Abstract
Heparin and the related glycosaminoglycan, heparan sulfate, are polydisperse linear polysaccharides that mediate numerous biological processes due to their interaction with proteins. Because of the structural complexity and heterogeneity of heparin and heparan sulfate, digestion to produce smaller oligosaccharides is commonly performed prior to separation and analysis. Current techniques used to monitor the extent of heparin depolymerization include UV absorption to follow product formation and size exclusion or strong anion exchange chromatography to monitor the size distribution of the components in the digest solution. In this study, we used 1H nuclear magnetic resonance (NMR) survey spectra and NMR diffusion experiments in conjunction with UV absorption measurements to monitor heparin depolymerization using the enzyme heparinase I. Diffusion NMR does not require the physical separation of the components in the reaction mixture and instead can be used to monitor the reaction solution directly in the NMR tube. Using diffusion NMR, the enzymatic reaction can be stopped at the desired time point, maximizing the abundance of larger oligosaccharides for protein-binding studies or completion of the reaction if the goal of the study is exhaustive digestion for characterization of the disaccharide composition. In this study, porcine intestinal mucosa heparin was depolymerized using the enzyme heparinase I. The unsaturated bond formed by enzymatic cleavage serves as a UV chromophore that can be used to monitor the progress of the depolymerization and for the detection and quantification of oligosaccharides in subsequent separations. The double bond also introduces a unique multiplet with peaks at 5.973, 5.981, 5.990, and 5.998 ppm in the 1H-NMR spectrum downfield of the anomeric region. This multiplet is produced by the proton of the C-4 double bond of the non-reducing end uronic acid at the cleavage site. Changes in this resonance were used to monitor the progression of the enzymatic digestion and compared to the profile obtained from UV absorbance measurements. In addition, in situ NMR diffusion measurements were explored for their ability to profile the different-sized components generated over the course of the digestion.

DOSY spectra of intact (blue) and digested (red) heparin illustrating the differences in their diffusion coefficients.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Heparin and heparan sulfate (HS) are long-chain unbranched linear polysaccharides implicated in a wide range of biological functions, including cell growth and proliferation, inhibition of angiogenesis, and tumor growth and metastasis, as well as viral activity [1–3]. Heparin and HS are members of the glycosaminoglycan (GAG) family of carbohydrates, which also includes chondroitin sulfate, dermatan sulfate, and keratan sulfate. Heparin is a highly sulfated polysaccharide while HS is comprised of domains of high and low sulfation partitioned by domains of variable sulfation. Both polysaccharides have molecular weights ranging from 5 to 70 kDa and consist of the same disaccharide building blocks, a uronic acid residue α-linked (1 → 4) to d-glucosamine (GlcN). While HS is predominantly found as a proteoglycan at the cell surface, heparin is stored as a polysaccharide in secretory granules along with histamine and mast cell proteases [4].
Although heparin and HS are biosynthesized from the same disaccharide building block of glucuronic acid (GlcA) and N-acetylglucosamine, as a result of modifications that occur during chain extension, the end products are heterogeneous with regard to chain length, charge, and substitution pattern. In these GAGs, the uronic acid residue may be either α-linked (1 → 4) l-iduronic acid (IdoA) or β-linked (1 → 4) d-GlcA, which can be O-sulfo substituted at the C-2 position. The α-linked (1 → 4) glucosamine can be N-sulfonated, N-acetylated, or unmodified, with O-sulfo substitution possible at the GlcN C-6 and, less commonly, at the C-3 position [5]. Heparin is structurally simpler than HS with approximately 70% of the disaccharide subunits being IdoA(2S)-(1 → 4)-GlcNS(6S), commonly labeled as disaccharide IS [4]. Because of their structural complexity and heterogeneity, prior to analysis, heparin and HS are typically exhaustively degraded enzymatically or chemically to their disaccharide building blocks, given in Fig. 1. However, much of the current understanding of heparin structure–activity relationships is obtained using larger oligosaccharides.
Much of the interest in characterizing heparin structure is focused on the determination of sequences that interact with proteins and peptides. The interaction responsible for heparin’s anticoagulant activity is the most studied and best understood [6]. Binding of a unique heparin pentasaccharide to antithrombin III (AT-III) induces a conformation change in AT-III enhancing its affinity for thrombin and coagulation factors IXa and Xa. The heparin pentasaccharide sequence responsible for binding to AT-III is present in only ∼30% of the intact heparin chains from porcine-derived heparin [7, 8]. Many other heparin and HS oligosaccharide sequences have been identified that bind site specifically to a variety of proteins and peptides modulating their function.
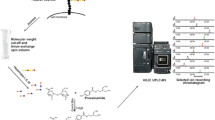
As mentioned above, a common approach to characterizing heparin structure and identifying binding motifs is to enzymatically or chemically depolymerize the heparin biopolymer into smaller oligosaccharides which are isolated, structurally characterized, and used in protein-binding experiments. The availability of the heparin lyase enzymes heparinase I, II, and III, isolated from Flavobacterium heparinum, has greatly facilitated the analysis of both heparin and HS [9]. The individual heparinase enzymes cleave unique glycosidic linkages, allowing heparin and HS to be degraded to specific oligosaccharide fragments [10]. Heparinase cleavage introduces a Δ4,5-unsaturated double bond via a β-elimination reaction at the non-reducing end uronic acid residue of the newly generated saccharide, generating a chromophore that can be detected by UV absorbance at 232 nm [11]. Chemical methods have also been established for the depolymerization of heparin. These methods, used commercially for the production of low molecular weight heparin, include nitrous acid treatment [12, 13] (dalteparin), base-catalyzed β-elimination [14, 15] (enoxaparin), peroxide treatment [16] (ardeparin), and periodate oxidation [17] (centaxarin).
Following either enzymatic or chemical depolymerization, analysis of GAG digests typically proceeds via the physical separation of the digest components based on differences in their size and charge. UV and fluorescence detection work well for the identification of GAGs at the disaccharide level where standards are available; however, they do not provide structural information for larger components. In cases where larger oligosaccharides are targeted, digest solutions are generally size-separated by size exclusion chromatography (SEC) to isolate corresponding disaccharide, tetrasaccharide, hexasaccharide, and larger oligosaccharide fractions which are further purified to individual oligosaccharides using strong anion exchange (SAX) [18–21] or reverse-phase ion-pair high-performance liquid chromatography (HPLC) [22, 23]. Purified oligosaccharides can then be structurally characterized by nuclear magnetic resonance (NMR) spectroscopy and mass spectrometry (MS).
In this study, we used 1H NMR spectra and diffusion NMR results in conjunction with UV absorbance at 232 nm to monitor the enzymatic depolymerization of heparin using heparinase I. Diffusion NMR, more specifically diffusion-ordered spectroscopy (DOSY) [24], has recently been used in the analysis of intact heparin samples to evaluate purity and to identify both natural and intentional contaminants of heparin [25, 26]. As demonstrated herein, diffusion NMR allows the investigator to follow the extent of the enzymatic reaction non-invasively and quench the digestion at the desired endpoint.
Experimental
Materials and reagents
Porcine intestinal mucosa heparin sodium salt grade 1-A, calcium acetate hydrate, ammonium bicarbonate, heparin disaccharide ΔUA(2S)-(1 → 4)-GlcNS(6S) (IS) sodium salt, and tris(hydroxymethyl)aminomethane (Tris) were obtained from Sigma Chemical Company (St. Louis, MO). Heparin disaccharide (IS) is the product of the digestion of heparin and heparan sulfate with heparinase I and II enzymes. Hydrochloric acid (HCl), tert-butanol, and sodium hydroxide were obtained from Fisher Scientific Co. (Fair Lawn, NJ). Heparinase I enzyme (EC 4.2.2.7) isolated from F. heparinum was purchased from IBEX Technologies Inc. (Montreal, Quebec). HPLC-grade water was purchased from Burdick and Jackson (Muskegon, MI). Deuterium oxide (D, 99.9%) was obtained from Cambridge Isotope Laboratories (Andover, MA).
Enzymatic depolymerization of heparin
Digestion of 1 g heparin was carried out in 50 mL of 100 mM Tris buffer, pH 6.8, containing 2.5 mM calcium acetate. The enzyme heparinase I (0.5 IU) was added to the mixture and incubated at 28°C in a water bath for 66 h. One international unit (IU) of the enzyme produces 1 μmol of unsaturated uronic acid from porcine mucosal heparin per min at 30°C and pH 7.0. The water bath temperature was controlled using a Fisher Scientific Isotemp 1013S scientific temperature regulator. The enzymatic reaction was quenched by placing the reaction vessel into boiling water for 5 min. The depolymerization solution was then lyophilized and reconstituted into 15 mL of the separation buffer (0.5 M NH4HCO3) prior to SEC separation.
Monitoring the progress of the enzymatic depolymerization with UV absorption
During the 66-h enzymatic depolymerization, UV absorption measurements were taken at regular intervals (ranging from 15 min to 6 h) to monitor the progress of the enzymatic reaction. UV measurements were performed at a wavelength of 232 nm using a Thermo Scientific NanoDrop 2000 spectrophotometer (Wilmington, DE). The NanoDrop has a 1-mm path length requiring only 1 μL of sample. For absorption measurements above 1 AU, samples were diluted by a factor of 2–4 times with the depolymerization buffer prior to analysis. At 232 nm, the molar extinction coefficient for the monounsaturated disaccharide is 5,500 M−1 cm−1 [11]. Graphs were prepared using Origin 7.5 (OriginLab Northampton, MA).
SEC fractionation
The heparin-derived oligosaccharides were size-fractionated on a column (200 cm × 3.2 cm) packed with Bio-Rad Bio-Gel P-10 resin fine (Bio-Rad Laboratories Hercules, CA) and eluted with 0.5 M NH4HCO3 at a flow rate of 0.08 mL/min. The total volume contained in each of the fractions collected was 4.5 mL. The progress of the separation was monitored offline by UV absorption measurements at 232 nm using the NanoDrop spectrophotometer as described above. Following SEC, similar-sized fractions were pooled and stored as a lyophilized powder at −20 °C for future use.
1H NMR experiments
Solutions for NMR analysis were prepared by adding 50 μL of D2O and 8 μL of 100 mM tert-butanol in D2O to 550 μL of the depolymerization solution in a 5-mm NMR tube. Solutions were mixed by gently tapping the tube. 1H NMR spectra were recorded using a Bruker Avance spectrometer operating at 599.84 MHz. All experiments were performed using a broadband inverse probe with x-, y-, and z-gradients. Chemical shifts were referenced to tert-butanol (1.253 ppm) added as an internal reference. For each 1H NMR spectrum, 64 transients were collected into 26,624 data points using a 6,009-Hz spectral window and a 2-s relaxation delay. NMR spectra were acquired using the 1D wet pulse program (WET) included with the standard Topspin release version 1.3 [27]. The automated Bruker shape tool was used to create the sinc pulse for selective excitation of the H2O resonance. 1H NMR spectra were processed using the academic edition of the ACD/Labs 1D NMR processing software, version 12.01. Prior to Fourier transformation, the free induction decays (FIDs) were zero filled to 65,536 points and apodized by multiplication with an exponential function equivalent to 1-Hz line broadening. The reaction solution was maintained at a temperature of 28 °C using the temperature control unit of the NMR spectrometer. The spectrometer was initially shimmed using a volume-matched solution of tert-butanol in the depolymerization buffer, with the reaction solution used for final optimization of the shims. The first spectrum was acquired at 10 min following a quick shim and optimization of the sinc-shaped pulses for WET solvent suppression. Subsequent spectra were manually initiated at intervals of 10 min for the first hour of the digestion to a total interval time of 8 h. From the 1H NMR spectra acquired throughout the course of the digestion, the peak area of the N-acetyl resonance was fit using the ACD NMR processing software and the calculated areas were normalized to the fitted area of the tert-butanol resonance. These normalized areas were used to compare the relative intensities between the subsequent spectra.
The calibration of the z-gradient coil used for the diffusion measurements was checked using the known diffusion coefficient for lysozyme (1.07 × 10−11 m2/s) at 20 °C and a concentration of 5 mg/mL prepared in D2O [28]. Diffusion spectra were acquired using the stimulated echo experiment with bipolar gradients (stebpgp1s). Diffusion (Δ) and gradient pulse times (δ) were optimized for each time point using a one-dimensional version of the stimulated echo pulse program, stepgp1s1d. Gradient amplitudes were incremented as a square dependence from 5% to 95% into 12 or 18 gradient increments. Following acquisition, the FIDs were apodized by multiplication with an exponential function equivalent to 1.5-Hz line broadening and baseline-corrected using a first-order polynomial prior to DOSY processing using the DOSY Toolbox software [29]. Reference deconvolution was performed on a few spectra but resulted in significant errors in the fitting of the decay plots and was therefore not used in the processing.
Because the diffusion spectra were acquired in a solution of 90% water, it was necessary to eliminate the first spectrum corresponding to a gradient field strength of 1.686 G cm−1 before DOSY processing. By eliminating the spectrum for the first gradient increment, we were able to drastically attenuate the water peak, permitting the measurement of the diffusion coefficients for the nearby H-4 and anomeric proton resonances. To simplify the DOSY analysis, processing was carried out on individual spectral regions of interest corresponding to the H-4, anomeric, and N-acetyl resonances. We found this helpful especially when processing later experiments, after the first day of the digestion, as these spectra were observed to be more complex in terms of the size distribution of the heparin oligosaccharides. Peaks above a manually set threshold level were automatically picked by the DOSY Toolbox software for DOSY processing. Peaks were then fit to a standard monoexponential decay (HR-DOSY) with a diffusion resolution of 256 points. Fitting statistics for the pure exponential fitting to the Stejskal–Tanner equation for the diffusion coefficients and fitting errors were determined and displayed following the DOSY processing.
Results and discussion
The commercially available heparinase enzymes I, II, and III, each with unique specificity, are commonly used for heparin and HS digestions. Heparinase I cleaves the GlcNS(6S) → IdoA(2S) glycosidic bond and is considered specific for the analysis of heparin [4, 6, 30]. Heparinase III is most active for 2-OH GlcA-containing glycosidic linkages and therefore is used for HS digestion [4]. Although heparinase III tolerates 6-O sulfonation, it does not act on IdoA residues containing 2-O-sulfo substituents [9]. Heparinase II has broader substrate specificity cleaving a wide range of disaccharide repeat units with either 2-O-sulfo or 2-OH at the uronic acid [4]. Because heparinase II has been reported to have two distinct active sites, one which is heparinase I like and the other heparinase III like, it can be used for the analysis of both heparin and HS [9]. Exhaustive depolymerization of heparin is often carried out using a cocktail containing all three enzymes.
Monitoring the enzymatic depolymerization with UV absorption
Cleavage of the heparin polysaccharide with heparinase enzymes introduces an unsaturated bond between C-4 and C-5 of the non-reducing end uronic acid, as shown in Fig. 1. By exploiting the UV absorbance of the double-bond-containing product, the progress of the reaction can be monitored as shown in Fig. 2a. In our hands, the results of heparinase digestions are highly variable and the extent of the reaction can be difficult to predict. The heparinase enzymes are not very stable and the point at which product formation plateaus in a digestion depends both on the consumption of reactant and the loss of enzymatic activity. If the 1 g heparin sample had been completely depolymerized to disaccharides by the 50 units of heparinase I used in this experiment, a product concentration of roughly 34 mM would be predicted. Obviously, the value expected would be less than this value, as some disaccharide sequences will not fit the consensus sequence required for heparinase I activity. The reaction shown in Fig. 2a was considered complete after 66 h as indicated by a plateau in the product concentration at 7.92 mM. If the goal of this digestion had been to produce mostly larger oligosaccharides for protein-binding studies, it could have been desirable to quench the reaction prior to its plateau.
Generation of heparin-derived oligosaccharides. a Progression of the enzymatic digestion measured at 232 nm using a 1-mm sample length. Aliquots were taken at regular intervals (15 min to 6 h). b Preparative SEC of heparin oligosaccharides following the enzymatic depolymerization of heparin with heparinase I. The labels represent the number of saccharide units in the depolymerization products with dp2 corresponding to disaccharides, dp4 to tetrasaccharides, etc. Product concentrations were calculated using the measured absorbance and the molar extinction coefficient for the monounsaturated disaccharide of 5,500 M−1 cm−1.
Following the enzymatic digestion, an SEC separation was performed to determine the size dispersion of the resulting heparin oligosaccharides and fractionate the products based on size. The 50-mL heparinase digest solution was lyophilized, reconstituted into 15 mL of the separation buffer (0.5 M NH4HCO3), and applied to a preparative-scale SEC column for separation. A column void volume of 346.5 mL was observed before the initial elution of the largest isolated oligosaccharide; the hexadecasaccharides (dp16) present in fractions 77–81.
The progress of the size separation was monitored by UV absorbance measurements plotted as effective double-bond concentration vs. SEC fraction in Fig. 2b. The resolution obtained provided an efficient separation even for the largest oligosaccharides. As can be seen from the SEC, completion of the reaction did not correspond to complete digestion of heparin to its corresponding disaccharide subunits as expected for heparinase I. The SEC peak with the highest concentration corresponds to the disaccharide component (dp2). Preceding peaks correspond to larger heparin oligosaccharides ranging from tetrasaccharides (dp4) to hexadecasaccharides (dp16), with each SEC fraction containing a heterogeneous mixture of similarly sized oligosaccharides varying in their disaccharide compositions.
Monitoring the enzymatic depolymerization of heparin with 1H NMR
In addition to following the enzymatic digestion using UV absorbance, 1H NMR spectra were acquired at various points throughout the reaction. Although NMR has been used extensively in the characterization of heparin-derived oligosaccharides [20, 31–33], to our knowledge, there has only been one study that used NMR to monitor heparin depolymerization [20]. Chuang et al. used 1H-NMR to monitor the kinetics of heparinase digestions with and without histamine present and to identify the oligosaccharide fragments that were bound to histamine in the digest solution.
In this study, 1H NMR was also used to follow the progress of the heparinase I enzymatic digestion. The digest solution was left in the NMR probe for the duration of the reaction to maintain temperature control and to avoid extensive shimming between acquisitions. Figure 3 shows a sample of the 1H NMR spectra measured as a function of reaction time. Near the start of the reaction (10 min), the spectra are essentially those of intact heparin. The heparin resonances are broad due in large part to the short T2 values of the large biopolymer (MW ∼ 17–19 kDa). Heparin resonance line widths are also affected by binding of trace paramagnetic impurities, and addition of small amounts of deuterated EDTA can significantly improve spectral quality [34]. However, because heparinase I requires Ca2+ for activity, EDTA could not be added to the reaction solution [35].
1H NMR spectra measured as a function of time for the depolymerization solution. Acetate was introduced into the solution as calcium acetate because the enzyme requires calcium for activity. Tris was added as the buffering salt at a pH of 6.8. Insets correspond to the H-4 resonances of the Δ4,5 double bond introduced during digestion. Arrows indicate the signature peaks used for the NMR diffusion measurements. Anomeric peaks at 5.460 and 5.514 ppm correspond to the formation of ΔUA(2S)-(1 → 4)-GlcNS(6S).
As the reaction proceeded, heparin oligosaccharide resonances corresponding to the digested heparin fragments were observed at chemical shifts of 5.973, 5.981, 5.990, and 5.998 ppm, consistent with results reported by Chuang [20]. These resonances correspond to oligosaccharides produced by the β-elimination reaction and containing a double-bond proton, H-4, at the C-4 position of the non-reducing end uronic acid at the cleavage site. As expected, the intensities of the H-4 resonances increased as the reaction progressed, although the rate of increase was different for the different peaks. The resonances at 5.990 and 5.998 ppm were observed to increase at a faster rate than the resonances at 5.973 and 5.981 ppm during the first 12 h of the digestion. As the reaction progressed, as indicated by the increase in UV absorption, the upfield resonances continued to increase and at 48 h the resonances at 5.973 and 5.981 ppm were similar in intensity to the resonances at 5.990 and 5.998 ppm as shown in the inset of Fig. 3. The two upfield resonances at 5.973 and 5.981 ppm are also more narrow, suggesting that these resonances may arise from smaller oligosaccharide fragments.
In addition to monitoring the H-4 protons, we were also interested in identifying additional resonances which act as signatures of heparinase I digestion, especially resonances for the H-1 protons of the heparin oligosaccharides. Two new resonances in the anomeric region of the spectrum at 5.460 and 5.514 ppm, indicated by the arrows in Fig. 3, were also observed to increase as the reaction progressed. In concert with the increase in intensity of these resonances, the anomeric resonances for the intact heparin at 5.230 and 5.415 ppm diminished over time. The increased intensity of the H-1 resonances at 5.460 and 5.514 ppm indicates a significant increase in the amount of ΔUA(2S)-(1 → 4)-GlcNS(6S), disaccharide IS, the primary disaccharide building block of heparin containing a double bond as a result of the enzymatic cleavage.
Monitoring the enzymatic depolymerization of heparin using DOSY
The NMR diffusion experiments were performed using dilute aqueous solutions of heparin. Because the diffusion behavior of solution components is related to hydrodynamic properties such as size, shape, and charge, solution components can be distinguished provided that the species have resolved resonances or significantly different diffusion coefficients [36–38]. DOSY is commonly used to process and display diffusion data. DOSY spectra are calculated using an approximation to the inverse Laplace transform from a series of pulsed-field gradient NMR spectra acquired using gradient pulses with varying amplitudes. DOSY processing produces a pseudo-2D spectrum with NMR chemical shifts on the horizontal axis and calculated self-diffusion coefficients along the vertical axis, resolving the spectra of the individual components based on their diffusion behavior [24, 39, 40].
DOSY NMR lends itself to monitoring reactions directly in the NMR tube. No physical separation of mixture components is required, and reaction components can be identified by virtue of their 1H NMR resonances and diffusion coefficients. These experiments used the bipolar pulse-pair-stimulated echo experiment (BPPSTE) [41]. The stimulated echo sequence avoids phase modulation of coupled resonances by storing the magnetization along the longitudinal axis during the diffusion delay, while the use of bipolar gradients reduces the effect of eddy currents and corresponding lock disturbances. Diffusion coefficients were calculated for individual resonances in the BPPSTE spectra by measuring signal attenuation as a function of gradient amplitude. Signal intensities were fit to the Stejskal–Tanner equation modified for the BPPSTE pulse sequence, as shown in Eq. 1,
where D is the diffusion coefficient, I is the resonance intensity measured for a given gradient amplitude, G, I 0 is the intensity in the absence of the gradient pulse, γ is the gyromagnetic ratio of the nucleus detected, δ is the duration of the bipolar gradient pulse pair, Δ is the diffusion delay time, and τ is a short gradient recovery delay during which relaxation and spin–spin coupling evolution are not significant.
Although interactions of heparin and HS with proteins and peptides is considered primarily electrostatic, involving interactions between the cationic ammonium, guanidinium, and imidazolium side chains of proteins and peptides with the anionic sites on heparin and HS, binding through unique heparin and HS structural motifs is also important for biological activity [4]. The exhaustive digestion of heparin significantly affects the biological activity of the oligosaccharides produced by eliminating binding specificity [42, 43]. Depolymerization reaction monitoring is currently performed using analytical-scale SEC or SAX separations with UV detection to determine the oligosaccharide size distribution as a function of time [18, 44]. These separations are time consuming and wasteful of sample as each measurement requires extraction of an aliquot from the depolymerization reaction solution.
DOSY has traditionally been used for the analysis and characterization of mixtures and aggregates [24, 39, 45–47]; however, the Morris group recently reported the use of DOSY to follow the acid hydrolysis of sucrose and maltotriose [48, 49]. To our knowledge, DOSY has not been used to monitor heparinase digestions; therefore, we designed experiments to test the ability of DOSY to follow the heparin depolymerization reaction in situ. If successful, DOSY could be used to monitor the size distribution of oligosaccharides produced as the enzymatic reaction proceeds. This would allow the investigator to quench the reaction at the optimum time to maximize the yield of the larger oligosaccharides most relevant for heparin-binding studies.
A portion of the DOSY spectra measured for the digest samples at 1 and 66 h are shown in Fig. 4a, b, respectively. At 1 h, only two major heparin anomeric resonances are observed in the DOSY spectrum (Fig. 4a). These peaks give rise to similar diffusion coefficients 1.03 ± 0.02 × 10−10 and 1.04 ± 0.02 × 10−10 m2 s−1 for the proton resonances at 5.230 and 5.415 ppm, respectively. The diffusion coefficients are larger than values reported previously for intact heparin (∼4.5 × 10−11 m2 s−1) [25]. This may be attributed to the slightly higher temperature used in the digestion as well as differences in the buffer concentration, heparin source, and the solution viscosities for water (1.00 mPa s) and deuterium oxide (1.25 mPa s). After 1 h, the H-4 resonances can barely be detected above the noise and are too weak to provide a reliable DOSY fit.
The DOSY spectrum measured after 66 h, Fig. 4b, shows much greater variability in the calculated diffusion coefficients. Regions of the spectrum were fit individually to reduce the complexity of the analysis. Peaks above a manually set threshold level were automatically selected and fit with a monoexponential decay using the DOSY Toolbox software. In addition to calculating diffusion coefficients and fitting errors, residual plots for each of the fitted peaks were also available to evaluate the fitting model. From the SEC separation obtained for the bulk digestion solution at the completion of the reaction, it is evident that the digestion solution is a heterogeneous mixture dominated by smaller disaccharide and tetrasaccharide components. Therefore, the resonances in the 1H NMR and DOSY spectra are produced by multiple oligosaccharide components with different molecular weights but similar structures. It is possible to use routines such as CONTIN [50, 51], CORE [52], and multivariate approaches [53, 54] to fit diffusion data measured for polydisperse components such as polymers or aggregates to a distribution of diffusion coefficients. However, for the diffusion spectra shown in Fig. 4, a monoexponential fit of Eq. 1 to the data provides acceptable fits yielding average diffusion coefficients and reasonable fitting errors from 1.5% to 6%.
As the digestion progressed, diffusion coefficients were monitored for the anomeric resonances of the intact heparin (5.230 and 5.415 ppm) and the H-4 resonances (5.973, 5.981, 5.990, and 5.998 ppm) of the cleaved product, as well as N-acetyl peaks (2.048 ppm) of the heparin oligosaccharides. The values obtained over the course of the experiments are consistent with what is expected for an enzymatic digestion; as the reaction proceeded, the measured diffusion coefficients for all peaks were observed to increase. This observation is expected as larger heparin oligosaccharides are continually cleaved into smaller fragments throughout the digestion, thus increasing their individual mobilities in the solution. Though the diffusion coefficient for the N-acetyl resonance changed over the course of the digestion from an initial value of 1.04 ± 0.01 × 10−10 m2 s−1 at 1 h to 4.24 ± 0.08 × 10−10 m2 s−1 at 66 h, its intensity remained constant throughout the digestion with an average intensity of 0.377 ± 0.03 normalized to the intensity of tert-butanol.
Diffusion coefficients measured for the H-4 and the H-1 resonances associated with the digested heparin oligosaccharides also increased over the course of the digestion. Four hours following the initiation of the reaction, integration of the H-4 resonances at 5.990 and 5.998 ppm was possible, producing diffusion coefficients of 1.681 ± 0.19 × 10−10 and 1.37 ± 0.17 × 10−10 m2 s−1, respectively. Fitting of the H-4 resonances at 5.973 and 5.981 ppm was possible only after 13 h, producing diffusion coefficients of 5.77 ± 0.76 × 10−10 and 3.14 ± 0.26 × 10−10 m2 s−1, respectively. The larger fitting errors obtained for the 4 and 13 h data reflect the greater heterogeneity of the product in these solutions and the fact that the enzymatic reaction is continuing during the 2 h required for acquisition of the diffusion data set. At the conclusion of the digestion, the measured diffusion coefficients for the H-4 resonances ranged from 4.87 ± 0.15 × 10−10 to 7.78 ± 0.46 × 10−10 m2 s−1 for the peaks at 5.990 and 5.973 ppm, respectively, as shown in Fig. 4b.
Differences observed between the diffusion coefficients determined for the N-acetyl, H-4, and anomeric resonances can be attributed to a reduced activity of the enzyme for heparin species low in sulfation, for example, heparin fragments containing an acetylated glucosamine residue [4]. Throughout the digestion, the components with upfield H-4 resonances (5.973 and 5.981 ppm) were observed to diffuse faster than components corresponding to the downfield resonances (5.990 and 5.998 ppm) as shown in Fig. 4b. The H-4 resonance is a doublet of doublets with a larger coupling of around 4 Hz, observed in these spectra due to coupling to the H-3 proton of the uronic acid, and a <1 Hz long-range coupling to H-2 not visible in these spectra due to the apodization function applied. The heterogeneity of the components giving rise to the H-4 peaks in the digest solution is also apparent from the varied intensities of the peaks. Clearly, even at 66 h, the spectrum contains H-4 resonances from more than two components. Because heparin is primarily composed of the disaccharide IdoA(2S)-(1 → 4)-GlcNS(6S), the 1H NMR spectrum of the IS standard in the depolymerization buffer was measured for comparison with that of the heparin depolymerization solution as shown in Fig. 5. From the overlay of the two spectra, it is evident that a major component of the digestion mixture is the disaccharide IS.
Figure 6 compares the diffusion coefficients measured using the H-4 resonances as a function of reaction time. Linear regression (R ≥ 0.93) of the data indicated a strong correlation between the diffusion coefficients and reaction time. Such a calibration plot could be useful for investigators wishing to quench the enzymatic digestion to maximize the yield of product with a desired size distribution. It is also interesting to note from the plot that, although the diffusion coefficients for the resonances at 5.981, 5.990, and 5.998 ppm appear to begin to level out toward the conclusion of the reaction, the diffusion coefficient measured for the resonance at 5.973 ppm continues to increase. These DOSY results may provide insight into the possible mode of action for heparinase I, as the digestion results from the repetitive binding and cleavage of larger fragments in the digest solution.
Conclusions
In this study, NMR provided a mechanism to monitor the enzymatic depolymerization of heparin in situ. In conjunction with UV absorption measurements, DOSY provided insight into the extent of reaction and the size distribution of the heparin oligosaccharides produced along the course of the digestion. The ability to monitor diffusion coefficients, which report on the size distribution of heparin-derived oligosaccharides, provides a means for the investigator to quench the reaction at an optimum time to maximize the yields of larger oligosaccharides most relevant for heparin-binding studies.
References
Gallagher JT, Lyon M, Steward WP (1986) Biochem J 236:313–325
Lever R, Page CR (2002) Nat Rev Drug Discovery 1:140–148
Coombe DR, Kett WC (2005) Cell Mol Life Sci 62:410–424
Rabenstein DL (2002) Nat Prod Rep 19:312–331
Whitelock JM, Iozzo RV (2005) Chem Rev 105:2745–2764
Capila I, Linhardt RJ (2002) Angew Chem Int Ed Engl 41:390–412
Shriver Z, Sundaram M, Venkataraman G, Fareed J, Linhardt R, Biemann K, Sasisekharan R (2000) Proc Natl Acad Sci U S A 97:10365–10370
Lindahl U, Backstrom G, Thunberg L, Leder IG (1980) Proc Natl Acad Sci U S A 77:6551–6555
Wei Z, Lyon M, Gallagher JT (2005) J Biol Chem 280:15742–15748
Linhardt RJ, Turnbull JE, Wang HM, Loganathan D, Gallagher JT (1990) Biochemistry 29:2611–2617
Rice KG, Linhardt RJ (1989) Carbohydr Res 190:219–233
Shively JE, Conrad HE (1976) Biochemistry 15:3932–3942
Bienkowski MJ, Conrad HE (1985) J Biol Chem 260:356–365
Mardiguian JS (1984) Heparin esters and processes for their preparation. United States Patent 4440926
Uzan A (1998) Sulfated polysaccharides obtained from heparin, preparation process, pharmaceutical composition and use thereof. United States Patent 5849721
Fussi F (1981) Process for obtaining low molecular weight heparins endowed with elevated pharmacological properties, and product so obtained. United States Patent 4281108
Islam T, Butler M, Sikkander SA, Toida T, Linhardt RJ (2002) Carbohydr Res 337:2239–2243
Chuang W-L, McAllister H, Rabenstein DL (2001) J Chromatogr A 932:65–74
Pervin A, Gallo C, Jandik KA, Han X-J, Linhardt RJ (1995) Glycobiology 5:83–95
Chuang W-L, McAllister H, Rabenstein DL (2002) Carbohydr Res 337:935–945
Imanari T, Toida T, Koshiishi I, Toyoda H (1996) J Chromatogr A 720:275–293
Thanawiroon C, Linhardt RJ (2003) J Chromatogr A 1014:215–223
Korir AK, Limtiaco JFK, Gutierrez SM, Larive CK (2008) Anal Chem 80:1297–1306
Morris KF, Johnson CS (1992) J Am Chem Soc 114:3139–3141
Sitkowski J, Bednarek E, Bocian W, Kozerski L (2008) J Med Chem 51:7663–7665
Bednarek E, Sitkowski J, Bocian W, Mulloy B, Kozerski L (2010) J Pharm Biomed Anal 53:302–308
Smallcombe SH, Patt SL, Keifer PA (1995) J Magn Reson Ser A 117:295–303
Henderson T (2010) Anal Bioanal Chem 396:1465–1471
Nilsson M (2009) J Magn Reson 200:296–302
Galliher PM, Cooney CL, Langer R, Linhardt RJ (1981) Appl Environ Microbiol 41:360–365
Korir A, Larive C (2007) Anal Bioanal Chem 388:1707–1716
Mikhailov D, Mayo KH, Vlahov IR, Toida T, Pervin A, Linhardt RJ (1996) Biochem J 318:93–102
Mikhailov D, Linhardt RJ, Mayo KH (1997) Biochem J 328:51–61
McEwen I (2009) J Pharm Biomed Anal 51:733–735
Shriver Z, Liu DF, Hu Y, Sasisekharan R (1999) J Biol Chem 274:4082–4088
Jayawickrama DA, Larive CK, McCord EF, DRoe C (1998) Magn Reson Chem 36:755–760
Derrick TS, Larive CK (1999) Appl Spectrosc 53:1595–1600
Antalek B (2002) Conc Magn Res 14:225–258
Morris KF, Stilbs P, Johnson CS (1994) Anal Chem 66:211–215
Barjat H, Morris GA, Smart S, Swanson AG, Williams SCR (1995) J Magn Reson B 108:170–172
Wu DH, Chen AD, Johnson CS (1995) J Magn Reson Ser A 115:260–264
Caldwell EEO, Nadkarni VD, Fromm JR, Linhardt RJ, Weiler JM (1996) Int J Biochem Cell Biol 28:203–216
Yu G, LeBrun L, Gunay NS, Hoppensteadt D, Walenga JM, Fareed J, Linhardt RJ (2000) Thromb Res 100:549–556
Ziegler A, Zaia J (2006) J Chromatogr B 837:76–86
Huo R, Wehrens R, Duynhoven Jv, Buydens LMC (2003) Anal Chim Acta 490:231–251
Balayssac S, Trefi S, Gilard V, Malet-Martino M, Martino R, Delsuc M-A (2009) J Pharm Biomed Anal 50:602–612
Politi M, Groves P, Chávez MI, Cañada FJ, Jiménez-Barbero J (2006) Carbohydr Res 341:84–89
Nilsson M, Khajeh M, Botana A, Bernstein MA, Morris GA (2009) Chem Commun 1252–1254
Khajeh M, Botana A, Bernstein MA, Nilsson M, Morris GA (2010) Anal Chem 82:2102–2108
Provencher SW (1982) Comput Phys Commun 27:229–242
Morris KF, Johnson CS (1993) J Am Chem Soc 115:4291–4299
Stilbs P, Paulsen K, Griffiths PC (1996) J Phys Chem 100:8180–8189
Schulze D, Stilbs P (1993) J Magn Reson Ser A 105:54–58
Windig W, Antalek B (1997) Chemom Intell Lab Syst 37:241–254
Acknowledgements
The authors gratefully acknowledge financial support from the National Science Foundation grant CHE 0848976. J.F.K.L. acknowledges support by a 2008–2010 US Pharmacopeia graduate fellowship. Sz.B. gratefully acknowledges a fellowship from the Hungarian American Enterprise Scholarship Fund and OTKA MB08A/80066.
Open Access
This article is distributed under the terms of the Creative Commons Attribution Noncommercial License which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and source are credited.
Author information
Authors and Affiliations
Corresponding author
Additional information
Published in the special issue Heparin Characterization with guest editor Cynthia K. Larive
Rights and permissions
Open Access This is an open access article distributed under the terms of the Creative Commons Attribution Noncommercial License (https://creativecommons.org/licenses/by-nc/2.0), which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and source are credited.
About this article
Cite this article
Limtiaco, J.F.K., Beni, S., Jones, C.J. et al. NMR methods to monitor the enzymatic depolymerization of heparin. Anal Bioanal Chem 399, 593–603 (2011). https://doi.org/10.1007/s00216-010-4132-7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00216-010-4132-7