Abstract
Key message
A novel QTL, qCIR9.1 , that controls callus induction rate in anther culture was identified on chromosome 9 in rice, and based on RNA-seq data, Os09g0551600 was the most promising candidate gene.
Abstract
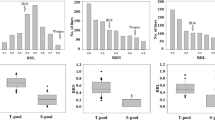
Anther culture, a doubled haploid (DH) technique, has become an important technology in many plant-breeding programmes. Although anther culturability is the key factor in this technique, its genetic mechanisms in rice remain poorly understood. In this study, we mapped quantitative trait loci (QTLs) responsible for anther culturability by using 192 recombinant inbred lines (RILs) derived from YZX (Oryza sativa ssp. indica) × 02428 (Oryza sativa ssp. japonica) and a high-density bin map. A total of eight QTLs for anther culturability were detected in three environments. Among these QTLs, a novel major QTL for callus induction rate (CIR) named qCIR9.1 was repeatedly mapped to a ~ 100 kb genomic interval on chromosome 9 and explained 8.39–14.14% of the phenotypic variation. Additionally, RNA sequencing (RNA-seq) was performed for the parents (YZX and 02428), low- (L-Pool) and high-CIR RILs (H-Pool) after 16 and 26 days of culture. By using the RNA of the bulked RILs for background normalization, the number of differentially expressed genes (DEGs) both between the parents and between the bulked RILs after 26 days of culture was drastically reduced to only 78. Among these DEGs, only one gene, Os09g0551600, encoding a high-mobility group (HMG) protein, was located in the candidate region of qCIR9.1. qRT-PCR analysis of Os09g0551600 showed the same results as RNA-seq, and the expression of this gene was decreased in the low-callus-induction parent (YZX) and L-Pool. Our results provide a foundational step for further cloning of qCIR9.1 and will be very useful for improving anther culturability in rice.







Similar content being viewed by others
References
Atanassov A, Zagorska N, Boyadjiev P, Djilianov D (1995) In vitro production of haploid plants. World J Microbiol Biotechnol 11:400–408. https://doi.org/10.1007/BF00364615
Bajaj YPS (1990) In vitro production of haploids and their use in cell genetics and plant breeding. In: Bajaj YPS (ed) Haploids in crop improvement I. Springer Berlin Heidelberg, Berlin, Heidelberg, pp 3–44. https://doi.org/10.1007/978-3-642-61499-6_1
Boutilier K, Offringa R, Sharma VK et al (2002) Ectopic expression of BABY BOOM triggers a conversion from vegetative to embryonic growth. Plant Cell 14:1737–1749. https://doi.org/10.1105/tpc.001941
Bregitzer P, Campbell R (2001) Genetic markers associated with green and albino plant regeneration from embryogenic barley callus. Crop Sci. https://doi.org/10.2135/cropsci2001.411173x
Chen T, Dent SYR (2014) Chromatin modifiers and remodellers: regulators of cellular differentiation. Nat Rev Genet 15:93–106. https://doi.org/10.1038/nrg3607
Chen L, Gao W, Chen S et al (2016) High-resolution QTL mapping for grain appearance traits and co-localization of chalkiness-associated differentially expressed candidate genes in rice. Rice (N Y) 9:48–48. https://doi.org/10.1186/s12284-016-0121-6
Chu C, Wang C, Sun C et al (1975) Establishment of an efficient medium for anther culture of rice through comparative experiments on the nitrogen sources. Sci Sin 18:659–668
Datta SJCS (2005) Androgenic haploids: factors controlling development and its application in crop improvement. Curr Sci 89:1870–1878
de Ronne M, Labbé C, Lebreton A et al (2020) Integrated QTL mapping, gene expression and nucleotide variation analyses to investigate complex quantitative traits: a case study with the soybean–Phytophthora sojae interaction. Plant Biotechnol J 18:1492–1494. https://doi.org/10.1111/pbi.13301
Debernardi JM, Tricoli DM, Ercoli MF et al (2020) A GRF–GIF chimeric protein improves the regeneration efficiency of transgenic plants. Nat Biotechnol 38:1274–1279. https://doi.org/10.1038/s41587-020-0703-0
Foroughi-Wehr B, Friedt W, Wenzel G (1982) On the genetic improvement of androgenetic haploid formation in Hordeum vulgare L. Theor Appl Genet 62:233–239. https://doi.org/10.1007/BF00276246
Germanà MA (2011a) Anther culture for haploid and doubled haploid production. Plant Cell Tiss Org (PCTOC) 104:283–300. https://doi.org/10.1007/s11240-010-9852-z
Germanà MA (2011b) Gametic embryogenesis and haploid technology as valuable support to plant breeding. Plant Cell Rep 30:839–857. https://doi.org/10.1007/s00299-011-1061-7
He P, Shen L, Lu C et al (1998) Analysis of quantitative trait loci which contribute to anther culturability in rice (Oryza sativa L.). Mol Breed 4:165–172. https://doi.org/10.1023/A:1009692221152
Hu L, Yang X, Yuan D et al (2011) GhHmgB3 deficiency deregulates proliferation and differentiation of cells during somatic embryogenesis in cotton. Plant Biotechnol J 9:1038–1048. https://doi.org/10.1111/j.1467-7652.2011.00617.x
Jan A, Kitano H, Matsumoto H, Komatsu S (2006) The rice OsGAE1 is a novel gibberellin-regulated gene and involved in rice growth. Plant Mol Biol 62:439–452. https://doi.org/10.1007/s11103-006-9030-1
Khanday I, Skinner D, Yang B et al (2019) A male-expressed rice embryogenic trigger redirected for asexual propagation through seeds. Nature 565:91–95. https://doi.org/10.1038/s41586-018-0785-8
Kong J, Martin-Ortigosa S, Finer J et al (2020) Overexpression of the transcription factor GROWTH-REGULATING FACTOR5 improves transformation of dicot and monocot Species. Front Plant Sci 11:572319–572319. https://doi.org/10.3389/fpls.2020.572319
Kumaravel M, Uma S, Backiyarani S et al (2020) Proteomic analysis of somatic embryo development in Musa spp. cv. Grand Naine (AAA). Sci Rep 10:4501. https://doi.org/10.1038/s41598-020-61005-2
Li S, Yan S, Wang Ah et al (2013) Identification of QTLs associated with tissue culture response through sequencing-based genotyping of RILs derived from 93–11 × Nipponbare in rice (Oryza sativa). Plant Cell Rep 32:103–116. https://doi.org/10.1007/s00299-012-1345-6
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCT method. Methods 25:402–408. https://doi.org/10.1006/meth.2001.1262
Love MI, Huber W, Anders S (2014) Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol 15:550. https://doi.org/10.1186/s13059-014-0550-8
Lu K, Li T, He J et al (2017) qPrimerDB: a thermodynamics-based gene-specific qPCR primer database for 147 organisms. Nucleic Acids Res 46:D1229–D1236. https://doi.org/10.1093/nar/gkx725
Luo G, Palmgren M (2020) GRF-GIF chimeras boost plant regeneration. Trends Plant Sci. https://doi.org/10.1016/j.tplants.2020.12.001E
Ma J, Deng M, Lv S-Y et al (2016) Identification of QTLs associated with tissue culture response of mature wheat embryos. Springerplus 5:1552–1552. https://doi.org/10.1186/s40064-016-3241-y
Mallik R, Kundu A, Chaudhuri S (2018) High mobility group proteins: the multifaceted regulators of chromatin dynamics. The Nucleus 61:213–226. https://doi.org/10.1007/s13237-018-0257-4
McCouch SR, Cgsnl, (2008) Gene nomenclature system for rice. Rice 1:72–84. https://doi.org/10.1007/s12284-008-9004-9
Mishra S, Sanyal I, Amla DV (2012) Changes in protein pattern during different developmental stages of somatic embryos in chickpea. Biol Plantarum 56:613–619. https://doi.org/10.1007/s10535-012-0124-0
Moleri S, Cappellano G, Gaudenzi G et al (2011) The HMGB protein gene family in zebrafish: evolution and embryonic expression patterns. Gene Expr Patterns 11:3–11. https://doi.org/10.1016/j.gep.2010.08.006
Niizeki H, Oono K (1968) Induction of haploid rice plant from anther culture. Proc Jpn Acad 44:554–557. https://doi.org/10.2183/pjab1945.44.554
Nishimura A, Ashikari M, Lin S et al (2005) Isolation of a rice regeneration quantitative trait loci gene and its application to transformation systems. Proc Natl Acad Sci USA 102:11940. https://doi.org/10.1073/pnas.0504220102
Pandit A, Rai V, Bal S et al (2010) Combining QTL mapping and transcriptome profiling of bulked RILs for identification of functional polymorphism for salt tolerance genes in rice (Oryza sativa L.). Mol Genet Genom MGG 284:121–136. https://doi.org/10.1007/s00438-010-0551-6
Park M, Lee J-H, Han K et al (2019) A major QTL and candidate genes for capsaicinoid biosynthesis in the pericarp of Capsicum chinense revealed using QTL-seq and RNA-seq. Theor Appl Genet 132:515–529. https://doi.org/10.1007/s00122-018-3238-8
Peng H, Wang K, Chen Z et al (2019) MBKbase for rice: an integrated omics knowledgebase for molecular breeding in rice. Nucleic Acids Res 48:D1085–D1092. https://doi.org/10.1093/nar/gkz921
Rajcan I, Boersma JG, Shaw EJ (2011) 4.11—Plant genetic techniques: plant breeder’s toolbox. In: Moo-Young M (ed) Comprehensive biotechnology, 2nd ed. Academic Press, Burlington, pp 133–147. https://doi.org/10.1016/B978-0-08-088504-9.00252-X
Smykal P (2000) Pollen embryogenesis—the stress mediated switch from gametophytic to sporophytic development. Current status and future prospects. Biol Plantarum 43:481–489. https://doi.org/10.1023/A:1002835330799
Suvorova G (2016) Chapter twenty nine—buckwheat tissue cultures and genetic transformation. In: Zhou M, Kreft I, Woo S-H, Chrungoo N, Wieslander G (eds) Molecular breeding and nutritional aspects of buckwheat. Academic Press, pp 365–375. https://doi.org/10.1016/B978-0-12-803692-1.00029-8
Taguchi-Shiobara F, Lin SY, Tanno K et al (1997) Mapping quantitative trait loci associated with regeneration ability of seed callus in rice, Oryza sativa L. Theor Appl Genet 95:828–833. https://doi.org/10.1007/s001220050632
Taguchi-Shiobara F, Yamamoto T, Yano M et al (2006) Mapping QTLs that control the performance of rice tissue culture and evaluation of derived near-isogenic lines. Theor Appl Genet 112:968–976. https://doi.org/10.1007/s00122-005-0200-3
Takeuchi Y, Abe T, Sasahara T (2000) RFLP Mapping of QTLs influencing shoot regeneration from mature seed-derived calli in rice. Crop Sci 40:245–247. https://doi.org/10.2135/cropsci2000.401245x
Tang K, Sun X, He Y et al (1998) Anther culture response of wild Oryza species. Plant Breed 117:443–446. https://doi.org/10.1111/j.1439-0523.1998.tb01970.x
Tsay SS, Tsay HS, Chao CY (1986) Cytochemical studies of callus development from microspore in cultured anther of rice. Plant Cell Rep 5:119–123. https://doi.org/10.1007/BF00269249
Wan X, Wu S, Li Z et al (2020) Lipid metabolism: critical roles in male fertility and other aspects of reproductive development in plants. Mol Plant 13:955–983. https://doi.org/10.1016/j.molp.2020.05.009
Wang M, van Bergen S, Van Duijn B (2000) Insights into a key developmental switch and its importance for efficient plant breeding. Plant Physiol 124:523. https://doi.org/10.1104/pp.124.2.523
Wang L, Xie W, Chen Y et al (2010) A dynamic gene expression atlas covering the entire life cycle of rice. Plant J 61:752–766. https://doi.org/10.1111/j.1365-313X.2009.04100.x
Wang C-C, Yu H, Huang J et al (2020) Towards a deeper haplotype mining of complex traits in rice with RFGB v2.0. Plant Biotechnol J 18:14–16. https://doi.org/10.1111/pbi.13215
Zhang D, Liang W, Yin C, Zong J, Gu F, Zhang D (2010) OsC6, encoding a lipid transfer protein, is required for postmeiotic anther development in rice. Plant Physiol 154:149–162. https://doi.org/10.1104/pp.110.158865
Zhang K, Su J, Xu M et al (2020) A common wild rice-derived BOC1 allele reduces callus browning in indica rice transformation. Nat Commun 11:443. https://doi.org/10.1038/s41467-019-14265-0
Zhang J, Guo T, Yang J et al (2020) QTL mapping and haplotype analysis revealed candidate genes for grain thickness in rice (Oryza sativa L.). Mol Breed 40:50. https://doi.org/10.1007/s11032-020-01128-6
Zhao L, Zhou H, Lu L et al (2009) Identification of quantitative trait loci controlling rice mature seed culturability using chromosomal segment substitution lines. Plant Cell Rep 28:247–256. https://doi.org/10.1007/s00299-008-0641-7
Zhao H, Yao W, Ouyang Y et al (2014) RiceVarMap: a comprehensive database of rice genomic variations. Nucleic Acids Res 43:D1018–D1022. https://doi.org/10.1093/nar/gku894
Acknowledgements
We are grateful for the financial support for this work by the Research and Development Plan for Key Areas in Guangdong province (No. 2018B020206002) and the earmarked fund for Modern Agro-Industry Technology Research System of the Ministry of Agriculture and Rural Affairs of the People’s Republic of China (No. CARS-01-17)
Author information
Authors and Affiliations
Contributions
TG and HW designed and supervised the study. CH performed the experiments (phenotypic assessments and qRT-PCR) and drafted the manuscript in Chinese, JZ, CH and LS analysed the data, JZ helpd to analysis (RNA-seq and haplotype analysis) and interpret the results, and write, translate and improve the manuscript, CH, DZ and YH participated in anther culture experiments and phenotypic assessments, ZC, GY and WL provided constructive comments, and all authors read and approved the manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Communicated by Mingliang Xu.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Huang, C., Zhang, J., Zhou, D. et al. Identification and candidate gene screening of qCIR9.1, a novel QTL associated with anther culturability in rice (Oryza sativa L.). Theor Appl Genet 134, 2097–2111 (2021). https://doi.org/10.1007/s00122-021-03808-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00122-021-03808-z