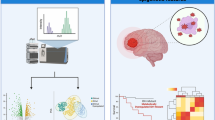
Abstract
Although anticancer drug resistance has been linked to high expression of P-glycoprotein and the enhanced DNA repair ability, the biochemical process and the underlying mechanisms of drug resistance are not clear. In order to clarify the biochemical mechanisms of drug resistance during anticancer drug treatment, we studied the metabolomics of MCF-7/S and MCF-7/Adr cell lines, the COC1 and COC1/DDP cell lines, including the metabolic pathways of multidrug-resistant tumor cells and the changes of endogenous substances in cells. The intracellular metabolites were profiled using ultra-performance liquid chromatography–tandem mass spectrometry (UPLC–MS/MS). In this study, 24 biomarkers in MCF-7/Adr cells and 15 biomarkers in COC1/DDP cells that are involved in some important metabolic pathways were putatively identified. Several metabolic pathways are changed in tumor cells showing drug resistance, such as protein synthesis pathways, cysteine synthesis, the glutamine metabolic pathway, and the ammonia cycle; the first of these are involved in the synthesis of some important proteins including membrane proteins, multidrug resistance-associated proteins, and P-glycoprotein (P-gp). Proteins related to drug resistance were overexpressed in multidrug-resistant tumor cells. These proteins depended on energy and play important roles in the emergence of drug resistance. The changes in glutathione and cysteine metabolic pathways showed that the cells can activate related metabolic pathways and reduce the cell apoptosis when they encounter oxidative damage. These findings indicate that drug resistance is likely associated with increased P-gp synthesis and reduced apoptosis of tumor cells.

Drug resistance was charactered in the changing of genomics and proteomics. Like enhancing DNA repair, reducing uptake, high P-g protein expression. Here, we studied the changes of metabolite pathway which could be also play an imported role in drug resistance








Similar content being viewed by others
References
Eiermann W, Pienkowski T, Crown J, Sadeghi S, Martin M, Chan A, et al. Phase III study of doxorubicin/cyclophosphamide with concomitant versus sequential docetaxel as adjuvant treatment in patients with human epidermal growth factor receptor 2-normal, node-positive breast cancer: BCIRG-005 trial. J Clin Oncol. 2011;29:3877–84.
Xu WH, Han M, Dong Q, Fu ZX, Diao YY, Liu H, et al. Doxorubicin-mediated radiosensitivity in multicellular spheroids from a lung cancer cell line is enhanced by composite micelle encapsulation. Int J Nanomed. 2012;7:2661–71.
Orban E, Mezo G, Schlage P, Csik G, Kulic Z, Ansorge P, et al. In vitro degradation and antitumor activity of oxime bond-linked daunorubicin-GnRH-III bioconjugates and DNA-binding properties of daunorubicin-amino acid metabolites. Amino Acids. 2011;41:469–83.
Susa M, Iyer AK, Ryu K, Hornicek FJ, Mankin H, Amiji MM, et al. Doxorubicin loaded polymeric nanoparticulate delivery system to overcome drug resistance in osteosarcoma. BMC Cancer. 2009;9:399.
Cao B, Li MJ, Zha WB, Zhao QJ, Gu RR, Liu LS, et al. Metabolomic approach to evaluating adriamycin pharmacodynamics and resistance in breast cancer cells. Metabolomics. 2013;9:960–73.
Scheffer GL, Wijngaard PL, Flens MJ, Izquierdo MA, Slovak ML, Pinedo HM, et al. The drug-resistance-related protein LRP is the human major vault protein. Nat Med. 1995;1:578–82.
Martin MB, Reiter R, Pham T, Avellanet YR, Camara J, Lahm M, et al. Estrogen-like activity of metals in MCF-7 breast cancer cells. Endocrinology. 2003;144:2425–36.
Juliano R, Ling V, Graves J. Drug-resistant mutants of Chinese hamster ovary cells possess an altered cell surface carbohydrate component. J Supramol Struc. 1976;4:521–6.
Alam J, Wicks C, Stewart D, Gong P, Touchard C, Otterbein S, et al. Mechanism of heme oxygenase-1 gene activation by cadmium in MCF-7 mammary epithelial cells role of p38 kinase and Nrf2 transcription factor. J Biol Chem. 2000;275:27694–702.
Pogribny IP, Filkowski JN, Tryndyak VP, Golubov A, Shpyleva SI, Kovalchuk O. Alterations ofmicroRNAs and their targets are associated with acquired resistance of MCF-7 breast cancer cells to cisplatin. Int J Cancer. 2010;127:1785–94.
Hill SM, Blask DE. Effects of the pineal hormone melatonin on the proliferation and morphological characteristics of human breast cancer cells (MCF-7) in culture. Cancer Res. 1998;48:6121–6.
Knowlden JM, Hutcheson IR, Jones HE, Madden T, Gee JM, Harper ME, et al. Elevated levels of epidermal growth factor receptor/c-erbB2 heterodimers mediate an autocrine growth regulatory pathway in tamoxifen-resistant MCF-7 cells. Endocrinology. 2003;144:1032–44.
O’Connell K, Prencipe M, O’Neill A, Corcoran C, Rani S, Henry M, et al. The use of LC-MS to identify differentially expressed proteins in docetaxel-resistant prostate cancer cell lines. Proteomics. 2012;12:2115–26.
Drabovich AP, Pavlou MP, Dimitromanolakis A, Diamandis EP. Quantitative analysis of energy metabolic pathways in MCF-7 breast cancer cells by selected reaction monitoring assay. Mol Cell Proteomics. 2012;11:422–34.
Kovalchuk O, Filkowski J, Meservy J, Ilnytskyy Y, Tryndyak VP, Chekhun CF, et al. Involvement of microRNA-451 in resistance of the MCF-7 breast cancer cells to chemotherapeutic drug doxorubicin. Mol Cancer Ther. 2008;7:2152–9.
Lutz NW, Franks SE, Frank MH, Pomer S, Hull WE. Investigation of multidrug resistance in cultured human renal cell carcinoma cells by 31P-NMR spectroscopy and treatment survival assays. Magma. 2005;18:144–61.
Klawitter J, Kominsky DJ, Brown JL, Klawitter J, Christians U, Leibfritz D, et al. Metabolic characteristics of imatinib resistance in chronic myeloid leukaemia cells. Br J Pharmacol. 2009;158:588–600.
Dewar BJ, Keshari K, Jeffries R, Dzeja P, Graves LM, Macdonald JM. Metabolic assessment of a novel chronic myelogenous leukemic cell line and an imatinib resistant subline by H NMR spectroscopy. Metabolomics. 2010;6:439–50.
Hirschhaeuser F, Sattler UG, Mueller-Klieser W. Lactate: a metabolic key player in cancer. Cancer Res. 2011;71:6921–5.
Wu G, Fang Y-Z, Yang S, Lupton JR, Turner ND. Glutathione metabolism and its implications for health. J Nutr. 2004;134:489–92.
Munoz-Pinedo C, El Mjiyad N, Ricci JE. Cancer metabolism: current perspectives and future directions. Cell Death Dis. 2012;3, e248.
Berg M, Vanaerschot M, Jankevics A, Cuypers B, Maes I, Mukherjee S, et al. Metabolic adaptations of Leishmania donovani in relation to differentiation, drug resistance, and drug pressure. Mol Microbiol. 2013;90:428–42.
Canuto GA, Castilho-Martins EA, Tavares M, Lopez-Gonzalvez A, Rivas L, Barbas C. CE-ESI-MS metabolic fingerprinting of Leishmania resistance to antimony treatment. Electrophoresis. 2012;33:1901–10.
Antti H, Fahlgren A, Nasstrom E, Kouremenos K, Sunden-Cullberg J, Guo Y, et al. Metabolic profiling for detection of staphylococcus aureus infection and antibiotic resistance. PLoS One. 2013;8, e56971.
Halama A, Moller G, Adamski J. Metabolic signatures in apoptotic human cancer cell lines. OMICS. 2011;15:325–35.
Zhao YY, Lei P, Chen DQ, Feng YL, Bai X. Renal metabolic profiling of early renal injury and renoprotective effects of Poria cocos epidermis using UPLC Q-TOF/HSMS/MSE. J Pharm Biomed. 2013;81-82:202-9.
Tedeschi P, Markert E, Gounder M, Lin H, Dvorzhinski D, Dolfi S, et al. Metabolic profiling for detection of Staphylococcus aureus infection and antibiotic resistance. Cell Death Dis. 2013;4, e877.
Caso G, McNurlan MA, McMillan ND, Eremin O, Garlick PJ. Tumour cell growth in culture: dependence on arginine. Clin Sci. 2004;107:371–9.
Brune DC, Hampton B, Kobayashi R, Leone JW, Linse KD, Pohl J, et al. ABRF ESRG 2006 study: Edman sequencing as a method for polypeptide quantitation. J Biomol Tech. 2007;18:306–20.
Brune K. Persistence of NSAIDs at effect sites and rapid disappearance from side-effect compartments contributes to tolerability. Curr Med Res Opin. 2007;23:2985–95.
Kalac P. Health effects and occurrence of dietary polyamines: a review for the period 2005-mid 2013. Food Chem. 2014;161:27–39.
Brzozowski T, Konturek PC, Konturek SJ, Brzozowska I, Kwiecien S, Hahn EG. Involvement of ornithine decarboxylase and polyamines in epidermal growth factor-induced recovery of gastric mucosa from gastric lesions provoked by stress. Regul Pept. 1998;74:73–84.
Ernestus RI, Rohn G, Schroder R, Els T, Klekner A, Paschen W, et al. Polyamine metabolism in brain tumours: diagnostic relevance of quantitative biochemistry. J Neurol Neurosurg Psychiatry. 2001;71:88–92.
Ekegren T, Gomes-Trolin C. Determination of polyamines in human tissues by precolumn derivatization with 9-fluorenylmethyl chloroformate and high-performance liquid chromatography. Anal Biochem. 2005;338:179–85.
Vuckovic D. Current trends and challenges in sample preparation for global metabolomics using liquid chromatography-mass spectrometry. Anal Bioanal Chem. 2012;403:1523–48.
Yao X, Lu C-D. Functional characterization of the potRABCD operon for spermine and spermidine uptake and regulation in Staphylococcus aureus. Curr Microbiol. 2014;69:75–81.
Xu L, Zhang XZ, Guo Y, Ren Q, Wu XX. Preparation and anti-oxidative effects of corn peptides. Chem Res Chin Univ. 2002;18:299–302.
Chen YP, Zhang J, Yang ZJ, Yu F, Li YQ. Relationship between nitric oxide accumulation, anti-oxidative system and freezing tolerance in the leaves of sabina during cold adaptation. J Anim Plant Sci. 2012;22:1133–41.
Swiderska-Kolacz G, Klusek J, Kolataj A. The effect of exogenous GSH, GSSG and GST-E on glutathione concentration and activity of selected glutathione enzymes in the liver, kidney and muscle of mice. Anim Sci Pap Rep. 2007;25:111–7.
Powis G. Anticancer drugs acting against signaling pathways. Curr Opin Oncol. 1995;7:554–9.
Pastore A, Federici G, Bertini E, Piemonte F. Analysis of glutathione: implication in redox and detoxification. Clin Chim Acta. 2003;333:19–39.
Dringen R, Pfeiffer B, Hamprecht B. Synthesis of the antioxidant glutathione in neurons: supply by astrocytes of CysGly as precursor for neuronal glutathione. J Neurosci. 1999;19:562–9.
Cullen KV, Davey RA, Davey MW. Verapamil-stimulated glutathione transport by the multidrug resistance-associated protein (MRP1) in leukaemia cells. Biochem Pharmacol. 2001;62:417–24.
Acknowledgements
This research was supported by the National Natural Science Foundation of China (No. 81373952, 81073040).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
No potential conflicts of interest were disclosed.
Additional information
Ruixing Zhang and Xiaoyu Zhuang contributed equally to this work.
Rights and permissions
About this article
Cite this article
Zhang, R., Zhuang, X., Zong, L. et al. Investigations on the cell metabolomics basis of multidrug resistance from tumor cells by ultra-performance liquid chromatography–mass spectrometry. Anal Bioanal Chem 408, 5843–5854 (2016). https://doi.org/10.1007/s00216-016-9696-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00216-016-9696-4