Abstract
Aims/hypothesis
Enteroviral infection has been implicated consistently as a key environmental factor correlating with the appearance of autoimmunity and/or the presence of overt type 1 diabetes, in which pancreatic insulin-producing beta cells are destroyed by an autoimmune response. Genetic predisposition through variation in the type 1 diabetes risk gene IFIH1 (interferon induced with helicase C domain 1), which encodes the viral pattern-recognition receptor melanoma differentiation-associated protein 5 (MDA5), supports a potential link between enterovirus infection and type 1 diabetes.
Methods
We used molecular techniques to detect enterovirus RNA in peripheral blood samples (in separated cellular compartments or plasma) from two cohorts comprising 79 children or 72 adults that include individuals with and without type 1 diabetes who had multiple autoantibodies. We also used immunohistochemistry to detect the enteroviral protein VP1 in the pancreatic islets of post-mortem donors (n=43) with type 1 diabetes.
Results
We observed enhanced detection sensitivity when sampling the cellular compartment compared with the non-cellular compartment of peripheral blood (OR 21.69; 95% CI 3.64, 229.20; p<0.0001). In addition, we show that children with autoimmunity are more likely to test positive for enterovirus RNA than those without autoimmunity (OR 11.60; 95% CI 1.89, 126.90; p=0.0065). Furthermore, we found that individuals carrying the predisposing allele (946Thr) of the common variant in IFIH1 (rs1990760, Thr946Ala) are more likely to test positive for enterovirus in peripheral blood (OR 3.07; 95% CI 1.02, 8.58; p=0.045). In contrast, using immunohistochemistry, there was no correlation between the common variant in IFIH1 and detection of enteroviral VP1 protein in the pancreatic islets of donors with type 1 diabetes.
Conclusions/interpretation
Our data indicate that, in peripheral blood, antigen-presenting cells are the predominant source of enterovirus infection, and that infection is correlated with disease stage and genetic predisposition, thereby supporting a role for enterovirus infection prior to disease onset.
Graphical abstract

Similar content being viewed by others
Avoid common mistakes on your manuscript.

Introduction
Type 1 diabetes is caused by progressive loss of the insulin-producing beta cells in pancreatic islets. Genetic factors are important in the predisposition to disease development [1]. However, a concordance rate of only around 50% in monozygotic twins [2] and the steadily increasing incidence rate [3], particularly in those individuals with lower genetic predisposition [3, 4], suggest that environmental factors also play a crucial role.
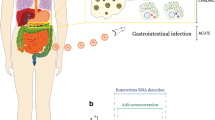
A prominent candidate environmental factor is virus infection [5], particularly infection with Coxsackievirus, a subgroup of the genus Enterovirus (EV) (Picornaviridae family) that has been extensively studied and linked to type 1 diabetes [6, 7]. EV is detectable at a higher frequency in stool samples [8, 9], pancreatic biopsies [10,11,12] and the peripheral blood [13, 14] of individuals with type 1 diabetes compared to those without, while the presence of neutralising antibodies against Coxsackievirus correlates with beta cell autoimmunity [15]. Studies have shown that EV is found more often in both the serum/plasma and peripheral blood mononuclear cells (PBMCs) [14, 16, 17] of individuals with type 1 diabetes and those with islet autoimmunity. Similarly, EV infection is detected in the pancreatic tissue of approximately 70% of post-mortem donors with recent-onset type 1 diabetes compared with less than 10% of similarly aged post-mortem donors without type 1 diabetes [10].
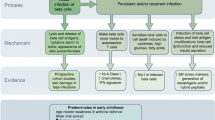
Mechanistically, there is an interaction between EV infection and the genetic variation that predisposes to type 1 diabetes. Several risk-determining variants have been identified in the gene IFIH1 (interferon induced with helicase C domain 1), which encodes the cytoplasmic viral pattern-recognition receptor melanoma differentiation-associated protein 5 (MDA5) [18, 19]. MDA5 is essential for the detection of members of the Picornaviridae family [20, 21], and its activation leads to production of type I IFN and proinflammatory cytokines [22]. Most informatively, four rare SNPs exist that reduce or abrogate the function of MDA5, and these variants all provide protection against type 1 diabetes [19, 23]. For the common variant SNP rs1990760 (Thr946Ala) in IFIH1, 946Thr is the predisposing allele [18]. In PBMCs, the disease-protective allele (946Ala) is associated with reduced expression of IFIH1 either under basal conditions [24] or after stimulation with IFNβ or polyinosinic-polycytidylic acid [25, 26]. Functionally, however, a greater degree of divergence has been reported, with one study finding that protection correlates with reduced type I IFN response [26], while this was not seen in other studies [23, 27]. Another study observed reduced type III IFN responses, but not reduced type I IFN responses, in virus-infected pancreatic islets from donors homozygous for the predisposing allele in IFIH1 [28].
Whether these functional consequences of variants in IFIH1 affect the rate of virus infection and clearance is still under investigation, and the studies that have investigated the relationship between detection of EV and variants in IFIH1 have yielded inconclusive results [8, 29]. Here we investigated whether detection of EV infection in peripheral blood and pancreatic tissue correlated with the predisposing allele (946Thr) of the common variant in IFIH1 (rs1990760, Thr946Ala).
Methods
Cohorts
We analysed two distinct cohorts: the ‘children cohort’, which included 79 children (median age 119 months, range 17–192 months, 54% female); and the ‘adult cohort’, which included 72 adults (median age 29 years, range 18–51 years, 61% female). Ethics approval was obtained from the Bromley National Research Ethics Service Committee (reference number 08/H0805/14) for the adult cohort, and from the Ethics Committee of Pirkanmaa Hospital District, Tampere, Finland, for the children cohort. Written informed consent was obtained from all participants or their legal guardians.
The children cohort included 49 case children who repeatedly tested positive for multiple biochemical islet autoantibodies (referred to as mAAb-positive) (i.e. combinations of insulin autoantibodies (IAA), GAD autoantibodies (GADA) and tyrosine phosphatase IA-2 autoantibodies (IA-2A)) and 30 autoantibody-negative control children who were matched for age (all <13 years old), sex and place of birth (city). Among the children who were positive for mAAb, 24 later progressed to type 1 diabetes, diagnosed according to the WHO recommendations [30]. Both case and control children carried HLA genotypes that confer increased risk for type 1 diabetes, and had been followed from birth in the Finnish Type 1 Diabetes Prediction and Prevention study described previously [31]. PBMCs and plasma were isolated by density gradient centrifugation (Ficoll-Paque PLUS, GE Healthcare BioSciences, Sweden). PBMCs were pelleted and stored in RLT buffer (Qiagen, Germany). Both PBMCs and plasma were stored at -80°C for subsequent RNA extraction.
The adult cohort (all >18 years old) included 37 individuals with recent-onset type 1 diabetes (within 3 months of diagnosis) and 35 individuals without type 1 diabetes, of similar age and matched for sex, and with no family history of autoimmune disease. PBMCs were isolated by density gradient centrifugation (Lymphoprep; Axis-Shield, Norway). PBMCs were treated with FcR blocking reagent (Miltenyi Biotec, Germany), and PBMC subsets were subsequently enriched using magnetic bead cell separation by autoMACS (Mitenyi Biotec) in the following order: B cells (using CD19 MicroBeads), monocytes (using CD14 MicroBeads), myeloid dendritic cells (mDCs) (using a CD1c [BDCA-1] dendritic cell isolation kit), plasmacytoid dendritic cells (pDCs) (using a CD304 [BDCA-4/neuropilin-1] MicroBead kit). All reagents for cell separation were obtained from Miltenyi Biotec, and the post-separation enrichment was >90%, according to the manufacturer. Samples were pelleted and stored at -80°C until RNA extraction.
For both cohorts, individuals who reported or showed symptoms of systemic ‘virus-like’ illness were not recruited to the study or did not undergo blood sampling. In the children cohort, none of the individuals were excluded from blood sampling due to ‘virus-like’ illness.
RNA extraction and detection of EV-RNA
RNA was extracted using a QIAamp viral RNA kit (Qiagen) and TRIzol reagent (Life Technologies, USA), in the adult and children cohorts, respectively, according to the manufacturer’s instructions. Detection of EV-RNA was performed by RT-PCR and liquid-phase hybridisation using a primer pair (forward: 5′-CGGCCCCTGAATGCGGCTAA-3′; reverse: 5′-GAAACACGGACACCCAAAGTA-3′) from the highly conserved 5′ non-coding region as previously described [32]. PCR amplicons were hybridised using a europium-labelled EV-specific probe (5′-TAITCGGTTCCGCTGC-3′) in a liquid-phase assay on a microtitre plate [33]. All positive samples were confirmed as positive by repeated RT-PCR and hybridisation assay.
IFIH1 genotyping
In the adult cohort, DNA was extracted from whole blood collected using the QIamp blood mini kit (Qiagen) according to the manufacturer’s instructions, and genotyping for the SNP rs1990760 was performed by TaqMan assay (Applied Biosystems, USA). In the children cohort, DNA was extracted from EDTA-treated blood samples by a salting-out protocol [34], and genotyping was performed either using a Sequenom platform (San Diego, USA) at the Genome Center of Eastern Finland, University of Eastern Finland (Kuopio), or by TaqMan assay (Applied Biosystems) in samples that were not included in the previous Sequenom-based study [35]. For each pancreas, sample DNA was extracted from 2 × 4 μm formalin-fixed, paraffin-embedded (FFPE) tissue curls using the QIAamp DNA FFPE tissue kit (Qiagen) according to the manufacturer’s instructions. SNP genotyping was performed by Kompetitive allele-specific PCR (KASP) (LGC Biosearch Technologies, UK) using 1 μl DNA amplified in a 5 μl KASP reaction. DNA was amplified and fluorescence detected using the QuantStudio 12K Flex Real-Time PCR system (ThermoFisher). Genotypes were called using QuantStudio 12K Flex software version 1.2.2 (ThermoFisher). We were unable to isolate pure and good-quality DNA from formalin-fixed, paraffin-embedded tissue for all donors, and therefore obtained IFIH1 genotypes for 43 of the previously reported 72 post-mortem donors with type 1 diabetes [10].
Immunohistochemistry
Formalin-fixed, paraffin-embedded pancreatic tissue from 43 individuals (median age 13.5 years, range 1–42 years, 69% female) with recent-onset type 1 diabetes, whose pancreatic histology has been described previously [36], was used for the immunohistochemical study. Data for the staining of the enteroviral protein VP1, and representative staining images, have been reported previously [10]. As previously described, VP1 positivity was assigned when at least one intensely stained endocrine cell was present in any islet within any given section [10]. All samples were used with ethical permission from the West of Scotland Research Ethics Committee (reference 20/WS/0074; Integrated Research Application System project ID 28362015/WS/0258). Sections were processed and labelled using a standard immunoperoxidase technique for paraffin sections, using heat-induced epitope retrieval. Sections to be labelled with Dako anti-vp1 (5D8/1; Dako Cytomation, UK) were heated in 1 mmol/l EDTA, pH 8.0. Primary antibodies were applied for 30 min at room temperature, and a Dako REAL EnVision detection system was used for antigen detection [10].
Statistical analysis
Sample size calculation with a power of 0.8 predicted that a sample size of 69 was required to detect a threefold increase in EV detection sensitivity from a proportion in population 1 (p1)=0.1 to p2=0.3. Statistical analysis was performed using GraphPad Prism (version 8, GraphPad Software, USA). Odds ratios and p values were calculated using two-sided Fisher’s exact test, and 95% confidence intervals were computed using the Baptista–Pike method [37]. A p value <0.05 was considered statistically significant. Power analysis (post hoc and a priori) was performed using G*Power (version 3.1.9.7) [38].
Results
Enhanced detection of EV-RNA in the cellular compartment of peripheral blood
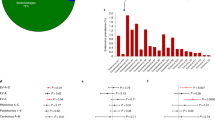
The presence of EV-RNA was evaluated in various peripheral blood fractions in individuals with type 1 diabetes, mAAb-positive individuals and individuals with neither type 1 diabetes nor autoantibody. We first aimed to establish which compartment in peripheral blood provides the highest sensitivity for detection of EV-RNA. We tested plasma and PBMCs isolated by density gradient centrifugation from the same blood drawn on 101 occasions from a total of 79 children in our children cohort. We found superior sensitivity to detect EV-RNA in the cellular compartment (i.e. PBMCs), in which 18 of 101 samples (17.8%) tested positive for EV-RNA, compared with the non-cellular compartment (i.e. plasma) in which 1 of 101 samples (1.0%) tested positive for EV-RNA (OR 21.69; 95% CI 3.64, 229.20; p<0.0001) (Fig. 1a). In the one instance where positivity was seen in the plasma sample, the PBMC sample also tested positive for EV-RNA. To further pinpoint the cellular compartment that harbours EV-RNA, we tested four immune cell subsets in addition to whole PBMCs for the presence of EV-RNA in a cohort of adults with and without type 1 diabetes. These subsets were B cells, monocytes, mDCs and pDCs, all representing antigen-presenting cells (APCs).
Detection of EV-RNA in peripheral blood. In the children cohort, the presence of EV-RNA was assessed in plasma and PBMCs (a) and in PBMCs from specific subgroups (d). In the adult cohort, the presence of EV-RNA was assessed in defined peripheral blood cell subsets (b) and in individuals with and without type 1 diabetes (c). Red shading indicates EV-RNA-positive; white indicates EV-RNA-negative. Differences between groups were statistically significant as indicated: *p<0.05; **p<0.01; ***p<0.001 (Fisher’s exact test, two-sided). T1D, type 1 diabetes
We detected EV-RNA in a higher proportion of individuals when analysing APC subsets combined (26.4%, 19/72) than when analysing whole PBMCs from the same individuals (5.6%, 4/72) (OR 6.09; 95% CI 2.10, 17,17; p=0.0011) (Fig. 1b). Individuals who tested positive for EV-RNA in whole PBMCs also tested positive for EV-RNA in at least one subset of APCs. Two of these individuals tested positive in the monocyte subset, one in the B cell subset, and one in all APC subsets. Among all individuals who tested positive for EV-RNA, EV-RNA was detected in the B cell subset for eight individuals, in the monocyte subset for eight individuals, in the mDC subset for four individuals, in the pDC subset for six individuals, and in whole PBMCs for four individuals. We did not find a difference in the sensitivity for detection of EV-RNA between the different subsets of APCs. Overall, we detected EV-RNA in the cellular compartment (i.e. PBMCs) of 15/79 individuals in the children cohort (19.0%) and 19/72 individuals in the adult cohort (26.4%) (Fig. 1d and b, respectively).
EV-RNA detection correlates with autoimmunity and disease in children but not in adults
Next we investigated whether positivity for EV-RNA in PBMCs correlates with defined stages of type 1 diabetes, i.e. adults with recently diagnosed type 1 diabetes (<3 months) and children positive for mAAb with an ongoing autoimmune reaction.
In the adult cohort, we detected EV-RNA in nine of 37 individuals with type 1 diabetes (24.3%) compared with 10 of 35 individuals without type 1 diabetes (28.6%) (p=0.79) (Fig. 1c). In the children cohort, EV-RNA was detected in PBMCs in 14 of 49 children with mAAb (with or without type 1 diabetes) (28.6%) compared with one of 30 matched control children (without autoantibody or type 1 diabetes) (3.3%) (OR 11.60; 95% CI 1.89, 126.90; p=0.0065) (Fig. 1d).
Increased detection of EV-RNA in peripheral blood but not tissue in individuals carrying the common type 1 diabetes-predisposing allele in IFIH1
We next investigated whether detection of EV infection (by detecting EV-RNA or VP1) in individuals correlates with the predisposing allele (946Thr) of the common variant in IFIH1 (rs1990760, Thr946Ala). The distribution of the common variant in IFIH1 in cohorts, and detection of EV-RNA according to subgroup and genotype, is summarised in Table 1. We found that homozygosity for the protective allele (946Ala) significantly reduced the OR to detect EV-RNA in both the recessive model (homozygous protective vs homozygous risk: OR 0.26; 95% CI 0.087, 0.84; reciprocal of OR 3.81; 95% CI 1.19, 11.46; p=0.031) and the additive protective model (homozygous protective vs homozygous risk and heterozygous: OR 0.33; 95% CI 0.12, 0.98; reciprocal of OR 3.07; 95% CI 1.02, 8.58; p=0.045), when analysing the children and adult cohorts in combination (Table 2). In the adult and children cohorts, respectively, EV-RNA was detected in 34.6% (9/26) and 25.0% (4/16) of individuals who were homozygous for the predisposing allele, 26.5% (9/34) and 21.6% (8/37) of individuals who were heterozygous, and 8.3% (1/12) and 11.5% (3/26) of individuals who were homozygous for the protective allele of the common variant in IFIH1 (Table 1).
We then explored whether the correlation between the protective allele (946Ala) and reduced detection of EV infection in the cellular compartment of peripheral blood also extends to pancreatic islets studied in situ. To this end, we assessed the presence of the EV capsid subunit viral protein 1 (VP1) in pancreatic tissue sections recovered from 43 donors with type 1 diabetes and held within the Exeter Archival Diabetes Biobank (data reported previously by Richardson et al [10]). VP1 was detected in the pancreatic islets of 72.1% of the donors (31/43). Detection of VP1 did not correlate with predisposing allele (946Thr) of the common variant (rs1990760, Thr946Ala) in IFIH1. VP1 was detected in the pancreatic islets of 70.0% (7/10), 76.2% (16/21) and 66.7% (8/12) of donors with the homozygous risk variant, those who were heterozygous, and those with the homozygous protective common variant (rs1990760, Thr946Ala) in IFIH1, respectively (Fig. 2).
Detection of EV capsid protein VP1 in pancreatic islet sections. EV capsid protein VP1 was detected by immunohistochemistry in tissues from 43 donors with type 1 diabetes, with the defined variant in IFIH1 (rs1990760, Thr946Ala). Red shading indicates EV-RNA-positive; white indicates EV-RNA-negative. Differences between groups were not statistically significant (Fisher’s exact test, two-sided)
Discussion
Our data from the children cohort show a significantly increased sensitivity for detection of EV-RNA within the cellular compartment of peripheral blood compared with plasma. Additionally, using the adult cohort, we found that EV infection was detected in more individuals when APC subsets (B cells, monocytes, mDCs and pDCs) were analysed for EV-RNA, compared with whole PBMCs. These observations had statistical power (post hoc) of >0.9. Hence, our data indicate that APCs are ‘carriers’ of EV-RNA in peripheral blood as every individual that tested positive for EV-RNA in the PBMC sample also tested positive for EV-RNA in at least one subset of APCs. Similar observations, that EV-RNA is found more frequently in PBMCs than serum, have been made previously, albeit in a smaller cohort [14]. We postulate that APCs are carriers of EV-RNA because they pick up enterovirus in infected tissues or because these cells are sites of active viral replication, as suggested previously [39, 40]. EV infection in APCs may markedly modulate their function and efficacy of viral and autoantigen presentation. Infected APCs may also serve as a carrier to transport virus to uninfected tissues.
Our analysis shows that positivity for EV-RNA is associated with islet autoimmunity. Children positive for mAAb were more likely to test positive for EV-RNA than those without mAAb (post hoc statistical power 0.85). In children positive for mAAb, we found a similar frequency of EV-RNA positivity among children who later progressed to type 1 diabetes and those who have not yet progressed. In the adult cohort, we did not detect a correlation (p=0.79) between positivity for EV-RNA and type 1 diabetes, potentially due to the increased sensitivity of detection of EV-RNA in PBMC subsets. Overall, our findings are in line with the results of previous studies summarised in meta-analyses by Yeung et al and Wang et al [6, 7], the majority of which reported increased detection of EV infection in individuals with autoimmunity and/or type 1 diabetes compared to those without.
Given the ‘snapshot’ nature of this and previous studies [14, 41] and the fact that EV viraemia lasts for only up to two weeks in peripheral blood [13], we suggest that larger study cohorts, longitudinal sampling, and improved sensitivity of viral detection (as shown here) are likely to be needed to reveal significant differences. This may be achieved using cohort studies such as the Finnish Type 1 Diabetes Prediction and Prevention study, which regularly sample children longitudinally. It is also probable that genetic variation, rather than disease stage, defines the effectiveness of the antiviral response, the rate of viral clearance and the level and spread of any EV infection, and therefore influences the detection of EV-RNA.
To obtain a larger cohort, we combined our two cohorts, and found that individuals carrying the predisposing allele (946Thr) of the common variant in IFIH1 (rs1990760, Thr946Ala) were more likely to test positive for EV-RNA than those without the predisposing allele (in both the additive and protective recessive models). However, our results are based on a limited sample size and low statistical power (post hoc) (0.62 and 0.54 for the recessive and additive protective models, respectively). In the few studies reported so far, no correlation was found between IFIH1 (rs1990760, Thr946Ala) homozygous genotypes and EV-RNA detection in peripheral blood [14, 29] or faecal samples [8]. Our results, and the proposed methodology for improved EV-RNA detection, suggests that further studies, with an increased sample size (power of 0.8 predicted at n=68 per homozygous group, based on our reported proportions of EV-RNA detection per group) should allow definition of the relationship between the IFIH1 Thr946Ala genotype and EV infection detected in peripheral blood.
A potential limitation of our study is that symptoms observed in individuals with recent-onset type 1 diabetes (particularly children) may overlap with those of a virus infection. This overlap in symptoms may introduce a sampling bias between study groups (i.e. individuals without type 1 diabetes and individuals with recent-onset type 1 diabetes) if symptoms observed in individuals with recent-onset type 1 diabetes are misinterpreted as the exclusion criterion, or vice versa. However, we did not observe a sampling bias with regard to the exclusion criterion ‘virus-like’ illness in the children cohort. In the adult cohort, none of the sampled participants exhibited symptoms of ‘virus-like’ illness at the time of recruitment and sampling. However, we do not have data on individuals that were not recruited to the study due to meeting the exclusion criterion. Therefore, we cannot state whether such a sampling bias occurred in the adult cohort. Thus, while we think it unlikely that a sampling bias occurred between the study groups of individuals without type 1 diabetes and with recent-onset type 1 diabetes in the adult cohort, we cannot exclude this.
We then further tested whether our finding of an association between the predisposing allele of the common variant and increased EV-RNA detection in peripheral blood extends to pancreatic tissue of post-mortem donors with type 1 diabetes. As previously reported, we detected the EV capsid protein VP1 in pancreatic islets in the majority (>70%) of donors with type 1 diabetes [10, 12]. Here we report that we did not detect a correlation between the predisposing allele (946Thr) of the common variant in IFIH1 (rs1990760, Thr946Ala) and the presence of EV infection (i.e. positivity for VP1) in pancreatic islets. This probably reflects the fact that most individuals with type 1 diabetes display signs of EV infection in pancreatic islets, and that the effect of the variant in IFIH1 may be more nuanced than simply the presence or absence of VP1 positivity in islets.
Our finding of a significantly increased prevalence of EV-RNA in children positive for mAAb, regardless of their IFIH1 genotype, suggests that a dysregulated immune response and ongoing autoimmunity may interfere with the control and/or clearance of EV infection. We cannot exclude the possibility that infection of pancreatic islet cells by enterovirus is influenced by genetic predisposition. Our analysis focused solely on detection of immunopositivity for the capsid protein VP1. Future analysis of the level of expression of VP1 within islet cells and/or the frequency of VP1-positive cells within pancreatic islets may provide further insights into the effects of genetic predisposition to type 1 diabetes by the common variant in IFIH1 (rs1990760, Thr946Ala).
In summary, our results indicate a correlation between the ability to detect EV-RNA in the cellular compartment of peripheral blood and the presence of the predisposing allele (946Thr) of the common variant (rs1990760, Thr946Ala) in the type 1 diabetes risk gene IFIH1. We also show that enterovirus is detected more often in children with islet autoimmunity compared to those without. Our data further support the view that analysis of APCs increases the sensitivity for detection of EV infection in peripheral blood, and that EV infection is part of the aetiology of type 1 diabetes. Ongoing studies for development of vaccines against Coxsackievirus strains to prevent type 1 diabetes will also be informative [42], and may require consideration of genotype/phenotype information for stratification of participants in trials.
Data availability
The datasets generated during and/or analysed during the current study are available from the corresponding author on reasonable request.
Abbreviations
- APC:
-
Antigen-presenting cell
- EV:
-
Enterovirus
- mAAb:
-
Multiple autoantibodie(s)
- MDA5:
-
Melanoma differentiation-associated protein 5
- mDC:
-
Myeloid dendritic cell
- PBMC:
-
Peripheral blood mononuclear cell
- pDC:
-
Plasmacytoid dendritic cell
- VP1:
-
Viral protein 1
References
Concannon P, Rich SS, Nepom GT (2009) Genetics of type 1A diabetes. N Engl J Med 360(16):1646–1654. https://doi.org/10.1056/NEJMra0808284
Redondo MJ, Jeffrey J, Fain PR, Eisenbarth GS, Orban T (2008) Concordance for islet autoimmunity among monozygotic twins. N Engl J Med 359(26):2849–2850. https://doi.org/10.1056/NEJMc0805398
Patterson CC, Dahlquist GG, Gyurus E, Green A, Soltesz G (2009) Incidence trends for childhood type 1 diabetes in Europe during 1989-2003 and predicted new cases 2005-20: a multicentre prospective registration study. Lancet 373(9680):2027–2033. https://doi.org/10.1016/S0140-6736(09)60568-7
Hermann R, Knip M, Veijola R et al (2003) Temporal changes in the frequencies of HLA genotypes in patients with Type 1 diabetes--indication of an increased environmental pressure? Diabetologia 46(3):420–425. https://doi.org/10.1007/s00125-003-1045-4
Hyoty H (2004) Environmental causes: viral causes. Endocrinol Metab Clin North Am 33(1):27–44. https://doi.org/10.1016/j.ecl.2003.12.004
Yeung WC, Rawlinson WD, Craig ME (2011) Enterovirus infection and type 1 diabetes mellitus: systematic review and meta-analysis of observational molecular studies. BMJ 342:d35. https://doi.org/10.1136/bmj.d35
Wang K, Ye F, Chen Y et al (2021) Association between enterovirus infection and type 1 diabetes risk: a meta-analysis of 38 case-control studies. Front Endocrinol 12:706964. https://doi.org/10.3389/fendo.2021.706964
Witso E, Tapia G, Cinek O, Pociot FM, Stene LC, Ronningen KS (2011) Polymorphisms in the innate immune IFIH1 gene, frequency of enterovirus in monthly fecal samples during infancy, and islet autoimmunity. PLoS ONE 6(11):e27781. https://doi.org/10.1371/journal.pone.0027781
Vehik K, Lynch KF, Wong MC et al (2019) Prospective virome analyses in young children at increased genetic risk for type 1 diabetes. Nat Med 25(12):1865–1872. https://doi.org/10.1038/s41591-019-0667-0
Richardson SJ, Willcox A, Bone AJ, Foulis AK, Morgan NG (2009) The prevalence of enteroviral capsid protein vp1 immunostaining in pancreatic islets in human type 1 diabetes. Diabetologia 52(6):1143–1151. https://doi.org/10.1007/s00125-009-1276-0
Krogvold L, Edwin B, Buanes T et al (2015) Detection of a low-grade enteroviral infection in the islets of langerhans of living patients newly diagnosed with type 1 diabetes. Diabetes 64(5):1682–1687. https://doi.org/10.2337/db14-1370
Richardson SJ, Leete P, Bone AJ, Foulis AK, Morgan NG (2013) Expression of the enteroviral capsid protein VP1 in the islet cells of patients with type 1 diabetes is associated with induction of protein kinase R and downregulation of Mcl-1. Diabetologia 56(1):185–193. https://doi.org/10.1007/s00125-012-2745-4
Oikarinen S, Martiskainen M, Tauriainen S et al (2010) Enterovirus RNA in blood is linked to the development of type 1 diabetes. Diabetes 60(1):276–279. https://doi.org/10.2337/db10-0186
Schulte BM, Bakkers J, Lanke KH et al (2010) Detection of enterovirus RNA in peripheral blood mononuclear cells of type 1 diabetic patients beyond the stage of acute infection. Viral Immunol 23(1):99–104. https://doi.org/10.1089/vim.2009.0072
Sioofy-Khojine AB, Lehtonen J, Nurminen N et al (2018) Coxsackievirus B1 infections are associated with the initiation of insulin-driven autoimmunity that progresses to type 1 diabetes. Diabetologia 61(5):1193–1202. https://doi.org/10.1007/s00125-018-4561-y
Yin H, Berg AK, Tuvemo T, Frisk G (2002) Enterovirus RNA is found in peripheral blood mononuclear cells in a majority of type 1 diabetic children at onset. Diabetes 51(6):1964–1971. https://doi.org/10.2337/diabetes.51.6.1964
Prather SL, Dagan R, Jenista JA, Menegus MA (1984) The isolation of enteroviruses from blood: a comparison of four processing methods. J Med Virol 14(3):221–227. https://doi.org/10.1002/jmv.1890140305
Smyth DJ, Cooper JD, Bailey R et al (2006) A genome-wide association study of nonsynonymous SNPs identifies a type 1 diabetes locus in the interferon-induced helicase (IFIH1) region. Nat Genet 38(6):617–619. https://doi.org/10.1038/ng1800
Nejentsev S, Walker N, Riches D, Egholm M, Todd JA (2009) Rare variants of IFIH1, a gene implicated in antiviral responses, protect against type 1 diabetes. Science 324(5925):387–389. https://doi.org/10.1126/science.1167728
Kato H, Takeuchi O, Sato S et al (2006) Differential roles of MDA5 and RIG-I helicases in the recognition of RNA viruses. Nature 441(7089):101–105. https://doi.org/10.1038/nature04734
Pichlmair A, Schulz O, Tan CP et al (2009) Activation of MDA5 requires higher order RNA structures generated during virus infection. J Virol 83(20):10761–10769. https://doi.org/10.1128/JVI.00770-09
Yoneyama M, Fujita T (2009) RNA recognition and signal transduction by RIG-I-like receptors. Immunol Rev 227(1):54–65. https://doi.org/10.1111/j.1600-065X.2008.00727.x
Shigemoto T, Kageyama M, Hirai R, Zheng J, Yoneyama M, Fujita T (2009) Identification of loss of function mutations in human genes encoding RIG-I and mda5: Implications for resistance to type I diabetes. J Biol Chem 284(20):13348–13354. https://doi.org/10.1074/jbc.M809449200
Liu S, Wang H, Jin Y et al (2009) IFIH1 polymorphisms are significantly associated with type 1 diabetes and IFIH1 gene expression in peripheral blood mononuclear cells. Hum Mol Genet 18(2):358–365. https://doi.org/10.1093/hmg/ddn342
Downes K, Pekalski M, Angus KL et al (2010) Reduced expression of IFIH1 is protective for type 1 diabetes. PLoS One 5(9):e12646. https://doi.org/10.1371/journal.pone.0012646
Gorman JA, Hundhausen C, Errett JS et al (2017) The A946T variant of the RNA sensor IFIH1 mediates an interferon program that limits viral infection but increases the risk for autoimmunity. Nat Immunol 18(7):744–752. https://doi.org/10.1038/ni.3766
Zouk H, Marchand L, Li Q, Polychronakos C (2014) Functional characterization of the Thr946Ala SNP at the type 1 diabetes IFIH1 locus. Autoimmunity 47(1):40–45. https://doi.org/10.3109/08916934.2013.832758
Domsgen E, Lind K, Kong L et al (2016) An IFIH1 gene polymorphism associated with risk for autoimmunity regulates canonical antiviral defence pathways in Coxsackievirus infected human pancreatic islets. Sci Rep 6(1):39378. https://doi.org/10.1038/srep39378
Cinek O, Tapia G, Witso E et al (2012) Enterovirus RNA in peripheral blood may be associated with the variants of rs1990760, a common type 1 diabetes associated polymorphism in IFIH1. PLoS One 7(11):e48409. https://doi.org/10.1371/journal.pone.0048409
Alberti KG, Zimmet PZ (1998) Definition, diagnosis and classification of diabetes mellitus and its complications. Part 1: diagnosis and classification of diabetes mellitus provisional report of a WHO consultation. Diabet Med 15(7):539–553. https://doi.org/10.1002/(SICI)1096-9136(199807)15:7<539::AID-DIA668>3.0.CO;2-S
Nanto-Salonen K, Kupila A, Simell S et al (2008) Nasal insulin to prevent type 1 diabetes in children with HLA genotypes and autoantibodies conferring increased risk of disease: a double-blind, randomised controlled trial. Lancet 372(9651):1746–1755. https://doi.org/10.1016/S0140-6736(08)61309-4
Lonnrot M, Knip M, Marciulionyte D et al (1999) Enterovirus antibodies in relation to islet cell antibodies in two populations with high and low incidence of type 1 diabetes. Diabetes Care 22(12):2086–2088. https://doi.org/10.2337/diacare.22.12.2086
Lonnrot M, Sjoroos M, Salminen K, Maaronen M, Hyypia T, Hyoty H (1999) Diagnosis of enterovirus and rhinovirus infections by RT-PCR and time-resolved fluorometry with lanthanide chelate labeled probes. J Med Virol 59(3):378–384. https://doi.org/10.1002/(SICI)1096-9071(199911)59:3<378::AID-JMV19>3.0.CO;2-I
Olerup O, Zetterquist H (1992) HLA-DR typing by PCR amplification with sequence-specific primers (PCR-SSP) in 2 hours: an alternative to serological DR typing in clinical practice including donor-recipient matching in cadaveric transplantation. Tissue Antigens 39(5):225–235. https://doi.org/10.1111/j.1399-0039.1992.tb01940.x
Lempainen J, Laine AP, Hammais A et al (2015) Non-HLA gene effects on the disease process of type 1 diabetes: from HLA susceptibility to overt disease. J Autoimmun 61:45–53. https://doi.org/10.1016/j.jaut.2015.05.005
Foulis AK, Liddle CN, Farquharson MA, Richmond JA, Weir RS (1986) The histopathology of the pancreas in type 1 (insulin-dependent) diabetes mellitus: a 25-year review of deaths in patients under 20 years of age in the United Kingdom. Diabetologia 29(5):267–274. https://doi.org/10.1007/BF00452061
Baptista J, Pike MC (1977) Algorithm AS 115: exact two-sided confidence limits for the odds ratio in a 2 × 2 table. Journal of the Royal Statistical Society Series C (Applied Statistics) 26(2):214–220. https://doi.org/10.2307/2347041
Faul F, Erdfelder E, Lang AG, Buchner A (2007) G*Power 3: a flexible statistical power analysis program for the social, behavioral, and biomedical sciences. Behav Res Methods 39(2):175–191. https://doi.org/10.3758/bf03193146
Vuorinen T, Vainionp R, Heino J, Hyypi T (1999) Enterovirus receptors and virus replication in human leukocytes. J Gen Virol 80(Pt 4):921–927. https://doi.org/10.1099/0022-1317-80-4-921
Vuorinen T, Vainionpaa R, Kettinen H, Hyypia T (1994) Coxsackievirus B3 infection in human leukocytes and lymphoid cell lines. Blood 84(3):823–829. https://doi.org/10.1182/blood.V84.3.823.823
Chistiakov DA, Voronova NV, Savost'Anov KV, Turakulov RI (2010) Loss-of-function mutations E6 27X and I923V of IFIH1 are associated with lower poly(I:C)-induced interferon-beta production in peripheral blood mononuclear cells of type 1 diabetes patients. Hum Immunol 71(11):1128–1134. https://doi.org/10.1016/j.humimm.2010.08.005
Hyoty H, Leon F, Knip M (2018) Developing a vaccine for type 1 diabetes by targeting coxsackievirus B. Expert Rev Vaccines 17(12):1071–1083. https://doi.org/10.1080/14760584.2018.1548281
Acknowledgements
We are grateful to all participants in the study and the clinical team involved in sample collection. We thank J. Allen (King’s College London) for conducting experiments for the adult cohort.
Authors’ relationships and activities
JAT is a member of the GlaxoSmithKline Human Genetics Advisory Board. All other authors declare that there are no relationships or activities that might bias, or be perceived to bias, this work.
Contribution statement
All authors made substantial contributions to the conception and design of the work and/or the acquisition, analysis or interpretation of the data, and drafted the article or revised it critically for important intellectual content. All authors approved the final submitted version and are accountable for their own contributions. As the corresponding author, ME accepts responsibility for the integrity of the data and work.
Funding
The study was supported by a centre grant Diabetes – Genes, Autoimmunity and Prevention (D-GAP) from the JDRF (1-2007-1803) to MP, JAT, P. Bingley (University of Bristol), T. Tree (King’s College London) and D. Dunger (University of Cambridge), a JDRF Career Development Award (5-CDA-2014-221-A-N) to SJR, a JDRF research grant awarded to the network of Pancreatic Organ Donors – Virus (nPOD-V) consortium (JDRF 25-2012-516), a Medical Research Council project grant (MR/P010695/1) awarded to SJR and NGM, a strategic award to JAT from the JDRF (4-SRA-2017-473-A-N) and the Wellcome Trust (07212/A/15/Z) and by the European Commission (Persistent Virus Infection in Diabetes Network [PEVNET] Frame Programme 7, coordinated by HH, contract number 261441). The virus analyses in Tampere University were additionally funded by grants to HH from the Sigrid Juselius Foundation and the Academy of Finland (grant number 288671). This study also reports independent research supported by the National Institute of Health Research Exeter Clinical Research Facility. Funders were not involved in the design and conduct of the study, the collection, analysis and interpretation of the data, or the preparation, review or approval of the manuscript. As such, the content is solely the responsibility of the authors and does not necessarily represent the official views of the funders. For the purpose of open access, the author has applied a Creative Commons Attribution (CC BY) licence to any Author Accepted Manuscript version arising.
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Sioofy-Khojine, AB., Richardson, S.J., Locke, J.M. et al. Detection of enterovirus RNA in peripheral blood mononuclear cells correlates with the presence of the predisposing allele of the type 1 diabetes risk gene IFIH1 and with disease stage. Diabetologia 65, 1701–1709 (2022). https://doi.org/10.1007/s00125-022-05753-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00125-022-05753-y