Abstract
Creatine kinase (CK) catalyzes the formation of phosphocreatine from adenosine triphosphate (ATP) and creatine. The highly reactive free cysteine residue in the active site of the enzyme (Cys283) is considered essential for the enzymatic activity. In previous studies we demonstrated that Cys283 is targeted by the alkylating chemical warfare agent sulfur mustard (SM) yielding a thioether with a hydroxyethylthioethyl (HETE)-moiety. In the present study, the effect of SM on rabbit muscle CK (rmCK) activity was investigated with special focus on the alkylation of Cys283 and of reactive methionine (Met) residues. For investigation of SM-alkylated amino acids in rmCK, micro liquid chromatography-electrospray ionization high-resolution tandem-mass spectrometry measurements were performed using the Orbitrap technology. The treatment of rmCK with SM resulted in a decrease of enzyme activity. However, this decrease did only weakly correlate to the modification of Cys283 but was conclusive for the formation of Met70-HETE and Met179-HETE. In contrast, the activity of mutants of rmCK produced by side-directed mutagenesis that contained substitutions of the respective Met residues (Met70Ala, Met179Leu, and Met70Ala/Met179Leu) was highly resistant against SM. Our results point to a critical role of the surface exposed Met70 and Met179 residues for CK activity.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Creatine kinase (CK, EC 2.7.3.2) belongs to an evolutionarily conserved group of enzymes. It catalyzes the reversible, magnesia-catalyzed (Mg) reaction between creatine (Cr) and adenosine triphosphate (ATP) forming phosphocreatine (PCr) and adenosine diphosphate (ADP) according to the reaction MgATP + Cr ⇆ MgADP + PCr (Schlattner et al. 2016), thus being a key player in maintaining cellular energy homeostasis. Four major CK isozymes, two cytosolic and two mitochondrial, which form dimers and octamers, respectively, have been described (McLeish and Kenyon 2005). The cytosolic subunits can be either B type (brain) or M type (muscle) resulting in either CK-MM, CK-BB or CK-MB dimers (Bais and Edwards 1982; Wallimann et al. 2011). The individual CK isoenzymes are encoded by four independent genes (Qin et al. 1998; McLeish and Kenyon 2005) with an overall conservation of the amino acid sequence between 78% to more than 99% and a highly preserved active site (Mühlebach et al. 1994; Qin et al. 1998; McLeish and Kenyon 2005). The first CK crystal structure was solved in 1996 (Fritz-Wolf et al. 1996). Since then, additional high-resolution crystal structures of different CK subtypes from various species were reported that allowed the assignment of enzyme function to certain amino acid motifs (Rao et al. 1998; Eder et al. 1999, 2000a; Bong et al. 2008). Several amino acid residues, including cysteine (Cys) (Reddy and Watts 1978; Furter et al. 1993; Reddy et al. 2000; Wang et al. 2006), arginine (Wood et al. 1998), histidine (Forstner et al. 1997) as well as tryptophan and aspartic acid (Gross et al. 1994; Cantwell et al. 2001) are considered to be important for substrate binding and conversion (Bickerstaff and Price 1978; Eder et al. 2000b). In particular, the highly reactive Cys283 residue (amino acid numbering does include the N-terminal methionine (Met) residue) in the active site of the enzyme is discussed to play a pivotal role in this context (Maggio and Kenyon 1977; Bickerstaff and Price 1978; Furter et al. 1993; Reddy et al. 2000; Wang et al. 2006) and is subject of current research.
It was shown that alkylation of CK with iodoacetamide (IAA) or iodomethane, which both target free Cys residues, reduced the enzyme activity (Atherton et al. 1970; Reddy and Watts 1978). Treatment of CK with the alkylating chemical warfare agent sulfur mustard (SM) resulted in the formation of the specific hydroxyethylthioethyl-(HETE)-moiety at Cys283 (Lüling et al. 2021; Steinritz et al. 2021) but its effect on enzyme activity has not been addressed so far. This was investigated in the present study with particular focus on Cys283 but also on reactive Met residues, that in principle might also be essential for the activity of certain enzymes (Rogers et al. 1976) and were shown to be a potential target of alkylation by SM (Siegert et al. 2019).
Materials and methods
Chemicals
SM and its eight-fold deuterated analog d8-SM (purity of SM and d8-SM > 99%, assessed in-house by nuclear magnetic resonance, NMR) were made available by the German ministry of Defense. Rabbit muscle CK (rmCK), IAA, ethanol (EtOH), 4-(2-hydroxyethyl)-piperazine-1-ethanesulfonic acid (HEPES), NaOH, 25% (w/v) polyethyleneglycol 4000, ammonium acetate, dithiothreitol (DTT), glycerol, sodium citrate, dimethylformamide, ethylenediaminetetraacetic acid (EDTA), ethyleneglycoltetraacetic acid (EGTA), Na2HPO4, NaH2PO4, protease-inhibitor mix, nuclease mix, and Tween20 were obtained from Sigma-Aldrich (Steinheim, Germany). HCl and NaCl were from Carl Roth (Karlsruhe, Germany) and phosphate-buffered saline (PBS) was from Life Technologies (Gibco, Karlsruhe, Germany). CK activity assay kit was purchased from Abcam (Cambridge, UK). Proteinase K (ProtK), water (LC–MS grade), acetonitrile (ACN, LC–MS grade) and formic acid (FA, > 98%) were purchased from Merck (Darmstadt, Germany). NH4HCO3 (ultra-grade, 99.5%) was from Fluka (Buchs, Switzerland) and three-fold deuterated atropine (d3-atr) from CDN Isotopes (Pointe Claire, Quebec, Canada).
CK activity assay
The effect of SM on rmCK enzyme activity was examined using a CK activity assay. RmCK was dissolved in PBS (pH 7.4, 10 mg/mL). This solution (49 µL) was mixed with 1 µL SM solution (200 mM in EtOH) in a 96-well plate with clear flat bottom (Greiner Bio-One, Frickenhausen, Germany) and incubated for 0, 5, 10, 15, 20, 25, and 30 min before starting the assay. After the respective incubation periods, 50 µL of the incubation solution was mixed immediately with 34 µL CK assay buffer, 2 µL CK enzyme mix, 2 µL CK developer, 2 µL ATP solution and 10 µL CK substrate solution (all included in the assay). The optical density (OD) at 450 nm was measured every minute for a total of 60 min at 25 °C using the TECAN infinite M200 PRO photometer (TECAN, Crailsheim, Germany). To characterize the effects on enzyme activity caused by the solvent or \({\mathrm{Cl}}^{-}\) as one hydrolysis product of SM, or \({\mathrm{H}}^{+}\), that is released during the alkylation reaction, rmCK was incubated with 1 µL of either EtOH or 8 mM NaCl or 0.2 mM HCl and measurements were performed as described above. The NaCl concentration of 8 mM mimics the theoretical maximum concentration of \({\mathrm{Cl}}^{-}\) resulting from complete hydrolysis of 4 mM SM.
Enzyme activities of mutant rmCK variants were analyzed after 15 min of incubation with different concentrations of SM (final concentrations: 0.04, 0.4, 1, 4, 10, 20, or 40 mM SM) using the same procedure.
µLC-ESI MS/HR MS analysis
Sample preparation
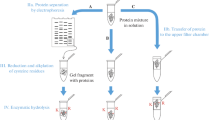
For investigation of SM-alkylated amino acids in rmCK, micro liquid chromatography-electrospray ionization high-resolution tandem-mass spectrometry (µLC-ESI MS/HR MS) measurements were performed using the Orbitrap technology. For this purpose, 198 µL of rmCK solution (10 mg/mL in PBS) was mixed with 2 µL solution of either ethanolic SM or d8-SM solution (final concentrations: 0.04, 0.4, 1, 4, 10, 20, and 40 mM, each) at room temperature (RT) for 15 min in an ultrafiltration (UF)-device (molecular weight cut-off, MWCO, 10 kDa, Amicon, Merck-Millipore, Darmstadt, Germany). After washing with 300 µL water and UF (15 min, 25 °C, 9,770 RCF), samples were diluted with PBS to a rmCK concentration of 2 mg/mL. Subsequently, 200 µL of each sample was incubated with 2 µL DTT solution (10 mg/mL in water) for 1 h at RT following incubation with 2 µL IAA solution (10 mg/mL in water) for 45 min at RT. Next, samples were subjected to UF (15 min, 25 °C, 9,770 RCF) and the retentate was diluted with PBS to a final rmCK concentration of 2 mg/mL. For proteolysis, 100 µL of each sample was incubated with 100 µL ProtK solution (15 mg/mL in 50 mM NH4HCO3) and 300 µL 50 mM NH4HCO3 buffer in an UF-device at 50 °C for 2 h. Filtrates obtained from subsequent UF were diluted 1:3 with d3-atr solution (3 ng/mL in 0.5% v/v FA) prior to µLC-ESI MS/HR MS.
Chromatographic separation
For chromatographic separation of 20 µL sample volume, a MicroPro pump (Eldex Laboratories, Napa, CA, USA) was used in combination with an INTEGRITY autosampler (sample tray kept at 15 °C) equipped with a 20 µL sample loop and a MISTRAL column oven (both Spark Holland, Emmen, The Netherlands). The chromatographic system was controlled by the Eldex MicroPro 1.0.54 control software (Eldex Laboratories). Samples were separated on an ACQUITY UPLC HSS T3 column (C18, 50 × 1.0 mm I.D., 1.8 µm, 100 Å, Waters, Eschborn, Germany) protected by a security guard ultra-cartridge (C18-peptide, Phenomenex, Aschaffenburg, Germany) with two gradients of solvent A (0.05% v/v FA) and solvent B (ACN/H2O 80:20 v/v, 0.05% v/v FA): µLC-gradient 1: t [min]/B [%]: 0/0; 3/0; 35/40; 35.5/95; 39.5/95; 40/0 including an initial equilibration period of 5 min and µLC-gradient 2: t [min]/B [%]: 0/0; 12/35; 12.5/95; 14.5/95; 15/0 including an initial equilibration period of 5 min.
MS/HR MS analysis
For identification and relative quantification of alkylated peptides originated from SM- and d8-SM-treated (40 mM, each) rmCK after cleavage with ProtK, a Q Exactive Plus (ThermoFisher Scientific, Waltham, USA) Orbitrap mass spectrometer was used. It was coupled online to the µLC system via the HESI-II probe. All MS experiments were performed in the positive mode using the following settings: sheath gas flow 23 arbitrary units (a.u.), auxillary (aux) gas flow 8 a.u., sweep gas flow 1 a.u., spray voltage 3.5 kV, capillary temperature 250 °C, S-lens RF level 50 a.u. and aux gas heater temperature 125 °C. The Orbitrap instrument was calibrated daily using the Pierce™ LTQ ESI positive ion calibration solution (ThermoFisher Scientific) according to the manufacturer’s protocol.
For identification of rmCK-derived alkylated peptides, data-dependent tandem-mass spectrometry (ddMS2) scans (top 10) in combination with µLC gradient 1 were used. A full-MS survey scan in the range of m/z 200–m/z 1000 was performed with a resolution of 70,000 full width at half maximum (FWHM), with an automatic gain control (AGC) target of 3 × 106 charges and a maximum injection time (IT) of 200 ms. For highest mass accuracy, the ubiquitous softener bis(2-ethylhexyl)terephthalate (single protonated, m/z 391.28429) was used as a lock mass. Identified precursor masses, which met the ddMS2 settings (minimum AGC target: 1 × 103 charges; intensity threshold: 1 × 104 charges; charge exclusion: z ≤ 4; peptide match: preferred; exclude isotopes: on) were measured by MS2 scans (17,500 FWHM; AGC target: 1 × 105 charges; maximum IT: 100 ms; loop count: 10; MSX count: 1; isolation window: ± 2 Th; fixed first mass: m/z 100; normalized collision energy (NCE): 25 V). In addition, peptides also showing precursor masses with the d8-mass shift (4 ppm tolerance interval) and similar retention times (tR, ± 0.5 min) were fixed and automatically assigned to SM-alkylated peptides derived from rmCK (2 ppm mass tolerance).
For relative quantification of alkylated peptides including ThrCys283(-HETE)ProSer (TC283(-HETE)PS), IleMet70(-HETE)ThrVal (IM70(-HETE)TV), LysSerMet179(-HETE)ThrGlu (KSM179(-HETE)TE) as well as of the internal standard d3-atr, parallel reaction monitoring (PRM) was performed in combination with µLC gradient 2. For all experiments, the resolution of the Orbitrap mass analyzer was set to 17,500 FWHM, the AGC target to 2 × 105 charges, the maximum IT to 80 ms, and the isolation window for precursor masses to Δm/z ± 2.0. Additional MS parameters depending on the analyte detected are summarized in Table 1.
Calculation of alkylation ratios
For calculation of the relative extent of alkylation by SM or d8-SM, the peak areas obtained from µLC-ESI MS/HR MS analysis of the corresponding non-alkylated peptides comprising IM70TV, KSM179TE and TC283PS were determined. Peak areas found after incubation with SM were related to those obtained without incubation to calculate the relative ratio of non-modified amino acids. By subtracting the latter values from 100%, the corresponding alkylation ratios were obtained.
Site-directed mutagenesis
For functional analysis of the residues Met70 and Met179 in rmCK, two recombinant single mutants (Met70Ala and Met179Leu) and one recombinant double mutant (Met70Ala/Met179Leu) were produced (Proteros biostructures, Martinsried, Germany). Three plasmids inducing the desired mutations were synthesized by Geneart (Regensburg, Germany): (i) HIS6-TEV-rabbitCK(1–381)-Met179Leu, (ii) HIS6-TEV-rabbitCK(1–381)-Met70Ala, and (iii) HIS6-TEV-rabbitCK(1–381)-Met70Ala/Met179Leu. Plasmids were transformed in E. coli NiCo21(DE3) obtained from New England Biolabs (Ipswich, MA, USA) for rmCK protein expression.
Protein expression and purification
E. coli NiCo21(DE3), transformed with wildtype rmCK (wt-rmCK) and rmCK mutant expression plasmids as described above, were shaken in terrific broth (TB)-medium (in-house preparation) at 37 °C. Expression was induced by adding 1 mM isopropyl ß-D-1-thiogalactopyranoside (Sigma-Aldrich, Taufkirchen, Germany). Three hours after induction, bacterial cells were harvested by centrifugation. Wt-rmCK and mutant proteins (Met70Ala, Met179Leu, Met70Ala/Met179Leu) were purified from the soluble fraction by Ni–NTA (nickel-nitrilotriacetic acid) affinity chromatography (Sigma-Aldrich). The N-terminal His6-fusion tag was removed by cleavage with tobacco etch virus (TEV) protease (in-house preparation) and wt-rmCK and mutant proteins were subsequently further purified to apparent homogeneity using size exclusion chromatography. The different rmCK proteins were lyophilized for storage.
Statistical analysis and graphical output
The statistics software R (R Core Team 2020) with the graphical user interface RStudio (version 1.1.383) (RStudio Team 2020) was used for statistical analysis and graphic presentation. The R package drc (Ritz et al. 2015) was used for non-linear curve fitting (four-parameter log-logistic function; LL.4). Group differences were calculated by using Student’s t test included in the ggpubr (Kassambara 2020) package. Graphical output was done by using ggplot from the tidyverse package (Wickham et al. 2019).
Results and discussion
Although various CK isoforms and their transition states revealed some insights into the communication between the subunits, substrate binding and conversion (Rao et al. 1998; Eder et al. 1999, 2000a; Lahiri et al. 2002), the exact catalytic mechanism of CK is not completely understood (Ohren et al. 2007; Shen et al. 2001; Tisi et al. 2001). CK possesses one highly reactive sulfhydryl-group (free cysteine residue) per subunit. This group can be modified by a number of sulfhydryl-specific reagents (Kenyon and Reed 1983), with impact on enzyme activity (Mahowald 1965; Buechter et al. 1992). This residue has been identified as Cys283 in the primary structure of rmCK (Mahowald 1965) and is conserved in all known CK sequences of other species (Buechter et al. 1992). Hence, rmCK (UniProtKB P00563) was used in the present study instead of human muscle CK (hmCK, UniProtKB P00563) due to economic reasons. Alignment of rmCK and hmCK using the Clustal Omega program (Sievers et al. 2011) revealed an overall identity of the amino acid sequences of 96.6% with a complete matching of all Cys and Met positions. Cys283 was recently identified as a target of alkylating agents including SM (Lüling et al. 2021; Steinritz et al. 2021). As Cys283 in the active site of CK (Wang et al. 2006) is supposed to bind creatine by electrostatic interactions between the free thiol-moiety of its side chain and the guanidine-group of creatine (Bong et al. 2008), we assumed that a loss of CK activity is due to the alkylation of this residue after treatment with SM.
CK activity after alkylation by SM
Treatment of rmCK with 4 mM SM for at least 10 min revealed a significant decrease of the enzyme activity compared to the non-treated enzyme, reaching its minimum after 15 min (Fig. 1A). Therefore, a 15 min incubation period with SM was chosen for the subsequent dose–response experiments.
Creatine kinase activity after alkylation by SM. Enzyme activity of rabbit muscle creatine kinase (rmCK) was assayed using a colorimetric assay to monitor ATP conversion as area under the curve (AUC). Normalized activity was calculated from values of the respective AUC of non-treated rmCK control (ctr, activity 1.0). A RmCK was incubated with 4 mM SM for different times before starting the activity measurements. B Activity of rmCK after incubation with different SM concentrations for 15 min, each. AUC normalized to control levels is illustrated. Asterisks indicate significant differences (* < 0.05, ** < 0.01, *** < 0.001, **** < 0.0001) between the control group and time (A) or ctr and SM concentration (B). Data are derived from 3 independent experiments (n = 3)
SM concentrations ≥ 4 mM significantly reduced the rmCK activity (Fig. 1B). In contrast, the solvent (2% v/v EtOH) or the SM reaction (\({\mathrm{H}}^{+}\)) and hydrolysis products (\({\mathrm{Cl}}^{-}\)) did not affect rmCK activity (Suppl. Fig. S1) thus confirming that alkylation caused the reduced enzyme activity. This is in accordance to previous studies reporting the loss or at least the decrease of CK activity after the treatment with alkylating IAA (Price 1979; Fossel and Hoefeler 1987).
Identification of alkylated Cys283 after SM treatment of rmCK and correlation to enzyme activity
We succeeded in the detection of diverse peptides comprising all cysteine residues (Cys74, Cys146, Cys254, and Cys283) of rmCK after proteolysis with ProtK by µLC-ESI MS/HR MS analysis (data not shown). Cys283 is located in the active site of CK and is susceptible by chemical modifications (Reddy et al. 2000). After treatment of rmCK with SM, alkylated Cys283 was detected present in the protonated tetrapeptide TC283(-HETE)PS ([M + H]+ m/z 511.2) which is characterized by its product ions at m/z 105.037 and m/z 137.009 obtained by MS/HR MS (Lüling et al. 2021; Steinritz et al. 2021). Alkylation of Cys74, Cys146, and Cys254 only occurred to a negligible extent (data not shown). After treatment with 1 mM SM, Cys283 was found to be alkylated to approx. 90% but rmCK activity was almost not affected (Fig. 2). Therefore, we concluded that alkylation of Cys283 by SM has no major impact on rmCK activity.
Correlation between alkylation ratios of Cys283, Met70, Met179 and rmCK activity. After 15 min treatment of wt-rmCK with different SM concentrations indicated, alkylation ratios (green line and diamonds for Cys283, red line & downwards triangles for Met70, and blue line and upright triangles for Met179) were calculated after µLC-ESI MS/HR MS analysis by normalizing the peak area of the respective precursor ions to control levels of non-treated rmCK. The rmCK activity is given as normalized AUC (squares with dashed gray line). Normalized alkylation ratios are given as green line and diamonds for Cys283(-HETE), red line and downward triangles for Met70(-HETE), and blue line and upward triangles for Met179(-HETE). The error bars display the standard deviation obtained from 3 independent experiments (n = 3) (color figure online)
Identification of alkylated Met70 and Met179 after SM treatment of rmCK and correlation to enzyme activity
RmCK contains 10 Met residues in total (Met30, Met70, Met179, Met207, Met240, Met246, Met272, Met360, Met363, and Met376). Following proteolysis of rmCK with Prot K, diverse peptides were identified by µLC-ESI MS/HR MS that contained these residues thus allowing monitoring of their chemical modification after SM treatment. The most prominent peptides proven to be alkylated at Met70 and Met179 are summarized in Table 1 with respect to their mass spectrometric detection.
Met residues in general represent important regulators of protein function and enzyme activity (Taylor et al. 2018; Aledo 2019; Lim et al. 2019; Valley et al. 2012). They interact with aromatic amino acid side chains (e.g. phenylalanine, tyrosine, tryptophan) thereby significantly increasing the stability of proteins (Valley et al. 2012). Met residues contain a single nucleophilic sulfur atom in the side chain that is accessible for covalent modifications. After alkylation, a Met sulfonium ion is formed containing a permanent positive charge at the sulfur atom (Kramer and Deming 2012) which may cause conformational changes in the protein tertiary structure or may even cause disturbance of the protein secondary structure (Kramer and Deming 2013). Recently, it was shown that Met residues may also be alkylated by SM as proven for Met329 in human serum albumin (Siegert et al. 2019).
The extent of SM-induced alkylation of Met272, Met360 and Met376 did not correlate with the impaired enzyme function (Suppl. Fig. S3). In contrast, the alkylation ratios of Met70 and Met179 increased in a concentration-dependent manner and the rmCK activity decreased reciprocally (Fig. 2). Considering the described three-dimensional structure of rmCK, these Met residues are exposed to the surface and thus allow a good accessibility by SM. In contrast, Met272, Met360 and Met376 are in positions of impaired accessibility which explain their low extent of alkylation. Therefore, our results suggest a causal relationship between Met70 and Met179 and enzyme activity. To prove this hypothesis, relevant mutants of rmCK were generated and their susceptibility towards SM was evaluated.
Activity of rmCK mutants (Met70Ala, Met179Leu) after SM treatment
To elucidate the role of Met70 and Met179 with regard to enzyme activity in more detail, rmCK mutants (Met70Ala, Met179Leu and Met70Ala/Met179Leu) were generated by recombinant expression. Ala or Leu were chosen as substitute amino acids because they preserve the environmental characteristics in the protein structure as close as possible albeit they cannot be alkylated. As analyzed from the database-accessible structure, the environment of Met70 in wt-rmCK is mainly hydrophilic, while that of Met179 is mainly hydrophobic. Therefore, Ala was used for substitution of Met70 and Leu for Met179. Mass spectrometric analysis (ddMS2) of the three mutants and the wt-protein after proteolysis using trypsin demonstrated the integrity of the four proteins (Suppl. Fig. S2A).
All mutants exhibited a relative mean enzyme activity (n = 3) similar to that of wt-rmCK (wt-rmCK: 1.0 ± 0.015, Met70Ala: 0.916 ± 0.077, Met179Leu: 1.0 ± 0.011 and Met70Ala/Met179Leu: 0.964 ± 0.057) thus indicating that the related non-modified Met residues are not essential for the enzyme function. In contrast, after treatment of the enzymes with SM, the activity of the mutants was obviously less diminished than that of the wt-rmCK showing remaining activities after incubation with 40 mM SM of 0.46 ± 0.03 for wt-rmCK, 0.81 ± 0.03 for Met70Ala, 0.96 ± 0.02 for Met179Leu, and 0.77 ± 0.01 for Met70Ala/ Met179Leu (Fig. 3). Especially, the activity of the Met179Leu mutant was shown to be highly resistant. Therefore, we concluded that the alkylation of the Met residues, introducing a permanent positive charge, was a major reason for the loss of CK activity.
Activity of wt-rmCK and rmCK mutants after SM treatment. Wt-rmCK and rmCK mutants (Met70Ala, Met179Leu and Met70Ala/Met179Leu) were incubated for 15 min with different SM concentrations indicated. Activity of rmCK was colorimetrically (450 nm) assayed using a commercial test kit monitoring ATP conversion. Normalized rmCK activity is illustrated as area under the curve (AUC) calculated from 3 independent experiments (n = 3). The error bars display the standard deviation and ribbons represent the 95% confidence intervals of the respective curve fits. Red line and downward triangles represent wt-rmCK, green line and upward triangles represent Met70Ala, blue line and diamonds represent Met179Leu, and yellow line and circles represent Met70Ala/Met179Leu. Activity levels (mean ± SD) of non-treated wt-rmCK are indicated by dotted lines (color figure online)
Conclusion
Our study confirms that Cys283 of rmCK as well as diverse Met residues were alkylated by SM. The use of IM70(-HETE)TV and KSM179(-HETE)TE as biomarkers for the verification of SM exposure, in addition to the already reported use of TC283(-HETE)PS (Steinritz et al. 2021), will be elaborated in future studies.
Alkylation of Cys283 was not suggested as primarily responsible for decreased rmCK enzyme activity, but, in contrast, the alkylation of Met70 and Met179 seemed to play a critical role in that context. Met residues were reported to be important for the conformational stabilization, high affinity ligand binding and function of proteins (Valley et al. 2012). Thus, it can be assumed that alkylation of Met-motifs in rmCK might cause a conformational change of rmCK. Initial experiments using the switchSENSE technique which allows determination of the hydrodynamic diameter (DH) of proteins (Blocquel et al. 2017; Cléry et al. 2017) were conducted to support our hypothesis (data not shown). The DH of untreated wt-rmCK was determined to be 5.5 ± 0.16 nm while that of the alkylated wt-rmCK had increased to 7.67 ± 0.51 nm. The rmCK mutants exhibited a less prominent increase of DH (6.5 ± 0.25 nm for Met70Ala, 5.65 ± 0.14 nm for Met179Leu, and 6.1 ± 0.23 nm for Met70Ala/ Met179Leu) suggesting smaller structural changes and thereby underlining the important role of Met70 and especially Met179 for stabilizing the rmCK protein structure. Additional crystallographic experiments were conducted (data not shown) to further prove the hypothesis of SM-induced conformational changes of the rmCK structure. The alkylated forms (wt and all mutants) exhibited a slower crystal growth than the non-alkylated forms, hinting towards a lower degree of conformational stability. Unfortunately, no adequate crystals of alkylated mutants suitable for synchrotron measurements were obtained, although various optimizations were applied. Future studies should thus investigate the protein structure of alkylated rmCK in more detail.
This is the first study showing that alkylation of Met residues by SM significantly impacts enzyme activity. It seems plausible that the activity of other enzymes might also be affected after alkylation of Met residues. This may help to understand the molecular toxicology of alkylating agents, especially SM, in more detail.
Abbreviations
- a.u.:
-
Arbitrary units
- d3-atr:
-
Three-fold deuterated atropine
- ACN:
-
Acetonitrile
- ADP:
-
Adenosine diphosphate
- AGC:
-
Automatic gain control
- ATP:
-
Adenosine triphosphate
- CE:
-
Collision energy
- CK:
-
Creatine kinase
- Cr:
-
Creatine
- d8-SM:
-
Eight-fold deuterated sulfur mustard
- ddMS2 :
-
Data-dependent tandem-mass spectrometry
- DH :
-
Hydrodynamic diameter
- DTT:
-
Dithiothreitol
- EDTA:
-
Ethylenediaminetetraacetic acid
- EGTA:
-
Ethyleneglycoltetraacetic acid
- ESI:
-
Electrospray ionization
- EtOH:
-
Ethanol
- FA:
-
Formic acid
- FWHM:
-
Full width at half maximum
- HEPES:
-
4-(2-Hydroxyethyl)-piperazine-1-ethanesulfonic acid
- IAA:
-
Iodoacetamide
- IEX:
-
Ion-exchange chromatography
- IT:
-
Injection time
- K:
-
Kelvin
- µLC:
-
Micro liquid chromatography
- MS2 :
-
See MS/MS
- MS/HR MS:
-
High-resolution tandem-mass spectrometry
- MS/MS:
-
Tandem-mass spectrometry
- NCE:
-
Normalized collision energy
- Ni-NTA:
-
Nickel-nitrilotriacetic acid
- NMR:
-
Nuclear magnetic resonance
- nt:
-
Nucleotide
- OD:
-
Optical density
- PBS:
-
Phosphate-buffered saline
- PCr:
-
Phosphocreatine
- PRM:
-
Parallel reaction monitoring
- ProtK:
-
Proteinase K
- RCF:
-
Relative centrifugal force
- rmCK:
-
Rabbit muscle CK
- RT:
-
Room temperature
- SM:
-
Sulfur mustard
- TB:
-
Terrific broth
- TEV:
-
Tobacco etch virus
- t R :
-
Retention time
- UF:
-
Ultrafiltration
- wt:
-
Wildtype
References
Aledo JC (2019) Methionine in proteins: the Cinderella of the proteinogenic amino acids. Protein Sci 28(10):1785–1796. https://doi.org/10.1002/pro.3698
Atherton RS, Laws JF, Thomson AR (1970) Alkylation of bovine brain creatine kinase. Biochem J 118(5):903–904. https://doi.org/10.1042/bj1180903
Bais R, Edwards JB (1982) Creatine kinase. Crit Rev Clin Lab Sci 16(4):291–335. https://doi.org/10.3109/10408368209107030
Bickerstaff GF, Price NC (1978) Creatine kinase: a review of some recent work on the mechanism and subunit behaviour of the enzyme. Int J Biochem 9(1):1–8. https://doi.org/10.1016/0020-711X(78)90128-3
Blocquel D, Li S, Wei N, Daub H, Sajish M, Erfurth M-L, Kooi G, Zhou J, Bai G, Schimmel P, Jordanova A, Yang X-L (2017) Alternative stable conformation capable of protein misinteraction links tRNA synthetase to peripheral neuropathy. Nucleic Acids Res 45(13):8091–8104. https://doi.org/10.1093/nar/gkx455
Bong SM, Moon JH, Nam KH, Lee KS, Chi YM, Hwang KY (2008) Structural studies of human brain-type creatine kinase complexed with the ADP-Mg2+-NO3−-creatine transition-state analogue complex. FEBS Lett 582(28):3959–3965. https://doi.org/10.1016/j.febslet.2008.10.039
Buechter DD, Medzihradszky KF, Burlingame AL, Kenyon GL (1992) The active site of creatine kinase. Affinity labeling of cysteine 283 with N-(2,3-epoxypropyl)-N-amidinoglycine. J Biol Chem 267(4):2173–2178
Cantwell JS, Novak WR, Wang PF, McLeish MJ, Kenyon GL, Babbitt PC (2001) Mutagenesis of two acidic active site residues in human muscle creatine kinase: implications for the catalytic mechanism. Biochemistry 40(10):3056–3061. https://doi.org/10.1021/bi0020980
Cléry A, Sohier TJM, Welte T, Langer A, Allain FHT (2017) switchSENSE: a new technology to study protein-RNA interactions. Methods (san Diego, CA) 118–119:137–145. https://doi.org/10.1016/j.ymeth.2017.03.004
Eder M, Schlattner U, Becker A, Wallimann T, Kabsch W, Fritz-Wolf K (1999) Crystal structure of brain-type creatine kinase at 1.41 A resolution. Protein Sci 8(11):2258–2269. https://doi.org/10.1110/ps.8.11.2258
Eder M, Fritz-Wolf K, Kabsch W, Wallimann T, Schlattner U (2000a) Crystal structure of human ubiquitous mitochondrial creatine kinase. Proteins 39(3):216–225. https://doi.org/10.1002/(sici)1097-0134(20000515)39:3%3c216:aid-prot40%3e3.0.co;2-#
Eder M, Stolz M, Wallimann T, Schlattner U (2000b) A conserved negatively charged cluster in the active site of creatine kinase is critical for enzymatic activity. J Biol Chem 275(35):27094–27099. https://doi.org/10.1074/jbc.M004071200
Forstner M, Müller A, Stolz M, Wallimann T (1997) The active site histidines of creatine kinase. A critical role of His 61 situated on a flexible loop. Protein Sci 6(2):331–339. https://doi.org/10.1002/pro.5560060208
Fossel ET, Hoefeler H (1987) Complete inhibition of creatine kinase in isolated perfused rat hearts. Am J Physiol 252(1 Pt 1):E124–E129. https://doi.org/10.1152/ajpendo.1987.252.1.E124
Fritz-Wolf K, Schnyder T, Wallimann T, Kabsch W (1996) Structure of mitochondrial creatine kinase. Nature 381(6580):341–345. https://doi.org/10.1038/381341a0
Furter R, Furter-Graves EM, Wallimann T (1993) Creatine kinase: the reactive cysteine is required for synergism but is nonessential for catalysis. Biochemistry 32(27):7022–7029. https://doi.org/10.1021/bi00078a030
Gross M, Furter-Graves EM, Wallimann T, Eppenberger HM, Furter R (1994) The tryptophan residues of mitochondrial creatine kinase: roles of Trp-223, Trp-206, and Trp-264 in active-site and quaternary structure formation. Protein Sci 3(7):1058–1068. https://doi.org/10.1002/pro.5560030708
Kassambara A (2020) ggpubr: ‘ggplot2’ based publication ready plots. https://CRAN.R-project.org/package=ggpubr
Kenyon GL, Reed GH (1983) Creatine kinase: structure-activity relationships. Adv Enzymol Relat Areas Mol Biol 54:367–426. https://doi.org/10.1002/9780470122990.ch6
Kramer JR, Deming TJ (2012) Preparation of multifunctional and multireactive polypeptides via methionine alkylation. Biomacromol 13(6):1719–1723. https://doi.org/10.1021/bm300807b
Kramer JR, Deming TJ (2013) Reversible chemoselective tagging and functionalization of methionine containing peptides. Chem Commun (camb) 49(45):5144–5146. https://doi.org/10.1039/c3cc42214c
Lahiri SD, Wang P-F, Babbitt PC, McLeish MJ, Kenyon GL, Allen KN (2002) The 2.1 A structure of Torpedo californica creatine kinase complexed with the ADP-Mg(2+)-NO(3)(−)-creatine transition-state analogue complex. Biochemistry 41(47):13861–13867. https://doi.org/10.1021/bi026655p
Lim JM, Kim G, Levine RL (2019) Methionine in proteins: it’s not just for protein initiation anymore. Neurochem Res 44(1):247–257. https://doi.org/10.1007/s11064-017-2460-0
Lüling R, Schmeißer W, Siegert M, Mückter H, Dietrich A, Thiermann H, Gudermann T, John H, Steinritz D (2021) Identification of creatine kinase and alpha-1 antitrypsin as protein targets of alkylation by sulfur mustard. Drug Test Anal 13(2):268–283. https://doi.org/10.1002/dta.2916
Maggio ET, Kenyon GL (1977) Properties of a CH3-blocked creatine kinase with altered catalytic activity Kinetic consequences of the presence of the blocking group. J Biol Chem 252(4):1202–1207
Mahowald TA (1965) The amino acid sequence around the “reactive” sulfhydryl groups in adenosine triphosphocreatine phosphotransferase. Biochemistry 4:732–740. https://doi.org/10.1021/bi00880a019
McLeish MJ, Kenyon GL (2005) Relating structure to mechanism in creatine kinase. Crit Rev Biochem Mol Biol 40(1):1–20. https://doi.org/10.1080/10409230590918577
Mühlebach SM, Gross M, Wirz T, Wallimann T, Perriard JC, Wyss M (1994) Sequence homology and structure predictions of the creatine kinase isoenzymes. Mol Cell Biochem 133–134:245–262. https://doi.org/10.1007/BF01267958
Ohren JF, Kundracik ML, Borders CL, Edmiston P, Viola RE (2007) Structural asymmetry and intersubunit communication in muscle creatine kinase. Acta Crystallogr Sect D Biol Crystallogr 63(Pt 3):381–389. https://doi.org/10.1107/S0907444906056204
Price NC (1979) The reaction of rabbit muscle creatine kinase with some derivatives of iodoacetamide. Biochem J 177(2):603–612. https://doi.org/10.1042/bj1770603
Qin W, Khuchua Z, Cheng J, Boero J, Payne RM, Strauss AW (1998) Molecular characterization of the creatine kinases and some historical perspectives. Mol Cell Biochem 184(1–2):153–167
R Core Team (2020) R: a language and environment for statistical computing. https://www.R-project.org/
Rao MJ, Bujacz G, Wlodawer A (1998) Crystal structure of rabbit muscle creatine kinase 1. FEBS Lett 439(1–2):133–137. https://doi.org/10.1016/s0014-5793(98)01355-6
Reddy SR, Watts DC (1978) Inhibition of rabbit muscle creatine kinase by iodomethane proceedings. Biochem Soc Trans 6(3):553–555. https://doi.org/10.1042/bst0060553
Reddy SR, Jones AD, Cross CE, Wong PS-Y, van der Vliet A (2000) Inactivation of creatine kinase by S-glutathionylation of the active-site cysteine residue. Biochem J 347(3):821–827. https://doi.org/10.1042/bj3470821
Ritz C, Baty F, Streibig JC, Gerhard D (2015) Dose-response analysis using R. PLoS ONE 10:e0146021
Rogers GA, Shaltiel N, Boyer PD (1976) Facile alkylation of methionine by benzyl bromide and demonstration of fumarase inactivation accompanied by alkylation of a methionine residue. J Biol Chem 251(18):5711–5717
RStudio Team (2020) RStudio: integrated development environment for R. http://www.rstudio.com/
Schlattner U, Klaus A, Ramirez Rios S, Guzun R, Kay L, Tokarska-Schlattner M (2016) Cellular compartmentation of energy metabolism: creatine kinase microcompartments and recruitment of B-type creatine kinase to specific subcellular sites. Amino Acids 48(8):1751–1774. https://doi.org/10.1007/s00726-016-2267-3
Shen YQ, Tang L, Zhou HM, Lin ZJ (2001) Structure of human muscle creatine kinase. Acta Crystallogr Sect D Biol Crystallogr 57(Pt 8):1196–1200. https://doi.org/10.1107/s0907444901007703
Siegert M, Gandor F, Kranawetvogl A, Börner H, Thiermann H, John H (2019) Methionine329 in human serum albumin: a novel target for alkylation by sulfur mustard. Drug Test Anal 11(5):659–668. https://doi.org/10.1002/dta.2548
Sievers F, Wilm A, Dineen DG, Gibson TJ, Karplus K, Li W, Lopez R, McWilliam H, Remmert M, Söding J, Thompson JD, Higgins DG (2011) Fast, scalable generation of high-quality protein multiple sequence alignments using Clustal Omega. Mol Syst Biol 7:539. https://doi.org/10.1038/msb.2011.75
Steinritz D, Lüling R, Siegert M, Herbert J, Mückter H, Taeger CD, Gudermann T, Dietrich A, Thiermann H, John H (2021) Alkylated epidermal creatine kinase as a biomarker for sulfur mustard exposure: comparison to adducts of albumin and DNA in an in vivo rat study. Arch Toxicol 95(4):1323–1333. https://doi.org/10.1007/s00204-021-03005-3
Taylor MT, Nelson JE, Suero MG, Gaunt MJ (2018) A protein functionalization platform based on selective reactions at methionine residues. Nature 562(7728):563–568. https://doi.org/10.1038/s41586-018-0608-y
Tisi D, Bax B, Loew A (2001) The three-dimensional structure of cytosolic bovine retinal creatine kinase. Acta Crystallogr Sect D Biol Crystallogr 57(Pt 2):187–193. https://doi.org/10.1107/s0907444900015614
Valley CC, Cembran A, Perlmutter JD, Lewis AK, Labello NP, Gao J, Sachs JN (2012) The methionine-aromatic motif plays a unique role in stabilizing protein structure. J Biol Chem 287(42):34979–34991. https://doi.org/10.1074/jbc.M112.374504
Wallimann T, Tokarska-Schlattner M, Schlattner U (2011) The creatine kinase system and pleiotropic effects of creatine. Amino Acids 40(5):1271–1296. https://doi.org/10.1007/s00726-011-0877-3
Wang P-F, Flynn AJ, Naor MM, Jensen JH, Cui G, Merz KM, Kenyon GL, McLeish MJ (2006) Exploring the role of the active site cysteine in human muscle creatine kinase. Biochemistry 45(38):11464–11472. https://doi.org/10.1021/bi0607002
Wickham H, Averick M, Bryan J, Chang W, McGowan L, François R, Grolemund G, Hayes A, Henry L, Hester J, Kuhn M, Pedersen T, Miller E, Bache S, Müller K, Ooms J, Robinson D, Seidel D, Spinu V, Takahashi K, Vaughan D, Wilke C, Woo K, Yutani H (2019) Welcome to the Tidyverse. JOSS 4(43):1686. https://doi.org/10.21105/joss.01686
Wood TD, Guan Z, Borders CL, Chen LH, Kenyon GL, McLafferty FW (1998) Creatine kinase: essential arginine residues at the nucleotide binding site identified by chemical modification and high-resolution tandem mass spectrometry. Proc Natl Acad Sci USA 95(7):3362–3365. https://doi.org/10.1073/pnas.95.7.3362
Acknowledgements
The authors thank Cornelia Muschik, Ram Prasad and Emine Cukur (all Bundeswehr Institute of Pharmacology and Toxicology) for their excellent technical support.
Funding
Open Access funding enabled and organized by Projekt DEAL. Part of the work was supported by the German Research Foundation (Deutsche Forschungsgemeinschaft, DFG, Research Training Group GRK 2338).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Harald Mückter deceased on May 07, 2020.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Steinritz, D., Lüling, R., Siegert, M. et al. Alkylation of rabbit muscle creatine kinase surface methionine residues inhibits enzyme activity in vitro. Arch Toxicol 95, 3253–3261 (2021). https://doi.org/10.1007/s00204-021-03137-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00204-021-03137-6