Abstract
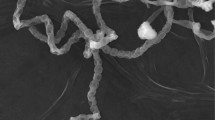
An actinomycete strain, AA8T, which produced a long straight chain of spores (verticillati type), was isolated from the rhizosphere soil of Mangifera indica in Bangkok, Thailand. A polyphasic taxonomic study was carried out to establish the taxonomic position of the strain. Strain AA8T formed a tight taxonomic position in the 16S rRNA gene tree with Streptomyces roseifaciens MBT76T. In contrast, the genome-based taxonomic analysis showed that strain AA8T shared low average nucleotide identity-BLAST (94.1%), the digital DNA–DNA hybridization (58.2%), and the average amino acid identity (93.6%) values with S. roseifaciens MBT76T. Moreover, a combination of physiological and biochemical properties indicated that strain AA8T was distinguished from all Streptomyces species with effectively published names. Strain AA8T, therefore, represents a novel species of Streptomyces, and the name Streptomyces telluris is proposed for the strain. The type strain is AA8T (= TBRC 8483T = NBRC 113461T). The chemical investigation led to the isolation of nine known compounds (compounds 1–9). Among these compounds, compound 7 (3,4-dihydroxybenzaldehyde) possesses strong antioxidant activity equal to ascorbic acid, a powerful antioxidative agent.





Similar content being viewed by others
Data availability
All the data generated in this study have been included in this published article and its supplementary information files. The 16S rRNA gene and the whole-genome sequences of strain AA8T have been deposited in the GenBank databases with the accession numbers LC377949 and JANIID000000000, respectively.
Abbreviations
- AAI:
-
Average amino acid identity
- ANIb:
-
Average nucleotide identity-BLAST
- DAP:
-
Diaminopimelic acid
- dDDH:
-
Digital DNA–DNA hybridization
- GBDP:
-
Genome blast distance phylogeny
- GGDC:
-
Genome-to-Genome Distance Calculator
- GL:
-
Glycolipid
- HPLC:
-
High-performance liquid chromatography
- ISCC-NBS:
-
Inter-Society Color Council-the National Bureau of Standards
- ISP:
-
International Streptomyces Project
- Ls:
-
Unidentified lipids
- MK:
-
Menaquinone
- ML:
-
Maximum-likelihood
- PE:
-
Phosphatidylethanolamine
- NPG:
-
Ninhydrin-positive lipid
- PI:
-
Phosphatidylinositol
- PLs:
-
Unidentified phospholipids
- NJ:
-
Neighbor-joining
- TLC:
-
Thin-layer chromatography
- TYGS:
-
Type strain genome server
References
Alanjary M, Steinke K, Ziemert N (2019) AutoMLST: an automated web server for generating multi-locus species trees highlighting natural product potential. Nucleic Acids Res 47:276–282
Ali MA, Punniyamurthy T (2010) Palladium-catalyzed one-pot conversion of aldehydes to amides. Adv Synth Catal 352:288–292
Arai T (1975) Culture media for actinomycetes. The Society for Actinomycetes, Tokyo
Arumugam M, Mitra A, Jaisankar P et al (2010) Isolation of an unusual metabolite 2-allyloxyphenol from a marine actinobacterium, its biological activities and applications. Appl Microbiol Biotechnol 86:109–117
Bankevich A, Nurk S, Antipov D et al (2012) SPAdes: a new genome assembly algorithm and its applications to single-cell sequencing. J Comput Biol 19:455–477
Blin K, Shaw S, Kloosterman AM et al (2021) antiSMASH 6.0: improving cluster detection and comparison capabilities. Nucleic Acids Res 49:W29–W35
Brand-Williams W, Cuvelier ME, Berset C (1995) Useofafree radical method to evaluate antioxidant activity. Leben Wissens Technol 28:25–30
Chang TS, Tseng M, Ding HY, Tai (2008) Isolation and characterization of Streptomyces hiroshimensis strain TI-C3 with anti-tyrosinase activity. J Cosmet Sci 59:33–40
Cheng C, Othman EM, Reimer A et al (2016) Ageloline A, new antioxidant and antichlamydial quinolone from the marine sponge-derived bacterium Streptomyces sp. SBT345. Tetrahedron Lett 57:2786–2789
Collins MD, Jones D (1980) Lipids in the classification and identification of coryneform bacteria containing peptidoglycans based on 2,4-diaminobutyric acid. J Appl Bacteriol 48:459–470
Collins MD, Pirouz T, Goodfellow M, Minnikin DE (1977) Distribution of menaquinones in actinomycetes and corynebacteria. J Gen Microbiol 100:221–230
Farris JS (1972) Estimating phylogenetic trees from distance matrices. Am Nat 106:645–667
Felsenstein J (1981) Evolutionary trees from DNA sequences: a maximum likelihood approach. J Mol Evol 17:368–376
Felsenstein J (1985) Confidence limits on phylogenies: an approach using the bootstrap. Evolution 39:783–791
Fitch WM (1971) Toward defining the course of evolution: minimum change for a specific tree topology. Syst Zool 20:406–416
Furutani Y, Naganawa H, Takeuchi T, Umezawa H (1977) Isolation and structure of new isocoumarins. Agric Biol Chem 41:1179–1183
Gordon RE, Barnett DA, Handerhan JE, Pang CHN (1974) Nocardia coeliaca, Nocardia autotrophica, and the nocardin strain. Int J Syst Bacteriol 24:54–63
Goris J, Konstantinidis KT, Klappenbach JA, Coenye T, Vandamme P, Tiedje JM (2007) DNA–DNA hybridization values and their relationship to whole-genome sequence similarities. Int J Syst Evol Microbiol 57:81–91
Ishida T, Yoshimura H, Takekawa M et al (2021) Discovery, characterization and functional improvement of kumamonamide as a novel plant growth inhibitor that disturbs plant microtubules. Sci Rep 11:6077
Itoh T, Kudo T, Parenti F, Seino A (1989) Amended description of the genus Kineosporia, based on chemotaxonomic and morphological studies. Int J Syst Bacteriol 39:168–173
Jaroszewski J, Stærk D, Holm-Møller SB et al (2005) Naravelia zeylanica: occurrence of primary benzamides in flowering plants. Nat Prod Res 19:291–294
Kämpfer P (2012) Genus Streptomyces. the actinobacteria. In: Goodfellow M et al (eds) Bergey’s manual of systematic bacteriology, 2nd edn. Springer, New York, pp 1455–1767
Kelly KL (1964) Inter-society color council: National Bureau of Standards Color name charts illustrated with centroid colors. US Government Printing Office, Washington
Kim WG, Kim JP, Kim CJ, Lee KH, Yoo ID (1996) Benzastatins A, B, C, and D: new free radical scavengers from Streptomyces nitrosporeus 30643. I. Taxonomy, fermentation, isolation, physico-chemical properties and biological activities. J Antibiot 49:20–25
Kim WG, Ryoo IJ, Park JS, Yoo ID (2001) Benzastatins H and I, new benzastatin derivatives with neuronal cell protecting activity from Streptomyces nitrosporeus. J Antibiot 54:513–516
Kimura M (1980) A simple method for estimating evolutionary rates of base substitutions through comparative studies of nucleotide sequences. J Mol Evol 16:111–120
Kolehmainen ET, Laihia KP, Hyötyläinen JMI, Kauppinen RT (1995) 1H, 13C and 17O NMR spectral study of chlorinated 3,4-dihydroxybenzaldehydes (protocatechualdehydes). Spectrochim Acta A Mol Biomol Spectrosc 51:419–427
Komagata K, Suzuki KI (1987) Lipid and cell-wall analysis in bacterial systematics. Methods Microbiol 19:161–207
Konstantinidis KT, Rosselló-Móra R, Amann R (2017) Uncultivated microbes in need of their own taxonomy. ISME J 11:2399–2406
Kumar S, Stecher G, Li M, Knyaz C, Tamura K (2018) MEGA X: molecular evolutionary genetics analysis across computing platforms. Mol Biol Evol 35:1547–1549
Lechevalier MP, De Bièvre C, Lechevalier HA (1977) Chemotaxonomy of aerobic actinomycetes: phospholipid composition. Biochem Syst Ecol 5:249–260
Lefort V, Desper R, Gascuel O (2015) FastME 2.0: a comprehensive, accurate, and fast distance-based phylogeny inference program. Mol Biol Evol 32:2798–2800
Li Y, Li XF, Kim DS et al (2003) Indolyl alkaloid derivatives, Nb-acetyltryptamine and oxaline from a marine-derived fungus. Arch Pharm Res 26:21–23
Meier-Kolthoff JP, Göker M (2019) TYGS is an automated high throughput platform for state- of-the-art genome-based taxonomy. Nat Commun 10:2182–2192
Meier-Kolthoff PJ, Alexander FA, Klenk HP, Göker M (2013) Genome sequence-based species delimitation with confidence intervals and improved distance functions. BMC Bioinform 14:60–73
Minnikin DE, O’Donnell AG, Goodfellow M et al (1984) An integrated procedure for the extraction of bacterial isoprenoid quinones and polar lipids. J Microbiol Methods 2:233–241
Parte AC, Sardà Carbasse J, Meier-Kolthoff JP, Reimer LC, Göker M (2020) List of prokaryotic names with standing in nomenclature (LPSN) moves to the DSMZ. Int J Syst Evol Microbiol 70:5607–5612
Phongsopitanun W, Thawai C, Suwanborirux K et al (2014) Streptomyces chumphonensis sp. nov., isolated from marine sediments. Int J Syst Evol Microbiol 64:2605–2610
Prior RL, Wu X, Schaich K (2005) Standardized methods for the determination of antioxidant capacity and phenolics in foods and dietary supplements. J Agric Food Chem 53:4290–4302
Quinn GA, Banat AM, Abdelhameed AM, Banat IM (2020) Streptomyces from traditional medicine: sources of new innovations in antibiotic discovery. J Med Microbiol 69:1040–1048
Richter M, Rosselló-Móra R (2009) Shifting the genomic gold standard for the prokaryotic species definition. Proc Natl Acad Sci U S A 106:19126–19131
Richter M, Rosselló-Móra R, Oliver Glöckner F, Peplies J (2016) J SpeciesWS: a web server for prokaryotic species circumscription based on pairwise genome comparison. Bioinfor 32:929–931
Rodriguez-R LM, Konstantinidis KT (2014) Bypassing cultivation to identify bacterial species. Microbe Mag 9:111–118
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425
Sasser M (1990) Identification of bacteria by gas chromatography of cellular fatty acids. MIDI technical note 101. MIDI Inc., Newark, pp 1–7
Schultz AW, Oh DC, Carney JR (2008) Biosynthesis and structures of cyclomarins and cyclomarazines, prenylated cyclic peptides of marine actinobacterial origin. J Am Chem Soc 130:4507–4516
Ser HL, Tan LTH, Palanisamy UD et al (2016) Streptomyces antioxidans sp. Nov., a novel mangrove soil actinobacterium with antioxidative and neuroprotective potentials. Front Microbiol 7:899
Seyoum A, Asres K, El-Fiky FK (2006) Structure-radical scavenging activity relationships of flavonoids. Phytochemistry 67:2058–2070
Shirling EB, Gottlieb D (1966) Methods for characterization of Streptomyces species. Int J Syst Bacteriol 16:313–340
Staneck JL, Roberts GD (1974) Simplified approach to identification of aerobic actinomycetes by thin-layer chromatography. Appl Microbiol 28:226–231
Supong K, Suriyachadkun C, Pittayakhajonwut P, Suwanborirux K, Thawai C (2013) Micromonospora spongicola sp. nov., an actinomycete isolated from a marine sponge in the Gulf of Thailand. J Antibiot 66:505–509
Supong K, Thawai C, Supothina S et al (2016) Antimicrobial and anti-oxidant activities of quinoline alkaloids from Pseudomonas aeruginosa BCC76810. Phytochem Lett 17:100–106
Tamaoka J (1994) Determination of DNA base composition. In: Goodfellow M, O’Donnell AG (eds) Chemical methods on prokaryotic systematics. Wiley, Chichester, pp 463–470
Tan LTH, Chan KG, Khan TM et al (2017) Streptomyces sp MUM212 as a source of antioxidants with radical scavenging and metal chelating properties. Front Pharmacol 8:276
Thawai C, Rungjindamai N, Klanbut K, Tanasupawat S (2017) Nocardia xestospongiae sp. nov., isolated from a marine sponge in the Andaman Sea. Int J Syst Evol Microbiol 67:1451–1456
Tietz JI, Schwalen CJ, Patel PS et al (2017) A new genome-mining tool redefines the lasso peptide biosynthetic landscape. Nat Chem Biol 13:470–478
van der Aart LT, Nouioui I, Kloosterman A et al (2019) Polyphasic classification of the gifted natural product producer Streptomyces roseifaciens sp. nov. Int J Syst Evol Microbiol 69:899–908
Williams ST, Cross T (1971) Chapter XI actinomycetes. In: Booth C et al (eds) Methods in microbiology. Academic Press, London, pp 295–334
Yang D, Fu HA (2010) Simple and practical copper-catalyzed approach to substituted phenols from aryl halides by using water as the solvent. Chemistry 16:2366–2370
Yoon SH, Ha SM, Kwon S et al (2017) Introducing EzBioCloud: a taxonomically united database of 16S rRNA and whole-genome assemblies. Int J Syst Evol Microbiol 67:1613–1617
Zdero C, Jakupovic J, Bohlmann F (1990) Diterpenes and other constituents from Pteronia species. Phytochemistry 29:1231–1245
Zinad DS, Shaaban KA, Abdalla MA et al (2011) Bioactive isocoumarins from a terrestrial Streptomyces sp. ANK302. Nat Prod Commun 6:1934578X1100600111
Acknowledgements
We thank the Actinobacterial Research Unit (ARU), School of Science, King Mongkut’s Institute of Technology Ladkrabang, for laboratory support.
Funding
The authors gratefully acknowledge the funding support from the National Science, Research and Innovation Fund (NSRF) (Grant number: RE-KRIS/FF65/23) to CT.
Author information
Authors and Affiliations
Contributions
OT: formal analysis, methodology, resources, writing original draft preparation. TD and ST: chemotaxonomic analyses and biological activity assay. PP and CI: chemotaxonomic analyses, and manuscript editing. CS: data curation, and resources. ST and Y-WH: manuscript editing. CT: conceptualization, data curation, funding acquisition, project administration, resources, validation, and manuscript editing. All authors have read and approved the final version of the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they do not have any conflicts of interest.
Additional information
Communicated by Wen-Jun Li.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Thayanuwadtanawong, O., Duangupama, T., Bunbamrung, N. et al. Streptomyces telluris sp. nov., a promising terrestrial actinobacterium with antioxidative potentials. Arch Microbiol 205, 247 (2023). https://doi.org/10.1007/s00203-023-03585-x
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00203-023-03585-x