Abstract
Aims/hypothesis
The incidence of Type 1 diabetes has increased 2.5 times during the time period from 1966 to 2000 in Finland—a general trend seen in almost all developed countries that can only be explained by environmental factors. The aim of this study was to test the possible effect of a changing environment on distribution of genotypes associated with disease susceptibility.
Methods
HLA DRB1-DQA1-DQB1 genes and two diabetes-associated polymorphisms at IDDM2 and IDDM12 were analyzed. The frequencies of genotypes were compared between cases diagnosed with childhood-onset Type 1 diabetes during the period of 1939–1965 (n=367) and those diagnosed between 1990 and 2001 (n=736). Chi-square statistics or the Fisher's Exact test were used for the comparison of frequencies of analyzed haplotypes and genotypes in the two groups.
Results
The frequencies of (DR3)-DQA1*05-DQB1*02 and (DR4)-DQB1*0302 risk haplotypes and the high risk (DR3)-DQA1*05-DQB1*02/DRB1*0401-DQB1*0302 genotype were higher while proportion of patients carrying protective haplotypes—(DR15)-DQB1*0602 and (DR1301)-DQB1*0603—or protective genotypes was lower in patients diagnosed before 1965 as compared to those who developed disease after 1990. No temporal variation was found in the frequencies of genotypes at IDDM2 and IDDM12.
Conclusion/interpretation
Our data suggest that the need for genetic susceptibility to develop Type 1 diabetes has decreased over time due to an increasing environmental pressure and this results in a higher disease progression rate especially in subjects with protective HLA genotypes.
Similar content being viewed by others
Development of Type 1 diabetes is the result of interactions between genetic susceptibility and environmental factors. Genes in the Human Leukocyte Antigen region (HLA; 6p21) are the major genetic determinants of Type 1 diabetes (T1D) susceptibility and resistance. HLA molecules play a key role in the initiation and regulation of immune processes that are induced as a response to contacts with foreign agents such as microbes or alimentary proteins. HLA molecules have an enormous diversity that enables individuals to develop specific responses against a multitude of agents. The population distribution of HLA alleles and haplotypes is subjected to selection determined by the surrounding environment. This phenomenon is reflected in variations in the background frequency of the HLA B8-DR3-DQ2 haplotype that is inversely related to the time length of wheat cultivation. This haplotype, being the major genetic factor predisposing to celiac disease, seems to be less frequent in populations, which have lived on wheat for a longer period of time indicating a negative selection [1]. Population distribution of HLA polymorphisms is also strongly influenced by infectious diseases. For example, the HLA-B53 allele and the DRB1*1302-DQB1*0501 haplotype, that are protective against severe malaria are common in West-Africa [2]. Similarly, other severe infectious diseases also show HLA association like typhoid fever where HLA DRB1*04 and DQB1*0401/02 alleles are associated with protection and DRB1*0301 and DQB1*02 alleles are associated with susceptibility to disease [3]. Distribution pattern of certain HLA haplotypes is influenced by the most prevalent infectious agents occurring in that population, and certain polymorphisms are maintained through balancing selections due to various diseases. Moreover, some evidence now exists to support an advantage conferred by HLA heterozygosity, which has been associated with protection against several viral diseases including resistance to hepatitis B carriage [4].
The incidence of T1D has continuously increased world-wide in the last decades, which can only be explained by environmental factors. The incidence rate has repeatedly been observed to be record high in Finland, amounting to 20 cases per year per 100 000 children in 1966, whereas it reached a figure close to 50 by the end of the 90's [5, 6]. During that period of time, substantial changes have occurred in eating habits, food constituents, vaccinations, general health care and personal hygiene that might have contributed to a rapidly increasing environmental pressure on individuals genetically susceptible to T1D. It is possible that the increased environmental pressure is reflected in the distribution of HLA genotypes among patients diagnosed during different time periods. There are very limited data on temporal changes in HLA genotype frequencies in patients with T1D [7, 8]. To further explore this issue we analyzed and compared frequencies of genotypes at HLA DR-DQ and two disease associated polymorphisms at IDDM2 and IDDM12 in patients who developed childhood onset T1D during different time periods.
Subjects and methods
Subjects
367 patients (200 boys and 167 girls) with T1D diagnosed between 1939 and 1965 (Patient Series 1-PS1), and 736 patients (376 boys and 360 girls) diagnosed between 1990 and 2001 (Patient Series 2-PS2) were studied.
Type 1 diabetes cases for PS1 participated in the ongoing Finnish Diabetic Nephropathy Study (FinnDiane), a nationwide comprehensive multi-centre study, with the aim to study genetic and clinical risk factors for the development of diabetic nephropathy and other diabetic complications. All adult Type 1 diabetic patients from a total of 91 referral centres throughout the country were invited to participate in FinnDiane. So far, 3200 patients have been recruited and 367 of them who were diagnosed between 1939 and 1965 were analyzed in the current study.
In PS2 all consecutive incident Type 1 diabetes cases who were diagnosed between 1990 and 2001 in three regions (Oulu, Turku and Helsinki areas) of Finland were included. The two patient series were ethnically and geographically matched. More than 99% of patients in both groups were of Finnish ethnic origin.
During the second half of the 20th century the average annual immigration rate to Finland was 9751 while the mean total population of Finland was about 4,700,000 in this period. Therefore, Finland has a very low (less than 2%) percentage of foreigners (data from Institute of Migration, Turku, Finland). The probability of annual internal migration between provinces within Finland was constantly at a low level (14.4–21.0‰) between the 1960's and 90's. These data confirm that neither population admixture from other ethnic groups nor internal migration could be a confounding factor that affects gene frequencies in the two groups. The geographical distribution of patients in PS1 and PS2 were as follows: Oulu region (Northern Finland) 99 (27.0%) and 223 (30.3%), Helsinki region (Southern Finland) 128 (34.8%) and 272 (37.0%), Turku region (South-Western Finland) 140 (38.1%) and 241 (32.7%). All patients in both groups were diagnosed with T1D below the age of 15 years and had classic symptoms of T1D such as hyperglycaemia, ketosis and dehydration.
The mean age of patients in PS1 and PS2 were 8.2±2.1 and 8.5±4.0 years, respectively. For some comparisons, both PS1 and PS2 were split into three groups according to age at diagnosis: 0 to 4.99 yrs (Group 1-G1), 5.0 to 9.99 yrs (Group 2-G2) and 10.0 to 14.99 yrs (Group 3-G3). An informed consent was obtained from all study subjects. The investigations have been carried out in accordance with the principles of the Declaration of Helsinki.
Genotyping
A low-resolution full house typing for the common Caucasian HLA DR-DQ haplotypes was carried out using a lanthanide labelled oligonucleotide hybridization method. Initial typing of DQB1 alleles was completed by DQA1 typing and DR4 subtyping in selected genotypes where informative [9, 10]. The MspI −2221 C/T polymorphism at the insulin gene region (IDDM2, 11p15.5) and the +49 A/G polymorphism at the CTLA4 gene (IDDM12, 2q33) were genotyped in all patients in PS1 and in 706 patients in PS2 using the same methodological approach.
Statistical analysis
Chi-square statistics or the Fisher's Exact test were used for the comparison of frequencies of analyzed haplotypes and genotypes. A p value of 0.05 or less was considered to be statistically significant. The size of patient series enabled us an 80% power to detect a 7.5% difference at an 0.05 significance level.
Results
The population frequencies of various HLA DRB1-DQA1-DQB1 haplotypes in the two time series are shown in Table 1. More patients carried the (DR3)-DQA1*05-DQB1*02 and (DR4)-DQB1*0302 risk haplotypes in PS1 than in PS2 (p=0.015 and p=0.04, respectively). Among the DRB1*04 subtypes associated with DQB1*0302, DRB1*0401 was less prevalent in PS2, but DRB1*0403 was increased in PS2 as compared to PS1, although these differences were not significant. Combined haplotype frequency of the four risk haplotypes—(DR3)-DQA1*05-DQB1*02, DRB1*0401/02/04–DQB1*0302—was higher in PS1 than in PS2 (68.5 vs. 62.3%, p=4.6×10−3). Protective haplotypes—(DR15)-DQB1*0602 and (DR1301)-DQB1*0603—were less prevalent in PS1 than in PS2 (p=3.2×10−3 and p=0.025, respectively). However, the protective (DR7)-DQA1*02-DQB1*0303 haplotype was decreased in PS2 as compared to PS1 (p=0.04). Combined frequencies of protective haplotypes(DR5)-DQA1*05-DQB1*0301, DRB1*0403-DQB1*0302, (DR7)-DQA1*02-DQB1*0303, (DR14)-DQB1*0503, (DR15)-DQB1*0602 and (DR1301)-DQB1*0603—showed a highly significant difference between PS1 and PS2 (5.2 vs. 9.7%, p=3.5×10−4). Similarly, differences in risk and protective haplotype frequencies observed between PS1 and PS2 were seen also when corresponding age at onset groups were compared, in addition to several age-dependent trends. Risk haplotypes—(DR3)-DQA1*05-DQB1*02 and DRB1*0401-DQB1*0302—tended to be most prevalent in the youngest (G1) age group as compared to G3 in both patient series (data not shown). The combined frequency of protective haplotypes—(DR5)-DQA1*05-DQB1*0301, DRB1*0403-DQB1*0302, (DR1)-DQB1*0501, (DR14)-DQB1*0503, (DR15)-DQB1*0602 and (DR1301)-DQB1*0603—was lower in the youngest age group in both time series (PS1 G1 9.7% vs. G2 17.5%, p=0.02; PS2 G1 13.6% vs. G3 21.1%, p=0.02). For analysis of genotype distributions the HLA DRB1-DQA1-DQB1 genotypes were grouped according to the diabetes risk they conferred in the Finnish population. Risk calculations were based on a comparison of diabetic children to affected family-based artificial controls in 622 nuclear families. The (DR3)-DQA1*05-DQB1*02/DRB1*0401-DQB1*0302 genotype carried the highest risk of diabetes with an odds ratio of more than 50 (high risk genotype). The moderate risk genotype group conferred odds ratios between 5.1–18.0 and comprised those homozygous for DR3-DQ2, HLA DQB1*0302, and genotypes where DRB1*0401-DQB1*0302 was associated with neutral haplotypes (neutral haplotypes (N): DQA1*0201-DQB1*02, DQA1*03-DQB1*0301, DQA1*03-DQB1*0303, DQB1*04, DQB1*0501, DQB1*0502, DQB1*0604).
Low risk genotypes comprised DRB1*0404-DQB1*0302/N, (DR3)-DQA1*05-DQB1*02/N and N/N combinations that conferred odds ratios between 1.6 to 2.4. Protective genotypes were homozygous or heterozygous for the protective haplotypes (DQA1*05-DQB1*0301, DRB1*0403-DQB1*0302, DQA1*0201-DQB1*0303, DQB1*0503, DQB1*0602, DQB1*0603) and conferred odds ratios below 1.
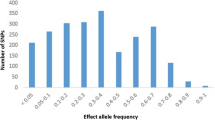
The high risk (DR3)-DQA1*05-DQB1*02/DRB1*0401-DQB1*0302 genotype was more common in PS1 than in PS2 (p=7.3×10−3) (Fig. 1). The difference in the frequency of the (DR3)-DQA1*05-DQB1*02/DRB1*0401-DQB1*0302 high risk genotype was even higher between cases who were diagnosed before 1960 (a subset of PS1) and patients diagnosed after 1990 (PS2) (28.8 vs. 18.2%; p=3.6×10−3). This high risk genotype was more prevalent in the younger age groups in both patient series, although the difference remained non-significant (Table 2). Frequencies of moderate and low risk HLA genotypes did not differ between the two time series; however, protective genotypes were less prevalent in PS1 than in PS2 (p=4.3×10–4) (Fig. 1). This difference was detected also between the corresponding age at onset groups (PS1 G1 vs. PS2 G1 p=0.04; PS1 G3 vs. PS2 G3 p=3×10−3) (Table 2.). Protective genotypes were less prevalent in the youngest group in both time series (PS1: G1 vs. G2, p=0.03 ; G1 vs. G3, p=0.03 and PS2 G1 vs. G3 p=1×10−4 and G1 vs. G2, p=0.03) (Table 2).
The distribution of alleles and genotypes at MspI −2221 C/T polymorphism at the insulin gene region (11p15.5) did not show a difference between the two patient series; TT/CT/CC genotype frequencies in the two groups were as follows; PS1: 0.8%, 16.9%, 82.3% vs. PS2: 0.7%, 15.7%, 83.6%, respectively. Similarly, no difference was seen between the two patient groups when allele and genotype frequencies at the +49 A/G polymorphism at the CTLA4 gene (2q33) were compared (AA/AG/GG genotype frequencies: PS1 19.1%, 52.0%, 28.9% vs. PS2 17.4%, 56.0, 26.6%, respectively).
Discussion
In this study, we documented that the distribution of HLA genotypes shows marked differences between patients with childhood onset T1D diagnosed about 50 years apart.
A critical point in this study was the possibility of a selection bias in collection of patients who were diagnosed with childhood onset T1D about 50 years ago (Patient Series 1). During World War 2 diabetes mortality was high all over Europe, however only 0.8% of our cases were diagnosed before 1946 and most of the cases in this patient group (94%) were diagnosed after 1954 which in fact excludes any major socio-economic confounders related to that period.
In addition, a considerable proportion of diabetes cases diagnosed before 1965 have died due to end stage renal disease or macrovascular complications. However, there is no evidence, that HLA genotype affects mortality from late diabetic complications, therefore we believe that our study was not confounded by population selection bias [11, 12].
We found that the proportion of patients with high-risk genotypes has decreased from 25.3 to 18.2% while the fraction of children carrying protective genotypes has increased to the same extent between the two observation periods. The DQB1*0602, *0503, and *0603 alleles were more protective in earlier years in the 20th century, whereas a higher proportion of individuals carrying these alleles developed diabetes during the more recent time period. There was no accumulation of any individual (specific) genotype, which could explain the increase observed in disease incidence. On the contrary, the data are suggesting a more general role for environmental factors. They also imply that dynamic changes have occurred in the natural history of T1D in a relatively short period of time. On this topic there are only a few data published. It has been found that the DR3 haplotype was more prevalent in patients diagnosed in the 1960's than in patients diagnosed in the 1980's, while no differences were observed in relation to the DR4 haplotype [8]. In another study, fluctuations were seen in the frequencies of the DR3 haplotype and the DR3/4 combination without any clear trend [7].
We observed that the HLA haplotypes and genotypes associated with increased diabetes risk were more prevalent while protective haplotypes and genotypes were more rare in the youngest age group that indicates a stronger HLA effect in very young children. These phenomena were equally pronounced in patients diagnosed before 1965 and in those diagnosed after 1990, which confirms a conserved age-dependent role for HLA DQ molecules in a changing environment. Similar age-dependent heterogeneity in frequencies of diabetes associated HLA haplotypes was reported earlier by our group and others [13, 14].
The increase in incidence of T1D diabetes is a general trend seen in almost all developed countries. In this respect, it is interesting to note that when comparing the frequencies of the HLA DR3-DQ2/DR4-DQ8 genotype in patients with T1D in Europe, we observed that this high risk genotype was more prevalent in patients from countries with a low disease incidence suggesting that these populations are at an earlier stage of the natural evolution of diabetes history [15, 16]. The declining protective efficiency of DQB1*0603 is reflected in fact in the behaviour of the DQB1*0302/DQB1*0603 genotype which was increased among patients diagnosed during the later time period and we observed this particular genotype to be associated with T1D in Finland, whereas in several other populations with lower diabetes incidence no such disease association has been detected [17].
The findings of this study indicate that the relative probability to develop disease in subjects with low or medium risk HLA genotypes has considerably increased during the last 30 years. We observed that the absolute risk of individuals with risk genotypes [(DR3)-DQA1*05-DQB1*02/v or DR4-DQB1*0302/s where v≠DQB1*0602, *0603, *0301 and s≠DQB1*0602] to develop disease has increased 2.3 times in the 1990' s as compared to the middle of the 20th century while absolute risk of individuals carrying protective genotypes has risen 7.1 times. The HLA high risk DR3-DQ2/DR4-DQ8 genotype had a stronger relative disease predisposing effect (OR=15.1, 95% CI 11.9–19.2) in the middle of the previous century than in the 1990's (OR=9.8, 95% CI: 8.24–11.7). Similarly, protective haplotypes seemed to have a stronger effect in the earlier time series (OR=0.02, 95% CI: 0.01–0.05 in patients diagnosed before 1965 vs. OR=0.07, 95% CI: 0.05–0.09 in patients diagnosed after 1990).
Temporal changes observed in the distribution of HLA genotypes in children with Type 1 diabetes indicate that importance of disease risk HLA genotypes, as being the major genetic disease determinant, in controlling initiation and/or progression of immune mediated beta-cell destruction has been gradually decreasing during the last decades. It is clear that several viruses—like rubella and mumps—that caused diabetes in a fraction of patients have practically been eradicated from Europe. Interestingly, frequency of infections caused by enteroviruses has also been rapidly decreasing in the Finnish population since the early 1980's [18]. Since the role of HLA in determining mechanisms by which viruses induce beta-cell damage has not been defined yet, the relation between temporal patterns of viral infections and our observations remains an enigma [19].
New food constituents and changes in our eating habits have affected the microbial colonization of the gut and have probably influenced development and maturation of mucosal immune system which seems to be important in the development of Type 1 diabetes [20] . Similarly, the role of HLA in controlling self tolerance in the gut associated lymphoid tissue remains to be clarified.
We tested the possibility of a time-dependent pattern in the genotype distribution of two non-HLA disease susceptibility loci, as well. IDDM2 locus that corresponds to the insulin gene region (11p15) and IDDM12-CTLA4 gene region are confirmed T1D susceptibility loci [21, 22, 23, 24, 25]. We studied MspI −2221 C/T polymorphism at IDDM2 and CTLA4 gene +49 A/G polymorphism that showed a disease association in the Finnish population (AP. Laine, unpublished observations). However, no significant temporal variation was found in the frequencies of genotypes at these disease polymorphisms. This might indicate that they do not interact with factors in the surrounding environment or disease associated polymorphisms at these loci are not under selection pressure; however, these results are not conclusive because of the weak contribution of these genes to disease susceptibility.
In conclusion, we have shown that considerable temporal changes have occurred in the distribution of T1D associated HLA genotypes over the last 50 years. This observation implies an increasing environmental pressure resulting in higher penetrance of disease especially in individuals with protective HLA genotypes.
Abbreviations
- T1D:
-
Type 1 diabetes mellitus
- CTLA4 :
-
cytotoxic T lymphocyte-associated protein 4
- HLA:
-
Human Leukocyte Antigen system
- IDDM2 :
-
Type 1 diabetes susceptibility locus No2
- IDDM12 :
-
Type 1 diabetes susceptibility locus No12
- PS1:
-
patient series 1
- PS2:
-
patient series 2
- G1:
-
age group 1
- G2:
-
age group 2
- G3:
-
age group 3
- N:
-
neutral haplotypes
- OR:
-
odds ratio
References
Piazza A, Menozzi P, Cavalli-Sforza LL (1980) The HLA-A,B gene frequencies in the world: migration or selection? Hum Immunol 1:297–304
Hill AV, Allsopp CE, Kwiatkowski D et al. (1991) Common west African HLA antigens are associated with protection from severe malaria. Nature 352:595–600
Dunstan SJ, Stephens HA, Blackwell JM et al. (2001) Genes of the class II and class III major histocompatibility complex are associated with typhoid fever in Vietnam. J Infect Dis 183:261–268
Thursz MR, Thomas HC, Greenwood BM, Hill AV (1997) Heterozygote advantage for HLA class-II type in hepatitis B virus infection. Nat Genet 17:11–12
Karvonen M, Viik-Kajander M, Moltchanova E, Libman I, LaPorte R, Tuomilehto J (2000) Incidence of childhood type 1 diabetes worldwide. Diabetes Mondiale (DiaMond) Project Group. Diabetes Care 23:1516–1526
Tuomilehto J, Rewers M, Reunanen A et al. (1991) Increasing trend in Type I (insulin-dependent) diabetes mellitus in childhood in Finland. Analysis of age, calendar time and birth cohort effects during 1965 to 1984. Diabetologia 34:282–287
Mäenpää A, Koskimies S, Scheinin T et al. (1991) Frequencies of HLA-DR3, -DR4, -B8 and -Bw62 in diabetic children diagnosed between 1960 and 1990. Diabetes Res 16:159–163
Kontiainen S, Scheinin T, Schlenzka A, Mäenpää J, Groop L, Koskimies S (1988) Differences in HLA types in children with insulin-dependent diabetes diagnosed in 1960s, 1970s, and 1980s. Lancet ii:219
Nejentsev S, Sjöroos M, Soukka T et al. (1999) Population-based genetic screening for the estimation of Type 1 diabetes mellitus risk in Finland: selective genotyping of markers in the HLA-DQB1, HLA-DQA1 and HLA-DRB1 loci. Diabet Med 16:985–992
Laaksonen M, Pastinen T, Sjöroos M et al. (2002) HLA class II associated risk and protection against multiple sclerosis—a Finnish family study. J Neuroimmunol 122:140–145
Chowdhury TA, Dyer PH, Mijovic CH, Dunger DB, Barnett AH, Bain SC (1999) Human leucocyte antigen and insulin gene regions and nephropathy in type I diabetes. Diabetologia 42:1017–1020
Ronningen KS, Bangstad HJ, Undlien DE, Thorsby E (1993) Influence of genetic factors (HLA class II genes, insulin-gene region polymorphisms) and metabolic control on the development of diabetic nephropathy. Diabetes Res 23:31–40
Caillat-Zucman S, Garchon HJ, Timsit J et al. (1992) Age-dependent HLA genetic heterogeneity of type 1 insulin-dependent diabetes mellitus. J Clin Invest 90:2242–2250
Komulainen J, Kulmala P, Savola K et al. (1999) Clinical, autoimmune, and genetic characteristics of very young children with type 1 diabetes. Childhood Diabetes in Finland (DiMe) Study Group. Diabetes Care 22:1950–1955
Hermann R, Mijovic CH, Rayner M et al. (2001) HLA alleles and IDDM in children in Hungary: a comparison with Finland. Hum Immunol 62:391–398
Nejentsev S, Koskinen S, Sjöroos M et al. (1998) Distribution of insulin-dependent diabetes mellitus (IDDM)-related HLA alleles correlates with the difference in IDDM incidence in four populations of the Eastern Baltic region. Tissue Antigens 52:473–477
Noble JA, Valdes AM, Cook M, Klitz W, Thomson G, Erlich HA (1996) The role of HLA class II genes in insulin-dependent diabetes mellitus: molecular analysis of 180 Caucasian, multiplex families. Am J Hum Genet 59:1134–1148
Viskari HR, Koskela P, Lönnrot M et al. (2000) Can enterovirus infections explain the increasing incidence of type 1 diabetes? Diabetes Care 23:414–416
Bach JF (2002) The effect of infections on susceptibility to autoimmune and allergic diseases. N Engl J Med 347:911–920
Vaarala O (2002) The gut immune system and type 1 diabetes. Ann NY Acad Sci 958:39–46
Lucassen AM, Julier C, Beressi JP et al. (1993) Susceptibility to insulin dependent diabetes mellitus maps to a 4.1 kb segment of DNA spanning the insulin gene and associated VNTR. Nat Genet 4:305–310
Bell GI, Horita S, Karam JH (1984) A polymorphic locus near the human insulin gene is associated with insulin-dependent diabetes mellitus. Diabetes 33:176–183
Nistico L, Buzzetti R, Pritchard LE et al. (1996) The CTLA-4 gene region of chromosome 2q33 is linked to, and associated with, type 1 diabetes. Belgian Diabetes Registry. Hum Mol Genet 5:1075–1080
Morahan G, Huang D, Tait BD, Colman PG, Harrison LC (1996) Markers on distal chromosome 2q linked to insulin-dependent diabetes mellitus. Science 272:1808–1810
Marron MP, Raffel LJ, Garchon HJ et al. (1997) Insulin-dependent diabetes mellitus (IDDM) is associated with CTLA4 polymorphisms in multiple ethnic groups. Hum Mol Genet 6:1275–1282
Acknowledgements
This study was supported by the JDRF, JDRF/EFSD/Novo Nordisk Focused Research Grant for Type 1 Diabetes, Novo Nordisk Foundation, Turku University Central Hospital and the Academy of Finland. We thank Professor O. Vaarala for the collection of samples in Helsinki and M. Karlson, R. Suominen and E. Nirhamo for their skilful technical assistance.
Author information
Authors and Affiliations
Consortia
Corresponding author
Rights and permissions
About this article
Cite this article
Hermann, R., Knip, M., Veijola, R. et al. Temporal changes in the frequencies of HLA genotypes in patients with Type 1 diabetes—indication of an increased environmental pressure?. Diabetologia 46, 420–425 (2003). https://doi.org/10.1007/s00125-003-1045-4
Received:
Revised:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00125-003-1045-4