Abstract
The Covid-19 pandemic is highly contagious and has spread rapidly across the globe. To date there have been no specific treatment options available for this life-threatening disease. During this medical emergency, target-based drug repositioning/repurposing with a continuous monitoring and recording of results is an effective method for the treatment and drug discovery. This review summarizes the recent findings on COVID-19, its genomic organization, molecular evolution through phylogenetic analysis and has recapitulated the drug targets by analyzing the viral molecular machinery as drug targets and repurposing of most frequently used drugs worldwide and their therapeutic applications in COVID-19. Data from solidarity trials have shown that the treatment with Chloroquine, hydroxychloroquine and lopinavir-ritonavir had no effect in reducing the mortality rate and also had adverse side effects. Remdesivir, Favipiravir and Ribavirin might be a safer therapeutic option for COVID-19. Recent clinical trial has revealed that dexamethasone and convalescent plasma treatment can reduce mortality in patients with severe forms of COVID-19.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Background information
Severe Acute Respiratory Syndrome coronavirus 2 (SARS-CoV-2), previously known as 2019 novel coronavirus (2019-nCoV), is a novel species of Coronaviridae family, which causes COVID-19, the ongoing pandemic outbreak across the globe. SARS-CoV-2 was first reported in Wuhan, Hubei Province, China in December 2019 as a novel pathogen causing pneumonia (Zhu et al. 2020). In comparison to other corona viruses (CoV) that caused deadly diseases in the past, SARS-CoV-2 has a stronger transmission capacity thus being highly contagious and causes rapid increase in the infection rate globally (Tang et al. 2020). As of 15th July about 13,070,095 positive cases and around 572,539 deaths were confirmed across the world (WHO 2020). In the past two decades, corona viruses have caused the SARS (Severe Acute Respiratory Syndrome) Outbreak in 2002 and the MERS (Middle East Respiratory Syndrome) in 2015 (Guo et al. 2020). The former has infected about 8000 positive cases with 9.6% fatality rate, while MERS had affected about 2500 cases with 34% fatality rate, respectively (Cascella et al. 2020; Raoult et al. 2020).
Corona viruses
Corona viruses were first identified in the 1960s and subsequently derived their name due to the presence of crown-like spikes on their surface. Corona viruses are of Coronaviridae family classified within the Nidovirales order with four genera (α, β, γ, and δ CoV) and may cause mild to severe respiratory or gastrointestinal infections in humans, animals and birds. Human corona viruses (HCoV) belong to α and β- CoV. To date seven known pathogenic HCoV have been documented. Among them, four [229E and NL63 (α-HCoV) and HKU1 and OC43(β-HCoV)] are causative agents of mild coryzal illness similar to the common cold while the remaining three are more virulent zoonotic strains of β-HCoV (SARS-CoV, MERS-CoV, and SARS-CoV-2) that cause severe acute respiratory syndrome (Ye et al. 2020; Schaffer et al. 2010).
Structure
In general, HCoV are spherical or pleomorphic enveloped non-segmented positive-sense single-stranded RNA viruses (Fig. 1) that measure about 80–200 nm in diameter with a genome size of 26–30kbp. The genome consist of untranslated region at both ends (5′-methylated cap and 3′-polyadenylated tail), with two large open reading frames, ORF1a and ORF1b, wherein the non-structural proteins (involved in genome replication such as replicase, proteolytic processing) are encoded within the 5′ region and the structural proteins (envelope protein S (spike), glycoprotein E (envelope) protein, M (membrane) protein and N (nucleocapsid) phosphoprotein) are encoded within the 3′ region. Certain types of corona viruses have additional special structural and accessory proteins such as HE (hemagglutinin-esterase) protein and I (internal) protein. The S protein forms the peplomers on the virion surface which is responsible for the crown-like morphology. It is involved in receptor binding and cell fusion and is a major inducer for neutralizing antibodies. The M protein is the most abundant structural glycoprotein, with three transmembrane domains that define the viral shape. The M protein can interact with all other structural proteins and thus play a key role in viral assembly. E proteins are small structural proteins involved in viral assembly and morphogenesis within the cell. N protein binds with the viral genomic RNA to form a helical capsid structure, thus playing a crucial role in structure, replication and transcription of corona viruses. The HE protein is a non-essential protein which makes smaller spikes on the membrane and acts as hemagglutinin. It binds to the sialic acid present on the spike proteins and enhances the viral entry into the host cell. HE proteins are absent in SARS-CoV (Schaffer et al. 2010; Cecil et al. 2012; Fehr and Perlman 2015).
Structure of corona virus, reproduced from Li 2020
Insight into SARS-CoV-2
The genome sequences of novel SARS-CoV-2 are 50% similar to MERS-CoV, 79.5% similar to SARS-CoV, and 96.2% similar to a bat corona virus (Bat-CoV) RaTG13. SARS-CoV-2 size measured 60–140 nm diameter with 8–12 nm long spikes (Guo et al. 2020; Yang and Wang 2020). The viral genome has 29,891 nucleotides with a GC content of 38%, encoding 9860 amino acids (Chan et al. 2020; Lu et al. 2020). The genomic encoded proteins are more similar to SARS-CoV which are arranged in the order of 5′—replicase (ORF1a/b)—structural proteins (S- E- M- N)—3′. As a typical corona virus, SARS-CoV-2 also has six ORFs. 5′-replicase genes contain two large open reading frames, ORF1a/b. The frame shift between ORF1a and ORF1b produces two polypeptides: pp1a and pp1ab. These replicase polypeptides are proteolytically processed by virally encoded chymotrypsin-like protease (3CLpro) or main protease (Mpro) and additionally cleaved by papain-like protease (PLpro) to generate 16 nonstructural proteins (NSPs 1–16) (Cascella et al. 2020). The other ORFs encode the structural proteins and accessory proteins. Spike (S) protein is divided into S1 and S2 subunits. The S1 subunit shares 70% homology with SARS-CoV and Bat-CoV, encompasses an N-terminal domain, a receptor binding domain and motif and are involved in receptor binding to the host cell. The S2 Subunit that share 99% homology with SARS-CoV as well as Bat-CoV, comprised of a fusion peptide, a transmembrane domain and a cytoplasmic domain, and are responsible for cell membrane fusion. Like SARS-CoV, the HE protein is absent in SARS-CoV-2 (Chan et al. 2020). Among the 16 non-structural proteins, NSP3 (papain-like protease), NSP5 (3-chymotrypsin-like protease), NSP12 (RNA-dependent RNA polymerase), and NSP13 (helicase) are key enzymes essential for viral transcription and replication process (Fig. 2). Therefore, these four nonstructural proteins and S protein would be attractive sites for antiviral agents (Chan et al. 2020; Lu et al. 2020). Compared to SARS-CoV, the transmembrane helical segments in the ORF1ab NSP2 and NSP3 in SARS-CoV-2 have no evident homologous structures. At 501, 723 and 1010 amino acid positions, glutamine, serine, and proline are present instead of alanine, glycine and isoleucine residues, respectively. The stabilizing mutations located within the endosome‐associated‐protein‐like domain of the NSP2 protein make the virus highly contagious. The destabilizing mutations in the phosphatase domain of the NSP3 proteins suggest differentiation mechanism of SARS-CoV-2 from SARS-CoV (Angeletti et al. 2020).
Genome organization of SARS-CoV-2, SARS-CoV, and MERS. Reproduced from Mousavizadeh and Ghasemi 2020
Phylogenetic analysis
Nearly 15,000 SRA files and 19,000 biosample datasets are available in the GenBank database in which 92 of them are assembled in comparison to SARS-CoV-2 genome. With the available assembled genome, we have tried to perform a phylogenetic analysis to find out the evolutionary distances between each sequence. Pair-wise comparisons of the nucleotide sequences were performed using the Genome-BLAST Distance Phylogeny (GBDP) method (Meier-Kolthoff et al. 2013) under the settings recommended for prokaryotic viruses in VICTOR tool (Meier-Kolthoff and Göker 2017). The resulting inter-genomic distances were used to infer a balanced minimum evolution tree with branch support via FASTME including SPR post processing (Lefort et al. 2015). Trees were rooted at the midpoint (Farris 1972) and visualized with FigTree (Rambaut 2006). Taxon boundaries were estimated with the OPTSIL program (Göker et al. 2009), with recommended clustering thresholds and an F value of 0.5 (Meier-Kolthoff et al. 2014). The phylogenomic tree clearly shows the difference between each genome (Fig. 3). The SARS-CoV-2 genome has just 15 genes, yet it mutates all the time when it spreads. Most of these modifications make very little difference but at times the virus becomes either more/ less contagious (Fell and Davis 2020). Studies on mutations of functional proteins between SARS-CoV-2 and SARS-CoV genomes revealed 996 mutational changes occurred in ORF 1a and 1b amino acid sequences, whereas the comparisons within SARS-CoV-2 genomes showed only 11 changes (Khan et al. 2020).
Pathogenesis
One could observe some basic structural similarities in all the three zoonotic HCoV; however, the genome sequence and phenotypic arrangements (length of spikes) differ in each case based on genetic and structural analyses. These differences influence their pathogenic and infectious nature. In all the three cases, the initial transmission occurred from animal to human through direct contact and rapid spread occurred among humans through respiratory droplets and direct contact routes. The SARS-CoV-2 viral genomic evolutionary analysis showed that bats are the natural host harboring the virus and the virus might have been transmitted to pangolins, the suspected intermediate host (Zheng 2020), before infecting humans. The SARS-CoV-2 enters the lungs through respiratory tract. The incubation period is 14 days with an average of 3–7 days. In some rare cases the incubation period is reported as long as 24 days (Table 1) (Chan et al. 2020).
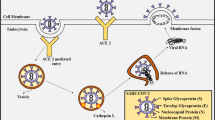
Proteolytic cleavage of the S glycoprotein at S1/S2 and S2′ is the critical step for viral attachment. Protease enzymes such as furin and TMPRSS2 are involved in proteolytic cleavage.
SARS-CoV-2 S glycoprotein has a furin cleavage site which is not present in other HCoV. This enables them to be more infectious and to spread efficiently among humans (Wu et al. 2020a). The furin activation site is targeted by furin protease found in the host cell and cleaves the S protein at S1/S2 site which is essential for cell–cell fusion. The cleaved S1 subunit thus recognizes the ACE2 receptor on the host cell membrane and binds to it. The S protein is activated by the cellular serine protease TMPRSS2 which cleaves at the S2′ site in order to bind to ACE2 receptor. The S2 subunit fuses the host and viral membranes. Viruses enter into the host cell via clathrin-mediated endocytosis and membrane fusion. After entry the virus uncoats and releases nucleocapsid and viral RNA into the cytoplasm, which is then transcribed and translated into structural proteins and two polyproteins. Viral genomic RNA (ssRNA) and subgenomic RNA (mRNA) are synthesized, replicated, and transcribed in the cytoplasm (Mousavizadeh and Ghasemi 2020). Following this, the viral structural proteins are inserted into endoplasmic reticulum and move to ERGIC (endoplasmic reticulum-Golgi intermediate compartment), where assembly of the viral genome, nucleocapsid proteins, and structural proteins occurs. This encapsidated viral genome matures and is multiplied, and released from the cell.
Variation in combating the virus by host immune system—key findings
Lymphocytes (T cells, B cells, and natural killer cells) play an important role in human immune system. During infection, the lymphocyte count increases to fight off the virus. But, in SARS-CoV-2 infection, the lymphocyte count gradually decreases with increasing severity. The mechanism of significant changes of lymphocyte and their correlation with the severity of the disease remains unclear. At early stage of infection peripheral blood lymphocyte count is normal or slightly low; however, lymphopenia is observed in patients. The lymphocyte count of less than 1.5 × 109/L is associated with the severity of COVID-19 (Zhao et al. 2020). B lymphocyte which is responsible for humoral immunity may also decrease in the early stage. This may affect antibody production and thus fail to prevent the viral multiplication (Lin et al. 2020). Another type of lymphocytes which play an important role in viral clearance is T cells (CD8+ and CD4+). Studies have shown that in COVID-19 patients, the total T cell, CD8+ , and CD4+ T cell counts are reduced dramatically, and the surviving T cells are functionally exhausted. This is due to the direct involvement of IL-10 (interleukin-10) which inhibits T cell proliferation and induces T cell exhaustion (Diao et al. 2020).
In some patients, an excessive inflammatory reaction takes place which leads to uncontrolled production of cytokines. These inflammatory responses are known as cytokine storm and cause damage to tissues and organs (Cascella et al. 2020). Another important immunological change in COVID-19 is abnormal increase of IL-6. It is produced by activated monocytes and macrophages which have a pivotal role in cytokine storm and functions through cis or trans signaling pathways. In the cis signal transduction pathway, the IL-6 complex with a transmembrane form of IL-6R (mlL-6R), and membrane bound gp130 activates downstream Janus kinase (JAKs), signal transducer, and activator of transcription 3 (STAT3). This activation leads to pleiotropic effects on immune cells which contribute to cytokine storm. In trans signal pathway, IL-6 forms complex with soluble form of IL-6R (sIL-6R) and gp130, and activates downstream JAKs and STAT3. The resultant IL-6- sIL-6R- JAK-STATA3 signaling is activated in immune cells leading to cytokine storm (Zhang et al. 2020; Liu et al. 2020a).
Clinical manifestations
Patients with COVID-19 initially have mild to severe respiratory illness with symptoms of fever, dry cough, sore throat, and fatigue. During the second phase of infection symptoms like headache, dyspnoea, haemoptysis, diarrhoea, lymphopenia, and difficulty in reading and in distinguishing smells have also been reported. The respiratory distress continues to deteriorate in some patients leading to acute respiratory distress syndrome (ARDS) and often requires medical aided respiration through mechanical ventilation by the third week of infection. RNAaemia and acute cardiac injury are other complications (Huang et al. 2020). Based on the currently available information, the severity rate is high in immunocompromised patients, patients with other co-morbid conditions, and elderly adults over 65, with an overall mortality rate of 3.4% (WHO 2020a).
The COVID-19 outbreak can be controlled through early detection of cases by screening and isolation of persons with symptoms. World Health Organization recommends contact and airborne precautions with N95 or higher respirator masks. Use of alcohol-based sanitizers is advisable to protect from COVID-19 (WHO 2020b). Routine cleaning and disinfection by lipid solvents like 75% ether, ethanol, peroxyacetic acid, chloroform, and chlorine-based disinfectants would help to decontaminate the surfaces and thus reduce the spread of COVID-19. SARS-CoV-2 is sensitive to heat and UV rays.
Potential therapeutics in COVID-19
To date, there is no specific anti-viral drug against COVID-19 which has been a major challenge for researchers trying to save lives and produce biological therapeutics to target SARS-CoV-2. Although little is known about the SARS-CoV-2, based on homology modeling and sequencing, it was found to have an 82% sequence similarity with the SARS-CoV and more than 90% sequence similarity in its essential enzymes (Morse et al. 2020). Therefore, knowing the viral-specific molecular machinery, researchers can effectively deal with SARS-CoV-2. From previous experiences in treating SARS-CoV and MERS-CoV, which may be directly helpful in treating the present outbreak of COVID-19, it is unlikely that any effort made at this point in a panic situation will benefit the patients. So, in this current review, we emphasize the various antiviral drug targets and the pharmacological therapeutic approaches in treating COVID-19.
Viral molecular machinery as drug targets
Isolated samples from the workers of the Wuhan seafood market revealed the 29.9-kb complete genome of the SARS-CoV-2 (Wu et al. 2020b). The CoV (coronavirus) genome has several open reading frames (ORFs) ranging from 6 to 11 (Song et al. 2019). About two-thirds of the RNA genome is occupied by ORF1a and 1b which translates into polyproteins (PP1a and PP1ab). The polyproteins further processed into 16 non-structural proteins (NSPs). The NSPs like RNA dependent RNA polymerase (RdRp), 3 C like protease (3CLpro), and papain-like protease (PLpro) are responsible for viral replication in the host cells by forming replicase transcriptase complex (Snijder et al. 2016). While, the rest of the RNA genome responsible for the formation of mRNA transcripts which produces structural proteins like spike glycoprotein (S), envelope protein (E), membrane protein (M), nucleocapsid protein (N) and other accessory proteins (Cui et al. 2019). So, the major targets for the anti-viral drugs are structural proteins like S along with the NSPs like RdRp, 3CLpro, and PLpro.
Structural spike glycoprotein (S)
The spike protein has three segments described as ectodomain, transmembrane anchor, and intracellular domain. Clove-shaped spike proteins gather in a trimeric form on the surface of the virus to form the distinctive crown-like appearance, so, it is named as coronavirus. The ectodomain consists of a receptor-binding domain (RBD) S1 and a membrane fusion domain (MFD) S2. These two domains are the main responsible elements in the viral entry into the host cells (Belouzard et al. 2012). SARS-CoV recognizes ACE-2 receptors as receptors for its association with the host cells (Li et al. 2003). Similar to the SARS-CoV, SARS-CoV-2 also recognizes the ACE-2 receptors for its binding to the host cells. Receptor binding domain S1 mediates association with the ACE-2 receptors of the host cells. The spike proteins of both origins have 76% similarity in sequence (Morse et al. 2020). Recent scientific reports revealed that the binding affinity of SARS-CoV-2 to the human ACE-2 receptors is ten times significantly higher than the SARS-CoV. Wrapp et al. also proved that the SARS-CoV RBD specific monoclonal antibodies have not shown the same binding ability to SARS-CoV-2 RBD (Wrapp et al. 2020). Hence, the cross-reactivity of the monoclonal antibodies related to the RBD of coronavirus was proved to be limited.
The importance of spike proteins in ACE-2 receptor binding and fusion of the membrane with the host cells makes it a potent drug target for the development of anti-viral drugs and vaccines. There are many scientific reports stating that the recombinant RBD binds strongly to the human and bat ACE-2 receptors. Further, the recombinant RBD has also prevented the entry of SARS-CoV-2 and SARS-CoV into the human ACE-2 receptor bearing cells, suggesting that the recombinant protein serves as a viral attachment inhibitor against the SARS-CoV. In contrast to the RBD specific monoclonal antibodies, Tai et al. emphasize that the SARS-CoV RBD specific polyclonal antibodies may cross neutralize the SARS-CoV-2 pseudovirus infection, suggesting the development of potent SARS-CoV RBD based vaccine for the prevention of CoV infections (Tai et al. 2020). Research groups have been trying to develop methods for building macrocyclic peptide libraries. Testing of these libraries for potent ligands for the SARS-CoV-2 RBD or to the ACE-2 peptide regions will lead to the development of anti-SARS-CoV-2 macrocyclic peptides.
The SARS-CoV spike protein, S2 domain plays a major role in the fusion of the virus with the host cells, in which the interaction of heptad repeat 1(HR1) and heptad repeat 2 (HR2) results in the formation of six helical bundles (6-HB), thereby promoting the fusion of the virus with the cellular membrane. Several researchers designed and developed fusion inhibitors against SARS-CoV (Liu et al. 2004) and MERS-CoV (Lu et al. 2014). Literature is scarce regarding whether the SARS-CoV-2 also has similar fusion and entry mechanisms to SARS-CoV and MERS-CoV and if it does, whether the S2 domain can also serve as a drug target for the development of fusion inhibitors for SARS-CoV-2. The S2 domain is the most conserved region in the CoV (Buchholz et al. 2004). According to the sequence alignment, both SARS-CoV and SARS-CoV-2 are highly conserved in HR1 and HR2 with 92.6% and 100% overall homology, respectively. Hence, scientific data suggested that SARS-CoV-2-HR2 derived peptide (SARS-CoV-2-HR2P) may act as a fusion inhibitor, indicating that the SARS-CoV-2-HR1 subunit could serve as a potent drug target for the inhibition of virus fusion with the host cells (Xia et al. 2020). Since the S2 domain is highly conserved in both SARS-CoV and SARS-CoV-2, neutralizing antibodies recognizing the RBD in the S protein S2 of the CoV could be a prominent treatment approach having cross-sensitivity (Yu et al. 2020).
In addition, after binding to the host cell ACE-2 receptor, before internalization, the S proteins of the virus are subjected to enzymatic modifications by several enzymes, especially transmembrane serine protease-2 (TMPRSS2). So protein priming is required for the cellular entry of the virus particle, which is a crucial step in the initiation of viral endocytosis (Hoffmann et al. 2020). After endocytosis, the S proteins are subjected to several enzymatic modifications by cathepsin L and B, lysosomal cysteine proteases in the endo-lysosomes in the acidic environment, in order to fuse the membranes and to release the viral RNA into the host cytosol. The lysosomal acidic environment proved to be an essential condition for the activation of several enzymes. Therefore, increased lysosomal cysteine proteases activity of cathepsin L and B, is highly dependent on lysosomal acidic pH (Ballout et al. 2020). Thus, it is reasonable to speculate that disrupting the binding and fusion of the virus with the host cells are two potential targets for the intervention against SARS-CoV-2.
Non-structural proteins
3Chymotrypsin-like cysteine protease (3CLpro) and papain-like protease (PLpro)
Non-structural proteins (NSPs) like 3CLpro and PLpro are responsible for the auto-proteolytic processing of replicase polyproteins encoded within NSP5 and NSP3, respectively. They are responsible for the processing of both structural and NSPs responsible for the replication and generation of new viruses. 3CLpro of SARS-CoV-2 showed 96% homology with the SARS-CoV (Morse et al. 2020). The 3CLpro processes the C-terminal of the PPs with 11 cleavage sites, while the PLpro Processes the N-terminal of the PPs with 3 cleavage sites (Fehr and Perlman 2015). The 3CLpro is in the form of dimers capable of generating mature proteins and anchors the transcription / functional replication complex during the replication of viral RNA (Barretto et al. 2005). Hence, small molecules that are targeting 3CLpro of SARS-CoV are expected to be the potential therapeutic agents for the treatment of SARS-CoV-2.
The PLpro of SARS-CoV has only 83% similarity with the SARS-CoV-2 in its sequence (Morse et al. 2020). However, the secondary structures showed the similarity in the active sites of the PLpro may lay the foundation for the development of SARS-CoV PLpro inhibitors that would also be expected to work for the SARS-CoV-2. PLpro also served as a deubiquitinase that is responsible for deubiquitylate host cell proteins, including NF-κB and interferon regulatory factor 3, which leads to immunosuppression in the virus-infected patients (Báez-Santos et al. 2015). So, therapeutic agents that can inhibit the PLpro activity might act as a potent drug molecule against SARS-CoV-2 and its associated immunosuppression in the viral-infected patients.
RNA dependent RNA polymerase (RdRp)
After entry into the host cells, the viral genome is released into the cytoplasm. The genome encodes for both structural and non-structural proteins. The RdRp is a product of a highly conserved region in NSP12 of the coronavirus polypeptide, which catalyzes the synthesis of viral RNA and plays a central role in the transcription process with the assistance of other NSP (Fehr and Perlman 2015). Though SARS-CoV and SARS-CoV-2 share 82% homology in their genetic sequence of RdRp, they share a remarkably high degree (96%) protein sequence in both origins (Morse et al. 2020; Gordon et al. 2020a). Apart from NSP12, other NSPs such as NSP13-helicase, NSP14-exoribonuclease, and N-methyl transferase, etc. too rely on the RdRp processivity, fidelity, and template switching, which cannot be ignored in the viral replication process (Fehr and Perlman 2015). Thus, NSP12 processed RdRp may be considered as a primary target for the anti-viral drugs in the treatment of COVID-19.
Treatment approaches based on molecular machinery
At present, there are no FDA approved drugs for the treatment of COVID-19. Therefore, identifying a potent antiviral agent to treat the disease is urgent. Currently, treatment is mainly focused on the symptomatic approach and respiratory support. According to WHO, refractory hypoxemia patients are recommended to provide extracorporeal membrane oxygenation despite lung-protective ventilation (WHO 2020c). At this point in a medical emergency, drug repositioning, or drug repurposing from existing drugs is an effective drug discovery process to shorten the time and reduce the cost of drug discovery (Zhou et al. 2020). Although it is essential to develop vaccination or potential therapeutic agents to specifically target the SARS-CoV-2, it is unlikely that any effort made at this moment will benefit the currently infected patients throughout the world.
Based on previous experiences with the SARS-CoV and MERS-CoV, we may learn some treatment approaches by targeting the common molecular machinery of the coronavirus. In the present review, we have focused mainly on the most frequently used anti-viral drugs all over the world and its potential application in COVID-19.
Viral entry inhibitors
Chloroquine and hydroxychloroquine
Chloroquine (CQ) and its derivative hydroxychloroquine (HCQ) are 4-aminoquinolines that have been known as safe and inexpensive drugs for the use of anti-malarial, anti-amoebic, and immunomodulatory activities (Schrezenmeier and Dörner 2020; Liu et al. 2020b). CQ and its analogue HCQ have been reported for their wide-spectrum of anti-viral activity against HIV, CoV, and Influenza viruses (Savarino et al. 2006; Chiang et al. 1996; Yazdany and Kim 2020). In recent in-vitro studies CQ and HCQ were found to inhibit SARS-CoV-2 (Liu et al. 2020b; Wang et al. 2020). This previous data and the current reports suggest that the CQ and HCQ may act as a potential pharmacological agent in preventing or treating the COVID-19.
The molecular mechanism in dealing with the CoV has not been fully elucidated and findings from the previous literature have suggested that the CQ and HCQ may act through a series of steps. Both CQ and HCQ are weak bases and they can penetrate the acidic cell organelles like endosomes and lysosomes, in non-protonated form. Once penetrated, the drug molecules convert into a protonated state to accumulate and alter the pH of the acidic cell organelles, which results in endosomal trafficking and prevents viral fusion into the cells. Further, the altered pH interfering with the cleavage of S proteins and glycosylation of cellular receptors of SARS-CoV results in impairment of viral entry into the host cells and prevents subsequent spread of infection (Vincent et al. 2005). The dual activity of anti-inflammatory and anti-viral activity of CQ and HCQ has been proposed to account for their utility in treating or preventing the SARS-CoV-2 and/or its related symptoms.
In vitro studies on SARS-CoV-2 exhibited a better activity of HCQ than the CQ in lowering the EC50 values (Wang et al. 2020). In addition, HCQ has fewer concerns about drug–drug interactions and better safety profile and allows higher daily dose in comparison to chloroquine, leading us to think HCQ may act as a promising drug for the treatment of COVID-19 (Marmor et al. 2016; Yao et al. 2020). Data supporting the treatment of COVID-19 with CQ and HCQ were quite scarce. A new brief report from China reported that the treatment with CQ enhanced viral clearance, improved radiological findings, and reduced disease progression (Gao et al. 2020). However, the data of study design and outcomes were not sent for peer review and published which limits the validity of the study. Another open-label nonrandomized trial conducted on 36 participants with COVID-19 reported improved virologic clearance in 6 participants given only HCQ compared to the 16-person control group of participants receive standard supportive care and the respective percentage of virologic clearance was 70% (14/20) Vs 12.5% (2/16). Further, the study also reported that the combination of Azithromycin to HCQ in 6 participants showed superior viral clearance 100% (6/6) in comparison with the HCQ 57% (8/14) monotherapy (Gautret et al. 2020). Despite the promising results, the study has several limitations such as low sample size in the combination therapy, early cessation of 6 participants from the CQ alone treatment group for intolerance towards medication and no safety outcome reported.
The dosing of the CQ and HCQ for the treatment of COVID-19 recommended based on the pharmacokinetic modeling studies and previous literature is 500 mg orally once or twice daily and 400 mg twice daily for one day, followed by twice daily 200 mg, respectively (Sanders et al. 2020). Both the drugs have a large volume of distribution, nearly every organ like eyes, kidney, lungs, and heart, etc., where retention is prolonged (Barlow et al. 2020).
CQ and HCQ are well-tolerated drugs with common adverse effects; gastrointestinal disturbances such as abdominal cramps, nausea, vomiting, and metallic taste. However, there are increasing safety concerns with both the drugs in increasing the QT interval leads to torsade’s de-pointes, drug-induced sudden cardiac deaths, hypoglycemia, neuropsychiatric effects, and retinopathy (Sanders et al. 2020; Barlow et al. 2020; Chen et al. 2006). Utmost care should be taken while prescribing the medication to patients with pre-existing cardiac diseases and those taking concomitant administration of drugs which prolongs the QT interval. Generally, both drugs are proved to be safe in pregnant women. There are contrary reports where both no significant and adverse effects were observed with doses and duration proposed for the SARS-CoV-2 (Sanders et al. 2020; Barlow et al. 2020).
Viral protease inhibitors
Lopinavir/ritonavir
Lopinavir along with Ritonavir (LPV/r) is FDA approved oral combination drug for the treatment of HIV. Lopinavir acts as an HIV-1 protease inhibitor and Ritonavir boosts the activity of Lopinavir by inhibiting both HIV-1 Protease and CYP enzymes’ metabolism of Lopinavir and leads to an increase in the bioavailability (Kempf et al. 1997; Sham et al. 1998). Studies have shown that 3-CLpro is a highly conserved region in different types of coronaviruses in terms of its sequence and 3D structure (Xue et al. 2008). Literature reveals that the HIV-1 protease inhibitors, LPV/r, are the potential drug candidates in inhibiting the 3-CLpro (Nukoolkarn et al. 2008). The protease enzymes are essential for the replication of the generation of new viruses (Morse et al. 2020). In accordance with the literature, LPV/r showed in vitro activity against SARS-CoV and MERS-CoV (Chu Study Group 2004; Wilde et al. 2014), which has promoted research interest in assessing the therapeutic potential of the combination for SARS-CoV-2.
Early reports comparing LPV/r along with the Ribavirin and a historical control group treated with Ribavirin alone revealed a decrease in ARDS caused by SARS-CoV but the historic, observational nature of the control arm prevents definitive conclusions (Chu SARS Study Group 2004). Studies on the SARS-CoV-2 in Vero E6 cells against Lopinovir exhibited anti-viral activity and the estimated EC50 at 26.63 µM (Choy et al. 2020), whereas the recent randomized, open-label, controlled trial including patients of COVID-19 received LPV/r (400 mg/100 mg) twice daily for 14 days compared with the standard care revealed no significant difference in viral clearance and mortality rate (Cao et al. 2020). The current data prove the limited therapeutic benefit of LPV/r in the treatment of COVID-19. Treatment with LPV/r may result in several gastrointestinal problems like nausea, vomiting sensation, and diarrhea. In addition, the patients in ICU care with pre-existing liver diseases and congenital cardiac disease are led to hepatic decompensation and QT prolongation, respectively (Barlow et al. 2020). These adverse effects of LPV/r may exacerbate the symptoms of COVID-19 (Sanders et al. 2020). Some other anti-retroviral drugs inhibiting protease and integrase were identified as potential drug molecules by observing their enzymatic activity. Darunavir proved its activity against SARS-CoV-2 in in vitro cell models (Sanders et al. 2020; Dong et al. 2020). Further clinical studies are needed for the clinical benefit of antiretroviral drugs in COVID-19 treatment.
Viral RNA polymerase inhibitors
Remdesivir and favipiravir
The molecular machinery, RNA-dependent RNA polymerase (RdRp) is a key enzyme responsible for the replication of viral RNA in the host cells. The high sequence conservation of the enzyme between the several CoV promotes the researchers to target the RdRp by developing the new drug molecules or repositioning of drug molecules. The currently used nucleotide analogues might provide the most promising research avenue towards disrupting the viral replication cycle. However, the exonuclease activity of NSP14 has been limiting the application of nucleotide analogues and the inactivation of the proof reading mechanism of NSP14 has been shown to improve the efficacy of the drugs in treating CoV infections. The current review mainly focuses on three drugs: Remdesivir, Favipiravir, and Ribavirin (Morse et al. 2020).
Remdesivir is an adenosine nucleotide analogue, originally developed for the treatment of Ebola and Marburg virus infections. However, it has not been approved by the FDA for any medical condition. It is a monophosphate prodrug that undergoes metabolism in the tissues into an active form C-adenosine nucleotide triphosphate. The active form of the drug inhibits the viral RNA-dependent RNA polymerase which prevents viral replication in the host cells. The drug proved its activity in several in vitro and in vivo animal studies against filoviridae (Ebola), paramyxoviridae (Nipah), and coronaviridae (MERS-CoV, SARS-CoV-1, and 2) (Huang et al. 2020; Lo et al. 2019; Sheahan et al. 2017; Gordon et al. 2020b). Studies revealed that the intact NSP14 proofreading activity of the virus, responsible for the resistance towards the nucleotide analogue, can be overcome by the administration of Remdesivir (Agostini et al. 2018). In another study, it was proved that the administration of Remdesivir and the combination of interferon β had better effect than the LPV/r and interferon β in both in vitro and a mouse model of MERS-CoV (Sheahan et al. 2020). Intravenous administration of Remdesivir improved the clinical condition of COVID-19 patients (Holshue et al. 2020). In another recent clinical study,COVID-19 patients, who underwent compassionate use of Remdesivir, showed clinical improvement in 36 (68%) out of 53 patients (Grein et al. 2020). But the study was not a controlled study and hence further randomized, placebo-controlled trials were required to draw concrete conclusions on its efficacy. The suppressive effect of Remdesivir but not the eradication of SARS-CoV 2 in immunocompromised patients was also documented in one recent study, thus suggesting the need to develop new therapies to enhance the outcome of long-term effect in immunocompromised patients (Helleberg et al. 2020). The ongoing clinical studies were conducted with a loading dose of 200 mg followed by 100 mg intravenously (https://clinicaltrials.gov/ct2/show/NCT04252664), which also demonstrated its linear pharmacokinetics between 3 and 225 mg, without any evidence of liver and kidney toxicity (Kujawski et al. 2020). Patients with COVID-19 reported having gastrointestinal disturbances and elevated levels of aminotransferase after the administration of Remdesivir therapy for 1–5 days (Kujawski et al. 2020). However, the clinical findings did not confirm whether the adverse effect is due to drug administration or SARS-CoV-2 infection. The current data on the adverse effect profile of the Remdesivir are sparse and mandate further clinical trials.
Favipiravir
Favipiravir is a licensed drug in Japan for the treatment of Influenza, which can be considered as a potential agent for the treatment of RNA-dependent viral infections. It is a purine nucleotide anti-viral prodrug which selectively inhibits the RdRp by metabolizing into an active form favipiravir ribofuranosyl-5B-triphosphate and also induces lethal viral mutagenesis, making it a virucidal drug. It showed RdRp enzyme inhibition of the Influenza virus at low concentrations (IC50 of 0.022 µg/mL), whereas it does not affect the human DNA polymerases even at a higher concentration of 100 µg/mL. Favipiravir exhibited a broad spectrum of antiviral activity against RNA viruses (Furuta et al. 2017). Furthermore, a recent study demonstrated that Favipiravir inhibited the SARS-CoV-2 infection in Vero E6 cells with EC50 of 61.88 µmol/L (Wang et al. 2020). Recent clinical data from an open-labeled, controlled clinical trial of Favipiravir showed a better therapeutic response in disease progression and viral clearance from the SARS-CoV-2 infected patients (Cai et al. 2020). A recent prospective, randomized, controlled, open-label multicenter clinical trial revealed the administration of Favipiravir (1600 mg twice daily on the first day and continued with 600 mg twice daily for another 9 days) did not lead to significant improvement when compared with Arbidol (200 mg thrice per day for 10 days), whereas it improved the latency to relieve clinical symptoms like pyrexia and cough. In the same study, the Favipiravir treatment group was observed to have elevated levels of serum uric acid, ALT/AST, and psychiatric symptoms (Chen et al. 2020). One of the recent case report studies also suggested the usage of Favipiravir for severe and critically ill patients in order to abate the progression of pneumonia and cytokine storm (Takahashi et al. 2020). Favipiravir also demonstrated rapid antiviral response in SARS-CoV-2 infected patients in the pilot stage study of phase II/III clinical trial and enabled viral clearance within 4 days in 62.5% of patients (Ivashchenko et al. 2020). Recent clinical evidences suggest the relative short-term safety and tolerability regarding total adverse events except that some adverse events like less serious gastrointestinal adverse events and increased blood uric acid levels remain a safety concern. As the pandemic spreads throughout the world, this drug received approval for emergency use in several countries like Italy, Japan, Russia, Saudi Arabia, and India, etc., (Agrawal et al. 2020). The current data are not sufficient to recommend the Favipiravir for the treatment of COVID-19, and several clinical trials are ongoing in an effort to confirm its clinical significance. However, the safety concerns like teratogenicity and QT prolongation have not yet been adequately studied. So, more evidence on the long-term effect of the treatment is needed for the safe and effective usage of the drug in the present pandemic situation (Ivashchenko et al. 2020).
Anti-inflammatory agents
The surge of Cytokines and inflammation induced by immunological responses towards the SARS-CoV-2 culminates in an acute respiratory distress syndrome. Evidence from animal studies also suggests that the inhibition of inflammation was proved to be effective against SARS-CoV and MERS-CoV infected animals in improving the outcome (Zha et al. 2020). Corticosteroids are the potent anti-inflammatory agents typically used for several inflammatory diseases like rheumatoid arthritis, asthma, and also for several acute respiratory viral infections. However, the clinical outcome of corticosteroids was disappointing and even delayed the viral clearance during the outbreak of SARS-CoV and MERS-CoV (Gangopadhyay et al. 2020). A recent study showed that the administration of corticosteroid (dexamethasone) showed a decreased mortality rate in patients receiving mechanical ventilation (Horby et al. 2020).
Dexamethasone
Dexamethasone is a corticosteroid with anti-inflammatory and immunosuppressant properties and has been used in the treatment of certain cancer, inflammatory disorders, severe influenza, SARS-CoV, MERS-CoV, and community acquired pneumonia. This drug was given to COVID-19 patients in the United Kingdom’s national clinical trial—Randomized Evaluation of COVID-19 Therapy (RECOVERY (https://www.recoverytrial.net/)), and demonstrated positive preliminary results. This is the first drug to be reported to reduce COVID-19 mortality rate. According to the RECOVERY team, treatment with dexamethasone with a dosage of 6 mg per day for up to 10 days reduced the mortality rate by about one-fifth and one-third in COVID-19 patients requiring oxygen and ventilator support, respectively. It also found that there were no risk-benefits for the patients who did not require respiratory assistance (Horby et al. 2020). WHO included the use of dexamethasone and other steroids in the COVID-19 treatment guidelines. However, further research is required to complement the usage of steroids in the development of new therapeutics in combination with other drugs for COVID-19.
Recent promising therapeutic approaches for COVID-19
New findings show that triple combination therapy of drugs interferon beta-1b, lopinavir–ritonavir, and ribavirin are safe and effective and reduce the risk of viral resistance. This treatment effectively suppresses viral load within an average of 7 days of treatment by lessening the symptoms, thereby improving the patient’s clinical conditions as well as curtailing the duration of hospitalization (Price 2020; Hung et al. 2020). Another promising therapeutic strategy is the passive immunization of COVID-19 patients using convalescent plasma of recovered COVID-19 donors. The neutralizing antibodies (Nabs), immunoglobulin G (IgG) and immunoglobulin M (IgM), present in the plasma, will inhibit viral entry, limiting viral amplification and promoting prompt therapeutic actions (Rojas et al. 2020).
Convalescent plasma as an alternative treatment
The current treatment approaches for critically ill COVID-19 patients are still under investigation; only limited evidence is available for a battery of antiviral, antibiotic, antimalarial, and anti-inflammatory agents along with the aggressive supportive care. In the current pandemic situation, multiple clinical trials are ongoing for the repurposing of various pharmacotherapeutic agents in order to prove their safety and efficacy against SARS-CoV-2. In the clinical trials, the anti-viral drugs like remdesivir, favipiravir and antimalarial drugs like hydroxychloroquine were reported to have efficacy in reducing the mortality while improving the health condition of COVID-19 patients; however, one should also consider its potential side effects (Duan et al. 2020). Moreover, the corticosteroid treatment for critically ill patients remains controversial, due to its complications and delayed viral clearance. Since there is no specific antiviral or vaccine for the critically ill patients, there is an urgent need for an alternative treatment strategy like the usage of convalescent plasma (Russell et al. 2020; Shang et al. 2020).
Concluding remarks
As COVID-19 is spreading rapidly throughout the world, it is necessary to re-purpose the existing approved drugs on an emergency basis in order to discover effective drugs for COVID-19. Several medicines have been given to the patients on a case by case basis with due consideration being given to potential risk and benefits. Current data prove that treatment with Chloroquine, hydroxychloroquine, and Lopinavir along with Ritonavir had adverse side effects. Remdesivir, Favipiravir, and Ribavirin might be safer therapeutics for COVID-19. Recent reports claim that the drug dexamethasone can cut mortality in patients with severe forms of COVID-19 by a third. Target-based drug repurposing and continuous monitoring and recording of results will accelerate drug discovery. Until we find a novel drug or vaccine, it is necessary to consider safe measures such as social distancing, wearing high-quality masks, sanitization, thus cutting-off the route of transmission.
References
Agostini ML, Andres EL, Sims AC, Graham RL, Sheahan TP, Lu X, Smith EC, Case JB, Feng JY, Jordan R, Ray AS, Cihlar T, Siegel D, Mackman RL, Clarke MO, Baric RS, Denison MR (2018) Coronavirus susceptibility to the antiviral remdesivir (GS-5734) is mediated by the viral polymerase and the proofreading exoribonuclease. MBio 9:2. https://doi.org/10.1128/mBio.00221-18
Agrawal U, Raju R, Udwadia ZF (2020) Favipiravir: a new and emerging antiviral option in COVID-19. Med J Arm Forc India 76(4):370–376. https://doi.org/10.1016/j.mjafi.2020.08.004
Angeletti S, Benvenuto D, Bianchi M, Giovanetti M, Pascarella S, Ciccozzi M (2020) COVID-2019: the role of the nsp2 and nsp3 in its pathogenesis. J Med Virol 92(6):584–588. https://doi.org/10.1002/jmv.25719
Báez-Santos YM, St John SE, Mesecar AD (2015) The SARS-coronavirus papain-like protease: structure, function and inhibition by designed antiviral compounds. Antiviral Res 115:21–38. https://doi.org/10.1016/j.antiviral.2014.12.015
Ballout RA, Sviridov D, Bukrinsky MI, Remaley AT (2020) The lysosome: A potential juncture between SARS-CoV-2 infectivity and Niemann-Pick disease type C, with therapeutic implications. FASEB J Off Publ Feder Am Soc Exp Biol 34(6):7253–7264. https://doi.org/10.1096/fj.202000654R
Barlow A, Landolf KM, Barlow B, Yeung SYA, Heavner JJ, Claassen CW, Heavner MS (2020) Review of emerging pharmacotherapy for the treatment of coronavirus disease 2019. Pharmacotherap J Hum Pharmacol Drug Therap 40(5):416–437. https://doi.org/10.1002/phar.2398
Barretto N, Jukneliene D, Ratia K, Chen Z, Mesecar AD, Baker SC (2005) The papain-like protease of severe acute respiratory syndrome coronavirus has deubiquitinating activity. J Virol 79(24):15189–15198. https://doi.org/10.1128/JVI.79.24.15189-15198.2005
Belouzard S, Millet JK, Licitra BN, Whittaker GR (2012) Mechanisms of coronavirus cell entry mediated by the viral spike protein. Viruses 4(6):1011–1033. https://doi.org/10.3390/v4061011
Buchholz UJ, Bukreyev A, Yang L, Lamirande EW, Murphy BR, Subbarao K, Collins PL (2004) Contributions of the structural proteins of severe acute respiratory syndrome coronavirus to protective immunity. Proc Natl Acad Sci 101(26):9804–9809. https://doi.org/10.1073/pnas.0403492101
Cai Q, Yang M, Liu D, Chen J, Shu D, Xia J, Liao X, Gu Y, Cai Q, Yang Y, Shen C, Li X, Peng L, Huang D, Zhang J, Zhang S, Wang F, Liu J, Chen L, Liu L (2020) Experimental treatment with favipiravir for COVID-19: an open-label control study. Engineering. https://doi.org/10.1016/j.eng.2020.03.007
Cao B, Wang Y, Wen D, Liu W, Wang J, Fan G, Ruan L, Song B, Cai Y, Wei M, Li X, Xia J, Chen N, Xiang J, Yu T, Bai T, Xie X, Zhang L, Li C, Wang C (2020) A trial of lopinavir-ritonavir in adults hospitalized with severe covid-19. N Engl J Med 382(19):1787–1799. https://doi.org/10.1056/NEJMoa2001282
Cascella M, Rajnik M, Cuomo A, Dulebohn SC, Di Napoli R. Features, evaluation and treatment coronavirus (COVID-19). In: StatPearls. StatPearls Publishing; 2020. http://www.ncbi.nlm.nih.gov/books/NBK554776/. Accessed 16 August 2020
Cecil R (2012) Goldman’s Cecil medicine. 24th edn. Philadelphia: Elsevier/Saunders/
Chan JF-W, Kok K-H, Zhu Z, Chu H, To KK-W, Yuan S, Yuen K-Y (2020) Genomic characterization of the 2019 novel human-pathogenic coronavirus isolated from a patient with atypical pneumonia after visiting Wuhan. Emerg Microb Infect 9(1):221–236. https://doi.org/10.1080/22221751.2020.1719902
Chen C-Y, Wang F-L, Lin C-C (2006) Chronic hydroxychloroquine use associated with QT prolongation and refractory ventricular arrhythmia. Clin Toxicol (Phila Pa) 44(2):173–175. https://doi.org/10.1080/15563650500514558
Chen C, Zhang Y, Huang J, Yin P, Cheng Z, Wu J, Chen S, Zhang Y, Chen B, Lu M, Luo Y, Ju L, Zhang J, Wang X (2020) Favipiravir versus Arbidol for COVID-19. Random Clin Trial. https://doi.org/10.1101/2020.03.17.20037432 (Preprint)
Chiang G, Sassaroli M, Louie M, Chen H, Stecher VJ, Sperber K (1996) Inhibition of HIV-1 replication by hydroxychloroquine: mechanism of action and comparison with zidovudine. Clin Ther 18(6):1080–1092. https://doi.org/10.1016/s0149-2918(96)80063-4
Choy K-T, Wong AY-L, Kaewpreedee P, Sia SF, Chen D, Hui KPY, Chu DKW, Chan MCW, Cheung PP-H, Huang X, Peiris M, Yen H-L (2020) Remdesivir, lopinavir, emetine, and homoharringtonine inhibit SARS-CoV-2 replication in vitro. Antiviral Res 178:104786. https://doi.org/10.1016/j.antiviral.2020.104786
Chu CM, Cheng VCC, Hung IFN, Wong MML, Chan KH, Chan KS, Kao RYT, Poon LLM, Wong CLP, Guan Y, Peiris JSM, Yuen KY, HKU/UCH SARS Study Group (2004) Role of lopinavir/ritonavir in the treatment of SARS: initial virological and clinical findings. Thorax 59(3):252–256. https://doi.org/10.1136/thorax.2003.012658
Cui J, Li F, Shi Z-L (2019) Origin and evolution of pathogenic coronaviruses. Nat Rev Microbiol 17(3):181–192. https://doi.org/10.1038/s41579-018-0118-9
de Wilde AH, Jochmans D, Posthuma CC, Zevenhoven-Dobbe JC, van Nieuwkoop S, Bestebroer TM, van den Hoogen BG, Neyts J, Snijder EJ (2014) Screening of an FDA-approved compound library identifies four small-molecule inhibitors of middle east respiratory syndrome coronavirus replication in cell culture. Antimicrob Agents Chemother 58(8):4875–4884. https://doi.org/10.1128/AAC.03011-14
Diao B, Wang C, Tan Y, Chen X, Liu Y, Ning L, Chen L, Li M, Liu Y, Wang G, Yuan Z, Feng Z, Zhang Y, Wu Y, Chen Y (2020) Reduction and functional exhaustion of t cells in patients with coronavirus disease 2019 (COVID-19). Front Immunol. https://doi.org/10.3389/fimmu.2020.00827
Dong L, Hu S, Gao J (2020) Discovering drugs to treat coronavirus disease 2019 (COVID-19). Drug Discov Therap 14(1):58–60. https://doi.org/10.5582/ddt.2020.01012
Duan K, Liu B, Li C, Zhang H, Yu T, Qu J, Zhou M, Chen L, Meng S, Hu Y, Peng C, Yuan M, Huang J, Wang Z, Yu J, Gao X, Wang D, Yu X, Li L, Yang X (2020) Effectiveness of convalescent plasma therapy in severe COVID-19 patients. Proc Natl Acad Sci USA 117(17):9490–9496. https://doi.org/10.1073/pnas.2004168117
Farris JS (1972) Estimating phylogenetic trees from distance matrices. Am Nat 106(951):645–668
Fehr AR, Perlman S (2015) Coronaviruses: an overview of their replication and pathogenesis. Coronaviruses 1282:1–23. https://doi.org/10.1007/978-1-4939-2438-7_1
Fell A, Davis UC. (2020). Genome variation gives insight into coronavirus spread. Retrieved 22 June 2020, from https://phys.org/news/2020-04-genome-variation-insight-coronavirus.html
Furuta Y, Komeno T, Nakamura T (2017) Favipiravir (T-705), a broad spectrum inhibitor of viral RNA polymerase. Proceedings of the Japan academy. Series B Phys Biol Sci 93(7):449–463. https://doi.org/10.2183/pjab.93.027
Gangopadhyay KK, Mukherjee JJ, Sinha B, Ghosal S (2020) The role of corticosteroids in the management of critically ill patients with coronavirus disease 2019 (COVID-19): a meta-analysis. MedRxiv. https://doi.org/10.1101/2020.04.17.20069773
Gao J, Tian Z, Yang X (2020) Breakthrough: Chloroquine phosphate has shown apparent efficacy in treatment of COVID-19 associated pneumonia in clinical studies. Biosci Trends 14(1):72–73. https://doi.org/10.5582/bst.2020.01047
Gautret P, Lagier J-C, Parola P, Hoang VT, Meddeb L, Mailhe M, Doudier B, Courjon J, Giordanengo V, Vieira VE, Tissot Dupont H, Honoré S, Colson P, Chabrière E, La Scola B, Rolain J-M, Brouqui P, Raoult D (2020) Hydroxychloroquine and azithromycin as a treatment of COVID-19: results of an open-label non-randomized clinical trial. Int J Antimicrob Agents 56(1):105949. https://doi.org/10.1016/j.ijantimicag.2020.105949
Göker M, García-Blázquez G, Voglmayr H, Tellería MT, Martín MP (2009) Molecular taxonomy of phytopathogenic fungi: a case study in peronospora. PLoS ONE 4(7):e6319. https://doi.org/10.1371/journal.pone.0006319
Gordon DE, Jang GM, Bouhaddou M, Xu J, Obernier K, White KM, O’Meara MJ, Rezelj VV, Guo JZ, Swaney DL, Tummino TA, Hüttenhain R, Kaake RM, Richards AL, Tutuncuoglu B, Foussard H, Batra J, Haas K, Modak M, Krogan NJ (2020a) A SARS-CoV-2 protein interaction map reveals targets for drug repurposing. Nature 583(7816):459–468. https://doi.org/10.1038/s41586-020-2286-9
Gordon CJ, Tchesnokov EP, Feng JY, Porter DP, Götte M (2020b) The antiviral compound remdesivir potently inhibits RNA-dependent RNA polymerase from Middle East respiratory syndrome coronavirus. J Biol Chem 295(15):4773–4779. https://doi.org/10.1074/jbc.AC120.013056
Grein J, Ohmagari N, Shin D, Diaz G, Asperges E, Castagna A, Feldt T, Green G, Green ML, Lescure F-X, Nicastri E, Oda R, Yo K, Quiros-Roldan E, Studemeister A, Redinski J, Ahmed S, Bernett J, Chelliah D, Flanigan T (2020) Compassionate use of remdesivir for patients with severe COVID-19. N Engl J Med 382(24):2327–2336. https://doi.org/10.1056/NEJMoa2007016
Guo Y-R, Cao Q-D, Hong Z-S, Tan Y-Y, Chen S-D, Jin H-J, Tan K-S, Wang D-Y, Yan Y (2020) The origin, transmission and clinical therapies on coronavirus disease 2019 (COVID-19) outbreak—An update on the status. Milit Med Res 7(1):11. https://doi.org/10.1186/s40779-020-00240-0
Helleberg M, Niemann CU, Moestrup KS, Kirk O, Lebech AM, Lane C, Lundgren J (2020) Persistent COVID-19 in an immunocompromised patient temporarily responsive to two courses of remdesivir therapy. J Infect Dis 222(7):1103–1107. https://doi.org/10.1093/infdis/jiaa446
Hoffmann M, Kleine-Weber H, Schroeder S, Krüger N, Herrler T, Erichsen S, Schiergens TS, Herrler G, Wu N-H, Nitsche A, Müller MA, Drosten C, Pöhlmann S (2020) SARS-CoV-2 Cell entry depends on ACE2 and TMPRSS2 and is blocked by a clinically proven protease inhibitor. Cell 181(2):271-280.e8. https://doi.org/10.1016/j.cell.2020.02.052
Holshue ML, DeBolt C, Lindquist S, Lofy KH, Wiesman J, Bruce H, Spitters C, Ericson K, Wilkerson S, Tural A, Diaz G, Cohn A, Fox L, Patel A, Gerber SI, Kim L, Tong S, Lu X, Lindstrom S, Pillai SK (2020) First case of 2019 novel coronavirus in the united states. N Engl J Med 382(10):929–936. https://doi.org/10.1056/NEJMoa2001191
Horby P, Lim WS, Emberson J, Mafham M, Bell J, Linsell L, Staplin N, Brightling C, Ustianowski A, Elmahi E, Prudon B, Green C, Felton T, Chadwick D, Rege K, Fegan C, Chappell L, Faust S, Jaki T, Landray M (2020) Effect of dexamethasone in hospitalized patients with COVID-19. Prelim Rep. https://doi.org/10.1101/2020.06.22.20137273
Huang C, Wang Y, Li X, Ren L, Zhao J, Hu Y, Zhang L, Fan G, Xu J, Gu X, Cheng Z, Yu T, Xia J, Wei Y, Wu W, Xie X, Yin W, Li H, Liu M, Cao B (2020) Clinical features of patients infected with 2019 novel coronavirus in Wuhan China. Lancet 395(10223):497–506. https://doi.org/10.1016/S0140-6736(20)30183-5
Hung IF-N, Lung K-C, Tso EY-K, Liu R, Chung TW-H, Chu M-Y, Ng Y-Y, Lo J, Chan J, Tam AR, Shum H-P, Chan V, Wu AK-L, Sin K-M, Leung W-S, Law W-L, Lung DC, Sin S, Yeung P, Yuen K-Y (2020) Triple combination of interferon beta-1b, lopinavir-ritonavir, and ribavirin in the treatment of patients admitted to hospital with COVID-19: an open-label, randomised, phase 2 trial. Lancet (Lond Engl) 395(10238):1695–1704. https://doi.org/10.1016/S0140-6736(20)31042-4
Ivashchenko AA, Dmitriev KA, Vostokova NV, Azarova VN, Blinow AA, Egorova AN, Gordeev IG, Ilin AP, Karapetian RN, Kravchenko DV, Lomakin NV (2020) AVIFAVIR for treatment of patients with moderate COVID-19: interim results of a phase II/III multicenter randomized clinical trial. medRxiv. https://doi.org/10.1093/cid/ciaa1176
Kempf DJ, Marsh KC, Kumar G, Rodrigues AD, Denissen JF, McDonald E, Kukulka MJ, Hsu A, Granneman GR, Baroldi PA, Sun E, Pizzuti D, Plattner JJ, Norbeck DW, Leonard JM (1997) Pharmacokinetic enhancement of inhibitors of the human immunodeficiency virus protease by coadministration with ritonavir. Antimicrob Agents Chemother 41(3):654–660. https://doi.org/10.1128/AAC.41.3.654
Khan MI, Khan ZA, Baig MH, Ahmad I, Farouk A-EAziem, Song YG, et al (2020) Comparative genome analysis of novel coronavirus (SARS-CoV-2) from different geographical locations and the effect of mutations on major target proteins: an in silico insight. PLoS ONE 15(9):e0238344. https://doi.org/10.1371/journal.pone.0238344
Kujawski SA, Wong KK, Collins JP, Epstein L, Killerby ME, Midgley CM, Abedi GR, Ahmed NS, Almendares O, Alvarez FN, Anderson KN, Balter S, Barry V, Bartlett K, Beer K, Ben-Aderet MA, Benowitz I, Biggs H, Binder AM, Zhang J (2020) First 12 patients with coronavirus disease 2019 (COVID-19) in the United States. MedRxiv. https://doi.org/10.1101/2020.03.09.20032896 (2020.03.09.20032896)
Lefort V, Desper R, Gascue O (2015) FastME 2.0: a comprehensive, accurate, and fast distance-based phylogeny inference program. Mol Biol Evol 32(10):2798–2800. https://doi.org/10.1093/molbev/msv150
Li YJ (2020) ScienCell Acts to help aid research for coronavirus SARS-CoV-2. ScienCell Research Laboratories. https://sciencellonline.com/blog/sciencell-acts-to-help-aid-research-for-coronavirus-sars-cov-2/ (Accessed on 18 June 2020)
Li W, Moore MJ, Vasilieva N, Sui J, Wong SK, Berne MA, Somasundaran M, Sullivan JL, Luzuriaga K, Greenough TC, Choe H, Farzan M (2003) Angiotensin-converting enzyme 2 is a functional receptor for the SARS coronavirus. Nature 426(6965):450–454. https://doi.org/10.1038/nature02145
Lin L, Lu L, Cao W, Li T (2020) Hypothesis for potential pathogenesis of SARS-CoV-2 infection-a review of immune changes in patients with viral pneumonia. Emerg Microb Infect 9(1):727–732. https://doi.org/10.1080/22221751.2020.1746199
Liu S, Xiao G, Chen Y, He Y, Niu J, Escalante CR, Xiong H, Farmar J, Debnath AK, Tien P, Jiang S (2004) Interaction between heptad repeat 1 and 2 regions in spike protein of SARS-associated coronavirus: Implications for virus fusogenic mechanism and identification of fusion inhibitors. Lancet (Lond Engl) 363(9413):938–947. https://doi.org/10.1016/S0140-6736(04)15788-7
Liu J, Cao R, Xu M, Wang X, Zhang H, Hu H, Li Y, Hu Z, Zhong W, Wang M (2020b) Hydroxychloroquine, a less toxic derivative of chloroquine, is effective in inhibiting SARS-CoV-2 infection in vitro. Cell Discov 6(1):1–4. https://doi.org/10.1038/s41421-020-0156-0
Liu B, Li M, Zhou Z, Guan X, Xiang Y (2020a) Can we use interleukin-6 (IL-6) blockade for coronavirus disease 2019 (COVID-19)-induced cytokine release syndrome (CRS)? J Autoimmun 111:102452. https://doi.org/10.1016/j.jaut.2020.102452
Lo MK, Feldmann F, Gary JM, Jordan R, Bannister R, Cronin J, Patel NR, Klena JD, Nichol ST, Cihlar T, Zaki SR, Feldmann H, Spiropoulou CF, De Wit E (2019) Remdesivir (GS-5734) protects African green monkeys from Nipah virus challenge. Sci Translat Med 11:494. https://doi.org/10.1126/scitranslmed.aau9242
Lu L, Liu Q, Zhu Y, Chan K-H, Qin L, Li Y, Wang Q, Chan JF-W, Du L, Yu F, Ma C, Ye S, Yuen K-Y, Zhang R, Jiang S (2014) Structure-based discovery of middle east respiratory syndrome coronavirus fusion inhibitor. Nat Commun 5(1):3067. https://doi.org/10.1038/ncomms4067
Lu R, Zhao X, Li J, Niu P, Yang B, Wu H, Wang W, Song H, Huang B, Zhu N, Bi Y, Ma X, Zhan F, Wang L, Hu T, Zhou H, Hu Z, Zhou W, Zhao L, Tan W (2020) Genomic characterisation and epidemiology of 2019 novel coronavirus: implications for virus origins and receptor binding. Lancet 395(10224):565–574. https://doi.org/10.1016/S0140-6736(20)30251-8
Marmor MF, Kellner U, Lai TYY, Melles RB, Mieler WF, American Academy of Ophthalmology (2016) Recommendations on screening for chloroquine and hydroxychloroquine retinopathy (2016 Revision). Ophthalmology 123(6):1386–1394. https://doi.org/10.1016/j.ophtha.2016.01.058
Meier-Kolthoff JP, Göker M (2017) VICTOR: Genome-based phylogeny and classification of prokaryotic viruses. Bioinformat (Oxf Engl) 33(21):3396–3404. https://doi.org/10.1093/bioinformatics/btx440
Meier-Kolthoff JP, Auch AF, Klenk H-P, Göker M (2013) Genome sequence-based species delimitation with confidence intervals and improved distance functions. BMC Bioinformatics 14(1):60. https://doi.org/10.1186/1471-2105-14-60
Meier-Kolthoff JP, Hahnke RL, Petersen J, Scheuner C, Michael V, Fiebig A, Rohde C, Rohde M, Fartmann B, Goodwin LA, Chertkov O, Reddy T, Pati A, Ivanova NN, Markowitz V, Kyrpides NC, Woyke T, Göker M, Klenk H-P (2014) Complete genome sequence of DSM 30083T, the type strain (U5/41T) of Escherichia coli, and a proposal for delineating subspecies in microbial taxonomy. Stand Gen Sci. https://doi.org/10.1186/1944-3277-9-2
Morse JS, Lalonde T, Xu S, Liu WR (2020) Learning from the past: possible urgent prevention and treatment options for severe acute respiratory infections caused by 2019-nCoV. Chembiochem Eur J Chem Biol 21(5):730–738. https://doi.org/10.1002/cbic.202000047
Mousavizadeh L, Ghasemi S (2020) Genotype and phenotype of COVID-19: their roles in pathogenesis. J Microbiol Immunol Infect. https://doi.org/10.1016/j.jmii.2020.03.022
Nukoolkarn V, Lee VS, Malaisree M, Aruksakulwong O, Hannongbua S (2008) Molecular dynamic simulations analysis of ritonavir and lopinavir as SARS-CoV 3CL(pro) inhibitors. J Theor Biol 254(4):861–867. https://doi.org/10.1016/j.jtbi.2008.07.030
Price S (2020). Drug combination shows promise for treating COVID-19. Health Europa. (Retreived May 11, 2020). https://www.healtheuropa.eu/drug-combination-shows-promise-for-treating-covid-19/99874/
Rambaut A (2006). FigTree 1.4.3: a graphical viewer of phylogenetic trees and a program for producing publication-ready figures. Retrieved 25 June 2020, from http://tree.bio.ed.ac.uk/software/figtree/
Raoult D, Zumla A, Locatelli F, Ippolito G, Kroemer G (2020) Coronavirus infections: Epidemiological, clinical and immunological features and hypotheses. Cell Stress 4(4):66–74. https://doi.org/10.15698/cst2020.04.216
Rojas M, Rodríguez Y, Monsalve DM, Acosta-Ampudia Y, Camacho B, Gallo JE, Rojas-Villarraga A, Ramírez-Santana C, Díaz-Coronado JC, Manrique R, Mantilla RD, Shoenfeld Y, Anaya J-M (2020) Convalescent plasma in Covid-19: possible mechanisms of action. Autoimmun Rev 19(7):102554. https://doi.org/10.1016/j.autrev.2020.102554
Russell CD, Millar JE, Baillie JK (2020) Clinical evidence does not support corticosteroid treatment for 2019-nCoV lung injury. Lancet 395(10223):473–475. https://doi.org/10.1016/S0140-6736(20)30317-2
Sanders JM, Monogue ML, Jodlowski TZ, Cutrell JB (2020) Pharmacologic treatments for coronavirus disease 2019 (COVID-19): a review. JAMA 323(18):1824–1836. https://doi.org/10.1001/jama.2020.6019
Savarino A, Di Trani L, Donatelli I, Cauda R, Cassone A (2006) New insights into the antiviral effects of chloroquine. Lancet Infect Dis 6(2):67–69. https://doi.org/10.1016/S1473-3099(06)70361-9
Schaffer K, La Rosa AM, Whimbey E (2010) Respiratory viruses. Infect Dis. https://doi.org/10.1016/B978-0-323-04579-7.00162-3
Schrezenmeier E, Dörner T (2020) Mechanisms of action of hydroxychloroquine and chloroquine: Implications for rheumatology. Nat Rev Rheumatol 16(3):155–166. https://doi.org/10.1038/s41584-020-0372-x
Sham HL, Kempf DJ, Molla A, Marsh KC, Kumar GN, Chen CM, Kati W, Stewart K, Lal R, Hsu A, Betebenner D, Korneyeva M, Vasavanonda S, McDonald E, Saldivar A, Wideburg N, Chen X, Niu P, Park C, Norbeck DW (1998) ABT-378, a highly potent inhibitor of the human immunodeficiency virus protease. Antimicrob Agents Chemother 42(12):3218–3224. https://doi.org/10.1128/AAC.42.12.3218
Shang L, Zhao J, Hu Y, Du R, Cao B (2020) On the use of corticosteroids for 2019-nCoV pneumonia. Lancet (Lond Engl) 395(10225):683–684. https://doi.org/10.1016/S0140-6736(20)30361-5
Sheahan TP, Sims AC, Graham RL, Menachery VD, Gralinski LE, Case JB, Leist SR, Pyrc K, Feng JY, Trantcheva I, Bannister R, Park Y, Babusis D, Clarke MO, Mackman RL, Spahn JE, Palmiotti CA, Siegel D, Ray AS, Baric RS (2017) Broad-spectrum antiviral GS-5734 inhibits both epidemic and zoonotic coronaviruses. Sci Translat Med 9:396. https://doi.org/10.1126/scitranslmed.aal3653
Sheahan TP, Sims AC, Leist SR, Schäfer A, Won J, Brown AJ, Montgomery SA, Hogg A, Babusis D, Clarke MO, Spahn JE, Bauer L, Sellers S, Porter D, Feng JY, Cihlar T, Jordan R, Denison MR, Baric RS (2020) Comparative therapeutic efficacy of remdesivir and combination lopinavir, ritonavir, and interferon beta against MERS-CoV. Nat Commun 11(1):222. https://doi.org/10.1038/s41467-019-13940-6
Snijder EJ, Decroly E, Ziebuhr J (2016) The nonstructural proteins directing coronavirus RNA synthesis and processing. Adv Virus Res 96:59–126. https://doi.org/10.1016/bs.aivir.2016.08.008
Song Z, Xu Y, Bao L, Zhang L, Yu P, Qu Y, Zhu H, Zhao W, Han Y, Qin C (2019) From SARS to MERS thrusting coronaviruses into the spotlight. Viruses 11:1
Tai W, He L, Zhang X, Pu J, Voronin D, Jiang S, Zhou Y, Du L (2020) Characterization of the receptor-binding domain (RBD) of 2019 novel coronavirus: implication for development of RBD protein as a viral attachment inhibitor and vaccine. Cell Mol Immunol 17(6):613–620. https://doi.org/10.1038/s41423-020-0400-4
Takahashi H, Iwasaki Y, Watanabe T, Ichinose N, Okada Y, Oiwa A, Kobayashi T, Moriya M, Oda T (2020) Case studies of SARS-CoV-2 treated with favipiravir among patients in critical or severe condition. Int J Infect Dis 100:283–285. https://doi.org/10.1016/j.ijid.2020.08.047
Tang B, Bragazzi NL, Li Q, Tang S, Xiao Y, Wu J (2020) An updated estimation of the risk of transmission of the novel coronavirus (2019-nCov). Infect Dis Model 5:248–255. https://doi.org/10.1016/j.idm.2020.02.001
Vincent MJ, Bergeron E, Benjannet S, Erickson BR, Rollin PE, Ksiazek TG, Seidah NG, Nichol ST (2005) Chloroquine is a potent inhibitor of SARS coronavirus infection and spread. Virol J 2(1):69. https://doi.org/10.1186/1743-422X-2-69
Wang M, Cao R, Zhang L, Yang X, Liu J, Xu M, Shi Z, Hu Z, Zhong W, Xiao G (2020) Remdesivir and chloroquine effectively inhibit the recently emerged novel coronavirus (2019-nCoV) in vitro. Cell Res 30(3):269–271. https://doi.org/10.1038/s41422-020-0282-0
WHO (2020). WHO Director-General’s opening remarks at the media briefing on COVID-19—11 March 2020. Retrieved 7 August 2020, from https://www.who.int/dg/speeches/detail/who-director-general-s-opening-remarks-at-the-media-briefing-on-covid-19---11-march-2020
WHO (2020). Advice for the public on COVID-19: World health organization. Retrieved 7 August 2020, from https://www.who.int/emergencies/diseases/novel-coronavirus-2019/advice-for-public
WHO (2020). Clinical management of COVID-19. Retrieved 5 July 2020, from https://www.who.int/publications-detail-redirect/clinical-management-of-covid-19
WHO, Coronavirus Disease (COVID-19) Situation Reports. (2020). Retrieved 7 August 2020, from https://www.who.int/emergencies/diseases/novel-coronavirus-2019/situation-reports
Wrapp D, Wang N, Corbett KS, Goldsmith JA, Hsieh C-L, Abiona O, Graham BS, McLellan JS (2020) Cryo-EM structure of the 2019-nCoV spike in the prefusion conformation. Science 367(6483):1260–1263. https://doi.org/10.1126/science.abb2507
Wu F, Zhao S, Yu B, Chen Y-M, Wang W, Song Z-G, Hu Y, Tao Z-W, Tian J-H, Pei Y-Y, Yuan M-L, Zhang Y-L, Dai F-H, Liu Y, Wang Q-M, Zheng J-J, Xu L, Holmes EC, Zhang Y-Z (2020) A new coronavirus associated with human respiratory disease in China. Nature 579(7798):265–269. https://doi.org/10.1038/s41586-020-2008-3
Wu C, Zheng M, Yang Y, Gu X, Yang K, Li M, Liu Y, Zhang Q, Zhang P, Wang Y, Wang Q, Xu Y, Zhou Y, Zhang Y, Chen L, Li H (2020) Furin: A Potential Therapeutic Target for COVID-19. IScience 23(10):101642. https://doi.org/10.1016/j.isci.2020.101642
Xia S, Zhu Y, Liu M, Lan Q, Xu W, Wu Y, Ying T, Liu S, Shi Z, Jiang S, Lu L (2020) Fusion mechanism of 2019-nCoV and fusion inhibitors targeting HR1 domain in spike protein. Cell Mol Immunol 17(7):765–767. https://doi.org/10.1038/s41423-020-0374-2
Xue X, Yu H, Yang H, Xue F, Wu Z, Shen W, Li J, Zhou Z, Ding Y, Zhao Q, Zhang XC, Liao M, Bartlam M, Rao Z (2008) Structures of two coronavirus main proteases: implications for substrate binding and antiviral drug design. J Virol 82(5):2515–2527. https://doi.org/10.1128/JVI.02114-07
Yang P, Wang X (2020) ‘COVID-19: a new challenge for human beings’, cellular and molecular immunology. Nat Publ Group 17(5):555–557. https://doi.org/10.1038/s41423-020-0407-x
Yao X, Ye F, Zhang M, Cui C, Huang B, Niu P, Liu X, Zhao L, Dong E, Song C, Zhan S, Lu R, Li H, Tan W, Liu D (2020) In vitro antiviral activity and projection of optimized dosing design of hydroxychloroquine for the treatment of severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2). Clin Infect Dis Off Publ Infect Dis Soc Am 71(15):732–739. https://doi.org/10.1093/cid/ciaa237
Yazdany J, Kim AHJ (2020) Use of Hydroxychloroquine and chloroquine during the COVID-19 pandemic: what every clinician should know. Ann Intern Med 172(11):754–755. https://doi.org/10.7326/M20-1334
Ye Z-W, Yuan S, Yuen K-S, Fung S-Y, Chan C-P, Jin D-Y (2020) Zoonotic origins of human coronaviruses. Int J Biol Sci 16(10):1686–1697. https://doi.org/10.7150/ijbs.45472
Yu F, Du L, Ojcius DM, Pan C, Jiang S (2020) Measures for diagnosing and treating infections by a novel coronavirus responsible for a pneumonia outbreak originating in Wuhan China. Microb Infect 22(2):74–79. https://doi.org/10.1016/j.micinf.2020.01.003
Zha L, Li S, Pan L, Tefsen B, Li Y, French N, Chen L, Yang G, Villanueva EV (2020) Corticosteroid treatment of patients with coronavirus disease 2019 (COVID-19). Med J Aust 212(9):416–420. https://doi.org/10.5694/mja2.50577
Zhang C, Wu Z, Li J-W, Zhao H, Wang G-Q (2020) Cytokine release syndrome in severe COVID-19: interleukin-6 receptor antagonist tocilizumab may be the key to reduce mortality. Int J Antimicrob Agents 55(5):105954. https://doi.org/10.1016/j.ijantimicag.2020.105954
Zhao Q, Meng M, Kumar R, Wu Y, Huang JF, Deng Y, LiYang ZW (2020) Lymphopenia is associated with severe coronavirus disease 2019 (COVID-19) infections: a systemic review and meta-analysis. Int J Infect Dis 96:131–135
Zheng J (2020) SARS-CoV-2: An Emerging Coronavirus that Causes a Global Threat. Int J Biol Sci 16(10):1678–1685. https://doi.org/10.7150/ijbs.45053
Zhou Y, Hou Y, Shen J, Huang Y, Martin W, Cheng F (2020) Network-based drug repurposing for novel coronavirus 2019-nCoV/SARS-CoV-2. Cell Discov 6(1):1–18. https://doi.org/10.1038/s41421-020-0153-3
Zhu N, Zhang D, Wang W, Li X, Yang B, Song J, Zhao X, Huang B, Shi W, Lu R, Niu P, Zhan F, Ma X, Wang D, Xu W, Wu G, Gao GF, Tan W (2020) A novel coronavirus from patients with pneumonia in China, 2019. N Engl J Med 382(8):727–733. https://doi.org/10.1056/NEJMoa2001017
Acknowledgements
The author (DK) acknowledges Academy of Medical Sciences, UK for the research grant funding (NIF\R1\181849).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
None of the authors have any conflict of interest to declare.
Additional information
Communicated by Erko Stackebrandt.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Raj, C.T.D., Kandaswamy, D.K., Danduga, R.C.S.R. et al. COVID-19: molecular pathophysiology, genetic evolution and prospective therapeutics—a review. Arch Microbiol 203, 2043–2057 (2021). https://doi.org/10.1007/s00203-021-02183-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00203-021-02183-z