Abstract
Integration of genomic and other data has begun to stratify type 2 diabetes in prognostically meaningful ways, but this has yet to impact on mainstream diabetes practice. The subgroup of diabetes caused by single gene defects thus provides the best example to date of the vision of ‘precision diabetes’. Monogenic diabetes may be divided into primary pancreatic beta cell failure, and primary insulin resistance. In both groups, clear examples of genotype-selective responses to therapy have been advanced. The benign trajectory of diabetes due to pathogenic GCK mutations, and the sulfonylurea-hyperresponsiveness conferred by activating KCNJ11 or ABCC8 mutations, or loss-of-function HNF1A or HNF4A mutations, often decisively guide clinical management. In monogenic insulin-resistant diabetes, subcutaneous leptin therapy is beneficial in some severe lipodystrophy. Increasing evidence also supports use of ‘obesity therapies’ in lipodystrophic people even without obesity. In beta cell diabetes the main challenge is now implementation of the precision diabetes vision at scale. In monogenic insulin-resistant diabetes genotype-specific benefits are proven in far fewer patients to date, although further genotype-targeted therapies are being evaluated. The conceptual paradigm established by the insulin-resistant subgroup with ‘adipose failure’ may have a wider influence on precision therapy for common type 2 diabetes, however. For all forms of monogenic diabetes, population-wide genome sequencing is currently forcing reappraisal of the importance assigned to pathogenic mutations when gene sequencing is uncoupled from prior suspicion of monogenic diabetes.
Graphical abstract

Similar content being viewed by others
Avoid common mistakes on your manuscript.

Background
Diabetes mellitus affects around 9.3% of the world population [1]. In the USA, 91% of diabetes is classified as type 2 diabetes, 6% as type 1 diabetes and 3% as ‘other forms’ [2]. Continuing efforts aim to stratify type 1 and type 2 diabetes into subtypes that inform therapy, the sine qua non of precision diabetes care; however, progress to date has not translated into significant changes in mainstream care. In contrast, study of diabetes caused by single gene mutations has transformed treatment for many patients, in a triumph of the precision medicine approach.
Dominantly inherited, early onset diabetes was first reported in the 1970s [3], a time when unusually severe forms of insulin resistance were also attracting scrutiny [4]. The first identification of a genetic basis for severe insulin resistance—mutations in the gene encoding the insulin receptor (INSR)—was in 1988 [5, 6], 3 years after sequencing of the INSR gene. GCK was the first gene established to cause so-called ‘MODY’, using traditional genetic mapping in 1992 [7, 8]. There has since been a rapid pace of discovery of further monogenic forms of diabetes, reinvigorated by application of next generation sequencing (NGS) over the past 12 years. For several subtypes of monogenic diabetes, distinct therapeutic responses have been demonstrated.
Despite these advances, there is still much work to be done to ensure that all patients with monogenic diabetes receive a timely diagnosis and, where appropriate, targeted therapy. The barriers to achieving this are paradoxically opposed. The high prevalence of diabetes means that diabetes service organisation has focused on efficient, high-throughput systems of care, built around algorithms derived from large clinical trials in type 1 and type 2 diabetes. This is at odds with the attention to detail and bespoke diagnostic testing required for monogenic diabetes, and leads to the problem of clinical underdiagnosis in many settings.
On the other hand, the rapid advance of NGS, including its deployment in population-based studies and its commercial provision directly to consumers, has yielded a surfeit of genetic information. This is often acquired without consideration of pretest probability of a monogenic condition. Attenuation of the traditional pathway from clinical assessment to targeted genetic analysis is sometimes compounded by insufficiently stringent algorithms to discriminate disease-causing mutations from multitudes of irrelevant variants. This risks genetic overdiagnosis in other settings [9,10,11]. In fact, ascertainment of cases purely through gene sequencing rather than by clinical assessment and targeted testing reduces penetrance and expressivity even for many established pathogenic variants [12, 13].
These developments pose challenges as well as opportunities for genetic stratification of diabetes, and demand regular re-evaluation of strategies for genetic testing in diabetes clinics. We now appraise the current implementation of precision medicine for monogenic diabetes, building on recent comprehensive treatment of monogenic beta cell diabetes in Diabetologia [14].
Monogenic pancreatic beta cell defects
Failure of pancreatic beta cells to secrete sufficient insulin to return high blood glucose rapidly to baseline is necessary and sufficient for diabetes to occur. Nearly all genes implicated in monogenic, non-autoimmune, insulin-deficient diabetes correspondingly encode proteins with roles in pancreatic beta cell development and/or function (Fig. 1). Monogenic, insulin-deficient diabetes encompasses rare neonatal diabetes (diagnosed under 6 months of age), more frequent MODY (usually presenting before 25 years of age) and rare recessive syndromes mostly observed in consanguineous families. Pathogenic mutations in more than 30 genes have been found to cause monogenic beta cell diabetes to date, initially using linkage or homozygosity analyses in families or Sanger sequencing of candidate genes, with NGS (whole-exome or whole-genome) [15] more recently dominating.
Genes linked with monogenic insulin-deficient diabetes encode proteins that play a key role in pancreatic beta cells. Genes marked with a light-blue star are actionable monogenic beta cell diabetes genes. The dashed line between G6P and pyruvate indicates that several steps are involved. G6P, glucose-6-phosphate. This figure is available as part of a downloadable slideset
Genes implicated in monogenic diabetes include several encoding transcription factors involved in pancreatic beta cell development (e.g. HNF1A, HNF1B, HNF4A, PDX1, GATA4, GATA6). Interestingly, diabetes is not simply a consequence of loss of beta cell differentiation in all cases, however. In people with MODY due to HNF4A mutations, early life hypoglycaemia due to insulin hypersecretion is also described, and may be sustained [16], while 7% of those deficient for HNF1A develop liver adenomatosis [17]. These observations suggest dysregulation rather than loss of endodermal development to beta cells.
Several more beta cell diabetes genes encode endoplasmic reticulum proteins that enable large-scale synthesis of insulin by beta cells (e.g. WFS1, DNAJC3, EIF2B1, EIF2AK3), and the gene encoding insulin itself (INS). Crucially for precision therapy, two further genes encode the core and regulatory subunits of a key ATP-sensitive potassium channel (KCNJ11 and ABCC8, respectively), while one encodes glucokinase (GCK). These three genes play vital roles in the sensing of blood glucose and the transducing of glucose concentration into insulin secretion (Fig. 1). Heterozygous mutations in GCK or HNF1A are the commonest causes of monogenic diabetes in European populations.
Autosomal recessive syndromes featuring insulin deficiency include Wolfram syndrome (WFS1 mutations), Mitchell–Riley Syndrome (RFX6) and Wolcott–Rallison syndrome (EIF2AK3) (Table 1). NGS in suspected MODY without syndromic features has shown some such people to harbour mutations in other genes implicated in syndromic diabetes [18,19,20,21]. Such blurring of the demarcation between syndromic and non-syndromic diabetes and between different Mendelian inheritance patterns is emerging across monogenic beta cell and insulin-resistant diabetes.
Precision therapy in monogenic beta cell diabetes
Identification of monogenic diabetes genes has allowed ground-breaking advances in precision medicine [14], many focused on the cheap sulfonylurea drugs long used in type 2 diabetes. Sulfonylureas bind and inhibit the ABCC8-encoded regulatory subunit of the hyperpolarising Kir6.2 potassium channel in beta cells. Mutations in KCNJ11 or ABCC8 that increase the opening probability of the channel hyperpolarise the beta cell membrane and inhibit insulin secretion, causing either neonatal diabetes or MODY. Affected patients, remarkably, can often stop insulin therapy on genetic diagnosis and transfer safely onto long-term sulfonylurea treatment [22,23,24,25,26].
People with MODY and heterozygous HNF1A or HNF4A mutations are also highly sensitive to sulfonylureas. Mouse studies suggested this is accounted for by delayed clearance of sulfonylureas [27], but this appears not to hold in humans [28]. Initial case reports were followed up by larger case series, and it is now established that people deficient for HNF1A or HNF4A are optimally treated with oral sulfonylureas, possibly with an adjunctive dipeptidyl peptidase 4 inhibitor or glucagon-like peptide 1 receptor agonist [29, 30].
Success in treatment changes after genetic diagnosis is neither certain nor always sustained. Standards for interpretation of variants in rare genetic disorders [31] must first be applied, so that treatment change is only attempted in appropriate genetic contexts. In a recent study, among 16 patients with MODY who were offered treatment switch after genetic diagnosis, nine (six HNF1A, three KCNJ11) were stably switched from insulin to oral sulfonylurea but seven (three HNF4A, two ABCC8 and one each HNF1A and KCNJ11) had to restart insulin [32]. The strongest predictor of stable transfer was a high plasma C-peptide concentration, with younger age, lower HbA1c and shorter duration of diabetes of lesser importance [32]. The last two factors, together with low BMI, predicted successful treatment change in another study focused on patients deficient for HNF1A or HNF4A [33]. These findings emphasise the importance of developing diagnostic pathways to identify patients potentially treatable with sulfonylureas early in disease progression.
Sometimes precision medicine involves avoidance of therapy in selected subgroups. Thus, people with MODY with mild to moderate hyperglycaemia solely due to GCK deficiency do not require glucose-lowering treatment. In this case the problem is altered beta cell glucose sensing, raising the homeostatic set point for glucose, and causing lifelong mild, non-progressive hyperglycaemia. People with GCK-MODY have been shown not to develop severe complications. Indeed, the prevalence of nephropathy and macrovascular complications in 99 patients with MODY deficient for GCK was found to be similar to the prevalence in people without diabetes despite nearly 50 years of hyperglycaemia [34]. While the prevalence of retinopathy was significantly increased in GCK-deficient individuals compared with control groups (30% vs 14%), they did not require laser therapy [34]. Hypoglycaemic agents are generally ineffective in GCK deficiency, likely due to the power of the homeostatic loop maintaining blood glucose concentration: among 799 patients with GCK-MODY, HbA1c was similar in patients on pharmacological treatment (oral agents or insulin) and those on no therapy [35]. Furthermore, among 16 patients, discontinuation of therapy for at least 3 months did not alter HbA1c [35].
Monogenic insulin-deficient diabetes as the sentinel feature of wider syndromes
A final important aspect of precision diabetes is that for some genetic alterations diabetes may be the sentinel feature of a wider spectrum of abnormalities [14]. This is unsurprising given the important roles in visceral development played by MODY transcription factors, and given fundamental cellular functions of recessive beta cell diabetes genes (Fig. 1). For example, patients with diabetogenic mutations in HNF1B, GATA4, GATA6, WFS1 or mitochondrial DNA often have developmental anomalies beyond the pancreatic islets, including pancreatic agenesis; kidney, genital tract and heart malformations; and deafness or hearing loss, all showing variable penetrance. Among 201 patients with an HNF1B mutation, one study showed that while 82% presented with diabetes, 44% also had stage 3–4 renal impairment and 21% end-stage renal disease at diagnosis [36]. Furthermore, among 102 patients who eventually developed diabetes and renal disease, kidney dysfunction was diagnosed before diabetes in 39%, diabetes and kidney dysfunction were diagnosed concomitantly in 24% and kidney dysfunction was diagnosed after a median diabetes duration of 11 years in 37% [36]. Patients with GATA6 or GATA4 mutations can present with a wide spectrum of diabetic presentations and/or cardiac malformation, even within the same family [37,38,39].
Variable penetrance for components of complex syndromes is common, and widely ascribed to differing genetic backgrounds. Indeed, a high genome-wide burden of common risk alleles from genome-wide association studies (GWAS) confers a risk of diseases including diabetes that approaches the risk of single pathogenic Mendelian mutations [40]. Moreover, the penetrance of diabetes in those with HNF1A mutations is significantly influenced by polygenic type 2 diabetes risk score [41, 42]. Interactions among rarer alleles are also likely, but adequately powered studies to identify genetic modifiers agnostically in rare disease are notoriously challenging or impossible for all but the least rare disease-causing mutations.
Cost-effectiveness of genetic testing in monogenic insulin-deficient diabetes
Genetic screening for monogenic diabetes has been found to be highly cost-effective in economic analyses in the USA based on contemporary incremental cost-effectiveness ratio thresholds [43, 44]. These studies, published before the advent of NGS, modelled: (1) testing for mutations in KCNJ11 or ABCC8 in hypothetical 6-year-old patients with permanent neonatal diabetes; or (2) testing for mutations in HNF1A, HNF4A or GCK in hypothetical newly diagnosed MODY patients at 25 to 40 years of age, otherwise presumed to have type 2 diabetes. More recently, simulation of routine screening for MODY based on NGS (targeting GCK, HNF1A, HNF4A, ABCC8 and KCNJ11) in all paediatric diabetes, including children with presumed type 1 diabetes, found that this could significantly reduce health system costs and improve patient quality of life in Australia [45].
Adipose tissue and insulin signalling defects
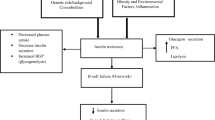
Single gene disorders causing severe insulin resistance are the complement of beta cell disorders. Unlike beta cell diabetes, where hyperglycaemia occurs early, it may be delayed or even absent in monogenic severe insulin resistance. This is because pancreatic islets have a large capacity to respond to severe insulin resistance through beta cell hyperplasia and insulin hypersecretion. It is not uncommon for people with insulin receptor (INSR) defects to maintain plasma insulin concentrations 1–2 orders of magnitude above normal for years before diabetes develops. Before this, however, morbidity is common, driven by effects of hyperinsulinaemia on ovaries, skin and other soft tissues. In lipodystrophy, ‘lipotoxic’ complications including fatty liver, dyslipidaemia and their sequelae also usually precede diabetes [46]. First presentation of severe insulin resistance may thus be to lipid, endocrinology, liver, gynaecology, dermatology, surgical or other services. All clinicians should be alert to the value of acanthosis nigricans, a velvety browning and thickening of flexural skin, as a sign of insulin resistance. Finding this in lean patients is an important indicator of possible monogenic insulin resistance.
Over 33 years since pathogenic mutations in the INSR gene were discovered in people with extreme insulin resistance, more than 25 monogenic disorders that feature insulin resistance disproportionate to body fat mass have been described (Table 1, Fig. 2). In this discussion, genetic disorders where insulin resistance is secondary to severe obesity, and appears wholly explained by the degree of obesity, will not be considered, as they have been well reviewed elsewhere [47].
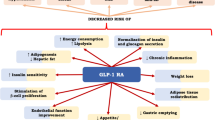
Monogenic insulin resistance subtypes and therapeutic strategies. (a) Clustering of severe insulin resistance subtypes according to severity of insulin resistance and lipotoxic features, with reference to health and type 2 diabetes (Type 2 DM). HDL-C, HDL-cholesterol; IR, insulin resistance; NAFLD, non-alcoholic fatty liver disease; PCOS, polycystic ovary syndrome; TG, triacylglycerol. (b) Genes implicated in monogenic severe insulin resistance of different subtypes and potential therapeutic strategies. Possible strategies, supported by case series and clinical experience for each subtype, are indicated in brackets, with reference to the Venn diagram (and labelled A–D). Senolytic therapies are a possible future prospect only. CGL, congenital generalised lipodystrophy; FPLD, familial partial lipodystrophy; FTIs, farnesyl transferase inhibitors; SGLT2i, sodium−glucose cotransporter 2 inhibitor. This figure is available as part of a downloadable slideset
When efforts began to establish the genetic aetiology of severe insulin resistance, the parsimonious assumption was that most causal genes would encode components of the insulin signalling pathway that was then being elucidated. However, such defects have proved surprisingly rare. Apart from INSR defects, only a handful of other bona fide signalling defects have been found, including a loss-of-function mutation in AKT2, a key intracellular mediator of insulin action, in a single family [48], and dominant mutations affecting another critical signalling enzyme, phosphoinositide 3-kinase (PI3K), in multiple families [49]. Rare heterozygous mutations in TBC1D4 [50], which regulates insulin-responsive glucose transport, also may cause severe insulin resistance.
Monogenic insulin resistance is now known to be caused much more commonly by defects in development or maintenance of adipose tissue [46], which produce lipodystrophy. In these clinically and genetically heterogeneous disorders, adipose defects may be generalised or partial, and inheritance may be recessive or dominant. Inherited lipodystrophies have recently been comprehensively reviewed elsewhere [51].
As well as relatively ‘clean’ defects in insulin action or adipose tissue development/function, a wide range of rare disorders have been described where severe insulin resistance is part of a more complex syndrome (Table 1, Fig. 1). Some recurring themes have emerged in cellular organelles or pathways affected in such disorders. More than one causal gene is involved in function of the nuclear lamina, an external scaffold for the nucleus (LMNA [52], ZMPSTE24 [53]), in formation of cell membrane invaginations known as caveoli (CAVIN1 [54], CAV1 [55]), in centrosomal function (e.g. ALMS1 [56], POC1A [57], PCNT [58]) or in DNA replication and/or repair (e.g. WRN [59], POLD1 [60], NSMCE2 [61]) (Table 1, Fig. 2). These findings imply vulnerability of adipose tissue to certain types of DNA damage or perturbation of cell division, in keeping with features of accelerated ageing seen in several of the syndromes (Table 1). Most of these syndromes are diagnosed early based on characteristic developmental abnormalities; however, for some disorders diabetes may be the sentinel presentation, either due to late onset of other syndromic features, as in Werner syndrome [59], or in formes frustes of classical syndromes. As for syndromic beta cell defects, it is important that diabetologists are vigilant for patterns of abnormalities associated with diabetes that give clues to a genetic cause.
Precision therapy in monogenic insulin resistance
Identifying monogenic insulin resistance has several implications for therapy, although evidence rests on uncontrolled case series and physiological rationales rather than randomised, controlled trials. In severely insulin-resistant diabetes plasma insulin concentrations are usually extremely high, and therapy with insulin-sensitising agents is strongly favoured over secretagogues such as sulfonylureas. Patients with monogenic insulin resistance also usually require more multidisciplinary care than is common in diabetes clinics. This may address subfertility and hyperandrogenism driven by insulin resistance and common in all monogenic insulin resistance [62], the severe fatty liver disease and dyslipidaemia seen in lipodystrophy, other manifestations of soft tissue overgrowth or syndromic features of complex insulin resistance. Some forms of precision therapy are specific to monogenic insulin resistance subtypes.
Insulin signalling defects
In extreme insulin resistance due to recessive INSR defects, uncontrolled case series indicate that recombinant human IGF-1 (rhIGF-1), a homologue of insulin, or leptin, discussed below, exerts acute and chronic benefits [63, 64]. Although rhIGF-1 has also occasionally been used in older patients with heterozygous INSR defects [63, 65], the long-term risks and benefits of these agents are unclear, and they have no place in therapy outside clinical trials in this group. Reactivation of some mutant INSR using monoclonal antibodies has been tested in models [66, 67], and has promise for the future, but remains experimental.
The lack of dyslipidaemia or fatty liver in severe insulin resistance due to INSR or PIK3R1 defects [68, 69] raises the possibility that these subtypes of insulin-resistant diabetes might confer lower macrovascular risk than type 2 diabetes, mandating a different, ‘precision’ approach to primary prevention; however, this has not been tested longitudinally (Fig. 2a).
Lipodystrophy
Lipodystrophic insulin resistance currently presents the greatest opportunity for precision therapy in monogenic insulin-resistant diabetes. Sustained positive energy balance, common in contemporary society, normally leads to sequestration of excess energy in adipose tissue as energy-dense triacylglycerol, and ultimately obesity. In lipodystrophy, however, adipose tissue is absent or has impaired ability to store energy. ‘Adipose failure’ thus occurs early when adipose storage is called upon [70, 71], and progression to insulin resistance and diabetes is dramatically accelerated, commonly being seen even in those with normal BMI. This is most severe in those with no adipose tissue (generalised lipodystrophy), who often develop extreme ‘lipotoxic’ insulin resistance, featuring hypertriacylglycerolaemia, pancreatitis, fatty liver disease and early atherosclerosis, despite being lean and often athletic in appearance.
This exquisite sensitivity to energy excess means that reversing positive energy balance in lipodystrophy is crucial. This is achieved by treating affected patients as having ‘obesity’ complications despite normal or low BMI. Adipose tissue offloading measures include hypoenergetic, low-fat diet, and therapies normally limited to obese patients, including glucagon-like peptide-1 (GLP-1) agonists or bariatric surgery [72] (Fig. 2b). The adipose offloading strategy is validated by observation of people with lipodystrophy who do sustain neutral energy balance, for example, through intensive endurance sport. Where no demand is placed on adipose tissue, such individuals can remain entirely metabolically healthy, in a state of latent lipodystrophy.
Lipotoxicity is particularly intractable when adipose tissue is absent or nearly absent, and blood concentrations of the adipose-derived hormone leptin are low or undetectable. This hyperactivates hypothalamic appetite centres, producing intense hunger and increased food intake. This targets the physiological weak point—impaired adipose storage capacity—in a vicious circle. Mitigating this drive to eat is likely to be the main reason for efficacy of recombinant human leptin injection in lipodystrophy when baseline leptin concentration is particularly low (e.g. [73,74,75]). Leptin has shown major metabolic benefits in uncontrolled trials in generalised lipodystrophy, and lesser benefit in severe partial lipodystrophy. It is licensed in Europe for both subtypes when metabolic control remains poor despite best available treatment, and in the USA for generalised lipodystrophy only. Even the lowest concentrations of leptin may be seen in metabolically healthy lean people, especially prepubertally, and so plasma leptin concentration alone cannot serve as an indication for replacement. Instead, it must be viewed in the context of overall metabolic state. In lipotoxic insulin resistance a plasma leptin threshold for replacement around 4 μg/l was initially suggested [76], but later analysis did not confirm a robust threshold [77]. Workup in an experienced centre is thus highly desirable before leptin therapy is considered.
A complementary strategy to offloading adipose tissue would, in principle, be to increase adipose storage capacity (Fig. 2b). The nuclear hormone receptor peroxisome proliferator-activated receptor-γ (PPARγ), encoded by PPARG, is the master regulator of adipose tissue biogenesis, and its pharmacological activation by pioglitazone or other thiazolidinediones (TZDs) seems an obvious strategy in lipodystrophy. Many reports attest to metabolic benefits of TZDs in partial lipodystrophies (e.g. [78, 79]), but they are often disliked by patients as they increase the size of non-affected adipose tissue depots [80], for example, around the head and neck or on the medial thighs. Moreover, the exquisite dietary sensitivity of people with lipodystrophy complicates efforts to discriminate beneficial effects of pharmacotherapy in uncontrolled observational studies, as detection of drug effects may be confounded by effects of concomitant behavioural alterations.
TZDs have attracted specific interest in familial partial lipodystrophy type 3 (FPLD3), the second commonest monogenic lipodystrophy, which is caused by mutations in the PPARG gene itself [81, 82]. A priori, it was thought that TZDs might either be particularly effective in FPLD3, by targeting the causal defect, or particularly ineffective, if causal PPARG mutations were unresponsive to agonist therapy. This question is not yet answered by trials, but at least some PPARG variants respond in cellular studies to potent exogenous TZDs, sometimes when they fail to respond to putative endogenous ligands [83].
Uniquely among genes causing monogenic insulin resistance, PPARG has been subject to ‘saturation’ mutagenesis coupled to massively parallel assay of mutation consequences. This has produced experimental evidence for the functional consequences of the large majority of possible PPARG missense mutations [84], and has suggested that up to one in 500 people harbour a PPARG loss-of-function mutation. This sets the stage for more formal testing at scale of precision TZD therapy in FPLD3, and provides a paradigm with potential applicability to many other forms of monogenic diabetes.
Complex insulin resistance syndromes
Several complex syndromes feature insulin-resistant diabetes with frank lipodystrophy, or resemble lipodystrophy metabolically with lesser degrees of abnormal adipose distribution. It is reasonable to assume that the principles of management of lipodystrophy apply to these disorders. No more precision therapies are yet proven to be efficacious for metabolic endpoints, although the US Food and Drug Administration (FDA) recently licensed lonafarnib, a farnesyltransferase inhibitor that reduces accumulation of damaging fragments of lamin proteins, in Hutchinson–Gilford syndrome, a rare premature ageing disorder caused by defects in the LMNA gene [85]. As lipodystrophy is a component of many laminopathies, it is plausible that this targeted therapy will also have metabolic benefits in selected patients. Other prospects include senolytic therapies that target senescent cells in adipose tissue [86]. These may be particularly worthy of evaluation in syndromes featuring defective DNA damage sensing and/or repair or impaired centrosomal function (Fig. 2b, Table 1).
Cost-effectiveness of testing for and treating monogenic insulin-resistant diabetes
Few health economic analyses of either screening for or treatment of monogenic insulin-resistant diabetes have been undertaken. The high cost of recombinant human leptin therapy (the UK National Health Service [NHS] indicative price for adults starts at £200,000 to 400,000 annually) meant that economic modelling was important prior to approval of leptin therapy in lipodystrophy by the UK National Institute for Health and Care Excellence [87]. This modelling included quality of life as well as metabolic control. Nevertheless, no economic evaluation of strategies to treat lipodystrophy has been undertaken that includes the full range of medical and surgical therapies. Given the cost of leptin, it is suggested that treatment should only be started in liaison with specialised lipodystrophy services, where accessible.
Improving diagnosis rates of monogenic diabetes
Many people with potentially treatable monogenic beta cell and insulin-resistant diabetes are currently misclassified as having type 2 diabetes. An immediate priority is thus to facilitate identification of patients for genetic testing. A widely used MODY probability calculator that incorporates disease biomarkers and simple clinical data [88, 89] has proven successful in such triage [14]. No such calculator yet exists for monogenic insulin-resistant diabetes, but ‘genetic’ lipodystrophy may occur in up to one in 7000 of the general population [90], suggesting a large unmet need. With the advance of artificial intelligence applied to healthcare data at scale, it is likely that digital approaches to predicting monogenic diabetes will soon become both more refined and ideally applicable to many more monogenic diabetes subtypes.
Any attempt to screen genetically for monogenic diabetes and to intervene based on results will need to take into account the clinical penetrance of gene variants. Fully penetrant, Mendelian, monogenic disease is to some extent a self-fulfilling construct arising from ascertainment of affected patients and families based on their phenotype (in this case diabetes). It has been of great value for early phases of study of genetic disease, but in some cases has become untenable as the spectrum of genetic variation has been revealed by NGS in large populations [91, 92]. In ‘monogenic’ diabetes, too, population studies ascertaining purely by genotype demonstrate weaker associations between genotype and disease than genetic testing with a clinical indication [12, 13]. This suggests that for some types of diabetes we may move away from the notion of mutations that confer disease risk deterministically (i.e. having the gene change always means having diabetes), to the concept that gene changes, even when implicated in monogenic disease, confer disease risk probabilistically (i.e. having the gene mutation increases diabetes probability/expressivity to different degrees) [93].
Significance of monogenic diabetes for ‘common’ type 2 diabetes
The importance of monogenic disease extends beyond clinically recognisable subtypes nested within diabetes populations, as NGS suggests overlap between monogenic and ‘common’ type 2 diabetes [92]. More than 5% of people with characteristics of type 2 diabetes have been shown to harbour mutations that cause monogenic diabetes (according to standards for interpretation of variants in rare genetic disorders [31]), and in particular MODY [10, 12, 94, 95]). Such people were leaner and developed diabetes earlier than non-carriers with type 2 diabetes [10], but would often not have been detected without genetic screening. None developed type 2 diabetes before 25 years of age, and the first-degree family history of diabetes was similar between carriers and non-carriers [10]. The finding of lipodystrophy causal variants in up to one in 7000 people [90] suggests that a similar paradigm may apply to insulin-resistant diabetes. These discoveries open a gateway for precision medicine among some newly diagnosed patients with apparent type 2 diabetes. Routine screening for monogenic diabetes genes warrants assessment; however, no modelling has been reported, and feasibility of screening followed by appropriate treatment changes requires testing in large-scale studies.
Taking one step further, the possibility that a significant portion of type 2 diabetes heritability may be attributable to rare, functional gene variants has long been mooted. This was encouraged by the limited early success of GWAS focused on commoner gene variants. Sequencing of large populations has indeed identified some rare variants (e.g. in MTNR1B, PPARG, SLC30A8, HNF1A, PDX1, PAM) that confer risk of type 2 diabetes that is intermediate between common SNPs and mutations causing monogenic diabetes [96]. Moreover, in isolated populations, even higher prevalence rates of loss-of-function mutations in AKT2 and TBC1D4, both involved in insulin signalling and implicated in monogenic insulin resistance, have been described [97, 98]. For TBC1D4 mutation carriers, some evidence for precision exercise intervention has recently been advanced [99].
Nevertheless, as GWAS have grown ever larger, several hundred common polymorphisms associated with type 2 diabetes risk [100] have been identified, mostly located in non-coding DNA regions, and NGS has not supported a major population-wide effect of rare alleles in monogenic diabetes genes [92]. Evidence thus now favours the notion that most type 2 diabetes risk is conferred by aggregated effect of large numbers of common variants of small effect size.
As genetic risk associations of ever smaller effect size proliferate, study of monogenic diabetes may become more important rather than less relevant, however. Understanding of monogenic diabetes syndromes and underpinning mechanisms can inform clustering of common polygenic risk alleles into functionally coherent subgroups. This is valuable given very small effect sizes of common variants, which are difficult to resolve experimentally. For example, learning from monogenic lipodystrophy has informed clustering of traits into lipodystrophic and non-lipodystrophic patterns [101,102,103], and has been used to suggest functionally distinct groups of common variants.
Conclusion
Monogenic diabetes is underdiagnosed, yet offers increasing opportunities for genotype-targeted behavioural and pharmacological therapy. Lessons from monogenic disorders are also likely applicable to a subset of people with type 2 diabetes with formes frustes of these conditions, which are difficult to discriminate clinically from type 2 diabetes. Whether systematic genetic screening is warranted remains to be determined. Monogenic diabetes may also be conceptually influential in understanding population propensity to obesity-related diabetes. Full realisation of the potential of genetic medicine in the diabetes clinic will require integration of expertise in rare disease genetics, population genetics, data science and clinical care, and breaking down of some traditional clinical silos. Thus, although genetically informed precision diabetes therapy is not yet quite ready for implementation in "common" type 2 diabetes, it now stands on the threshold. The fruits of more than 30 years of genetic research may soon be harvested for wider benefits in routine practice.
Abbreviations
- FPLD3:
-
Familial partial lipodystrophy type 3
- GLP-1:
-
Glucagon-like peptide-1
- GWAS:
-
Genome-wide association study
- NGS:
-
Next generation sequencing
- PI3K:
-
Phosphoinositide 3-kinase
- rhIGF-1:
-
Recombinant human IGF-1
- TZDs:
-
Thiazolidinediones
References
Saeedi P, Petersohn I, Salpea P et al (2019) Global and regional diabetes prevalence estimates for 2019 and projections for 2030 and 2045: Results from the International Diabetes Federation Diabetes Atlas, 9(th) edition. Diabetes Res Clin Pract 157:107843. https://doi.org/10.1016/j.diabres.2019.107843
Bullard KM, Cowie CC, Lessem SE et al (2018) Prevalence of Diagnosed Diabetes in Adults by Diabetes Type - United States, 2016. MMWR Morb Mortal Wkly Rep 67(12):359–361. https://doi.org/10.15585/mmwr.mm6712a2
Tattersall RB (1974) Mild familial diabetes with dominant inheritance. Q J Med 43(170):339–357
Kahn CR, Podskalny JM (1980) Demonstration of a primary (? genetic) defect in insulin receptors in fibroblasts from a patient with the syndrome of insulin resistance and acanthosis nigricans type A. J Clin Endocrinol Metab 50(6):1139–1141. https://doi.org/10.1210/jcem-50-6-1139
Kadowaki T, Bevins CL, Cama A et al (1988) Two mutant alleles of the insulin receptor gene in a patient with extreme insulin resistance. Science 240(4853):787–790. https://doi.org/10.1126/science.2834824
Yoshimasa Y, Seino S, Whittaker J et al (1988) Insulin-resistant diabetes due to a point mutation that prevents insulin proreceptor processing. Science 240(4853):784–787. https://doi.org/10.1126/science.3283938
Hattersley AT, Turner RC, Permutt MA et al (1992) Linkage of type 2 diabetes to the glucokinase gene. Lancet 339(8805):1307–1310. https://doi.org/10.1016/0140-6736(92)91958-b
Froguel P, Vaxillaire M, Sun F et al (1992) Close linkage of glucokinase locus on chromosome 7p to early-onset non-insulin-dependent diabetes mellitus. Nature 356(6365):162–164. https://doi.org/10.1038/356162a0
Ellard S, Colclough K, Patel KA, Hattersley AT (2020) Prediction algorithms: pitfalls in interpreting genetic variants of autosomal dominant monogenic diabetes. J Clin Invest 130(1):14–16. https://doi.org/10.1172/JCI133516
Bonnefond A, Boissel M, Bolze A et al (2020) Pathogenic variants in actionable MODY genes are associated with type 2 diabetes. Nat Metab 2(10):1126–1134. https://doi.org/10.1038/s42255-020-00294-3
Philippe J, Derhourhi M, Durand E et al (2015) What Is the Best NGS Enrichment Method for the Molecular Diagnosis of Monogenic Diabetes and Obesity? PLoS One 10(11):e0143373. https://doi.org/10.1371/journal.pone.0143373
Goodrich JK, Singer-Berk M, Son R et al (2021) Determinants of penetrance and variable expressivity in monogenic metabolic conditions across 77,184 exomes. Nat Commun 12(1):3505. https://doi.org/10.1038/s41467-021-23556-4
Flannick J, Beer NL, Bick AG et al (2013) Assessing the phenotypic effects in the general population of rare variants in genes for a dominant Mendelian form of diabetes. Nat Genet 45(11):1380–1385. https://doi.org/10.1038/ng.2794
Hattersley AT, Patel KA (2017) Precision diabetes: learning from monogenic diabetes. Diabetologia 60(5):769–777. https://doi.org/10.1007/s00125-017-4226-2
Bonnefond A, Shuldiner AR, Froguel P (2016) Historical Overview of Gene Discovery Methodologies in Type 2 Diabetes. In: Florez JC (ed) The Genetics of Type 2 Diabetes and Related Traits: Biology, Physiology and Translation. Springer International Publishing, Cham, pp 3–12
Flanagan SE, Kapoor RR, Mali G et al (2010) Diazoxide-responsive hyperinsulinemic hypoglycemia caused by HNF4A gene mutations. Eur J Endocrinol 162(5):987–992. https://doi.org/10.1530/EJE-09-0861
Haddouche A, Bellanne-Chantelot C, Rod A et al (2020) Liver adenomatosis in patients with hepatocyte nuclear factor-1 alpha maturity onset diabetes of the young (HNF1A-MODY): Clinical, radiological and pathological characteristics in a French series. J Diabetes 12(1):48–57. https://doi.org/10.1111/1753-0407.12959
Patel KA, Kettunen J, Laakso M et al (2017) Heterozygous RFX6 protein truncating variants are associated with MODY with reduced penetrance. Nat Commun 8(1):888. https://doi.org/10.1038/s41467-017-00895-9
Bonnycastle LL, Chines PS, Hara T et al (2013) Autosomal dominant diabetes arising from a Wolfram syndrome 1 mutation. Diabetes 62(11):3943–3950. https://doi.org/10.2337/db13-0571
Colclough K, Ellard S, Hattersley A, Patel K (2021) Syndromic Monogenic Diabetes Genes Should be Tested in Patients With a Clinical Suspicion of MODY. Diabetes 71(3):530–537. https://doi.org/10.2337/db21-0517
Saint-Martin C, Bouvet D, Bastide M, Chantelot CB, Monogenic Diabetes Study Group of the Societe Francophone du Diabète (2021) Gene Panel Sequencing of Patients With Monogenic Diabetes Brings to Light Genes Typically Associated With Syndromic Presentations. Diabetes 71(3):578–584. https://doi.org/10.2337/db21-0520
Gloyn AL, Pearson ER, Antcliff JF et al (2004) Activating mutations in the gene encoding the ATP-sensitive potassium-channel subunit Kir6.2 and permanent neonatal diabetes. N Engl J Med 350(18):1838–1849. https://doi.org/10.1056/NEJMoa032922
Babenko AP, Polak M, Cave H et al (2006) Activating mutations in the ABCC8 gene in neonatal diabetes mellitus. N Engl J Med 355(5):456–466. https://doi.org/10.1056/NEJMoa055068
Bowman P, Mathews F, Barbetti F et al (2021) Long-term Follow-up of Glycemic and Neurological Outcomes in an International Series of Patients With Sulfonylurea-Treated ABCC8 Permanent Neonatal Diabetes. Diabetes Care 44(1):35–42. https://doi.org/10.2337/dc20-1520
Bowman P, Sulen A, Barbetti F et al (2018) Effectiveness and safety of long-term treatment with sulfonylureas in patients with neonatal diabetes due to KCNJ11 mutations: an international cohort study. Lancet Diabetes Endocrinol 6(8):637–646. https://doi.org/10.1016/S2213-8587(18)30106-2
Lanning MS, Carmody D, Szczerbinski L, Letourneau LR, Naylor RN, Greeley SAW (2018) Hypoglycemia in sulfonylurea-treated KCNJ11-neonatal diabetes: Mild-moderate symptomatic episodes occur infrequently but none involving unconsciousness or seizures. Pediatr Diabetes 19(3):393–397. https://doi.org/10.1111/pedi.12599
Boileau P, Wolfrum C, Shih DQ, Yang TA, Wolkoff AW, Stoffel M (2002) Decreased glibenclamide uptake in hepatocytes of hepatocyte nuclear factor-1alpha-deficient mice: a mechanism for hypersensitivity to sulfonylurea therapy in patients with maturity-onset diabetes of the young, type 3 (MODY3). Diabetes 51(Suppl 3):S343–S348. https://doi.org/10.2337/diabetes.51.2007.s343
Urbanova J, Andel M, Potockova J et al (2015) Half-Life of Sulfonylureas in HNF1A and HNF4A Human MODY Patients is not Prolonged as Suggested by the Mouse Hnf1a(-/-) Model. Curr Pharm Des 21(39):5736–5748. https://doi.org/10.2174/1381612821666151008124036
Pearson ER, Pruhova S, Tack CJ et al (2005) Molecular genetics and phenotypic characteristics of MODY caused by hepatocyte nuclear factor 4alpha mutations in a large European collection. Diabetologia 48(5):878–885. https://doi.org/10.1007/s00125-005-1738-y
Pearson ER, Starkey BJ, Powell RJ, Gribble FM, Clark PM, Hattersley AT (2003) Genetic cause of hyperglycaemia and response to treatment in diabetes. Lancet 362(9392):1275–1281. https://doi.org/10.1016/S0140-6736(03)14571-0
Richards S, Aziz N, Bale S et al (2015) Standards and guidelines for the interpretation of sequence variants: a joint consensus recommendation of the American College of Medical Genetics and Genomics and the Association for Molecular Pathology. Genet Med 17(5):405–424. https://doi.org/10.1038/gim.2015.30
Stankute I, Verkauskiene R, Blouin JL et al (2020) Systematic Genetic Study of Youth With Diabetes in a Single Country Reveals the Prevalence of Diabetes Subtypes, Novel Candidate Genes, and Response to Precision Therapy. Diabetes 69(5):1065–1071. https://doi.org/10.2337/db19-0974
Shepherd MH, Shields BM, Hudson M et al (2018) A UK nationwide prospective study of treatment change in MODY: genetic subtype and clinical characteristics predict optimal glycaemic control after discontinuing insulin and metformin. Diabetologia 61(12):2520–2527. https://doi.org/10.1007/s00125-018-4728-6
Steele AM, Shields BM, Wensley KJ, Colclough K, Ellard S, Hattersley AT (2014) Prevalence of vascular complications among patients with glucokinase mutations and prolonged, mild hyperglycemia. JAMA 311(3):279–286. https://doi.org/10.1001/jama.2013.283980
Stride A, Shields B, Gill-Carey O et al (2014) Cross-sectional and longitudinal studies suggest pharmacological treatment used in patients with glucokinase mutations does not alter glycaemia. Diabetologia 57(1):54–56. https://doi.org/10.1007/s00125-013-3075-x
Dubois-Laforgue D, Cornu E, Saint-Martin C et al (2017) Diabetes, Associated Clinical Spectrum, Long-term Prognosis, and Genotype/Phenotype Correlations in 201 Adult Patients With Hepatocyte Nuclear factor 1B (HNF1B) Molecular Defects. Diabetes Care 40(11):1436–1443. https://doi.org/10.2337/dc16-2462
Bonnefond A, Sand O, Guerin B et al (2012) GATA6 inactivating mutations are associated with heart defects and, inconsistently, with pancreatic agenesis and diabetes. Diabetologia 55(10):2845–2847. https://doi.org/10.1007/s00125-012-2645-7
De Franco E, Shaw-Smith C, Flanagan SE et al (2013) GATA6 mutations cause a broad phenotypic spectrum of diabetes from pancreatic agenesis to adult-onset diabetes without exocrine insufficiency. Diabetes 62(3):993–997. https://doi.org/10.2337/db12-0885
Shaw-Smith C, De Franco E, Lango Allen H et al (2014) GATA4 mutations are a cause of neonatal and childhood-onset diabetes. Diabetes 63(8):2888–2894. https://doi.org/10.2337/db14-0061
Khera AV, Chaffin M, Aragam KG et al (2018) Genome-wide polygenic scores for common diseases identify individuals with risk equivalent to monogenic mutations. Nat Genet 50(9):1219–1224. https://doi.org/10.1038/s41588-018-0183-z
Kettunen JLT, Rantala E, Dwivedi OP et al (2021) A multigenerational study on phenotypic consequences of the most common causal variant of HNF1A-MODY. Diabetologia 65(4):632–643. https://doi.org/10.1007/s00125-021-05631-z
Lango Allen H, Johansson S, Ellard S et al (2010) Polygenic risk variants for type 2 diabetes susceptibility modify age at diagnosis in monogenic HNF1A diabetes. Diabetes 59(1):266–271. https://doi.org/10.2337/db09-0555
Naylor RN, John PM, Winn AN et al (2014) Cost-effectiveness of MODY genetic testing: translating genomic advances into practical health applications. Diabetes Care 37(1):202–209. https://doi.org/10.2337/dc13-0410
Greeley SA, John PM, Winn AN et al (2011) The cost-effectiveness of personalized genetic medicine: the case of genetic testing in neonatal diabetes. Diabetes Care 34(3):622–627. https://doi.org/10.2337/dc10-1616
Johnson SR, Carter HE, Leo P et al (2019) Cost-effectiveness Analysis of Routine Screening Using Massively Parallel Sequencing for Maturity-Onset Diabetes of the Young in a Pediatric Diabetes Cohort: Reduced Health System Costs and Improved Patient Quality of Life. Diabetes Care 42(1):69–76. https://doi.org/10.2337/dc18-0261
Semple RK, Savage DB, Cochran EK, Gorden P, O'Rahilly S (2011) Genetic syndromes of severe insulin resistance. Endocr Rev 32(4):498–514. https://doi.org/10.1210/er.2010-0020
van der Klaauw AA, Farooqi IS (2015) The hunger genes: pathways to obesity. Cell 161(1):119–132. https://doi.org/10.1016/j.cell.2015.03.008
George S, Rochford JJ, Wolfrum C et al (2004) A family with severe insulin resistance and diabetes due to a mutation in AKT2. Science 304(5675):1325–1328. https://doi.org/10.1126/science.1096706
Avila M, Dyment DA, Sagen JV et al (2016) Clinical reappraisal of SHORT syndrome with PIK3R1 mutations: toward recommendation for molecular testing and management. Clin Genet 89(4):501–506. https://doi.org/10.1111/cge.12688
Dash S, Sano H, Rochford JJ et al (2009) A truncation mutation in TBC1D4 in a family with acanthosis nigricans and postprandial hyperinsulinemia. Proc Natl Acad Sci U S A 106(23):9350–9355. https://doi.org/10.1073/pnas.0900909106
Lim K, Haider A, Adams C, Sleigh A, Savage DB (2021) Lipodistrophy: a paradigm for understanding the consequences of "overloading" adipose tissue. Physiol Rev 101(3):907–993. https://doi.org/10.1152/physrev.00032.2020
Shackleton S, Lloyd DJ, Jackson SN et al (2000) LMNA, encoding lamin A/C, is mutated in partial lipodystrophy. Nat Genet 24(2):153–156. https://doi.org/10.1038/72807
Agarwal AK, Fryns JP, Auchus RJ, Garg A (2003) Zinc metalloproteinase, ZMPSTE24, is mutated in mandibuloacral dysplasia. Hum Mol Genet 12(16):1995–2001. https://doi.org/10.1093/hmg/ddg213
Hayashi YK, Matsuda C, Ogawa M et al (2009) Human PTRF mutations cause secondary deficiency of caveolins resulting in muscular dystrophy with generalized lipodystrophy. J Clin Invest 119(9):2623–2633. https://doi.org/10.1172/JCI38660
Karhan AN, Zammouri J, Auclair M et al (2021) Biallelic CAV1 null variants induce congenital generalized lipodystrophy with achalasia. Eur J Endocrinol 185(6):841–854. https://doi.org/10.1530/EJE-21-0915
Dassie F, Favaretto F, Bettini S et al (2021) Alstrom syndrome: an ultra-rare monogenic disorder as a model for insulin resistance, type 2 diabetes mellitus and obesity. Endocrine 71(3):618–625. https://doi.org/10.1007/s12020-021-02643-y
Chen JH, Segni M, Payne F et al (2015) Truncation of POC1A associated with short stature and extreme insulin resistance. J Mol Endocrinol 55(2):147–158. https://doi.org/10.1530/JME-15-0090
Huang-Doran I, Bicknell LS, Finucane FM et al (2011) Genetic defects in human pericentrin are associated with severe insulin resistance and diabetes. Diabetes 60(3):925–935. https://doi.org/10.2337/db10-1334
Raffan E, Hurst LA, Turki SA et al (2011) Early Diagnosis of Werner's Syndrome Using Exome-Wide Sequencing in a Single, Atypical Patient. Front Endocrinol (Lausanne) 2:8. https://doi.org/10.3389/fendo.2011.00008
Weedon MN, Ellard S, Prindle MJ et al (2013) An in-frame deletion at the polymerase active site of POLD1 causes a multisystem disorder with lipodystrophy. Nat Genet 45(8):947–950. https://doi.org/10.1038/ng.2670
Payne F, Colnaghi R, Rocha N et al (2014) Hypomorphism in human NSMCE2 linked to primordial dwarfism and insulin resistance. J Clin Invest 124(9):4028–4038. https://doi.org/10.1172/JCI73264
Huang-Doran I, Kinzer AB, Jimenez-Linan M et al (2021) Ovarian Hyperandrogenism and Response to Gonadotropin-releasing Hormone Analogues in Primary Severe Insulin Resistance. J Clin Endocrinol Metab 106(8):2367–2383. https://doi.org/10.1210/clinem/dgab275
McDonald A, Williams RM, Regan FM, Semple RK, Dunger DB (2007) IGF-I treatment of insulin resistance. Eur J Endocrinol 157(Suppl 1):S51–S56. https://doi.org/10.1530/EJE-07-0271
Okawa MC, Cochran E, Lightbourne M, Brown RJ (2021) Long-term effects of metreleptin in Rabson-Mendenhall Syndrome on glycemia, growth, and kidney function. J Clin Endocrinol Metab 107(3):e1032–e1046. https://doi.org/10.1210/clinem/dgab782
Regan FM, Williams RM, McDonald A et al (2010) Treatment with recombinant human insulin-like growth factor (rhIGF)-I/rhIGF binding protein-3 complex improves metabolic control in subjects with severe insulin resistance. J Clin Endocrinol Metab 95(5):2113–2122. https://doi.org/10.1210/jc.2009-2088
Brierley GV, Siddle K, Semple RK (2018) Evaluation of anti-insulin receptor antibodies as potential novel therapies for human insulin receptoropathy using cell culture models. Diabetologia 61(7):1662–1675. https://doi.org/10.1007/s00125-018-4606-2
Brierley GV, Webber H, Rasijeff E, Grocott S, Siddle K, Semple RK (2020) Anti-Insulin Receptor Antibodies Improve Hyperglycemia in a Mouse Model of Human Insulin Receptoropathy. Diabetes 69(11):2481–2489. https://doi.org/10.2337/db20-0345
Huang-Doran I, Tomlinson P, Payne F et al (2016) Insulin resistance uncoupled from dyslipidemia due to C-terminal PIK3R1 mutations. JCI Insight 1(17):e88766. https://doi.org/10.1172/jci.insight.88766
Semple RK, Sleigh A, Murgatroyd PR et al (2009) Postreceptor insulin resistance contributes to human dyslipidemia and hepatic steatosis. J Clin Invest 119(2):315–322. https://doi.org/10.1172/JCI37432
Kim JK, Gavrilova O, Chen Y, Reitman ML, Shulman GI (2000) Mechanism of insulin resistance in A-ZIP/F-1 fatless mice. J Biol Chem 275(12):8456–8460. https://doi.org/10.1074/jbc.275.12.8456
Danforth E Jr (2000) Failure of adipocyte differentiation causes type II diabetes mellitus? Nat Genet 26(1):13. https://doi.org/10.1038/79111
Melvin A, Adams C, Flanagan C et al (2017) Roux-en-Y Gastric Bypass Surgery in the Management of Familial Partial Lipodystrophy Type 1. J Clin Endocrinol Metab 102(10):3616–3620. https://doi.org/10.1210/jc.2017-01235
Chong AY, Lupsa BC, Cochran EK, Gorden P (2010) Efficacy of leptin therapy in the different forms of human lipodystrophy. Diabetologia 53(1):27–35. https://doi.org/10.1007/s00125-009-1502-9
Oral EA, Simha V, Ruiz E et al (2002) Leptin-replacement therapy for lipodystrophy. N Engl J Med 346(8):570–578. https://doi.org/10.1056/NEJMoa012437
Petersen KF, Oral EA, Dufour S et al (2002) Leptin reverses insulin resistance and hepatic steatosis in patients with severe lipodystrophy. J Clin Invest 109(10):1345–1350. https://doi.org/10.1172/JCI15001
Diker-Cohen T, Cochran E, Gorden P, Brown RJ (2015) Partial and generalized lipodystrophy: comparison of baseline characteristics and response to metreleptin. J Clin Endocrinol Metab 100(5):1802–1810. https://doi.org/10.1210/jc.2014-4491
Meral R, Malandrino N, Walter M et al (2021) Endogenous Leptin Concentrations Poorly Predict Metreleptin Response in Patients with Partial Lipodystrophy. J Clin Endocrinol Metab. https://doi.org/10.1210/clinem/dgab760
Luedtke A, Boschmann M, Colpe C et al (2012) Thiazolidinedione response in familial lipodystrophy patients with LMNA mutations: a case series. Horm Metab Res 44(4):306–311. https://doi.org/10.1055/s-0031-1301284
Arioglu E, Duncan-Morin J, Sebring N et al (2000) Efficacy and safety of troglitazone in the treatment of lipodystrophy syndromes. Ann Intern Med 133(4):263–274. https://doi.org/10.7326/0003-4819-133-4-200008150-00009
McLaughlin PD, Ryan J, Hodnett PA, O'Halloran D, Maher MM (2012) Quantitative whole-body MRI in familial partial lipodystrophy type 2: changes in adipose tissue distribution coincide with biochemical improvement. AJR Am J Roentgenol 199(5):W602–W606. https://doi.org/10.2214/AJR.11.8110
Barroso I, Gurnell M, Crowley VE et al (1999) Dominant negative mutations in human PPARgamma associated with severe insulin resistance, diabetes mellitus and hypertension. Nature 402(6764):880–883. https://doi.org/10.1038/47254
Hegele RA, Cao H, Frankowski C, Mathews ST, Leff T (2002) PPARG F388L, a transactivation-deficient mutant, in familial partial lipodystrophy. Diabetes 51(12):3586–3590. https://doi.org/10.2337/diabetes.51.12.3586
Agostini M, Schoenmakers E, Beig J et al (2018) A Pharmacogenetic Approach to the Treatment of Patients With PPARG Mutations. Diabetes 67(6):1086–1092. https://doi.org/10.2337/db17-1236
Majithia AR, Tsuda B, Agostini M et al (2016) Prospective functional classification of all possible missense variants in PPARG. Nat Genet 48(12):1570–1575. https://doi.org/10.1038/ng.3700
Dhillon S (2021) Lonafarnib: First Approval. Drugs 81(2):283–289. https://doi.org/10.1007/s40265-020-01464-z
Palmer AK, Xu M, Zhu Y et al (2019) Targeting senescent cells alleviates obesity-induced metabolic dysfunction. Aging Cell 18(3):e12950. https://doi.org/10.1111/acel.12950
National Institute for Health and Care Excellence (2021) Metreleptin for treating lipodystrophy; Highly specialised technologies guidance. Reference number: HST14. Available from https://www.nice.org.uk/guidance/hst14. Accessed 5 Mar 2022
Carlsson A, Shepherd M, Ellard S et al (2020) Absence of Islet Autoantibodies and Modestly Raised Glucose Values at Diabetes Diagnosis Should Lead to Testing for MODY: Lessons From a 5-Year Pediatric Swedish National Cohort Study. Diabetes Care 43(1):82–89. https://doi.org/10.2337/dc19-0747
Delvecchio M, Salzano G, Bonura C et al (2018) Can HbA1c combined with fasting plasma glucose help to assess priority for GCK-MODY vs HNF1A-MODY genetic testing? Acta Diabetol 55(9):981–983. https://doi.org/10.1007/s00592-018-1179-y
Gonzaga-Jauregui C, Ge W, Staples J et al (2020) Clinical and Molecular Prevalence of Lipodystrophy in an Unascertained Large Clinical Care Cohort. Diabetes 69(2):249–258. https://doi.org/10.2337/db19-0447
Katsanis N (2016) The continuum of causality in human genetic disorders. Genome Biol 17(1):233. https://doi.org/10.1186/s13059-016-1107-9
Fuchsberger C, Flannick J, Teslovich TM et al (2016) The genetic architecture of type 2 diabetes. Nature 536(7614):41–47. https://doi.org/10.1038/nature18642
Lupski JR, Belmont JW, Boerwinkle E, Gibbs RA (2011) Clan genomics and the complex architecture of human disease. Cell 147(1):32–43. https://doi.org/10.1016/j.cell.2011.09.008
Bansal V, Gassenhuber J, Phillips T et al (2017) Spectrum of mutations in monogenic diabetes genes identified from high-throughput DNA sequencing of 6888 individuals. BMC Med 15(1):213. https://doi.org/10.1186/s12916-017-0977-3
Donath X, Saint-Martin C, Dubois-Laforgue D et al (2019) Next-generation sequencing identifies monogenic diabetes in 16% of patients with late adolescence/adult-onset diabetes selected on a clinical basis: a cross-sectional analysis. BMC Med 17(1):132. https://doi.org/10.1186/s12916-019-1363-0
Flannick J, Florez JC (2016) Type 2 diabetes: genetic data sharing to advance complex disease research. Nat Rev Genet 17(9):535–549. https://doi.org/10.1038/nrg.2016.56
Moltke I, Grarup N, Jorgensen ME et al (2014) A common Greenlandic TBC1D4 variant confers muscle insulin resistance and type 2 diabetes. Nature 512(7513):190–193. https://doi.org/10.1038/nature13425
Manning A, Highland HM, Gasser J et al (2017) A Low-Frequency Inactivating AKT2 Variant Enriched in the Finnish Population Is Associated With Fasting Insulin Levels and Type 2 Diabetes Risk. Diabetes 66(7):2019–2032. https://doi.org/10.2337/db16-1329
Schnurr TM, Jorsboe E, Chadt A et al (2021) Physical activity attenuates postprandial hyperglycaemia in homozygous TBC1D4 loss-of-function mutation carriers. Diabetologia 64(8):1795–1804. https://doi.org/10.1007/s00125-021-05461-z
Mahajan A, Taliun D, Thurner M et al (2018) Fine-mapping type 2 diabetes loci to single-variant resolution using high-density imputation and islet-specific epigenome maps. Nat Genet 50(11):1505–1513. https://doi.org/10.1038/s41588-018-0241-6
Lotta LA, Gulati P, Day FR et al (2017) Integrative genomic analysis implicates limited peripheral adipose storage capacity in the pathogenesis of human insulin resistance. Nat Genet 49(1):17–26. https://doi.org/10.1038/ng.3714
Scott RA, Fall T, Pasko D et al (2014) Common genetic variants highlight the role of insulin resistance and body fat distribution in type 2 diabetes, independent of obesity. Diabetes 63(12):4378–4387. https://doi.org/10.2337/db14-0319
Yaghootkar H, Scott RA, White CC et al (2014) Genetic evidence for a normal-weight "metabolically obese" phenotype linking insulin resistance, hypertension, coronary artery disease, and type 2 diabetes. Diabetes 63(12):4369–4377. https://doi.org/10.2337/db14-0318
Acknowledgements
We are grateful to D. Savage, University of Cambridge, UK, for review of the manuscript.
Authors’ relationships and activities
AB is a member of the Editorial Board of Diabetologia. The authors declare that there are no other relationships or activities that might bias, or be perceived to bias, their work.
Contribution statement
Both authors were responsible for drafting the article and revising it critically for intellectual content. Both authors approved the version to be published.
Funding
RKS is funded by the Wellcome Trust (grant 210752/Z/18/Z). AB is supported by the European Research Council (ERC Reg-Seq: 715575).
Author information
Authors and Affiliations
Corresponding authors
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
ESM 1
(PPTX 568 kb)
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Bonnefond, A., Semple, R.K. Achievements, prospects and challenges in precision care for monogenic insulin-deficient and insulin-resistant diabetes. Diabetologia 65, 1782–1795 (2022). https://doi.org/10.1007/s00125-022-05720-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00125-022-05720-7