www.ncbi.nlm.nih.gov/books/NBK459357/.
World Health Organization, Obesity and overweight 2020. https://www.who.int/news-room/fact-sheets/detail/obesity-and-overweight. Accessed 6 Dec 2020.
Liu TT, Liu XT, Chen QX, Shi Y. Lipase inhibitors for obesity: a review. Biomed Pharmacother. 2020;128:1–29. https://doi.org/10.1016/j.biopha.2020.110314.
Bonamichi B, Parente EB, dos Santos RB, Beltzhoover R, Lee J, Salles JEN. The challenge of obesity treatment: a review of approved drugs and new therapeutic targets. J Obes Eat Disord. 2018;4:1–10. https://doi.org/10.21767/2471-8203.100034.
Solomon LR, Nixon AC, Ogden L, Nair B. Orlistat-induced oxalate nephropathy: an under-recognised cause of chronic kidney disease. BMJ Case Rep. 2017. https://doi.org/10.1136/bcr-2016-218623.
Beyea MM, Garg AX, Weir MA. Does orlistat cause acute kidney injury ? Ther Adv Drug Saf. 2012;3:53–7. https://doi.org/10.1177/2042098611429985.
Sridhar SNC, Palawat S, Paul AT. Design, synthesis, biological evaluation and molecular modelling studies of indole glyoxylamides as a new class of potential pancreatic lipase inhibitors. Bioorg Chem. 2019;85:373–81. https://doi.org/10.1016/j.bioorg.2019.01.012.
Huang R, Zhang Y, Shen S, Zhi Z, Chen H, Chen S, et al. Antioxidant and pancreatic lipase inhibitory effects of flavonoids from different citrus peel extracts: an in vitro study. Food Chem. 2020;126785. https://doi.org/10.1016/j.foodchem.2020.126785.
Hu Q, Guan XQ, Song LL, Wang HN, Xiong Y, Liu JL, et al. Inhibition of pancreatic lipase by environmental xenoestrogens. Ecotoxicol Environ Saf. 2020;192:110305. https://doi.org/10.1016/j.ecoenv.2020.110305.
Segura RL, Betancor L, Palomo JM, Hidalgo A, Fernández-Lorente G, Terreni M, et al. Purification and identification of different lipases contained in PPL commercial extracts: a minor contaminant is the main responsible of most esterasic activity. Enzym Microb Technol. 2006;39:817–23. https://doi.org/10.1016/j.enzmictec.2006.01.007.
Gammacurta M, Waffo-Teguo P, Winstel D, Dubourdieu D, Marchal A. Isolation of taste-active triterpenoids from Quercus robur: sensory assessment and identification in wines and spirit. J Nat Prod. 2020;83:1611–22. https://doi.org/10.1021/acs.jnatprod.0c00106.
Marchal A, Prida A, Dubourdieu D. New approach for differentiating sessile and pedunculate oak: development of a LC-HRMS method to quantitate triterpenoids in wood. J Agric Food Chem. 2016;64:618–26. https://doi.org/10.1021/acs.jafc.5b05056.
Anthis NJ, Clore GM. Sequence-specific determination of protein and peptide concentrations by absorbance at 205 nm. Protein Sci. 2013;22:851–8. https://doi.org/10.1002/pro.2253.
Shahani KM, Khan IM, Chandan RC. Bovine pancreatic lipase. I. isolation, homogeneity, and characterization. J Dairy Sci. 1976;59:369–75. https://doi.org/10.3168/jds.S0022-0302(76)84214-2.
Algar WR. A brief introduction to traditional bioconjugate chemistry. In: Algar WR, Dawson P, Medintz I, editors. Chemoselective and bioorthogonal ligation reactions: concepts and applications, vol. 1. Weinheim, Germany: Wiley-VCH Verlag GmbH & Co. KGaA; 2017. p. 3–36.
Roy I, Gupta MN. Freeze-drying of proteins : some emerging concerns. Biotechnol Appl Biochem. 2004;39:165–77. https://doi.org/10.1042/BA20030133.
Heck AM, Yanovski JA, Calis KA. Orlistat, a new lipase inhibitor for the management of obesity. Pharmacotherapy. 2000;20:270–9. https://doi.org/10.1592/phco.20.4.270.34882.
Wingfield P. Protein precipitation using ammonium sulfate. Curr Protoc Protein Sci Appendix. 2001;3:1–10. https://doi.org/10.1002/0471140864.psa03fs13.
Polgár L. Basic kinetic mechanisms of proteolytic enzymes. In: Sterchi E, Stöcker W, editors. Proteolytic enzymes: tools and targets. Berlin Heidelberg: Springer-Verlag; 1999. p. 148–66. https://doi.org/10.1007/978-3-642-59816-6.
Acknowledgements
The authors would like to thank Dr. B. Claude for her aid and advices in lipase purification experiments, Dr. D. Da Silva for his aid and advices for MS analyses and lipase desalting, F. Coudray for the design and construction of the 3D printed scaffold for the C4D cell utilized in CE assays, and C. Maffre for technical support.
Funding
This work has been financially supported by the Région Centre Val de Loire (PhD fellowship of G. Al Hamoui Dit Banni) and the Labex SynOrg (ANR-11- LABX-0029).
Author information
Authors and Affiliations
Contributions
G. Al Hamoui Dit Banni and R. Nasreddine planned and performed MST and CE-based enzymatic experiments and analyzed the corresponding data; G. Al Hamoui Dit Banni and C. Colas carried out the HRMS assessment of PL purification; S. Fayad and A. Marchal planned and performed purifications and characterization of oakwood and wine extracts. R. Nehmé managed the project and data analysis. All the authors discussed the results and contributed to the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
ESM 1
(PDF 522 kb)
Rights and permissions
About this article
Cite this article
Al Hamoui Dit Banni, G., Nasreddine, R., Fayad, S. et al. Investigation of lipase-ligand interactions in porcine pancreatic extracts by microscale thermophoresis. Anal Bioanal Chem 413, 3667–3681 (2021). https://doi.org/10.1007/s00216-021-03314-7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00216-021-03314-7