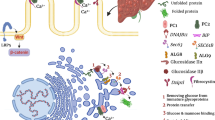
Abstract
Autosomal-recessive polycystic kidney disease (ARPKD; MIM #263200) is a severe, hereditary, hepato-renal fibrocystic disorder that causes early childhood morbidity and mortality. Mutations in the polycystic kidney and hepatic disease 1 (PKHD1) gene, which encodes the protein fibrocystin/polyductin complex (FPC), cause all typical forms of ARPKD. Several mouse lines carrying diverse, genetically engineered disruptions in the orthologous Pkhd1 gene have been generated, but none expresses the classic ARPKD renal phenotype. In the current study, we characterized a spontaneous mouse Pkhd1 mutation that is transmitted as a recessive trait and causes cysticliver (cyli), similar to the hepato-biliary disease in ARPKD, but which is exacerbated by age, sex, and parity. We mapped the mutation to Chromosome 1 and determined that an insertion/deletion mutation causes a frameshift within Pkhd1 exon 48, which is predicted to result in a premature termination codon (UGA). Pkhd1cyli/cyli (cyli) mice exhibit a severe liver pathology but lack renal disease. Further analysis revealed that several alternatively spliced Pkhd1 mRNA, all containing exon 48, were expressed in cyli kidneys, but in lower abundance than in wild-type kidneys, suggesting that these transcripts escaped from nonsense-mediated decay (NMD). We identified an AAAAAT motif in exon 48 upstream of the cyli mutation which could enable ribosomal frameshifting, thus potentially allowing production of sufficient amounts of FPC for renoprotection. This mechanism, expressed in a species-specific fashion, may help explain the disparities in the renal phenotype observed between Pkhd1 mutant mice and patients with PKHD1-related disease.
Key messages
-
The Pkhd1cyli/cyli mouse expresses cystic liver disease, but no kidney phenotype.
-
Pkhd1 mRNA expression is decreased in cyli liver and kidneys compared to wild-type.
-
Ribosomal frameshifting may be responsible for Pkhd1 mRNA escape from NMD.
-
Pkhd1 mRNA escape from NMD could contribute to the absent kidney phenotype.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Autosomal-recessive polycystic kidney disease (ARPKD; MIM #263200) is a hereditary hepato-renal fibrocystic disorder with an estimated incidence of 1 in 26,500 live births [1]. In most patients, ARPKD is characterized by the cystic dilatation of renal collecting ducts, causing progressive renal insufficiency and ultimately end-stage kidney disease [2, 3]. In the liver, the ductal plate malformation gives rise to congenital hepatic fibrosis, which often leads to portal hypertension [2, 3]. Virtually, all cases of typical ARPKD are caused by mutations within the polycystic kidney and hepatic disease 1 (PKHD1) gene located on chromosome 6p21.1 [4,5,6]. The full-length PKHD1 transcript is composed of 67 exons with the longest open reading frame (ORF) encoding a 4074 amino acid protein called fibrocystin or fibrocystin/polyductin complex (FPC) [4, 5]. Despite the passage of two decades since the identification of PKHD1 as the genetic determinant of ARPKD, the function of FPC remains poorly understood beyond its suggested roles in E3 ubiquitin ligase complex and SRC/STAT3 signaling [7, 8].
Several orthologous mouse models of ARPKD have been described (Table 1), primarily generated through random mutagenesis or targeted genetic engineering of the Pkhd1 gene [9,10,11,12,13,14,15,16]. Virtually, all mutant Pkhd1 mice exhibit a liver phenotype resembling human disease, but renal cystic disease is either absent or very mild and slowly progressive [3]. The mouse Pkhd1 locus, located on Chromosome 1qA3-4, consists of 67 non-overlapping exons encoding a protein of 4059 amino acids [17]. Human and mouse FPC share 87% overall identity across domains encompassing a predicted N-terminal signal peptide, multiple immunoglobulin-like plexin domains, multiple parallel β-helix 1 repeats, and a single transmembrane domain. In contrast, the short C-terminal cytoplasmic domains of mouse and human FPC are only 40% identical [17, 18].
In mice, in addition to the full-length Pkhd1 mRNA, multiple alternatively spliced Pkhd1 transcripts have been reported [17,18,19], whereas in humans, PKHD1 appears to be less transcriptionally complex [20]. Our previous work identified 22 alternative Pkhd1 transcripts in wild-type/normal (WT) mice [17, 18]. This is consistent with the presence of several intronic and exonic splicing enhancers (ISE, ESE) that are likely to play roles in the canonical as well as alternative splicing of the Pkhd1 mRNA [18]. In comparison, human PKHD1 is much less transcriptionally complex, with only 2 validated alternatively spliced isoforms (https://www.ncbi.nlm.nih.gov/gene/5314).
Here, we report the identification of a new, spontaneously occurring Pkhd1 mutation, that causes cysticliver (cyli), similar to the hepato-biliary disease in ARPKD. We describe the liver phenotype of cyli mice, mapping of the cyli mutation to the Pkhd1 gene, and comparative studies of Pkhd1 transcript profiles and abundance in WT and cyli mice. The phenotype of Pkhd1cyli/cyli (cyli) mice is similar to other Pkhd1 mutant models with liver cystic disease and no/minimal renal involvement. The insertion/deletion (indel) mutation in exon 48 results in a frameshift and a predicted premature termination codon (PTC) (UGA). In characterizing the Pkhd1-derived transcripts, we noted multiple differences in WT and cyli kidney-derived mRNAs, but observed exon 48 containing transcripts from both WT and mutant kidneys.
Materials and methods
Mice
All mouse experiments were approved by the Institutional Animal Care and Use Committees at The Jackson Laboratory, University of Alabama at Birmingham (UAB) and Children’s National Research Institute (CNRI). The study was conducted in accordance with relevant guidelines and regulations in the Guide for the Care and Use of Laboratory Animals of the National Institutes of Health. The Jackson Laboratory, UAB, and the Comparative Medicine Unit (previously named Research Animal Facility) at Children’s National Medical Center are fully accredited by the AAALAC.
The D.B/11Ei congenic inbred mouse strain was generated by introgression of a segment of distal Chromosome 4 from C57BL/6J (B6) onto the DBA/2J background. The first affected mouse noted was a 5-month-old (22-week-old) D.B/11Ei female breeder, generation N11F13. This female had successfully raised two litters, was pregnant with a third litter, and appeared sick. Further investigation revealed a hard, distended abdomen containing an enlarged liver with yellow, fluid-filled cysts. Liver disease was not evident in the male breeder. Offspring from this pair and closely related mice were monitored for signs of liver disease. These offspring were used to establish the D.B/11Ei strain carrying the recessive cyli allele and the results reported here are derived from the original breeding pair.
The D.B/11Ei-cyli strain used in this study was first transferred to UAB and subsequently to Children’s National Research Institute (CNRI). Because affected mice survive into adulthood and are capable of reproducing, the cyli mutation can be maintained using homozygous breeding of the D.B/11Ei-cyli strain.
Locus mapping, gene identification, and mutation sequencing
A standard backcross mating scheme was used to identify the chromosomal location of the cyli gene [21,22,23]. F1 females, produced by mating B6 to D.B/11Ei-cyli mice, were backcrossed to D.B/11Ei-cyli males. The backcross offspring (n = 221) was evaluated at 22 weeks of age for the presence of cystic liver disease. An initial genetic variant mapping approach and subsequent fine-mapping studies were performed using MIT microsatellite markers [24]. The introgressed B6 segment from Chromosome 4 was excluded as a candidate disease interval. A disease-associated interval that contained the Pkhd1 locus was identified on Chromosome 1. DNA sequencing of Pkhd1 exons and flanking intronic sequences were amplified by PCR and the amplicons sequenced in both directions using primer sets (Supplementary Table 1) designed based on the published Pkhd1 gene sequence [17].
Mouse genotyping
DNA for genotyping was isolated from biopsied tail tissue. Tissue was lysed at 55 °C in Cell Lysis Solution (Qiagen, # 158116) containing Proteinase K (Qiagen, # 19133), followed by protein precipitation with the Protein Precipitation Solution (Qiagen, # 158126) for 10 min at −20 °C. Sample was then centrifuged at 16,000 × g for 10 min at 4 °C. Genomic DNA was precipitated from the supernatant with ethanol, pelleted by centrifugation at 16,000 × g for 5 min at 4 °C, air dried, and resuspended in water. PCR-based genotyping was performed using primers 5′-TGG CTA TAC TGT GAA GAC CAG GCA-3′ (forward) and 5′-AAG CTT GGG CCT ATC TGA ATG GCA-3′ (reverse) and the following conditions: 15 min at 95 °C initial denaturation, followed by 35 cycles of 45-s denaturation at 94 °C, 45-s annealing at 52 °C, and 1-min extension at 72 °C, with a 10-min final extension at 72 °C. PCR products were digested with BsaI, and the products were resolved by agarose gel electrophoresis. Bands of 126 bp and 359 bp were diagnostic of the WT gene and a 484 bp band identified the cyli mutant allele.
Tissue histology and morphometric analysis
Kidneys and livers were harvested from male and female WT and cyli mice, paraffin embedded, sectioned, deparaffinized, rehydrated, and stained with hematoxylin and eosin (H&E) according to standard protocols [25,26,27,28]. Stained tissue sections were examined by light microscopy using an Olympus CX41 microscope equipped with a Leica DX320 color camera and Leica software. Histomorphometry was performed on blinded experimental specimens by a veterinary pathologist according to previously described protocols [29]. Images were collected using a Nikon E600 microscope equipped with a SPOT Insight digital camera (Diagnostic Instruments) and analyzed using Image-Pro Plus v6.2 image analysis software (Media Cybernetics Inc). Cyst and tissue areas were quantified by converting images to grayscale and thresholding them to produce a black image on a white background. Cysts were represented as white objects within the image. Cystic and total (including cysts) areas were determined automatically using the count/size and macro functions of Image-Pro Plus. The results were expressed as % of cyst area relative to total area.
For the initial evaluation of disease course in D.B/11Ei-cyli line, mice were weaned at 3 weeks of age and assigned to a specific age group for timed tissue harvesting. To investigate liver disease progression in females as a function of parity, 3 sib-mated pairs were assigned for tissue harvesting after the birth of their first litter, and 3 sib-mated pairs were assigned for tissue harvesting after the birth of their second litter.
Reverse transcription (RT)–PCR and quantitative (q)RT-PCR
Total RNA samples from kidneys and livers harvested from 4.5- and 7-week-old WT and cyli mice were prepared using RNeasy Mini kit (Qiagen, # 74104), treated with RQ1 RNase-Free DNase (Promega, # M6101), and then re-purified using the RNeasy Mini kit. For RT-PCR, RNA samples were reverse transcribed using SuperScript III First-Strand Synthesis SuperMix (Thermo Fisher Scientific, # 18080400) and oligo dT primers. RT-PCR was performed to compare Pkhd1 transcripts in WT and cyli kidney and liver tissue using primers specific for Pkhd1 exons 1 (forward: 5′-CAT TTG AGG CAC AAG GCT GAC ACA-3′) and 67 (reverse: 5′-CTG AGG TCT GGG CGT AAC AG-3′) sequences. Relative Pkhd1 transcript abundance in WT vs. cyli kidneys was determined by quantitative real-time PCR performed on a QuantStudio 7 Flex Real-Time PCR System (Thermo Fisher Scientific) using the default program. PCR was performed with cDNA templates using Power SYBR Green PCR Master Mix (Thermo Fisher Scientific, # 4368706) and primers specific for sequences of Pkhd1 exons 2–5 (forward: 5′-ATG ATG CTT GCC TGG CTG GTC-3′; reverse: 5′-TAT GGC CCT GCA TCT GCT TCT GAT-3′), Pkhd1 exons 5–6 (forward: 5′-GTC TCT TCC ATC AGA AGC AGA TGC-3′; reverse: 5′-GGG TAA ACT TGA TAT AAA ACA GG-3′), Pkhd1 exons 37 (forward: 5′-CAT GGA TCC AGG ACC CAT TG-3′; reverse: 5′-AGG GAA AGA AGG GAG TGG AA-3′), Pkhd1 exons 48–49 (forward: 5′-TGG CTA TAC TGT GAA GAC CA-3′; reverse: 5′-GAT CCA AGA GCA GAG CCA TC-3′), Pkhd1 exons 61–62 (forward: 5′-TCA CTC TTG AGA TGC CTG GC-3′; reverse: 5′-AGG TTC CCA GTT ATT AAA CTA C-3′), and Pkhd1 exons 66–67 (forward: 5′-CCA GAA GAC ATA TCT GAA TCC CAG GC-3′; reverse: 5′-AGC AAG AGA TCC TGG AAC ACA GGT-3′). Peptidylprolyl Isomerase A (Ppia) was used for normalization (forward: 5′-AGC ACT GGA GAG AAA GGA TT-3′; reverse: 5′-ATT ATG GCG TGT AAA GTC ACC A-3′) [30]. Results were analyzed using QuantStudio Real-Time PCR Software and the ΔΔCt method [31]. The results were graphed with GraphPad Prism version 9.1.2 for Windows, GraphPad Software, San Diego, CA, USA, www.graphpad.com. Expression levels in cyli mice were normalized to their respective WT values. Comparison of expression levels in cyli mice was compared to a constant value of 1.0 using a two-sided Wilcoxon sign rank test.
Immunoblotting
Kidneys were collected from 2-week-old WT and cyli mice and immediately snap frozen in liquid nitrogen. Kidneys were homogenized on ice for 20 s in 1mL ice-cold RIPA buffer (Sigma-Aldrich, # R0278) containing proteinase inhibitors (Protease Inhibitor Cocktail Mini-Tablet EDTA-free, Bimake, # B14012). Homogenates were centrifuged for 10 min at 15,000 × g at 4 °C. Protein concentration in supernatants was determined using BCA protein assay kit (Thermo Scientific, # 23227). Twenty micrograms of total protein was mixed with NuPAGE LDS sample buffer (Life Technologies, # NP0007) containing sample reducing agent (Life Technologies, # NP0009). Samples were heated at 100 °C for 10 min prior to electrophoresis through a Novex NuPAGE 4–12% Bis-Tris gel (Life Technologies, # NP0335BOX) in MES SDS running buffer (Life Technologies, # NP0002) for 30 min at 200 V. Proteins were transferred to a polyvinylidene fluoride membrane using Bio-Rad Trans-Blot Turbo Transfer System. The membrane was incubated with rat anti-mouse FPC monoclonal antibody PD1E1, which was obtained from the Baltimore PKD Center [16], in 1× PBS plus 0.1% Tween-20 (PBST) with 5% bovine serum albumin overnight at 4 °C. The membrane was washed 3 times 10 min with 1× PBST, then incubated with goat anti-rat secondary antibody (Thermo Fisher Scientific, # 31475) 1:5000 dilution in 1× PBST with 5% non-fat dry milk for 1 h at room temperature, followed by 3 washes with 1× PBST. Immunoreactive bands were detected using SuperSignal West Dura chemiluminescent substrate (Thermo Fisher Scientific, # 34076) and imaged using Bio-Rad ChemiDoc MP Imaging System. Densitometry of FPC bands was performed with Bio-Rad Image Lab Software 6.1 and normalized to GAPDH. The relative intensity was calculated with WT as 1.00. The result was graphed with GraphPad Prism version 9.1.2 for Windows. Two-tailed Student’s t-test was used for statistical analysis.
Targeted long-read cDNA sequencing
Approximately 1μg of PCR-amplified kidney cDNA from 7-week-old WT and cyli mice was used to perform nanopore-based long-read sequencing. SQK-LSK109 Amplicon sequencing kit [Oxford Nanopore Technologies (ONT)] was used for library preparation. Briefly, DNA repair and end-prep were accomplished with NEBNext FFPE DNA Repair and Ultra II End-Prep reaction and enzyme mixes (New England Biolabs). AMPure XP beads were then used to bind DNA and prepared for adapter ligation with NEBNext Quick t4 DNA ligase and adapter mix (ONT). Long fragment buffer was used to select for reads greater than 3 kb in size. The resultant DNA was primed and loaded onto the ONT Flongle flowcell. Each sample was run for 24 h, generating approximately 300 Mbp of raw reads. Quality estimation and visualization of the raw fastq files were performed using NanoPlot and nanoQC functions of the nanoPack suite of tools [32]. The fastq reads were then mapped to the mouse reference genome (GRCm39) (https://www.ncbi.nlm.nih.gov/assembly/GCF_000001635.27/) using minimap2 [33]. To estimate the read coverage per exon, the aligned bam files were first sorted and indexed using samtools [34], followed by read coverage estimation per kilobase using bamCoverage [35]. The output bed files were then read and merged with sample metadata using LoadTrackFile function of ggcoverage [36], followed by mapping to the 67 exon locations using a custom R script. Integrative Genomics Viewer (IGV) and Adobe Illustrator were used to visualize Pkhd1 splicing patterns in WT and cyli kidneys after exclusion of reads shorter than 1 kb.
Results
Discovery of the cyli mutant mouse
Investigation of a sick D.B/11Ei female mouse revealed an enlarged liver containing multiple fluid-filled cysts. Subsequently, we determined that this disorder was transmitted as a recessive trait. Homozygous cyli mutant mice developed cystic liver disease around 4 weeks of age. Histopathological analysis demonstrated biliary dysgenesis characterized by ductal plate malformation phenocopying the liver lesion characteristic of human ARPKD, with portal tracts of affected livers exhibiting multiple irregularly shaped and variably dilated bile ducts lined with hyperplastic epithelium (Fig. 1A). However, there was no histopathologic evidence of renal cystic disease in cyli mutants examined between 8 and 11 weeks of age (Fig. 1A).
Initial characterization of the cyli mouse model. A Kidney (top) and liver (middle and bottom) tissue sections stained with hematoxylin and eosin (H&E) from 8-week-old WT and cyli mice. Bottom panels are higher magnification views of the boxed areas in middle panels (BD, bile duct; C, cortex; M, medulla; PV, portal vein). B Schematic illustrating the position of the cyli mutation (affecting Pkhd1) on Chromosome 1 between the genetic markers D1Mit168 and D1Mit231 (genetic distances are in centiMorgan (cM) units). C Sequence comparison of Pkhd1 from WT and cyli mice (NM_153179.3: c.7588_7589delGGinsT (p.G2530VfsTer15). D Comparison of WT and cyli reading frames. The cyli mutation leads to the formation of a PTC that is predicted to cause premature protein truncation
Gene identification and mutation analysis
The disease-causing locus was positioned to Chromosome 1 between markers D1Mit168 and D1Mit231 (D1Mit168 - 0.9 cM – cyli - 1.3 cM – D1Mit231), the region containing the Pkhd1 gene (Fig. 1B). Sequence analysis of cyli mice identified an indel mutation NM_153179.3:c.7588_7589delGGinsT (p.G2530VfsTer15) in Pkhd1 exon 48 (Fig. 1C) leading to a frameshift and predicted PTC within exon 48, 44 bp downstream of the T insertion (Fig. 1D).
Progressive liver disease modulated by sex, age, and parity
Morphometric analysis of liver sections from female and male cyli mice at multiple time points revealed an age-associated increase in the severity of liver disease. Younger cyli mice (less than 11 weeks old) displayed a liver phenotype characterized by dilated bile ducts radiating from the portal region into the parenchyma (Fig. 2 and Table 2). By week 14, mice displayed coalescing cysts progressively replacing areas of the liver parenchyma. Beyond 20 weeks of age, female mice tended to exhibit more extensive cystic lesions than male mice of comparable age (Table 2). Comparing liver cyst formation at 20–26 weeks of age between mice with two or three pregnancies (P2, P3) suggested that in addition to age and sex, parity may also contribute to the severity of the cystic liver disorder (Fig 2, left, bottom two images and Table 2). The study did not extend phenotypic examination of mice beyond 26 weeks of age or 3 pregnancies. A detailed longitudinal comparison of age-matched nulliparous female mice (separated from male littermates prior to the first pregnancy) to female mice with specific number of pregnancies was beyond the scope of the current project.
Progressive cystic liver phenotype in cyli mutant female mice. Representative hematoxylin and eosin (H&E) stained liver sections from female cyli mice between 4 and 26 weeks of age demonstrate progressive liver cyst formation and severely reduced typical liver parenchyma. P0 (10–11 weeks), P1 (14–16 weeks), P2 (20–26 weeks), and P3 (20–26 weeks) denote numbers of litters produced and the age of cyli females (left). The significantly larger cysts in P3, 20–26 weeks old mice (bottom left), suggest that in addition to age, parity also contributes to the severity of the disorder. Age-matched WT liver sections are shown on the right. Scale bar = 200 µm
Differential Pkhd1 transcript profiles in WT vs. cyli mice
We previously demonstrated that WT Pkhd1 mRNA is subject to extensive alternative splicing in the kidney but not in the liver and that the longer transcripts typically include exon 48, where the cyli indel mutation occurs [18]. To evaluate whether the cyli mutation altered the alternative splicing products, we compared Pkhd1 transcripts from WT and cyli kidneys as well as the corresponding livers (Fig. 3A). The Pkhd1 cDNA was amplified using primers specific for Pkhd1 exons 1 and 67. Consistent with previously published findings [18], we identified four major products of 12, 6.5, 4.5, and 2.5 kb, as well as some additional minor bands, representing the full-length and major alternatively spliced Pkhd1 mRNAs from WT kidneys (Fig. 3A, lane 3) [18]. In WT liver, we observed a 12 kb product (Fig. 3A, lane 4) while in cyli liver, we identified a 12 kb and a 4.5 kb band (Fig. 3A, lane 2). As in previous studies, we were able to reproducibly amplify a 4.5 kb product from WT liver RNA [18], but the 4.5 kb band was not reproducibly detectable in liver RNA from cyli mice. This variability in detection may reflect the very low transcript abundance in total liver RNA as Pkhd1 is expressed only in cholangiocytes which represent approximately 5% of the total liver cellular mass [37].
Pkhd1 expression in WT and cyli mice. A Pkhd1 transcript (mRNA) profiles amplified from total kidney and liver RNAs of 7-week-old WT and cyli male mice. PCR products (amplicons) were generated from oligo dT primed template cDNA using primers specific for Pkhd1 exons 1 and 67. In lane 9 (Plasmid), the band at 2000 bp represents either a non-specific or plasmid recombination-derived amplification product. B Visualization of Pkhd1 mRNA splicing patterns in the kidneys of 7-week-old WT and cyli mice, respectively. Gray – forward and reverse RNA-seq reads spanning adjacent/consecutive Pkhd1 exons. Cyan – forward and reverse RNA-seq reads spanning alternatively spliced Pkhd1 exons in WT kidneys. Purple – forward and reverse RNA-seq reads spanning alternatively spliced Pkhd1 exons in cyli kidneys. Arrows indicate the direction of transcription. Exons 10, 20, 30, 40, 50, and 60 are indicated in red. Exon 48 is shown in orange. C Relative expression of Pkhd1 mRNA containing exons 2–5, 5–6, 37, 48–49, 61–62, and 66–67 in the kidneys and liver of 4.5-week-old WT and cyli female mice. Data were normalized to Ppia mRNA; expression of Pkhd1 mRNA in WT mouse kidneys (left y-axis) and liver (right y-axis) was set as 1.00. Data were expressed as mean ± S.E; n=3 or 5 per group. D Immunoblot of FPC protein in kidney lysates from 2-week-old WT and cyli male mice using a rat monoclonal primary antibody specific for Pkhd1 exon 67-encoded amino acid sequences [16]. Data were expressed as mean ± SEM; n=3 mice per group
Next, we used ONT long-read sequencing technology to examine the splicing patterns of Pkhd1 transcripts in WT and cyli kidneys. Abundance of Pkhd1 transcripts in the liver was too low for ONT sequencing. Distribution of read lengths from sequencing WT Pkhd1 transcripts showed maxima at approximately 2.5, 4.5, 6.5, and 12 kb (Supplementary Fig. S1) which corresponded to the sizes of WT Pkhd1 cDNA amplification products (Fig. 3A). Distribution of read lengths from sequencing cyli Pkhd1 transcripts had maxima at approximately 2.5 and 4.5 kb, and included long reads that corresponded to full-length Pkhd1 transcripts (Supplementary Fig. S1). Again, these data correspond to sizes of cyli Pkhd1 cDNA amplification products (Fig. 3A). Visualization of Pkhd1 splicing patterns showed the presence of transcripts containing all 67 exons in both WT and cyli kidneys (Fig. 3B). Splicing of Pkhd1 exons 6 and 37 has been observed previously in the kidneys from WT mice [18]. Other alternative slicing events in WT Pkhd1 transcripts are novel (Fig. 3B) and underscore the complex pattern of the Pkhd1 splicing in mouse kidneys. While patterns of alternatively spliced Pkhd1 exons differed in WT and cyli kidneys, exon 48 was present in the cyli Pkhd1 transcripts, indicating that full-length FPC could be translated in mutant kidneys (Fig. 3B).
We then used qRT-PCR and primer-pairs specific for junctions spanning exons 2–5, 5–6, 37, 48–49, 61–62, and 66–67 to examine differential amplicon profiles and confirm the presence of exon 48 in the population of cyli derived mRNAs. We observed significantly lower levels of amplification products in both WT and cyli livers compared to kidneys for all targeted amplicons (Fig. 3C, right y-axis for the kidney and left y-axis for the liver). The abundance of amplicons spanning exon 37, which is alternatively spliced in WT kidneys [18], was lower in cyli than in the WT kidneys and livers (Fig. 3C). The abundance of amplicons spanning exons 48–49, 61–62, and 66–67 was lower in cyli than in the WT kidneys (Fig. 3C). Unlike in kidneys, the differences in abundance of amplicons spanning exons 48–49, 61–62, and 66–67 were not statistically significant in the liver samples, perhaps because Pkhd1 transcripts were in very low abundance due to their expression in cholangiocytes. Lower abundance of amplicons spanning exons 48, 61–62, and 66–67 in cyli than in the WT kidneys and livers suggests that a fraction of Pkhd1 transcripts in cyli kidneys escape NMD.
In addition, we investigated expression of FPC in WT and cyli kidneys and livers using an antibody generated against an exon 67–encoded epitope [16]. We detected full-length FPC in WT kidney protein extracts (Fig. 3D). In contrast, full-length FPC was below the detection limit in extracts prepared from cyli kidneys (Fig. 3D). Full-length FPC was not detected in lysates from either WT or cyli livers (Supplementary Fig. S2). Again, this is likely due to the low abundance of FPC-expressing cholangiocytes in the liver compared to hepatocytes, which do not express FPC. The lower (20–171 kD) molecular weight bands observed in lysates from both WT and mutant livers at the same intensity likely represent non-specific cross-reacting proteins (Supplementary Fig. S2).
Discussion
The mouse Pkhd1cyli mutation arose spontaneously due to a de novo indel mutation, NM_153179.3:c.7588_7589delGGinsT, in exon 48 of the Pkhd1 gene, which causes a frameshift and results in a predicted PTC. Similar to other Pkhd1 mutant models, cyli mice express a hepato-biliary phenotype that progresses with age. Our longitudinal analysis revealed that female sex and pregnancy exacerbate liver disease severity. This sexual dimorphism in cyli-related liver disease severity has not previously been reported for human PKHD1-related disease or in other genetically engineered Pkhd1 mouse mutants, possibly due to the paucity of longitudinal studies in both human patients and previously reported mouse models. Our data suggests that, at least in cyli mutants, hepato-biliary disease progression may be hormonally modulated, as has been observed in human ADPKD and mouse Pkd1 models [38]. Further studies will be required to explore this intriguing pathogenic mechanism in both human patients and the multiple Pkhd1 mouse models.
The most striking feature of all mouse Pkhd1 models described to date is the absent or minimal renal cystic phenotype [9,10,11,12,13,14, 16, 19]. In contrast, truncating human PKHD1 mutations typically cause severe renal cystic disease during fetal life [3]. In the current study, we show that cyli mice with an indel mutation in exon 48 do not express renal cystic disease when aged to at least 26 weeks. However, patients with frameshift mutations involving PKHD1 exon 48 express the classic ARPKD phenotype with severe renal cystic disease expressed in infancy (http://www.humgen.rwth-aachen.de) [39, 40].
The mouse Pkhd1 gene, unlike its human orthologue, is transcriptionally complex in the WT kidney with a number of alternatively spliced isoforms. In cyli mutant kidneys, the majority of the transcripts were 2.5 and 4.5 kb. In our previous study, we demonstrated that these lower molecular weight amplicons contain isoforms that result from exon-skipping events, but maintain the FPC open reading frame (ORF) [18]. For example, the 2.5 kb amplicon contains an isoform resulting exon 6–61 splicing. Similarly, among the transcripts in the 4.5 kb amplicon, some of the products originate from exon 4–49 splicing and exon 6 can be spliced to exons 51, 52, or 53 as well (Supplementary Fig. S3). The long-read sequencing of Pkhd1 transcripts from WT kidneys detected previously observed splicing of exons 6 and 37 [18] in both WT and cyli kidneys. Therefore, a diverse set of alternative Pkhd1 splice variants observed in the cyli kidneys could encode multiple novel isoforms of FPC with sufficient residual function to attenuate cystogenesis in mutant kidneys. In contrast, Pkhd1 in the WT liver has minimal transcriptional complexity and the full-length Pkhd1 transcript is predominant [4, 18]. Therefore, the cyli liver with limited alternatively spliced Pkhd1 mRNAs would not have functional redundancy, resulting in the development of the hepato-biliary lesion.
Since the transcripts from the cyli kidney did not fully exclude exon 48, we sought mechanisms that could explain this apparent evasion from NMD. Ribosomal frameshifting, which controls polyamine biosynthesis in mammalian cells [41], is one such mechanism. Indeed, sequence analysis of exon 48 revealed an AAAAAT motif upstream of the cyli indel. This motif has been shown to promote ribosome shifting on the reading frame by +1 or −1 [42]. For example, if ribosomes shift −1, instead of reading GGG, coding for Gly2530, the ribosome would read TGT, coding for Cys, and proceed with in-frame translation of the FPC sequence.
At the protein level, immunoblotting detected only full-length FPC in WT kidneys. In contrast, bands corresponding to full-length FPC were not observed in immunoblots of cyli mouse kidney protein extracts. This finding is consistent with the reduced levels of Pkhd1 gene expression demonstrated by qRT-PCR and our previous analysis [18] and suggests the possibility that variant transcripts and/or ribosomal frameshifting give rise to very low abundance isoforms of functional FPC, which are capable of abrogating cystogenesis in this model.
While the basis for PKHD1/Pkhd1 species–specific differences in renal disease expression is not understood, differences in the phenotypic severity between orthologous human disease and mouse models are not uncommon. For example, the mdx mouse model of Duchenne muscular dystrophy (DMD) has a mild phenotype compared to human DMD patients [43]. Although mdx mice exhibit muscle histopathology, elevated plasma pyruvate kinase and creatine kinase levels, and muscle weakness similar to DMD patients, mdx mice are viable and fertile [43], whereas human DMD is a fatal degenerative muscle disorder [44]. Similarly, mouse models of cystic fibrosis (CF) also do not fully recapitulate the human disease (for review: [45]). The numerous mouse CF models exhibit defects in fluid secretion and develop severe intestinal and mild pancreatic disease, but fail to develop the signature lung infections that are the major cause of mortality in human CF [46,47,48,49]. The basis for this species-specific difference in lung phenotype appears to involve anatomical differences between the mouse and human airways [50] as well as pH differences in the airway surface liquid in human CF patients vs. mouse models [40].
We speculate that the minimal renal cystic disease in mouse Pkhd1 models reflects a combination of mechanisms. Based on our data in this specific model, ribosomal frameshifting may allow alternatively spliced mutant transcripts to escape NMD. More broadly, variations in molecular interactions which may be dictated by genetic background rather than Pkhd1 genotype per se might act to modulate the degree to which a renal cystic phenotype is expressed in mouse models [11, 51, 52]. Finally, we speculate that FPC functions differently in species-specific molecular pathways. Informatic analyses indicate that the short C-terminal cytoplasmic domains of mouse and human FPC are only 40% identical [17, 18], raising the possibility that these protein domains have different intracellular binding partners and participate in different molecular pathways. To this point, we note that mice homozygous for the Pkhd1 exon 67 deletion (Pkhd1del67), which removes most of the FPC carboxy terminus domain, have no renal or biliary phenotype [16] and mice lacking virtually the entire Pkhd1 locus (exons 3 through 67) express the hepato-biliary lesion, but only a minor renal phenotype in older mice [19, 53]. In contrast, the corresponding defects in human patients, which involvines loss of a functional carboxy terminus [39, 54] or large intragenic deletions [55, 56] are associated with both the hepato-biliary lesion and severe renal cystic disease.
As noted above for CF, identifying specific mechanisms that underlie the discordant phenotypes between human genetic disease and orthologous mouse mutant models may yield valuable insights into potential treatment strategies [40]. Although the physiological function of FPC remains undefined, we anticipate that continued study of Pkhd1 mutant mouse models will expand our understanding of the mechanism(s) underlying mouse resistance to the severe renal disease that characterizes human ARPKD. Defining such mechanisms, in turn, could yield potential new targets for therapies that mitigate the full expression of this devastating human disease.
Data availability
The datasets generated during and/or analyzed during the current study are included in this manuscript and its supplementary information files.
References
Alzarka B, Morizono H, Bollman JW, Kim D, Guay-Woodford LM (2017) Design and implementation of the hepatorenal fibrocystic disease core center clinical database: a centralized resource for characterizing autosomal recessive polycystic kidney disease and other hepatorenal fibrocystic diseases. Frontiers in pediatrics 5:80. https://doi.org/10.3389/fped.2017.00080
Guay-Woodford LM, Bissler JJ, Braun MC, Bockenhauer D, Cadnapaphornchai MA, Dell KM, Kerecuk L, Liebau MC, Alonso-Peclet MH, Shneider B et al (2014) Consensus expert recommendations for the diagnosis and management of autosomal recessive polycystic kidney disease: report of an international conference. The Journal of pediatrics 165:611–617. https://doi.org/10.1016/j.jpeds.2014.06.015
Bergmann C, Guay-Woodford LM, Harris PC, Horie S, Peters DJM, Torres VE (2018) Polycystic kidney disease. Nat Rev Dis Primers 4:50. https://doi.org/10.1038/s41572-018-0047-y
Onuchic LF, Furu L, Nagasawa Y, Hou X, Eggermann T, Ren Z, Bergmann C, Senderek J, Esquivel E, Zeltner R et al (2002) PKHD1, the polycystic kidney and hepatic disease 1 gene, encodes a novel large protein containing multiple immunoglobulin-like plexin-transcription-factor domains and parallel beta-helix 1 repeats. American journal of human genetics 70:1305–1317. https://doi.org/10.1086/340448
Ward CJ, Hogan MC, Rossetti S, Walker D, Sneddon T, Wang X, Kubly V, Cunningham JM, Bacallao R, Ishibashi M et al (2002) The gene mutated in autosomal recessive polycystic kidney disease encodes a large, receptor-like protein. Nature genetics 30:259–269. https://doi.org/10.1038/ng833
Xiong H, Chen Y, Yi Y, Tsuchiya K, Moeckel G, Cheung J, Liang D, Tham K, Xu X, Chen XZ et al (2002) A novel gene encoding a TIG multiple domain protein is a positional candidate for autosomal recessive polycystic kidney disease. Genomics 80:96–104. https://doi.org/10.1006/geno.2002.6802
Kaimori JY, Lin CC, Outeda P, Garcia-Gonzalez MA, Menezes LF, Hartung EA, Li A, Wu G, Fujita H, Sato Y et al (2017) NEDD4-family E3 ligase dysfunction due to PKHD1/Pkhd1 defects suggests a mechanistic model for ARPKD pathobiology. Scientific reports 7:7733. https://doi.org/10.1038/s41598-017-08284-4
Dafinger C, Mandel AM, Braun A, Gobel H, Burgmaier K, Massella L, Mastrangelo A, Dotsch J, Benzing T, Weimbs T et al (2020) The carboxy-terminus of the human ARPKD protein fibrocystin can control STAT3 signalling by regulating SRC-activation. Journal of cellular and molecular medicine 24:14633–14638. https://doi.org/10.1111/jcmm.16014
Moser M, Matthiesen S, Kirfel J, Schorle H, Bergmann C, Senderek J, Rudnik-Schoneborn S, Zerres K, Buettner R (2005) A mouse model for cystic biliary dysgenesis in autosomal recessive polycystic kidney disease (ARPKD). Hepatology (Baltimore, Md) 41:1113–1121
Woollard JR, Punyashtiti R, Richardson S, Masyuk TV, Whelan S, Huang BQ, Lager DJ, vanDeursen J, Torres VE, Gattone VH et al (2007) A mouse model of autosomal recessive polycystic kidney disease with biliary duct and proximal tubule dilatation. Kidney international 72:328–336. https://doi.org/10.1038/sj.ki.5002294
Garcia-Gonzalez MA, Menezes LF, Piontek KB, Kaimori J, Huso DL, Watnick T, Onuchic LF, Guay-Woodford LM, Germino GG (2007) Genetic interaction studies link autosomal dominant and recessive polycystic kidney disease in a common pathway. Human molecular genetics 16:1940–1950. https://doi.org/10.1093/hmg/ddm141
Gallagher AR, Esquivel EL, Briere TS, Tian X, Mitobe M, Menezes LF, Markowitz GS, Jain D, Onuchic LF, Somlo S (2008) Biliary and pancreatic dysgenesis in mice harboring a mutation in Pkhd1. The American journal of pathology 172:417–429. https://doi.org/10.2353/ajpath.2008.070381
Kim I, Fu Y, Hui K, Moeckel G, Mai W, Li C, Liang D, Zhao P, Ma J, Chen XZ et al (2008) Fibrocystin/polyductin modulates renal tubular formation by regulating polycystin-2 expression and function. J Am Soc Nephrol 19:455–468. https://doi.org/10.1681/ASN.2007070770
Williams SS, Cobo-Stark P, James LR, Somlo S, Igarashi P (2008) Kidney cysts, pancreatic cysts, and biliary disease in a mouse model of autosomal recessive polycystic kidney disease. Pediatric nephrology (Berlin, Germany) 23:733–741. https://doi.org/10.1007/s00467-007-0735-4
Hu B, He X, Li A, Qiu Q, Li C, Liang D, Zhao P, Ma J, Coffey RJ, Zhan Q et al (2011) Cystogenesis in ARPKD results from increased apoptosis in collecting duct epithelial cells of Pkhd1 mutant kidneys. Experimental cell research 317:173–187. https://doi.org/10.1016/j.yexcr.2010.09.012
Outeda P, Menezes L, Hartung EA, Bridges S, Zhou F, Zhu X, Xu H, Huang Q, Yao Q, Qian F et al (2017) A novel model of autosomal recessive polycystic kidney questions the role of the fibrocystin C-terminus in disease mechanism. Kidney international 92:1130–1144. https://doi.org/10.1016/j.kint.2017.04.027
Nagasawa Y, Matthiesen S, Onuchic LF, Hou X, Bergmann C, Esquivel E, Senderek J, Ren Z, Zeltner R, Furu L et al (2002) Identification and characterization of Pkhd1, the mouse orthologue of the human ARPKD gene. J Am Soc Nephrol 13:2246–2258
Boddu R, Yang C, O’Connor AK, Hendrickson RC, Boone B, Cui X, Garcia-Gonzalez M, Igarashi P, Onuchic LF, Germino GG et al (2014) Intragenic motifs regulate the transcriptional complexity of Pkhd1/PKHD1. J Mol Med (Berl) 92:1045–1056. https://doi.org/10.1007/s00109-014-1185-7
Bakeberg JL, Tammachote R, Woollard JR, Hogan MC, Tuan HF, Li M, van Deursen JM, Wu Y, Huang BQ, Torres VE et al (2011) Epitope-tagged Pkhd1 tracks the processing, secretion, and localization of fibrocystin. J Am Soc Nephrol 22:2266–2277. https://doi.org/10.1681/ASN.2010111173
Frank V, Zerres K, Bergmann C (2014) Transcriptional complexity in autosomal recessive polycystic kidney disease. Clinical journal of the American Society of Nephrology : CJASN 9:1729–1736. https://doi.org/10.2215/cjn.00920114
Guay-Woodford LM, Bryda EC, Christine B, Lindsey JR, Collier WR, Avner ED, D’Eustachio P, Flaherty L (1996) Evidence that two phenotypically distinct mouse PKD mutations, bpk and jcpk, are allelic. Kidney international 50:1158–1165
Guay-Woodford LM, Wright CJ, Walz G, Churchill GA (2000) Quantitative trait loci modulate renal cystic disease severity in the mouse bpk model. J Am Soc Nephrol 11:1253–1260
Mrug M, Zhou J, Yang C, Aronow BJ, Cui X, Schoeb TR, Siegal GP, Yoder BK, Guay-Woodford LM (2015) Genetic and informatic analyses implicate Kif12 as a candidate gene within the Mpkd2 locus that modulates renal cystic disease severity in the Cys1cpk mouse. PloS one 10: e0135678. https://doi.org/10.1371/journal.pone.0135678
Rhodes M, Straw R, Fernando S, Evans A, Lacey T, Dearlove A, Greystrong J, Walker J, Watson P, Weston P et al (1998) A high-resolution microsatellite map of the mouse genome. Genome Res 8:531–542. https://doi.org/10.1101/gr.8.5.531
Mrug M, Zhou J, Woo Y, Cui X, Szalai AJ, Novak J, Churchill GA, Guay-Woodford LM (2008) Overexpression of innate immune response genes in a model of recessive polycystic kidney disease. Kidney international 73:63–76. https://doi.org/10.1038/sj.ki.5002627
Chase TH, Cox GA, Burzenski L, Foreman O, Shultz LD (2009) Dysferlin deficiency and the development of cardiomyopathy in a mouse model of limb-girdle muscular dystrophy 2B. Am J Pathol 175:2299–2308. https://doi.org/10.2353/ajpath.2009.080930
Zhou J, Ouyang X, Cui X, Schoeb TR, Smythies LE, Johnson MR, Guay-Woodford LM, Chapman AB, Mrug M (2010) Renal CD14 expression correlates with the progression of cystic kidney disease. Kidney international 78:550–560. https://doi.org/10.1038/ki.2010.175
Shan D, Rezonzew G, Mullen S, Roye R, Zhou J, Chumley P, Revell DZ, Challa A, Kim H, Lockhart ME et al (2019) Heterozygous Pkhd1(C642*) mice develop cystic liver disease and proximal tubule ectasia that mimics radiographic signs of medullary sponge kidney. American journal of physiology Renal physiology 316:F463–F472. https://doi.org/10.1152/ajprenal.00181.2018
Zhou J, Ouyang X, Schoeb TR, Bolisetty S, Cui X, Mrug S, Yoder BK, Johnson MR, Szalai AJ, Mrug M (2012) Kidney injury accelerates cystogenesis via pathways modulated by heme oxygenase and complement. J Am Soc Nephrol 23:1161–1171. https://doi.org/10.1681/asn.2011050442
Arensdorf AM, Rutkowski DT (2013) Endoplasmic reticulum stress impairs IL-4/IL-13 signaling through C/EBPβ-mediated transcriptional suppression. J Cell Sci 126:4026–4036. https://doi.org/10.1242/jcs.130757
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods 25:402–408. https://doi.org/10.1006/meth.2001.1262
De Coster W, D’Hert S, Schultz DT, Cruts M, Van Broeckhoven C (2018) NanoPack: visualizing and processing long-read sequencing data. Bioinformatics 34:2666–2669. https://doi.org/10.1093/bioinformatics/bty149
Li H (2018) Minimap2: pairwise alignment for nucleotide sequences. Bioinformatics 34:3094–3100. https://doi.org/10.1093/bioinformatics/bty191
Li H, Handsaker B, Wysoker A, Fennell T, Ruan J, Homer N, Marth G, Abecasis G, Durbin R, Genome Project Data Processing S (2009) The Sequence Alignment/Map format and SAMtools. Bioinformatics 25:2078–2079. https://doi.org/10.1093/bioinformatics/btp352
Ramírez F, Ryan DP, Grüning B, Bhardwaj V, Kilpert F, Richter AS, Heyne S, Dündar F, Manke T (2016) deepTools2: a next generation web server for deep-sequencing data analysis. Nucleic Acids Res 44:W160-165. https://doi.org/10.1093/nar/gkw257
Song Y, Wang J (2022) <em>ggcoverage</em>: an R package to visualize and annotate genome coverage for various NGS data. bioRxiv: 2022.2009.2001.503744. https://doi.org/10.1101/2022.09.01.503744
Banales JM, Huebert RC, Karlsen T, Strazzabosco M, LaRusso NF, Gores GJ (2019) Cholangiocyte pathobiology. Nat Rev Gastroenterol Hepatol 16:269–281. https://doi.org/10.1038/s41575-019-0125-y
Chebib FT, Jung Y, Heyer CM, Irazabal MV, Hogan MC, Harris PC, Torres VE, El-Zoghby ZM (2016) Effect of genotype on the severity and volume progression of polycystic liver disease in autosomal dominant polycystic kidney disease. Nephrology, dialysis, transplantation : official publication of the European Dialysis and Transplant Association - European Renal Association 31:952–960. https://doi.org/10.1093/ndt/gfw008
Sharp AM, Messiaen LM, Page G, Antignac C, Gubler MC, Onuchic LF, Somlo S, Germino GG, Guay-Woodford LM (2005) Comprehensive genomic analysis of PKHD1 mutations in ARPKD cohorts. J Med Genet 42:336–349. https://doi.org/10.1136/jmg.2004.024489
Shah VS, Meyerholz DK, Tang XX, Reznikov L, Abou Alaiwa M, Ernst SE, Karp PH, Wohlford-Lenane CL, Heilmann KP, Leidinger MR et al (2016) Airway acidification initiates host defense abnormalities in cystic fibrosis mice. Science (New York, NY 351: 503-507. https://doi.org/10.1126/science.aad5589
Bekaert M, Ivanov IP, Atkins JF, Baranov PV (2008) Ornithine decarboxylase antizyme finder (OAF): fast and reliable detection of antizymes with frameshifts in mRNAs. BMC Bioinformatics 9:178. https://doi.org/10.1186/1471-2105-9-178
Ketteler R (2012) On programmed ribosomal frameshifting: the alternative proteomes. Front Genet 3:242. https://doi.org/10.3389/fgene.2012.00242
Bulfield G, Siller WG, Wight PA, Moore KJ (1984) X chromosome-linked muscular dystrophy (mdx) in the mouse. Proc Natl Acad Sci U S A 81:1189–1192. https://doi.org/10.1073/pnas.81.4.1189
Emery AEH (1993) Duchenne Muscular Dystrophy Oxford University Press, Oxford
McCarron A, Donnelley M, Parsons D (2018) Airway disease phenotypes in animal models of cystic fibrosis. Respir Res 19:54. https://doi.org/10.1186/s12931-018-0750-y
Guilbault C, Saeed Z, Downey GP, Radzioch D (2007) Cystic fibrosis mouse models. Am J Respir Cell Mol Biol 36:1–7. https://doi.org/10.1165/rcmb.2006-0184TR
Fisher JT, Zhang Y, Engelhardt JF (2011) Comparative biology of cystic fibrosis animal models. Methods Mol Biol 742:311–334. https://doi.org/10.1007/978-1-61779-120-8_19
Lavelle GM, White MM, Browne N, McElvaney NG, Reeves EP (2016) Animal models of cystic fibrosis pathology: phenotypic parallels and divergences. Biomed Res Int 2016:5258727. https://doi.org/10.1155/2016/5258727
Semaniakou A, Croll RP, Chappe V (2018) Animal models in the pathophysiology of cystic fibrosis. Front Pharmacol 9:1475. https://doi.org/10.3389/fphar.2018.01475
Keiser NW, Engelhardt JF (2011) New animal models of cystic fibrosis: what are they teaching us? Curr Opin Pulm Med 17:478–483. https://doi.org/10.1097/MCP.0b013e32834b14c9
Lager DJ, Qian Q, Bengal RJ, Ishibashi M, Torres VE (2001) The pck rat: a new model that resembles human autosomal dominant polycystic kidney and liver disease. Kidney international 59:126–136. https://doi.org/10.1046/j.1523-1755.2001.00473.x
O’Meara CC, Hoffman M, Sweeney WE Jr, Tsaih SW, Xiao B, Jacob HJ, Avner ED, Moreno C (2012) Role of genetic modifiers in an orthologous rat model of ARPKD. Physiological genomics 44:741–753. https://doi.org/10.1152/physiolgenomics.00187.2011
Olson RJ, Hopp K, Wells H, Smith JM, Furtado J, Constans MM, Escobar DL, Geurts AM, Torres VE, Harris PC (2019) Synergistic genetic interactions between Pkhd1 and Pkd1 result in an ARPKD-like phenotype in murine models. J Am Soc Nephrol 30:2113–2127. https://doi.org/10.1681/ASN.2019020150
Burgmaier K, Brinker L, Erger F, Beck BB, Benz MR, Bergmann C, Boyer O, Collard L, Dafinger C, Fila M et al (2021) Refining genotype-phenotype correlations in 304 patients with autosomal recessive polycystic kidney disease and PKHD1 gene variants. Kidney Int 100:650–659. https://doi.org/10.1016/j.kint.2021.04.019
Bergmann C, Senderek J, Windelen E, Kupper F, Middeldorf I, Schneider F, Dornia C, Rudnik-Schoneborn S, Konrad M, Schmitt CP et al (2005) Clinical consequences of PKHD1 mutations in 164 patients with autosomal-recessive polycystic kidney disease (ARPKD). Kidney international 67:829–848. https://doi.org/10.1111/j.1523-1755.2005.00148.x
Zvereff V, Yao S, Ramsey J, Mikhail FM, Vijzelaar R, Messiaen L (2010) Identification of PKHD1 multiexon deletions using multiplex ligation-dependent probe amplification and quantitative polymerase chain reaction. Genetic testing and molecular biomarkers 14:505–510. https://doi.org/10.1089/gtmb.2009.0188
Acknowledgements
The authors thank the members of the Guay-Woodford laboratory for helpful advice. Maryanne C. Odinakachukwu (Center for Translational Research) assisted with the mouse phenotyping, genotyping, and colony maintenance. Adam M. Richman (Center for Translational Research) and Amber K. O’Connor (akoWriting LLC) provided editorial assistance. We thank Trenton R. Schoeb, DVM, PhD (UAB Comparative Pathology Laboratory) for assisting with the histopathological analysis. The authors thank the members of the Eicher laboratory, including Linda L. Washburn, for the help throughout the project, Lisa Somes for maintaining the D.B/11Ei-cyli strain and providing mice for pathology studies, and Leona Gagnon and Andrew Rechnagle for isolating and establishing DNA plates for mapping. In addition, we thank Douglas McMinimy from The Jackson Laboratory for conducting a genome scan to determine the chromosomal position of cyli.
Funding
This work was supported by NIH grants RR01183 and GM20919 to EME; NIH grant DK121530 and a gift from the Moran Family Foundation to LGW.
Author information
Authors and Affiliations
Contributions
All authors contributed to the study conception and design. Eva Eicher identified the founder mutant mouse, determined the trait inheritance, described the initial phenotype, and directed the genetic mapping. Subsequent material preparation, data collection, and analysis were performed by Chaozhe Yang, Naoe Harafuji, Weiying Yu, Ravindra Boddu, Zsuzsa Bebok, and Oded Foreman. The data analysis of the ONT study was performed by Surajit Bhattacharya, Hayk Barseghyan, and Ljubica Caldovic. Heather Gordish-Dressman performed the statistical analysis. The first draft of the manuscript was written by Ravindra Boddu, Ljubica Caldovic, and Naoe Harafuji. Ljubica Caldovic, Eva M. Eicher, and Lisa M. Guay-Woodford revised the manuscript. All authors commented on previous versions of the manuscript. All authors also read and approved the final manuscript.
Corresponding author
Ethics declarations
Ethics approval
All mouse experiments were approved by the Institutional Animal Care and Use Committees at the Jackson Laboratory, University of Alabama at Birmingham (UAB) and Children’s National Research Institute. The study was conducted in accordance with relevant guidelines and regulations in the Guide for the Care and Use of Laboratory Animals of the National Institutes of Health. The Jackson Laboratory, UAB, and the Comparative Medicine Unit (previously named Research Animal Facility) at Children’s National Medical Center are fully accredited by the AAALAC.
Competing interests
The authors declare no competing interests.
Additional information
Communicating author: Naoe Harafuji
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Yang, C., Harafuji, N., Caldovic, L. et al. Pkhd1cyli/cyli mice have altered renal Pkhd1 mRNA processing and hormonally sensitive liver disease. J Mol Med 101, 1141–1151 (2023). https://doi.org/10.1007/s00109-023-02351-2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00109-023-02351-2