Abstract
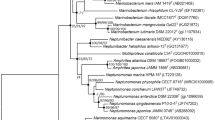
The bacterial strain AP-MA-4T isolated from the marine dinoflagellate Alexandrium pacificum (KCTC AG60911), was subjected to a taxonomic analysis. Cells of strain AP-MA-4T were Gram-stain-negative, aerobic, rod-shaped, optimum growth at 20 °C, pH 7.0, in the presence of 5% (w/v) NaCl. Strain AP-MA-4T shared the highest 16S rRNA gene sequence similarity to Pseudosulfitobacter pseudonitzschiae DSM 26824T (98.5%), followed by Ascidiaceihabitans donghaensis RSS1-M3T (96.3%), Pseudoseohaeicola caenipelagi BS-W13T (95.7%), and Sulfitobacter pontiacus CHLG 10T (95.3%). Based on 16S rRNA phylogeny, strain AP-MA-4T is phylogenetically closely related to Pseudosulfitobacter pseudonitzschiae (type species of Pseudosulfitobacter) and could be distinguished from the type species based on their phenotypic properties. The genome length of strain AP-MA-4T was 3.48 Mbp with a 62.9% G + C content. The average nucleotide identity (ANI) and digital DNA–DNA hybridization (dDDH) values between strain AP-MA-4 T and its closely related type strains were 72.2–83.3 and 18.2–27.6%, respectively. Summed feature 8 (C18:1ω7c and/or C18:1ω6c) was identified the major fatty acids (> 10%). Phosphatidylglycerol (PG), phosphatidylethanolamine (PE), and phospholipid (PL) were demonstrated as the major polar lipids. The major respiratory quinone is ubiquinone-10 (Q-10). Based on genotypic and phenotypic features, strain AP-MA-4T (= KCTC 92289T = GDMCC 1.3585T) represents a new Pseudosulfitobacter species, in which the name Pseudosulfitobacter koreense sp. nov. is proposed.


Similar content being viewed by others
Data availability
The GenBank accession numbers of the 16S rRNA gene and the genome sequences of the strain AP-MA-4T are ON979826 and JANKJG000000000, respectively. This strain can be obtained from the Korea Collection for Type Cultures (KCTC 92289T) and the Guangdong Microbial Culture Collection Center (GDMCC 1.3585T).
Abbreviations
- ANI:
-
Average nucleotide identity
- dDDH:
-
Digital DNA-DNA hybridization
- GGDC:
-
Genome-to-genome distance calculation
- LBA:
-
Luria–Bertani agar
- R2A:
-
Reasoner’s 2A agar
- MA:
-
Marine agar 2216
- TSA:
-
Tryptic soy agar
- PDA:
-
Potato dextrose agar
- NA:
-
Nutrient agar
References
Arndt D, Marcu A, Liang Y, Wishart DS (2019) PHAST, PHASTER and PHASTEST: tools for finding prophage in bacterial genomes. Brief Bioinform 20:1560–1567. https://doi.org/10.1093/bib/bbx121
Asnicar F et al (2020) Precise phylogenetic analysis of microbial isolates and genomes from metagenomes using PhyloPhlAn 3.0. Nat Commun 11:2500. https://doi.org/10.1038/s41467-020-16366-7
Cho S-H et al (2020) Elucidation of the biosynthetic pathway of vitamin B groups and potential secondary metabolite gene clusters via genome analysis of a marine bacterium Pseudoruegeria sp. M32A2M. J Microbiol 30(4):505–514. https://doi.org/10.4014/jmb.1911.11006
Chun J et al (2018) Proposed minimal standards for the use of genome data for the taxonomy of prokaryotes. Int J Syst Evol Microbiol 68:461–466. https://doi.org/10.1099/ijsem.0.002516
Collins MD, Jones D (1981) Distribution of isoprenoid quinone structural types in bacteria and their taxonomic implication. Clin Microbiol Rev 45:316–354. https://doi.org/10.1128/MMBR.45.2.316-354.1981
De Swaaf ME, Sijtsma L, Pronk JT (2003) High-cell-density fed-batch cultivation of the docosahexaenoic acid producing marine alga Crypthecodinium cohnii. Front Bioeng Biotechnol 81:666–672. https://doi.org/10.1002/bit.10513
Du Y et al (2019) Comparative genomic analysis of Bacillus paralicheniformis MDJK30 with its closely related species reveals an evolutionary relationship between B. paralicheniformis and B. licheniformis. BMC Genom 20:283. https://doi.org/10.1186/s12864-019-5646-9
Felsenstein J (1981) Evolutionary trees from DNA sequences: a maximum likelihood approach. J Mol Evol 17:368–376. https://doi.org/10.1007/BF01734359
Felsenstein J (1985) Confidence limits on phylogenies: an approach using the bootstrap. Evolution 39:783–791. https://doi.org/10.2307/2408678
Galperin MY, Wolf YI, Makarova KS, Vera Alvarez R, Landsman D, Koonin EV (2021) COG database update: focus on microbial diversity, model organisms, and widespread pathogens. Nucleic Acids Res 49:D274–D281. https://doi.org/10.1093/nar/gkaa1018
Graham ED, Heidelberg JF, Tully BJ (2018) Potential for primary productivity in a globally-distributed bacterial phototroph. ISME J 12:1861–1866. https://doi.org/10.1038/s41396-018-0091-3
Hall TA (1999) BioEdit: a user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucleic Acids Symp Ser 41:95–98. https://doi.org/10.1017/CBO9780511790492.007
Hong Z et al (2015) Sulfitobacter pseudonitzschiae sp. nov., isolated from the toxic marine diatom Pseudo-nitzschia multiseries. Int J Syst Evol Microbiol 65:95–100. https://doi.org/10.1099/ijs.0.064972-0
Jagessar KL, Mchaourab HS, Claxton DP (2019) The N-terminal domain of an archaeal multidrug and toxin extrusion (MATE) transporter mediates proton coupling required for prokaryotic drug resistance. J Biol Chem 294:12807–12814. https://doi.org/10.1074/jbc.RA119.009195
Jiang L et al (2020) Saccharibacillus brassicae sp. nov., an endophytic bacterium isolated from kimchi cabbage (Brassica rapa subsp. pekinensis) seeds. J Microbiol 58:24–29. https://doi.org/10.1007/s12275-020-9346-6
Kellmann R, Mihali TK, Jeon YJ, Pickford R, Pomati F, Neilan BA (2008) Biosynthetic intermediate analysis and functional homology reveal a saxitoxin gene cluster in cyanobacteria. Appl Environ Microbiol 74:4044–4053. https://doi.org/10.1128/AEM.00353-08
Kim Y-O et al (2014) Ascidiaceihabitans donghaensis gen. nov., sp. nov., isolated from the golden sea squirt Halocynthia aurantium. Int J Syst Evol Microbiol 64:3970–3975. https://doi.org/10.1099/ijs.0.066399-0
Kimura M (1980) A simple method for estimating evolutionary rates of base substitutions through comparative studies of nucleotide sequences. J Mol Evol 16:111–120. https://doi.org/10.1007/BF01731581
Konstantinidis KT, Tiedje JM (2005) Towards a genome-based taxonomy for prokaryotes. J Bacteriol 187:6258–6264. https://doi.org/10.1128/JB.187.18.6258-6264.2005
Kumar S, Stecher G, Tamura K (2016) MEGA7: molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol Biol Evol 33:1870–1874. https://doi.org/10.1093/molbev/msw054
Li Z et al (2019) Discovery of a new clade nested within the genus Alexandrium (Dinophyceae): morpho-molecular characterization of Centrodinium punctatum (Cleve) F.J.R. Taylor. Protist 170:168–186. https://doi.org/10.1016/j.protis.2019.02.003
Liang KY, Orata FD, Boucher YF, Case RJ (2021) Roseobacters in a sea of poly-and paraphyly: whole genome-based taxonomy of the family Rhodobacteraceae and the proposal for the split of the “Roseobacter clade” into a novel family. Roseobacteraceae Fam Nov Front Microbiol 12:1635. https://doi.org/10.3389/fmicb.2021.683109
Luo C, Rodriguez-r LM, Konstantinidis KT (2014) MyTaxa: an advanced taxonomic classifier for genomic and metagenomic sequences. Nucleic Acids Res 42:e73–e73. https://doi.org/10.1093/nar/gku169
Manni M, Berkeley MR, Seppey M, Simão FA, Zdobnov EM (2021) BUSCO update: novel and streamlined workflows along with broader and deeper phylogenetic coverage for scoring of eukaryotic, prokaryotic, and viral genomes. Mol Biol Evol 38:4647–4654. https://doi.org/10.1093/molbev/msab199
Meier-Kolthoff JP, Auch AF, Klenk HP, Goker M (2013) Genome sequence-based species delimitation with confidence intervals and improved distance functions. BMC Bioinform 186:1471–2105. https://doi.org/10.1186/1471-2105-14-60
Meier-Kolthoff JP, Göker M (2019) TYGS is an automated high-throughput platform for state-of-the-art genome-based taxonomy. Nat Commun 10:283. https://doi.org/10.1038/s41467-019-10210-3
Minnikin D et al (1984) An integrated procedure for the extraction of bacterial isoprenoid quinones and polar lipids. J Microbiol Methods 2:233–241. https://doi.org/10.1016/0167-7012(84)90018-6
Moran M et al (2007) Ecological genomics of marine Roseobacters. Appl Environ 73:4559–4569. https://doi.org/10.1128/AEM.02580-06
Murray SA, Diwan R, Orr RJ, Kohli GS, John U (2015) Gene duplication, loss and selection in the evolution of saxitoxin biosynthesis in alveolates. Mol Phylogenet Evol 92:165–180. https://doi.org/10.1016/j.ympev.2015.06.017
Na SI, Kim YO, Yoon SH, Ha SM, Baek I, Chun J (2018) UBCG: Up-to-date bacterial core gene set and pipeline for phylogenomic tree reconstruction. J Microbiol 56:280–285. https://doi.org/10.1007/s12275-018-8014-6
Nei M, Kumar S (2000) Molecular evolution and phylogenetics. Mol Biol Evol. https://doi.org/10.1016/S1055-7903(02)00247-6
Orr RJ, Stüken A, Murray SA, Jakobsen KS (2013) Evolution and distribution of saxitoxin biosynthesis in dinoflagellates. Mar Drugs 11:2814–2828. https://doi.org/10.3390/md11082814
Overbeek R et al (2014) The SEED and the Rapid Annotation of microbial genomes using Subsystems Technology (RAST). Nucleic Acids Res 42:D206-214. https://doi.org/10.1093/nar/gkt1226
Park S, Park J-M, Kang C-H, Kim S-G, Yoon J-H (2015) Pseudoseohaeicola caenipelagi gen. nov., sp. nov., isolated from a tidal flat. Int J Syst Evol Microbiol 65:1819–1824. https://doi.org/10.1099/ijs.0.000182
Rodriguez-R LM, Konstantinidis KT (2014) Bypassing cultivation to identify bacterial species. Microbe 9:111–118. https://doi.org/10.1128/microbe.9.111.1
Rojas-Garbanzo C, Gleichenhagen M, Heller A, Esquivel P, Schulze-Kaysers N, Schieber A (2017) Carotenoid profile, antioxidant capacity, and chromoplasts of pink guava (Psidium guajava L. Cv. ‘Criolla’) during fruit ripening. Food Chem 65:3737–3747. https://doi.org/10.1021/acs.jafc.6b04560
Rzhetsky A, Nei M (1992) A simple method for estimating and testing minimum-evolution trees. Mol Biol Evol. https://doi.org/10.1093/oxfordjournals.molbev.a040771
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425. https://doi.org/10.1093/oxfordjournals.molbev.a040454
Sasser M (2006) Bacterial identification by gas chromatographic analysis of fatty acids methyl esters (GC-FAME). J Chromatogr Sci. https://doi.org/10.1093/chromsci/8.5.279
Seemann T (2014) Prokka: rapid prokaryotic genome annotation. Bioinformatics 30:2068–2069. https://doi.org/10.1093/bioinformatics/btu153
Sorokin DY (1995) Sulfitobacter pontiacus gen. nov., sp. nov.-a new heterotrophic bacterium from the Black Sea, specialized on sulfite oxidation. Syst Appl Microbiol 64:295–295. https://doi.org/10.1016/j.syapm.2005.05.006
Stüken A, Orr RJ, Kellmann R, Murray SA, Neilan BA, Jakobsen KS (2011) Discovery of nuclear-encoded genes for the neurotoxin saxitoxin in dinoflagellates. PLoS One 6:e20096. https://doi.org/10.1371/journal.pone.0020096
Tamura K, Stecher G, Kumar S (2021) MEGA11: molecular evolutionary genetics analysis version 11. Mol Biol Evol 38:3022–3027. https://doi.org/10.1093/molbev/msab120
Tatusova T et al (2016) NCBI prokaryotic genome annotation pipeline. Nucleic Acids Res 44:6614–6624. https://doi.org/10.1093/nar/gkw569
Ventosa A, Marquez MC, Kocur M, Tindall BJ (1993) Comparative study of “Micrococcus sp”. strains CCM 168 and CCM 1405 and members of the genus Salinicoccus. Int J Syst Evol Microbiol 43:245–248. https://doi.org/10.1099/00207713-43-2-245
Wagner-Döbler I, Biebl H (2006) Environmental biology of the marine Roseobacter lineage. Annu Rev Microbiol 60:255–280. https://doi.org/10.5846/stxb201310172507
Wang H, Guo R, Lim W-A, Allen AE, Ki J-S (2020) Comparative transcriptomics of toxin synthesis genes between the non-toxin producing dinoflagellate Cochlodinium polykrikoides and toxigenic Alexandrium pacificum. Harmful Algae 93:101777. https://doi.org/10.1016/j.hal.2020.101777
Weisburg WG, Barns SM, Pelletier DA, Lane DJ (1991) 16S ribosomal DNA amplification for phylogenetic study. J Bacteriol 173:697–703. https://doi.org/10.1128/jb.173.2.697-703.1991
Wirth JS, Whitman WB (2018) Phylogenomic analyses of a clade within the roseobacter group suggest taxonomic reassignments of species of the genera Aestuariivita, Citreicella, Loktanella, Nautella, Pelagibaca, Ruegeria, Thalassobius, Thiobacimonas and Tropicibacter, and the proposal of six novel genera. Int J Syst Evol Microbiol 68:2393–2411. https://doi.org/10.1099/ijsem.0.002833
Xu L et al (2019) OrthoVenn2: a web server for whole-genome comparison and annotation of orthologous clusters across multiple species. Nucleic Acids Res 47:W52–W58. https://doi.org/10.1093/nar/gkz333
Yoon SH et al (2017a) Introducing EzBioCloud: a taxonomically united database of 16S rRNA gene sequences and whole-genome assemblies. Int J Syst Evol Microbiol 67:1613–1617. https://doi.org/10.1099/ijsem.0.001755
Yoon SH, Ha SM, Lim J, Kwon S, Chun J (2017b) A large-scale evaluation of algorithms to calculate average nucleotide identity. Antonie Van Leeuwenhoek 110:1281–1286. https://doi.org/10.1007/s10482-017-0844-4
Yu G, Lam TT-Y, Zhu H, Guan Y (2018) Two methods for mapping and visualizing associated data on phylogeny using ggtree. Mol Biol Evol 35:3041–3043. https://doi.org/10.1093/molbev/msy194
Funding
This work was supported by grant from the Korea Research Institute of Bioscience and Biotechnology (KRIBB) Research Initiative Program (KGM5232322) and by the National Research Foundation of Korea (NRF) grant funded by the Korea government (MSIT) (No. 2021R1C1C1008377).
Ethics declarations
Conflict of interest
The authors declare that there are no conflicts of interest.
Additional information
Communicated by Wen-Jun Li.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Jiang, Y., Li, Z. Identification and genomic analysis of Pseudosulfitobacter koreense sp. nov. isolated from toxin-producing dinoflagellate Alexandrium pacificum. Arch Microbiol 205, 245 (2023). https://doi.org/10.1007/s00203-023-03583-z
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00203-023-03583-z