Abstract
In the genome of Aspergillus nidulans, a defensin-like protein, Anisin1, was annotated with high homology to the mosquito defensin AaDefA1. So far, no studies exist on defensins from filamentous ascomycetes. Therefore, we characterized the anisin1 gene in A. nidulans and generated a deletion mutant, which suffered from a defect in mitospore development and produced less conidia at 42°C compared to the reference strain. In surface cultures of A. nidulans wild type, the anisin1 expression correlated with that of the central regulator for asexual development, brlA, and with the major scavanger of H2O2 stress, catB, which is indicative for cell differentiation in developing fungi. Interestingly, brlA and anisin1 expressions were deregulated in a ΔsrrA strain that covers a central role in the histidine-to-aspartate (His-Asp) phosphorelay signaling pathway and shows impaired asexual development and H2O2 detoxification. In submers cultures of A. nidulans wild type and other mutants of the His-Asp phosphorelay signaling pathway, anisin1 was repressed, but derepressed in a ΔsrrA background, and anisin1 transcription was further increased in this mutant by H2O2 addition. We therefore conclude that the secreted protein Anisin1 contributes to the optimal development of A. nidulans and we further propose that it has a sensing/signaling function for elevated H2O2 levels.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Antimicrobial peptides (AMPs), such as defensins, are widely distributed in nature as they can be found in organisms ranging from bacteria, plants, insects to humans where they are the product of gene transcription and translation of a single gene (Raj and Dentino 2002; Aerts et al. 2008). In higher eukaryotes, the synthesis of AMPs occurs constitutively or after infection or injury and forms a first line of defense against invading pathogens. Importantly, AMPs represent inducible effector molecules that act as modulators of the innate immune system beside their antimicrobial toxicity (Gallo et al. 2002; Hancock and Patrzykat 2002; Raj and Dentino 2002; Toke 2005; Aerts et al. 2008).
The role of AMPs of prokaryotes and lower eukaryotes is less well understood. Lacking an immune system comparable to that of higher eukaryotes, AMPs could contribute to an ecological advantage of these microorganisms when exposed to unfavorable environmental conditions (Marx 2004). Indeed, bacterial AMPs are part of the quorum sensing mechanism, which plays an important role in microorganisms to communicate and coordinate their behavior by accumulating diffusible signaling molecules in the extracellular environment. This enables, for example, bacteria to chemically measure their cell density and to regulate numerous cellular processes such as secretion of virulence factors, production of biofilms, transcriptional activation of AMPs and oxidative stress resistance (Raina et al. 2009). In analogy, AMPs of filamentous ascomycetes, such as PAF from Penicillium chrysogenum or AFP from Aspergillus giganteus, have antifungal activities (Kaiserer et al. 2003; Theis et al. 2003) and are developmentally regulated (Meyer et al. 2002; Hegedüs et al. 2011b). We could recently show that PAF exhibits an important function in the modulation of asexual development (Hegedüs et al. 2011b).
Defensins represent a unique family of small, cysteine-rich and structured peptides (Raj and Dentino 2002). They consist of about 28–44 amino acids (aa) and have a low molecular mass (3–5 kDa). Up to eight cysteine residues participate in intramolecular disulfide bonds that stabilize an antiparallel β-sheet and an α-helix, (Dimarcq et al. 1998; Gallo et al. 2002; Thevissen et al. 2007; Verma et al. 2007; Aerts et al. 2008). In silico analysis indicated that putative defensin orthologous genes are present also in the genomes of filamentous ascomycetes (Zhu 2008), but they have not been characterized in this fungal group so far.
In this study, we started to investigate the function of a defensin-like encoding gene from the model organism Aspergillus nidulans, anisin1 (AN5046.3; Zhu 2008). The gene product Anisin1 shows high homology to AaDefA1, a defensin from the blood feeding mosquito Aedes aegypti.
By the characterization of an anisin1 deletion mutant in A. nidulans, we provide for the first time evidence that anisin1 plays an important role in the asexual development of A. nidulans by supporting conidiation, especially at elevated cultivation temperatures. Expression analyses in wild-type cells and mutants of the histidine-to-aspartate (His-Asp) phosphorelay signaling cascade indicate that anisin1 transcription is derepressed in a mutant (ΔsrrA) that suffers from impaired asexual development, which further underlines a developmental function of anisin1. Moreover, the expression pattern of anisin1 in ΔsrrA, which also suffers from defective H2O2 detoxification, provides first data that Anisin1 may be involved in sensing/signaling elevated H2O2 levels.
Materials and methods
Strains, media and culture methods
The A. nidulans strains used in this work are listed in Table S1 (Supplements). Generally, fungal strains were grown at 37°C on solid complete medium (CM; 2% d-glucose, 0.2% peptone, 0.1% yeast extract, 0.1% NZ-amine A, 0.05% KCl, 0.04% MgSO4·7H2O, 0.15% KH2PO4, pH 6.5) with the appropriate supplements. Alternatively, minimal medium (MM) according to Pontecorvo et al. (1953) with appropriate supplements was used. To synchronize surface cultures, 18-h-old submers precultures (200 ml MM) were harvested by filtration, transferred to solid MM and further incubated at 37°C. To test oxidative stress response, 18-h-old submers cultures were exposed to 20 mM H2O2 for 2 h before harvesting. For crossfeeding experiments, 5 × 105 conidia (in 50 μl) of A. nidulans Δku70 and Δanisin1 were spotted on 30-ml solid MM and incubated for 38–48 h at 42°C.
Deletion of anisin1 and complementation
To delete the anisin1 gene in A. nidulans, the plasmid pSK275 was used, containing the pyrithiamine (ptrA) expression cassette (Kubodera et al. 2002). According to the splitmarker method (Nielsen et al. 2006), A. nidulans TN02A3 (Δku70) was co-transformed with two DNA constructs, each containing an incomplete fragment of the ptrA expression cassette fused to 1.168 and 1.199 bp of the anisin1 gene flanking sequences, respectively. Homologous integration of each fragment into the flanking sites of anisin1 allowed recombination of the ptrA fragments, which shared a 560 nucleotide overlap, and resulted in the generation of the intact ptrA resistance gene at the site of recombination.
The two DNA constructs for the co-transformation were prepared as follows: the 5′-UTR fragment and 3′-UTR fragment of anisin1 were PCR amplified from genomic DNA (gDNA) with the specific primers 5′ UTR forward B and 5′ UTR reverse and 3′ UTR forward and 3′ UTR reverse (Supplements, Table S2). The PCR fragments of 1.629 bp (5′-UTR fragment) and 1.596 bp (3′-UTR fragment) in size were digested with SacI (5′-UTR fragment) and HindIII (3′-UTR fragment), respectively. The ptrA selection marker was released from the plasmid pSK275 by digestion with SacI and HindIII and ligated to the 5′-UTR fragment and to the 3′-UTR. The two overlapping fragments were amplified from the ligation product using the primers 5′ UTR forward nested and ptrA reverse generating splitmarker I and ptrA forward and 3′ UTR reverse nested generating splitmarker II (Supplements, Table S2).
For complementation, a 4.114-bp anisin1 gene construct was amplified from gDNA using the specific primers 5′ UTR forward A and 3′ UTR reverse and then ligated into pGEM-T (Promega). The anisin1 fragment was released by NdeI digestion and ligated into the NdeI linearized pAN8.1 vector that carries the phleomycin (ble) resistance marker (Punt and van den Hondel 1992). The plasmid pAN8.1/anisin1 was linearized by BsiWI and used for complementation of the deletion strain ∆anisin1.
Transformation of A. nidulans was carried out as previously described by Tilburn et al. (1983). anisin1-depleted transformants were prescreened three times for their ability to grow on MM containing 100 μg/l pyrithiamine hydrobromide. Transformants complemented with the anisin1 wild-type gene were prescreened three times for their ability to grow on MM containing 40 mg/l phleomycin.
Verification of gene deletion and complementation by Southern blot analysis
Site-specific, single-copy integration of transforming DNA was verified by Southern blot analysis according to the protocol of Sambrock et al. (1989). Digested gDNA was separated on a 0.7% agarose gel and blotted onto Hybond N membranes (Amersham, Biosciences). For verification of anisin1 deletion, gDNA was restricted with NdeI and hybridized with the digoxigenin-dUTP-labeled ptrA and anisin1 probes amplified by PCR with the respective oligonucleotides (Supplements, Table S2). For verification of the complementation, gDNA was digested with EcoRI and hybridized with a digoxigenin-dUTP-labeled probe spanning the 5′-UTR and half of the anisin1 gene. Probes were detected with anti-digoxigenin-dUTP antibodies conjugated to alkaline phosphatase according to the manufacturer’s instructions (Roche).
Viability of conidia and vegetative growth tests
Conidia were harvested from 48-h-old surface cultures and resuspended in spore suspension buffer (0.9% NaCl, 0.01% Tween 80). The spore suspension was then diluted to 1 × 105 conidia/ml and stored at 4°C with or without 15% glycerol for up to 12 days. Every 3 days (day 0, 3, 6, 9, 12), aliquots were diluted appropriately, spread on MM agar plates and further incubated at 37°C for 48 h. The number of colonies was determined and those at day 0 were used as a reference to calculate the survival rates (%).
Growth tests were performed in 6-well plates containing 5 ml appropriately supplemented solid MM (nitrogen sources: 20 mM ammoniumtartrate or NaNO2, NaNO3; carbon sources: 10% glucose, 50 or 200 mM sodium acetate). 2 × 103 conidia were point inoculated and incubated at 37°C. Growth was monitored by determining the colony diameters in a time course over 72 h. Growth was also monitored by determining the dry weight of the fungal strains when grown in liquid medium. To this end, 100 ml of CM and MM was inoculated with 1 × 108 conidia and incubated for 24 h at 37°C before the mycelia were harvested. All experiments were performed in duplicates and repeated twice.
Application of oxidative stress in agar plates
To determine the sensitivity of the A. nidulans Δanisin1 deletion mutant to oxidative stress–inducing compounds, agar diffusion assays were performed as described by Eisendle et al. (2006).
Briefly, 1 × 104 conidia were spread on solid MM. Then, holes of 0.5 cm in size were punched in every agar plate, and 50 μl of 20–100 mM H2O2, 1–10 mM menadione and 100–500 mM paraquat were applied. The plates were incubated at 37°C for 48 h, and the diameter of the growth inhibition zone was determined. All experiments were performed in duplicates and repeated twice.
Determination of conidial counts
To investigate mitospore production of the anisin1 deletion strain, 2 × 103 conidia were point inoculated in 6-well plates containing 5-ml solid MM and incubated at 30, 37 and 42°C. Growth was monitored by determining the colony diameters in a time course over 72 h. Conidia were harvested by vortexing the excised surface area in spore suspension buffer and counted. The amount of conidiospores/cm2 was calculated according to the formula: conidia/cm2 = conidia per ml × volume/A whereby A = r 2 · π. Mycelium from equivalent cultures was harvested and subjected to RNA isolation for Northern blot analysis. All experiments were set up in duplicates and were repeated twice.
Northern blot analysis
Total RNA was isolated using TRI reagent (Sigma-Aldrich). 5–10 μg of total RNA was separated on 1.2% formaldehyde agarose gels and blotted onto Hybond N membranes (Amersham, Biosciences) according to the protocols of Fourney et al. (1988) and Kroczek and Siebert (1990). The hybridization probes were generated and labeled with digoxigenin-dUTP (Roche) by PCR using gene-specific primers (Supplements, Table S2). Hybridization probes were detected with anti-digoxigenin-dUTP antibodies conjugated to alkaline phosphatase according to the manufacturer’s instructions (Roche). The quantification of signal intensities of mRNAs on Northern blots was calculated and compared using the Image Quant 5.2 Software (Molecular Dynamics). 18S and 26S rRNA were used as internal standards. All experiments were repeated twice.
Results and discussion
In silico analysis
A tblastn search in the genome of A. nidulans identified two candidate genes with high homology to the insect defensin gene AaDefA1 of the mosquito A. aegypti. One was annotated by Sanger, Broad Institute, as a conserved hypothetical protein on chromosome 3, which contains an arthropod defensin protein domain (contig 84; AN5046.3). This protein was named Anisin1 by Zhu (2008). The other was assigned as Atesin3 (Sanger, Broad Institute) and is localized on chromosome 1 (contig 108; AN11510.3). This protein corresponds to Anisin2 in Zhu (2008).
A ClustalW multiple alignment of the aa sequence of the premature protein forms of Anisin1 and AaDefA1 revealed that both exhibit 35% identity (Supplements, Figure S1). A signal peptide, a propeptide and the N-terminus of the putative mature protein of Anisin1 were identified (Kyte and Doolittle 1982; iPSORT, http://ipsort.hgc.jp/), indicating that Anisin1 is an extracellular protein. The mature proteins Anisin1 and AaDefA1 contain 41 and 40 aa, respectively, and exhibit 51% aa identity (Fig. 1). The postulated positions of the disulfide bonds between the six cysteine residues of the AaDefA1 also fit to Anisin1, which underlines the structural similarity between both the A. nidulans and the mosquito defensin (Zhu 2008) (Fig. 1). The aa sequence of Atesin3 and AaDefA1 showed an identity of 25% in their premature form and 46% identity after maturation (data not shown). Anisin1 and Anisin2/Atesin3 exhibit 69% identity as premature and 70% identity as mature proteins (data not shown).
Alignment of the aa sequences of the mature A. nidulans Anisin1 and A. aegypti insect defensin AaDefA1. Putative structural motifs (γ-core, the α-helix and the β-strands) and the proposed disulfide bond pattern of Anisin1 and AaDefA1 are indicated at the top. The cysteins involved in the formation of disulfide bridges (S–S) are underlined. Identical residues are marked with an asterisk, conserved substitutions with a colon and semi-conserved substitutions with a dot
In this study, we concentrated on the characterization of anisin1 that showed the highest aa identity to the mosquito defensin AaDefA1. Analysis of the promoter region of anisin1 revealed several putative regulatory elements that could be involved in the transcriptional regulation, for example, by the HAP-like complex (Brakhage et al. 1999), CREA (Kulmburg et al. 1993) and GATA factors (Scazzocchio 2000). Besides, four putative stress–responsive elements (STREs) are present (5′-CCCCT-3′ or 5′-AGGGG-3′; position −113, −816, −988 and −998), which might be involved in the activation of anisin1 expression in response to signals related to oxidative stress, nutrient starvation, osmotic stress or low pH (Toone and Jones 1998). A TATAA box is missing (Supplements, Figure S2).
Anisin1 balances asexual development
To investigate the role of anisin1 in A. nidulans, an anisin1 knockout strain was constructed in a Δku70 background (Supplements, Figure S3A–C), and growth and spore viability tests were performed to characterize the phenotype of the Δanisin1 mutant. No difference between the mutant strain and the reference strain Δku70 could be detected in respect to the viability of conidia and the growth rate in liquid or on solid medium. Neither secondary carbon nor secondary nitrogen sources affected the growth of the deletion mutant compared to the control strain under the conditions tested (see “Materials and methods”; data not shown).
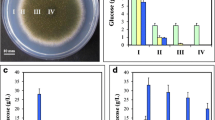
Next, we characterized the mitospore production in the anisin1 deletion strain compared to Δku70 on solid MM at 30, 37 and 42°C, respectively. After 48 h of incubation, there was little phenotypical difference between the mutant and the reference strain when grown at 30 and 37°C, with the exception that at 37°C the Δku70 colony was covered with aerial hyphae which may represent the second generation (Fig. 2a; Supplements Table S3). A severe defect in mitospore development, however, was evident in the Δanisin1 strain at 42°C (Fig. 2a). To assess this phenomenon, the conidial counts were determined (Table 1). At 30°C, the number of conidia of the Δanisin1 strain was reduced by 10% after 48 h and 1% after 72 h and further decreased at 37°C by 15% after 48 h and 21% after 72 h. However, at 42°C the defect in conidia formation of the mutant culminated after 48 h in a reduction by 93% compared to Δku70. This defect in asexual development slightly ameliorated to −65% after 72 h of incubation compared to the reference strain. The complementation of anisin1 rescued the defect in mitospore production (Table 1; Fig. 2a, Supplements Figure S4A, B) and confirmed that the identified phenotype resulted from the deletion of the anisin1 gene. Furthermore, crossfeeding of Δanisin1 by co-cultivation with Δku70 restored the defect in mitospore development at the point of contact of both colonies (Fig. 2b, left panel). This effect was less pronounced when Δanisin1 was co-cultivated with Δanisin1 (Fig. 2b, right panel). This suggests that Anisin1 is secreted by Δku70 into the agar and rescues at least in part the sporulation defect of the Δanisin1 strain. However, other so far not defined factors support a slight sporulation independently from Anisin1 under the conditions tested.
Impact of anisin1 deletion in asexual development of A. nidulans. a Surface cultures of A. nidulans reference strain ∆ku70, the anisin1 deletion strain and ∆anisin1::anisin1 complemented strain. 2 × 103 conidia were point inoculated on solid MM and incubated for 48 h at 30, 37 and 42°C, respectively. b Co-cultivation of A. nidulans Δku70/Δanisin1 (left panel) and Δanisin1/Δanisin1 (right panel). 5 × 105 conidia were point inoculated on solid MM and incubated for 40 h at 42°C
The role of anisin1 in asexual development was further corroborated by Northern blot analysis where we monitored the expression of anisin1 and the central regulator for asexual development in A. nidulans, brlA (Adams et al. 1998), in point inoculated surface cultures of the mutant and the reference strain. In a time course over 48 h, anisin1 and brlA were similarly expressed in a Δku70 surface culture grown at 37 and 42°C (Fig. 3). The anisin1 and brlA transcription was strongest after 24 h and decreased by 48 h. At 30°C, however, the transcription of anisin1 and brlA was delayed, reflecting the lower proliferation and conidiation rate of A. nidulans at this temperature. The anisin1 and brlA mRNAs were most abundant after 36 h of incubation and the transcription level of anisin1 declined rapidly by 48 h of incubation whereas the level of brlA transcripts decreased only moderately at this time point. In the Δanisin1 mutant, the brlA expression at 30°C paralleled the expression in the reference strain (Fig. 3). The quantification of the Northern blot signal intensities of the respective mRNAs revealed, however, that at 37°C the brlA expression was moderately decreased in the mutant by 40% after 24 h, 63% after 36 h and 60% after 48 h compared to the reference strain (Fig. 3; Table 2). Note that the signal intensities of all samples were first corrected according to the respective loading controls (18S and 26S rRNA) and then the values of the Δanisin1 mutant were correlated with the Δku70 strain. Strikingly, at 42°C the brlA expression was severely retarded in the Δanisin1 mutant and the amount of brlA mRNA was reduced by 70% after 24 h. At later time points, brlA expression increased (−18% after 36 h) and finally reached +87% after 48 h compared to the control strain (Fig. 3; Table 2). This may reflect the amelioration in the production of conidia in the Δanisin1 mutant with later incubation times (Table 1). Therefore, we conclude that the defect in mitospore production of Δanisin1 at 42°C results from a delayed and reduced expression of brlA.
Northern blot analysis of the expression of anisin1, brlA and catB in surface cultures of the A. nidulans reference strain Δku70 and the Δanisin1 mutant, respectively, after 24, 36 and 48 h incubation at 30, 37 and 42°C. 10 μg of total RNA was loaded into each well, blotted and hybridized with the respective gene-specific digoxigenin-labeled probes. Ethidiumbromide-stained 26S and 18S rRNA are shown as loading controls
It is known that the intracellular level of reactive oxygen species (ROS) regulates most diverse cellular functions in Aspergillus sp., among others also hyphal differentiation and mitospore production (Hansberg and Aguirre 1990; Aguirre et al. 2005; Reverberi et al. 2008). As such, hyperoxidant conditions trigger microbial differentiation (Kawasaki et al. 1997). The increase in catB expression, which is one of the major scavengers against H2O2, is indicative for elevated endogenous ROS levels during cell differentiation as well as for the response to oxidative stress–inducing agents such as H2O2 (Hansberg et al. 1993; Kawasaki et al. 1997). Therefore, it is crucial for microorganisms, such as A. nidulans, that the oxidative stress response is tightly regulated (Pócsi et al. 2005; Miskei et al. 2009; Balázs et al. 2010).
To answer the question whether Anisin1 function may be interconnected with oxidative stress response in asexual development, we investigated the expression pattern of catB in the surface cultures of Δku70 and Δanisin1 (Fig. 3). At 30°C, the catB expression pattern of Δanisin1 resembled that of Δku70. Instead, the intensity of the catB signals was moderately, but consistently higher in the Δanisin1 strain grown at 37 and 42°C compared to the reference strain (Fig. 3; Table 2). At 37°C, the catB expression increased by 66% after 24 h, 84% after 36 h and 71% after 48 h in the mutant strain compared to the control (Δku70). At 42°C, the amount of catB mRNA increased by 78% after 24 h, 44% after 36 h and 182% after 48 h in the Δanisin1 strain compared to the Δku70.
This indicates that depletion of anisin1 not only leads to the deregulation of brlA, but also moderately triggers catB expression in developing A. nidulans. Possibly, the developmental response to hyperoxidant conditions at elevated temperatures is impaired in the anisin1-depleted strain, and a ROS threshold higher than in the Δku70 is needed for the onset of differentiation. Importantly, except for the Δanisin1 mutant cultivated at 42°C, the catB transcription pattern correlated with that of brlA and anisin1 in the Δku70 and with that of brlA in the Δanisin1 at the growth temperatures of 30 and 37°C, respectively. Thus, our observations further support the role of anisin1 in the developmental regulation of A. nidulans.
Finally, to confirm or dismiss the hypothesis that the Δanisin1 mutant suffers from an impaired oxidative stress response, we tested the sensitivity of Δanisin1 to 20–100 mM H2O2 (hydroxyl radical generator), 1–10 mM menadione and 0.1–0.5 M paraquat (both superoxide radical generators) by agar diffusion tests. No difference in the susceptibility between the anisin1 deletion mutant and the reference strain Δku70 was detectable (data not shown). This result let us conclude that Anisin1 is not primarily involved in oxidative stress response such as the detoxification of oxidative radicals.
anisin1 expression is deregulated in a ΔsrrA mutant
To investigate in depth the role of anisin1 in asexual development, we monitored the anisin1 transcription in an A. nidulans mutant defective in SrrA function compared to the wild-type strain BPU1. SrrA is a Skn7-type response regulator (RR) in the His-Asp phosphorelay signaling pathway, which supports the ability of growth and survival under unfavorable environmental conditions. In the last years, several studies contributed to the characterization of some of the major players of this pathway in yeasts and filamentous fungi (Maeda et al. 1994; Posas et al. 1996; Hagiwara et al. 2007a, b; Izumitsu et al. 2007; Vargas-Pérez et al. 2007; Ruprich-Robert et al. 2008; Zhang et al. 2010). In A. nidulans, one of several histidine kinases (HK), namely NikA, senses environmental signals and transmits it to RRs. The transfer of phosphoryl groups (phosphorelay system) renders the RRs functional, generally as transcriptional regulators (SrrA, NapA) or as intermediate regulators (SskA) of mitogen-activated kinases (Mpk), like HogA, which in turn activate specific transcriptional regulators (Asano et al. 2007; Vargas-Pérez et al. 2007; Hagiwara et al. 2009). Apart from osmotic adaptation, fungicide inhibition and cell wall resistance, SrrA has been implicated in asexual development and oxidative stress response (Hagiwara et al. 2007a; Vargas-Pérez et al. 2007). Notably, the ΔsrrA mutant produces less conidia compared to the wild-type strain and is defective in catB induction. Thus, the detoxification of ROS, especially H2O2, is impaired (Hagiwara et al. 2007a; Vargas-Pérez et al. 2007). Due to these phenotypical features, the ΔsrrA mutant represented a valuable A. nidulans strain to study the function of anisin1.
First, we performed Northern blot analysis to monitor the transcription levels of anisin1, brlA and catB for 24 h in synchronized surface cultures of the ΔsrrA mutant compared to the wild-type strain (Fig. 4). In the wild-type strain, anisin1 transcripts could be detected 6–24 h after the hyphae were exposed to the air (Fig. 4). This correlated with the expression of brlA and the induction of catB transcription. In contrast, anisin1, brlA and catB expressions were deregulated in the ΔsrrA mutant (Fig. 4). The anisin1 gene was derepressed already in the 18 h old submers preculture before the hyphae were exposed to the air (0 h), and the transcription decreased after 12 h of synchronization. Only low amounts of brlA mRNA could be detected 6 h after synchronization, and the induction of brlA expression was limited to a very short time (12 h after synchronization), which reflects the reported conidiation defect of this A. nidulans mutant (Hagiwara et al. 2007a; Vargas-Pérez et al. 2007). Importantly, this result points toward a BrlA-independent induction of anisin1 transcription.
Northern blot analysis of the expression of anisin1, brlA and catB in synchronized surface cultures of A. nidulans wild-type BPU1 and ΔsrrA grown at 37°C under isotonic and hypertonic (0.6 M sucrose) conditions. Total RNA was extracted at the indicated time points (0, 6, 12 and 24 h) with time point 0 h referring to the liquid preculture before exposure to the air. 5 μg of total RNA was loaded into each well, blotted and hybridized with the respective gene-specific digoxigenin-labeled probes. Ethidiumbromide-stained 26S and 18S rRNA are shown as loading controls
The expression of catB, however, was not triggered by the exposure of the ΔsrrA mutant to the air. The catB expression was similarly expressed at the time points 0 and 24 h and was even slightly decreased at the time points 6 and 12 h (Fig. 4). This indicates that ΔsrrA is unable to adequately respond to the modulation of endogenously generated H2O2 levels when exposed to the air, and this might contribute at least in part to the defect in mitospore production.
Vargas-Pérez et al. (2007) documented that mild hyperosmotic conditions rescued the conidiation defect of the ΔsrrA mutant possibly by channeling oxidative stress toward SskA activation. Therefore, we tested the effect of 0.6 M sucrose on the expression of anisin1, brlA and catB in the ΔsrrA mutant and used the wild type as a control (Fig. 4). The supplementation of the growth medium with 0.6 M sucrose repressed catB transcription in the wild-type strain, and the transcription of anisin1 and brlA was delayed by 6 h compared to normal growth conditions. Furthermore, the amount of anisin1 mRNA was significantly less under mild hyperosmotic growth conditions. The repression of catB transcription in the wild-type strain implies that high osmolarity lowers the oxidative stress burden of the developing cells.
Hyperosmotic growth conditions also repressed catB transcription in the ΔsrrA mutant when compared to normal growth conditions. However, the expression of anisin1 decreased already after 6 h of exposure to the air, and minimal amounts of brlA mRNA were detectable over the time course investigated. This result points toward a severely impaired asexual development of the ΔsrrA at least within the first 24 h. Thus, a remediation of the sporulation defect of this mutant by high osmolarity could not be observed at the molecular level within 24 h after exposition to the air and contrasts to the report of Vargas-Pérez et al. (2007). This discrepancy might be explained by the different experimental setup that we applied in our study: we used synchronized surface cultures and monitored the gene expression over 24 h whereas Vargas-Pérez and coworkers analyzed the number of conidia of point inoculated surface cultures incubated for 6 days (Vargas-Pérez et al. 2007). It is conceivable to assume that the formation of conidia in the ΔsrrA strain ameliorates during successive developmental generations incubated for longer times on hyperosmotic medium.
Further evidence that hyperosmotic growth conditions do not trigger anisin1 expression in a submers A. nidulans wild-type culture is provided by the transcriptome analysis for the A. nidulans HogA Mpk signaling pathway in response to osmotic stress that has been recently published by Hagiwara et al. (2009) (Gene Expression Omnibus (GEO) from NCBI, accession number GSE16566; data not shown).
Next, we wanted to investigate in more detail the deregulation of anisin1 in a ΔsrrA submers culture. To this end, we compared the anisin1 expression pattern of the ΔsrrA and other mutants of the His-Asp phosphorelay pathway. In contrast to the situation in the ΔsrrA mutant, anisin1 was repressed in the wild-type strain and in the mutants ΔsskA and ΔhogA (Fig. 5). This was also true for the deletion mutants ΔnikA and ΔnapA (data not shown). The derepression of anisin1 in ΔsrrA was even more pronounced under H2O2 (20 mM for 2 h) stress (Fig. 5). To monitor reduced CatB function in the ΔsrrA strain, we determined the catB expression as well. H2O2 triggered catB transcription in the wild type and in the ΔsskA and ΔhogA mutants, but not in the ΔsrrA mutant when compared to the respective untreated controls (Fig. 5). This clearly shows that CatB-mediated detoxification of H2O2 was impaired in this strain. These data further indicate that anisin1 expression is derepressed in a ΔsrrA mutant that suffers from defective H2O2 detoxification. The ROS burden of the cell increases even more upon H2O2 challenge, which is accompanied by a substantial increase in anisin1 expression.
Northern blot analysis of the expression of anisin1 and catB in liquid cultures of A. nidulans wild-type BPU1 and single mutants of the His-Asp phosphorelay pathway ΔsrrA, ∆sskA, ∆hogA challenged with 20 mM H2O2 for 2 h. Controls (Co) were left untreated. 5 μg of total RNA was loaded into each well, blotted and hybridized with the respective gene-specific digoxigenin-labeled probes. Ethidiumbromide-stained 26S and 18S rRNA are shown as loading controls
Conclusion
Our results indicate that anisin1 codes for a secreted protein that supports optimal conidiation of A. nidulans, and we propose that Anisin1 directly or indirectly modulates the expression of brlA and/or the activity of BrlA, possibly by interacting with other factors that are involved in developmental regulation of A. nidulans. These factors could be, for example, StuA, AbaA (Adams et al. 1998) or other so far unidentified players. This assumption, however, remains to be evidenced at the molecular level, and it is crucial to identify the interacting partner of Anisin1.
Importantly, anisin1 is not directly involved in the detoxification of ROS, because (1) depletion of anisin1 did not increase the susceptibility of A. nidulans to externally applied ROS-inducing agents and (2) anisin1 expression was not triggered by externally applied H2O2 in A. nidulans wild type. In a ΔsrrA mutant, however, anisin1 was derepressed under standard growth conditions and its expression was strongly induced by externally applied H2O2. Further studies are needed to address the question whether SrrA directly or rather indirectly regulates anisin1 expression and which signal transduction pathways are involved in anisin1 regulation.
For completion, it has to be noted that no expression of the anisin1-related anisin2/atesin3 gene could be detected neither in the Δanisin1 mutant and the respective reference strain Δku70 nor in the mutants of the His-Asp phosphorelay signaling pathway and the wild-type strain BPU1 under all conditions tested (data not shown). This excludes a redundant function of this gene and indicates that it plays a different role than anisin1.
The existence of various endogenous secreted molecules that regulate growth and conidiation point to the possibility that autoinducer-mediated mechanisms are widespread among filamentous fungi (Lee and Adams 1994; Roncal et al. 2002; Tsitsigiannis and Keller 2007). Based on our results, some conditional parallels between the effect of Anisin1 and other modulators of growth and development can be drawn. For example, the cysteine-rich secreted antifungal protein PAF of P. chrysogenum helps the producing mold to adjust to variable environmental conditions by balancing asexual development (Hegedüs et al. 2011a, b). This function, in addition to its antifungal activity, provides P. chrysogenum an ecological advantage over concurring organisms. Work is in progress, however, to verify a putative antimicrobial function of Anisin1.
Interestingly, defensins of higher eukaryotes are central players in the modulation of the innate immune system and in stress signaling. Both functions have been proposed to be even more relevant than their antimicrobial activities (Lowenberger 2001). Notably, cysteine-rich small molecular proteins are perfect targets for oxidative or reductive protein transformation resulting in a conformational change that could influence and modulate their activity and explain their variable function in response to environmental stimuli, such as hyperoxidant states or hypoxic conditions (Hansberg and Aguirre 1990; Aguirre et al. 2005; Batta et al. 2009). In this respect, our results let us propose that anisin1 plays a role in sensing intracellular H2O2, which arises either in response to developmental signals or due to defective detoxification. The presence of putative regulatory elements, like STRE, in the anisin1 promoter could further indicate that anisin1 is a stress-responsive gene. This, however, needs to be clarified in further experiments.
References
Adams TH, Wieser JK, Yu JH (1998) Asexual sporulation in Aspergillus nidulans. Microbiol Mol Biol Rev 62:35–54
Aerts AM, Francois IE, Cammue BP, Thevissen K (2008) The mode of antifungal action of plant, insect and human defensins. Cell Mol Life Sci 65:2069–2079
Aguirre J, Rios-Momberg M, Hewitt D, Hansberg W (2005) Reactive oxygen species and development in microbial eukaryotes. Trends Microbiol 13:111–118
Asano Y, Hagiwara D, Yamashino T, Mizuno T (2007) Characterization of the bZip-type transcription factor NapA with reference to oxidative stress response in Aspergillus nidulans. Biosci Biotechnol Biochem 71:1800–1803
Balázs A et al (2010) AtfA bZIP-type transcription factor regulates oxidative and osmotic stress responses in Aspergillus nidulans. Mol Genet Genomics 283:289–303
Batta G et al (2009) Functional aspects of the solution structure and dynamics of PAF—a highly-stable antifungal protein from Penicillium chrysogenum. FEBS J 276:2875–2890
Brakhage AA et al (1999) HAP-Like CCAAT-binding complexes in filamentous fungi: implications for biotechnology. Fungal Genet Biol 27:243–252
Dimarcq JL, Bulet P, Hetru C, Hoffmann J (1998) Cysteine-rich antimicrobial peptides in invertebrates. Biopolymers 47:465–477
Eisendle M, Schrettl M, Kragl C, Müller D, Illmer P, Haas H (2006) The intracellular siderophore ferricrocin is involved in iron storage, oxidative-stress resistance, germination, and sexual development in Aspergillus nidulans. Eukaryot Cell 5:1596–1603
Fourney RM, Miyakoshi J, Day RS, Paterson MC (1988) Northern blotting: efficient RNA staining and transfer. Focus 10:5–7
Gallo RL, Murakami M, Ohtake T, Zaiou M (2002) Biology and clinical relevance of naturally occurring antimicrobial peptides. J Allergy Clin Immunol 110:823–831
Hagiwara D et al (2007a) The SskA and SrrA response regulators are implicated in oxidative stress responses of hyphae and asexual spores in the phosphorelay signaling network of Aspergillus nidulans. Biosci Biotechnol Biochem 71:1003–1014
Hagiwara D et al (2007b) Characterization of the NikA histidine kinase implicated in the phosphorelay signal transduction of Aspergillus nidulans, with special reference to fungicide responses. Biosci Biotechnol Biochem 71:844–847
Hagiwara D, Asano Y, Marui J, Yoshimi A, Mizuno T, Abe K (2009) Transcriptional profiling for Aspergillus nidulans HogA MAPK signaling pathway in response to fludioxonil and osmotic stress. Fungal Genet Biol 46:868–878
Hancock RE, Patrzykat A (2002) Clinical development of cationic antimicrobial peptides: from natural to novel antibiotics. Curr Drug Targets Infect Disord 2:79–83
Hansberg W, Aguirre J (1990) Hyperoxidant states cause microbial cell differentiation by cell isolation from dioxygen. J Theor Biol 142:201–221
Hansberg W, de Groot H, Sies H (1993) Reactive oxygen species associated with cell differentiation in Neurospora crassa. Free Radic Biol Med 14:287–293
Hegedüs N et al (2011a) The small molecular mass antifungal protein of Penicillium chrysogenum—a mechanism of action oriented review. J Basic Microbiol 51:1–11
Hegedüs N, Sigl C, Zadra I, Pócsi I, Marx F (2011b) The paf gene product modulates asexual development in Penicillium chrysogenum. J Basic Microbiol 51:253–262
Izumitsu K, Yoshimi A, Tanaka C (2007) Two-component response regulators Ssk1p and Skn7p additively regulate high-osmolarity adaptation and fungicide sensitivity in Cochliobolus heterostrophus. Eukaryot Cell 6:171–181
Kaiserer L, Oberparleiter C, Weiler-Görz R, Burgstaller W, Leiter É, Marx F (2003) Characterization of the Penicillium chrysogenum antifungal protein PAF. Arch Microbiol 180:204–210
Kawasaki L, Wysong D, Diamond R, Aguirre J (1997) Two divergent catalase genes are differentially regulated during Aspergillus nidulans development and oxidative stress. J Bacteriol 179:3284–3292
Kroczek RA, Siebert E (1990) Optimization of northern analysis by vacuum-blotting, RNA-transfer visualization, and ultraviolet fixation. Anal Biochem 184:90–95
Kubodera T, Yamashita N, Nishimura A (2002) Transformation of Aspergillus sp. and Trichoderma reesei using the pyrithiamine resistance gene (ptrA) of Aspergillus oryzae. Biosci Biotechnol Biochem 66:404–406
Kulmburg P, Mathieu M, Dowzer C, Kelly J, Felenbok B (1993) Specific binding sites in the alcR and alcA promoters of the ethanol regulon for the CREA repressor mediating carbon catabolite repression in Aspergillus nidulans. Mol Microbiol 7:847–857
Kyte J, Doolittle RF (1982) A simple method for displaying the hydropathic character of a protein. J Mol Biol 157:105–132
Lee BN, Adams TH (1994) The Aspergillus nidulans fluG gene is required for production of an extracellular developmental signal and is related to prokaryotic glutamine synthetase I. Genes Dev 8:641–651
Lowenberger C (2001) Innate immune response of Aedes aegypti. Insect Biochem Mol Biol 31:219–229
Maeda T, Wurgler-Murphy SM, Saito H (1994) A two-component system that regulates an osmosensing MAP kinase cascade in yeast. Nature 369:242–245
Marx F (2004) Small, basic antifungal proteins secreted from filamentous ascomycetes: a comparative study regarding expression, structure, function and potential application. Appl Microbiol Biotechnol 65:133–142
Meyer V, Wedde M, Stahl U (2002) Transcriptional regulation of the antifungal protein in Aspergillus giganteus. Mol Genet Genomics 266:747–757
Miskei M, Karányi Z, Pócsi I (2009) Annotation of stress-response proteins in the aspergilli. Fungal Genet Biol 46(Suppl 1):S105–S120
Nayak T, Szewczyk E, Oakley CE, Osmani A, Ukil L, Murray SL, Hynes MJ, Osmani SA, Oakley BR (2006) A versatile and efficient gene-targeting system for Aspergillus nidulans. Genetics 172:1557–1566
Nielsen ML, Albertsen L, Lettier G, Nielsen JB, Mortensen UH (2006) Efficient PCR-based gene targeting with a recyclable marker for Aspergillus nidulans. Fungal Genet Biol 43:54–64
Pócsi I et al (2005) Comparison of gene expression signatures of diamide, H2O2 and menadione exposed Aspergillus nidulans cultures—linking genome-wide transcriptional changes to cellular physiology. BMC Genomics 6:182
Pontecorvo G, Roper JA, Hemmons LM, Macdonald KD, Bufton AW (1953) The genetics of Aspergillus nidulans. Adv Genet 5:141–238
Posas F, Wurgler-Murphy SM, Maeda T, Witten EA, Thai TC, Saito H (1996) Yeast HOG1 MAP kinase cascade is regulated by a multistep phosphorelay mechanism in the SLN1-YPD1-SSK1 “two-component” osmosensor. Cell 86:865–875
Punt PJ, van den Hondel CA (1992) Transformation of filamentous fungi based on hygromycin B and phleomycin resistance markers. Methods Enzymol 216:447–457
Raina S, De Vizio D, Odell M, Clements M, Vanhulle S, Keshavarz T (2009) Microbial quorum sensing: a tool or a target for antimicrobial therapy? Biotechnol Appl Biochem 54:65–84
Raj PA, Dentino AR (2002) Current status of defensins and their role in innate and adaptive immunity. FEMS Microbiol Lett 206:9–18
Reverberi M et al (2008) Modulation of antioxidant defense in Aspergillus parasiticus is involved in aflatoxin biosynthesis: a role for the ApyapA gene. Eukaryot Cell 7:988–1000
Roncal T, Cordobes S, Sterner O, Ugalde U (2002) Conidiation in Penicillium cyclopium is induced by conidiogenone, an endogenous diterpene. Eukaryot Cell 1:823–829
Ruprich-Robert G, Chapeland-Leclerc F, Boisnard S, Florent M, Bories G, Papon N (2008) Contributions of the response regulators Ssk1p and Skn7p in the pseudohyphal development, stress adaptation, and drug sensitivity of the opportunistic yeast Candida lusitaniae. Eukaryot Cell 7:1071–1074
Sambrock J, Fritsch EF, Maniatis T (1989) Molecular cloning: a laboratory manual, 2nd edn. Cold spring Harbour Laboratory Press, Cold Spring Harbor
Scazzocchio C (2000) The fungal GATA factors. Curr Opin Microbiol 3:126–131
Theis T, Wedde M, Meyer V, Stahl U (2003) The antifungal protein from Aspergillus giganteus causes membrane permeabilization. Antimicrob Agents Chemother 47:588–593
Thevissen K, Kristensen HH, Thomma BP, Cammue BP, Francois IE (2007) Therapeutic potential of antifungal plant and insect defensins. Drug Discov Today 12:966–971
Tilburn J, Scazzocchio C, Taylor GG, Zabicky-Zissman JH, Lockington RA, Davies RW (1983) Transformation by integration in Aspergillus nidulans. Gene 26:205–221
Toke O (2005) Antimicrobial peptides: new candidates in the fight against bacterial infections. Biopolymers 80:717–735
Toone WM, Jones N (1998) Stress-activated signalling pathways in yeast. Genes Cells 3:485–498
Tsitsigiannis DI, Keller NP (2007) Oxylipins as developmental and host-fungal communication signals. Trends Microbiol 15:109–118
Vargas-Pérez I, Sánchez O, Kawasaki L, Georgellis D, Aguirre J (2007) Response regulators SrrA and SskA are central components of a phosphorelay system involved in stress signal transduction and asexual sporulation in Aspergillus nidulans. Eukaryot Cell 6:1570–1583
Verma C et al (2007) Defensins: antimicrobial peptides for therapeutic development. Biotechnol J 2:1353–1359
Zhang H et al (2010) A two-component histidine kinase, MoSLN1, is required for cell wall integrity and pathogenicity of the rice blast fungus, Magnaporthe oryzae. Curr Genet 56:517–528
Zhu S (2008) Discovery of six families of fungal defensin-like peptides provides insights into origin and evolution of the CSalphabeta defensins. Mol Immunol 45:828–838
Acknowledgments
We want to express our special thanks to Dr. D. Hagiwara (Tohoku University, Japan) for providing the A. nidulans mutant strains of the histidine-to-aspartate signaling pathway. We are grateful to Doris Bratschun for technical assistance. This work was supported by the Austrian Science Fund FWF (grant no. P19970) to F. M. and by the Hungarian Scientific Research Fund (grant no. 77515) to I. P.
Open Access
This article is distributed under the terms of the Creative Commons Attribution Noncommercial License which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and source are credited.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by Axel Brakhage.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This is an open access article distributed under the terms of the Creative Commons Attribution Noncommercial License (https://creativecommons.org/licenses/by-nc/2.0), which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and source are credited.
About this article
Cite this article
Eigentler, A., Pócsi, I. & Marx, F. The anisin1 gene encodes a defensin-like protein and supports the fitness of Aspergillus nidulans . Arch Microbiol 194, 427–437 (2012). https://doi.org/10.1007/s00203-011-0773-y
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00203-011-0773-y