Abstract
We describe a new species of purple sulfur bacteria (Chromatiaceae, anoxygenic phototrophic bacteria) isolated from a microbial mat in the sulfidic geothermal outflow of a hot spring in Rotorua, New Zealand. This phototroph, designated as strain NZ, grew optimally near 45 °C but did not show an absorption maximum at 915 nm for the light-harvesting-reaction center core complex (LH1-RC) characteristic of other thermophilic purple sulfur bacteria. Strain NZ had a similar carotenoid composition as Thermochromatium tepidum, but unlike Tch. tepidum, grew photoheterotrophically on acetate in the absence of sulfide and metabolized thiosulfate. The genome of strain NZ was significantly larger than that of Tch. tepidum but slightly smaller than that of Allochromatium vinosum. Strain NZ was phylogenetically more closely related to mesophilic purple sulfur bacteria of the genus Allochromatium than to Tch. tepidum. This conclusion was reached from phylogenetic analyses of strain NZ genes encoding 16S rRNA and the photosynthetic functional gene pufM, from phylogenetic analyses of entire genomes, and from a phylogenetic tree constructed from the concatenated sequence of 1090 orthologous proteins. Moreover, average nucleotide identities and digital DNA:DNA hybridizations of the strain NZ genome against those of related species of Chromatiaceae supported the phylogenetic analyses. From this collection of properties, we describe strain NZ here as the first thermophilic species of the genus Allochromatium, Allochromatium tepidum NZT, sp. nov.






Similar content being viewed by others
Code availability
Not applicable.
Change history
11 March 2022
A Correction to this paper has been published: https://doi.org/10.1007/s00203-022-02827-8
Abbreviations
- Tch. :
-
Thermochromatium
- Alc.:
-
Allochromatium
- Ccm. :
-
Caldichromatium
- Cba.:
-
Chlorobaculum
- Bchl:
-
Bacteriochlorophyll
- LH:
-
Light-harvesting
References
Achenbach LA, Carey J, Madigan MT (2001) Photosynthetic and phylogenetic primers for detection of anoxygenic phototrophs in natural environments. Appl Environ Microbiol 67:2922–2926. https://doi.org/10.1128/AEM.67.7.2922-2926.2001
Brito JA, Denkmann K, Pereira IA, Archer M, Dahl C (2015) Thiosulfate dehydrogenase (TsdA) from Allochromatium vinosum: structural and functional insights into thiosulfate oxidation. J Biol Chem 291:24804–24818. https://doi.org/10.1074/jbc.M114.623397
Castenholz RW, Bauld J, Jørgensen BB (1990) Anoxygenic microbial mats of hot springs: thermophilic Chlorobium sp. FEMS Microbiol Ecol 72:325–336. https://doi.org/10.1111/j.1574-6968.1990.tb04079.x
Chin CS, Alexander DH, Marks P, Klammer AA, Drake J, Heiner C, Clum A, Copeland A, Huddleston J, Eichler EE, Turner SW, Korlach J (2013) Nonhybrid, finished microbial genome assemblies from long-read SMRT sequencing data. Nat Methods 10:563–569. https://doi.org/10.1038/nmeth.2474
Dahl C, Franz B, Hensen D, Kesselheim A, Zigann R (2013) Sulfite oxidation in the purple sulfur bacterium Allochromatium vinosum: identification of SoeABC as a major player and relevance of SoxYZ in the process. Microbiology 159:2626–2638. https://doi.org/10.1099/mic.0.071019-0
Dewey ED, Stokes LM, Burchell BM et al (2020) Analysis of the complete genome of the alkaliphilic and phototrophic Firmicute Heliorestis convoluta strain HHT. Microorganisms 8:313. https://doi.org/10.3390/microorganisms8030313
Emms DM, Kelly S (2019) OrthoFinder: phylogenetic orthology inference for comparative genomics. Genome Biol 20:238. https://doi.org/10.1186/s13059-019-1832-y
Finney AJ, Sargent F (2019) Formate hydrogenlyase: a group 4 [NiFe]-hydrogenase in tandem with a formate dehydrogenase. Adv Microb Physiol 74:465–486. https://doi.org/10.1016/bs.ampbs.2019.02.004
Garcia D, Parot P, Verméglio A, Madigan MT (1986) The light-harvesting complexes of a thermophilic purple sulfur photosynthetic bacterium Chromatium tepidum. Biochim Biophys Acta Bioenerg 850:390–395. https://doi.org/10.1016/0005-2728(86)90195-7
Gardiner AT, Nguyen-Phan TC, Cogdell RJ (2020) A comparative look at structural variations among RC–LH1 ‘core’ complexes present in anoxygenic phototrophic bacteria. Photosyn Res 145:83–96. https://doi.org/10.1007/s11120-020-00758-3
Ghosh W, Dam B (2009) Biochemistry and molecular biology of lithotrophic sulfur oxidation by taxonomically and ecologically diverse bacteria and archaea. FEMS Microbiol Revs 33:999–1043. https://doi.org/10.1111/j.1574-6976.2009.00187.x
Hashwa F (1975) Thiosulfate metabolism in some red phototrophic bacteria. Plant Soil 43:41–47. https://doi.org/10.1007/BF01928475
Heda GD, Madigan MT (1988) Thermal properties and oxygenase activity of ribulose 1,5 bisphosphate carboxylase from the thermophilic purple bacterium, Chromatium tepidum. FEMS Microbiol Letts 51:45–50. https://doi.org/10.1111/j.1574-6968.1988.tb02966.x
Heda GD, Madigan MT (1989) Purification and characterization of the thermostable ribulose-1,5 bisphosphate carboxylase/oxygenase from the thermophilic purple bacterium Chromatium tepidum. Eur J Biochem 184:313–319. https://doi.org/10.1111/j.1432-1033.1989.tb15021.x
Hernandez JA, Curatti L, Aznar CP, Perova Z, Britt RD, Rubio LM (2008) Metal trafficking for nitrogen fixation: NifQ donates molybdenum to NifEN/NifH for the biosynthesis of the nitrogenase FeMo-cofactor. Proc Natl Acad Sci U S A 105:11679–11684. https://doi.org/10.1073/pnas.0803576105
Holm HW, Vennes JW (1970) Occurrence of purple sulfur bacteria in a sewage treatment lagoon. Appl Microbiol 19:988–996. https://doi.org/10.1128/am.19.6.988-996.1970
Imhoff JF (2005) Genus II. Allochromatium. In: Brenner DJ, Krieg NR, Staley JT, Garrity GM (eds) Bergey’s manual of systematic bacteriology, vol 2B, 2nd edn. Springer, New York, pp 12–14
Kämpf C, Pfennig N (1980) Capacity of Chromatiaceae for chemotrophic growth. Specific respiration rates of Thiocystis violacea and Chromatium vinosum. Arch Microbiol 127:125–135. https://doi.org/10.1007/BF00428016
Katoh K, Standley DM (2014) MAFFT: iterative refinement and additional methods. Methods Mol Biol 1079:131–146. https://doi.org/10.1007/978-1-62703-646-7_8
Kim M, Oh H-Y, Park S-C, Chun J (2014) Towards a taxonomic coherence between average nucleotide identity and 16S rRNA gene sequence similarity for species demarcation of prokaryotes. Intl J Syst Evol Microbiol 64:346–351. https://doi.org/10.1099/ijs.0.059774-0
Kimble LK, Mandelco L, Woese CR, Madigan MT (1995) Heliobacterium modesticaldum, sp. nov., a thermophilic heliobacterium of hot springs and volcanic soils. Arch Microbiol 163:259–267. https://doi.org/10.1007/BF00393378
Kimura Y, Hirano Y, Yu L-J, Suzuki H, Kobayashi M, Wang Z-Y (2008) Calcium ions are involved in the unusual red shift of the light-harvesting 1 Qy transition of the core complex in thermophilic purple sulfur bacterium Thermochromatium tepidum. J Biol Chem 283:13867–13873. https://doi.org/10.1074/jbc.M800256200
Kimura Y, Yu L-J, Hirano Y, Suzuki H, Wang Z-Y (2009) Calcium ions are required for the enhanced thermal stability of the light-harvesting-reaction center core complex from thermophilic purple sulfur bacterium Thermochromatium tepidum. J Biol Chem 284:93–99. https://doi.org/10.1074/jbc.M806840200
Kimura Y, Lyu S, Okoshi A, Okazaki K, Nakamura N, Ohashi A, Ohno T, Kobayashi M, Imanishi M, Takaichi S, Madigan MT, Wang-Otomo Z-Y (2017) Effects of calcium ions on the thermostability and spectroscopic properties of the LH1-RC complex from a new thermophilic purple bacterium Allochromatium tepidum. J Phys Chem B 121:5025–5032. https://doi.org/10.1021/acs.jpcb.7b03341
Kishi R, Imanishi M, Kobayashi M, Takenaka S, Madigan MT, Wang-Otomo Z-Y, Kimura Y (2021) Quinone transport in the closed light-harvesting 1 reaction center complex from the thermophilic purple bacterium Thermochromatium tepidum. Biochim Biophys Acta Bioenerg 1862:148307. https://doi.org/10.1016/j.bbabio.2020.148307
Kumar S, Stecher G, Li M, Knyaz C, Tamura K (2018) MEGA X: molecular evolutionary genetics analysis across computing platforms. Mol Biol Evol 35:1547–1549. https://doi.org/10.1093/molbev/msy096
Lefort V, Desper R, Gascuel O (2015) FastME 2.0: a comprehensive, accurate, and fast distance-based phylogeny inference program. Mol Biol Evol 32:2798–2800. https://doi.org/10.1093/molbev/msv150
Long M, Liu J, Chen Z, Bleijlevens B, Roseboom W, Albracht SPJ (2007) Characterization of a HoxEFUYH type of [NiFe] hydrogenase from Allochromatium vinosum and some EPR and IR properties of the hydrogenase module. J Biol Inorg Chem 12:62–78. https://doi.org/10.1007/s00775-006-0162-1
Madigan MT (1984) A novel photosynthetic purple bacterium isolated from a Yellowstone hot spring. Science 225:313–315. https://doi.org/10.1126/science.225.4659.313
Madigan MT (1986) Chromatium tepidum sp. nov., a thermophilic photosynthetic bacterium of the family Chromatiaceae. Int J Syst Bacteriol 36:222–227. https://doi.org/10.1099/00207713-36-2-222
Madigan MT (1995) Microbiology of nitrogen fixation by anoxygenic photosynthetic bacteria. In: Blankenship RE, Madigan MT, Bauer CE (eds) anoxygenic photosynthetic bacteria. Kluwer, Dordrecht, pp 915–928
Madigan MT, Takigiku R, Lee RG, Gest H, Hayes JM (1989) Carbon isotope fractionation by thermophilic phototrophic sulfur bacteria: evidence for autotrophic growth in natural populations. Appl Environ Microbiol 55:639–644. https://doi.org/10.1128/aem.55.3.639-644.1989
Madigan MT, Jung DO, Woese CR, Achenbach LA (2000) Rhodoferax antarcticus sp. nov., a moderately psychrophilic purple nonsulfur bacterium isolated from an Antarctic microbial mat. Arch Microbiol 173:269–277. https://doi.org/10.1007/s002030000140
Meier-Kolthoff JP, Auch AF, Klenk H-P, Göker M (2013) Genome sequence-based species delimitation with confidence intervals and improved distance functions. BMC Bioinformatics 14:60. https://doi.org/10.1186/1471-2105-14-60
Meier-Kolthoff JP, Carbasse JS, Peinado-Olarte RL, Göker M (2021) TYGS and LPSN: a database tandem for fast and reliable genome-based classification and nomenclature of prokaryotes. Nucl Acids Res. https://doi.org/10.1093/nar/gkab902
Nagashima KVP, Sasaki M, Hashimoto K, Takaichi S, Nakashima S, Yu LJ, Abe Y, Gotou K, Kawakami T, Takenouchi M, Shibuya Y, Yamaguchi A, Ohno T, Shen JR, Inoue K, Madigan MT, Kimura Y, Wang-Otomo Z-Y (2017) Probing structure-function relationships in early events in photosynthesis using a chimeric photocomplex. Proc Nat Acad Sci U S A 114:10906–10911. https://doi.org/10.1073/pnas.1703584114
Nagatsuma S, Gotou K, Yamashita T, Yu L-J, Shen J-R, Madigan MT, Kimura Y, Wang-Otomo Z-Y (2019) Phospholipid distributions in purple phototrophic bacteria and LH1-RC core complexes. Biochim Biophys Acta Bioenerg 1860:461–468. https://doi.org/10.1016/j.bbabio.2019.04.001
Pfennig N, Trüper HG (2013) The family Chromatiaceae. In: Balows A, Trüper HG, Dworkin M, Harder W, Schleifer K-H (eds) The prokaryotes: a handbook on the biology of bacteria: ecophysiology, isolation, identification, applications. Springer, New York, pp 3200–3221
Revsbech NP, Ward DM (1984) Microelectrode studies of interstitial water chemistry and photosynthetic activity in a hot spring microbial mat. Appl Environ Microbiol 48:270–275. https://doi.org/10.1128/aem.48.2.270-275.1984
Saini MK, ChihChe W, Soulier N, Sebastian A, Albert I, Thiel V, Bryant DA, Hanada S, Tank M (2020) Caldichromatium japonicum gen. nov., sp. nov., a novel thermophilic phototrophic purple sulphur bacterium of the Chromatiaceae isolated from Nakabusa hot springs, Japan. Int J Syst Evol Microbiol 70:5701–5710. https://doi.org/10.1099/ijsem.0.004465
Sattley WM, Madigan MT, Swingley WD et al (2008) The genome of Heliobacterium modesticaldum, a phototrophic representative of the Firmicutes containing the simplest photosynthetic apparatus. J Bacteriol 190:4687–4696. https://doi.org/10.1128/JB.00299-08
Sattley WM, Swingley WD, Burchell BM, Dewey ED, Hayward MK, Renbarger TL, Shaffer KN, Stokes LM, Gurbani SA, Kujawa CM, Nuccio DA, Schladweiler J, Touchman JW, Wang-Otomo Z-Y, Blankenship RE, Madigan MT (2021) Complete genome sequence of the thermophilic purple sulfur bacterium Thermochromatium tepidum compared to Allochromatium vinosum and other Chromatiaceae. Photosyn Res. https://doi.org/10.1007/s11120-021-00870-y
Stamatakis A (2014) RAxML version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 30:1312–1313. https://doi.org/10.1093/bioinformatics/btu033
Stecher G, Tamura K, Kumar S (2020) Molecular evolutionary genetics analysis (MEGA) for macOS. Mol Biol Evol 37:1237–1239. https://doi.org/10.1093/molbev/msz312
Stockdreher Y, Sturm M, Josten M, Sahl H-G, Dobler N, Zigann R, Dahl C (2014) New proteins involved in sulfur trafficking in the cytoplasm of Allochromatium vinosum. J Biol Chem 289:12390–12403. https://doi.org/10.1074/jbc.M113.536425
Stoffels L, Krehenbrink M, Berks BC, Unden G (2012) Thiosulfate reduction in Salmonella enterica is driven by the proton motive force. J Bacteriol 194:475–485. https://doi.org/10.1128/JB.06014-11
Suzuki H, Hirano Y, Kimura Y, Takaichi S, Kobayashi M, Miki K, Wang Z-Y (2007) Purification, characterization and crystallization of the core complex from thermophilic purple sulfur bacterium Thermochromatium tepidum. Biochim Biophys Acta Bioenerg 1767:1057–1063. https://doi.org/10.1016/j.bbabio.2007.06.002
Tabita FR, Hanson TE, Li H, Satagopan S, Singh J, Chan S (2020) Function, structure and evolution of the RubisCO-like proteins and their RubisCO homologs. Microbiol Mol Biol Rev 74:576–599. https://doi.org/10.1128/MMBR.00015-07
Takaichi S (1999) Carotenoids and carotenogenesis in anoxygenic photosynthetic bacteria. In: Frank HA, Young A, Britton G, Cogdell RJ (eds) The photochemistry of carotenoids. Kluwer, Dordrecht, pp 39–69
Takaichi S, Wang Z-Y, Umetsu M, Nozawa T, Shimada K, Madigan MT (1997) New carotenoids from the thermophilic green sulfur bacterium Chlorobium tepidum: 1′2′-dihydro-γ-carotene, 1′2′-dihydrochlorobactene, and OH-chlorobactene glucoside ester, and the carotenoid composition of different strains. Arch Microbiol 168:270–276. https://doi.org/10.1007/s0020300504
Talavera G, Castresana J (2007) Improvement of phylogenies after removing divergent and ambiguously aligned blocks from protein sequence alignments. Syst Biol 56:564–577. https://doi.org/10.1080/10635150701472164
Tani K, Kobayashi K, Hosogi N, Ji XC, Nagashima S, Nagashima KVP, Tsukatani Y, Kanno R, Hall M, Yu, LJ, Ishikawa I, Okura Y, Madigan MT, Mizoguchi A, Humbel BM, Kimura Y, Wang-Otomo ZY (2021) A Ca2+-binding motif underlies the unusual properties of certain photosynthetic bacterial core light-harvesting complexes. In review at Sci Adv (AAAS)
Tanizawa Y, Fujisawa T, Nakamura Y (2018) DFAST: a flexible prokaryotic genome annotation pipeline for faster genome publication. Bioinformatics 34:1037–1039. https://doi.org/10.1093/bioinformatics/btx713
Vignais PM, Billoud B (2007) Occurrence, classification, and biological function of hydrogenases: an overview. Chem Rev 107:4206–4272. https://doi.org/10.1021/cr050196r
Wahlund TM, Madigan MT (1993) Nitrogen fixation by the thermophilic green sulfur bacterium Chlorobium tepidum. J Bacteriol 175:474–478. https://doi.org/10.1128/jb.175.2.474-478.1993
Wahlund TM, Woese CR, Castenholz RW, Madigan MT (1991) A thermophilic green sulfur bacterium from New Zealand hot springs Chlorobium tepidum sp. nov. Arch Microbiol 156:81–90. https://doi.org/10.1007/BF00290978
Walker BJ, Abeel T, Shea T, Priest M, Abouelliel A, Sakthikumar S, Cuomo CA, Zeng Q, Wortman J, Young SK, Earl AM (2014) Pilon: an integrated tool for comprehensive microbial variant detection and genome assembly improvement. PLoS ONE 9:e112963. https://doi.org/10.1371/journal.pone.0112963
Weissgerber T, Zigann R, Bruce D et al (2011) Complete genome sequence of Allochromatium vinosum DSM 180T. Stand Genomic Sci 5:311–330. https://doi.org/10.4056/sigs.2335270
Yoon S-H, Ha S-M, Lim J, Kwon S, Chun J (2017) A large-scale evaluation of algorithms to calculate average nucleotide identity. Antonie Van Leeuwenhoek 110:1281–1286. https://doi.org/10.1007/s10482-017-0844-4
Yu L-J, Suga M, Wang-Otomo Z-Y, Shen J-R (2018) Structure of the photosynthetic LH1-RC supercomplex at 1.9 Å resolution. Nature 556:209–213. https://doi.org/10.1038/s41586-018-0002-9
Zhao D, Curatti L, Rubio LM (2007) Evidence for nifU and nifS participation in the biosynthesis of the iron-molybdenum cofactor of nitrogenase. J Biol Chem 282:37016–37025. https://doi.org/10.1074/jbc.M708097200
Acknowledgements
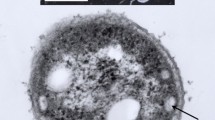
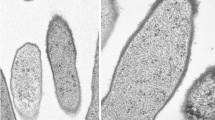
We thank Steven Schmitt, SIUC Electron Microscopy Center, for the electron micrographs, and Dr. Scott Hamilton-Brehm for supplying supplemental H2 for growth experiments.
Funding
This work was supported in part by NASA Exobiology grant 80NSSC17M0071 to MTM and by JSPS KAKENHI grant number 19H02018 to YT.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflicts of interest.
Additional information
Communicated by Erko Stackebrandt.
This article is dedicated to the memory of Thomas D. Brock who first introduced microbiology to the fascinating microbial diversity of geothermal environments.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Allochromatium tepidum strain NZT 16S rRNA gene sequence, AF384209; complete genome sequence, AP024563, AP024564; culture accession numbers, DSM 26924 and JCM 33379.
Rights and permissions
About this article
Cite this article
Madigan, M.T., Absher, J.N., Mayers, J.E. et al. Allochromatium tepidum, sp. nov., a hot spring species of purple sulfur bacteria. Arch Microbiol 204, 115 (2022). https://doi.org/10.1007/s00203-021-02715-7
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00203-021-02715-7