Abstract
Transition from hormone-sensitive to hormone-refractory metastatic tumor types poses a major challenge for prostate cancer treatment. Tumor antigens that are differentially expressed during this transition are likely to play important roles in imparting prostate cancer cells with the ability to grow in a hormone-deprived environment and to metastasize to distal sites such as the bone and thus, are likely targets for therapeutic intervention. To identify those molecules and particularly cell surface antigens that accompany this transition, we studied the changes in cell surface antigenic profiles between a hormone-sensitive prostate cancer line LNCaP and its hormone-refractory derivative C4-2B, using an antibody library-based affinity proteomic approach. We selected a naïve phage antibody display library to identify human single-chain antibodies that bind specifically to C4-2B but not LNCaP. Using mass spectrometry, we identified one of the antibody-targeted antigens as the ICAM-1/CD54/human rhinovirus receptor. Recombinant IgG1 derived from this single-chain antibody binds to a neutralizing epitope of ICAM-1 and blocks C4-2B cell invasion through extracellular matrix in vitro. ICAM-1 is thus differentially expressed during the transition of the hormone-sensitive prostate cancer cell line LNCaP to its hormone-refractory derivative C4-2B, plays an important role in imparting the C4-2B line with the ability to invade, and may therefore be a target for therapeutic intervention.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
A major challenge of prostate cancer treatment is the transition from a hormone-sensitive localized form of the disease to a hormone-refractory metastatic tumor. Bone is one of the most common sites for prostate cancer metastasis [1, 2]. The exact reason for this bone-tropism is under active investigation and several molecules/pathways have been identified as involved in this process [3–8]. Despite this progress, a complete understanding of this behavior of prostate cancer cells is lacking.
We sought to identify cell surface antigens differentially expressed by a hormone-refractory metastatic prostate cancer cell line but not by an isotype-matched hormone-sensitive line with less metastatic potential. We hypothesized that those differentially expressed cell surface antigens may impart prostate cancer cells with the ability to thrive in a hormone-deprived environment and to colonize at distal organ sites such as the bone and may thus be targets for therapeutic intervention. As a model system for the transition, we studied the hormone-sensitive prostate cancer line LNCaP that does not metastasize to the bone and its hormone-refractory derivative C4-2B that metastasizes to the bone. The C4-2B line was derived from LNCaP following serial passages in vivo and selective recovery of cells from the bone [9, 10] and has been widely used to study hormone-sensitivity/refractory status and bone tropism of prostate cancer [8, 10–12].
We have previously developed and utilized a naïve phage antibody display library to identify internalizing human single-chain antibodies targeting several solid human tumors including prostate tumors [13–21]. We report in this study the use of the phage antibody display approach to probe the altered cell surface epitope space associated with the metastatic, hormone-refractory C4-2B line. We identified a panel of human scFvs that bind preferentially to the C4-2B cell line but not the parental LNCaP line, suggesting that a molecular signature exists that accompanies the transition of prostate cancer cells from a hormone-sensitive to a hormone-refractory bone metastatic form, and that this signature can be identified by our antibody library-based approach. Using mass spectrometric analysis, we identified ICAM-1/CD54/human rhinovirus receptor precursor as the target antigen of one of our C4-2B-targeting scFvs. We have further demonstrated that this antibody binds to a neutralizing epitope of ICAM-1 and that a human IgG1 derived from this scFv effectively blocks prostate cancer invasions in vitro through extracellular matrix components.
Materials and methods
Reagents
For scFv and IgG purification and characterization: CHO-S-SFM II media and G418 (Invitrogen, Carlsbad, CA, USA), 1 L spinner flask (Corning, Corning, NY, USA), CellgroTM Stirrers (Barnstead, Dubuque, IA, USA), HisTrapTMHP, HiTrap protein A HP and HiTrap desalting columns (GE Healthcare, Piscataway, NJ, USA). For FACS: streptavidin-phycoerythrin (SA-PE) (Invitrogen/BioSource, Camarillo, CA, USA), PE-conjugated anti-human Fc-specific mAb (Jackson ImmunoResearch, West Grove, PA, USA), murine anti-ICAM-1 mAb (clone BBIG-l1) (R&D System, Minneapolis, MN, USA), and biotin-labeled rabbit anti-fd bacteriophage (Sigma-Aldrich, St. Louis, MO, USA). For immunoprecipitation: mammalian protease inhibitor cocktails (Sigma-Aldrich), NP40 and protein A agarose beads (Pierce, Rockford, IL, USA). For cell invasion assay: Matrigel Basement Membrane Mix (BD Biosciences, San Jose, CA, USA), Transwell (CorningCostar, Corning, NY, USA), and Diff-Quik (Dade Behring Inc, Newark, DE, USA).
Cell lines
LNCaP was purchased from American Type Culture Collection (ATCC) and C4-2B from DIANON/Urocor (Oklahoma City, OK, USA). The CHO-ICAM-1 line, a stably transfected CHO cell line expressing the full-length human ICAM-1 protein, was purchased from the ATCC. The CHO DG44 cell line (dhfr-) was purchased from Invitrogen. The BPH-1 line was originally obtained from Dr. Gerald Cunha [22] at UCSF and maintained in the lab.
Selection of C4-2B-specific phage antibodies
For counter-selection, a naïve library containing 500 million members was incubated with LNCaP cells at 4°C for 8 h, centrifuged to collect the supernatants, and the counter-selected library was further incubated with C4-2B cells at 37°C for 2 h. The cells were then washed three times with ice-cold RPMI media containing 0.5% fetal bovine serum (FBS), once with 100 mM glycine containing 150 mM NaCl, pH 2.8, and once with phosphate-buffered saline (PBS), and lysed with 300 µl 100 mM triethylamine at RT for 5 min, neutralized with 150 µl 1 M Tris–HCl, and used to infect exponentially growing TG1 bacteria. The infected TG1 were plated on LB-agar plates containing tetracycline (50 µg/ml) and incubated at 37°C overnight. Colonies on the plate were scraped off and used to inoculate 2× YT media containing tetracycline (50 µg/ml) at 37°C for 12 h, the supernatants were collected by centrifugation and phage prepared by polyethylene glycol precipitation as described [16–18]. The polyclonal phage antibodies were used as the input for the next round of selection using the same counter-selection then selection procedure described above.
Analysis of binding phage antibodies
FACS was used to screen for monoclonal phage antibodies that bind specifically to C4-2B but not LNCaP. Phage-infected bacteria grown on agar plates were picked into 96-well U bottom culture plates in 2× YT media containing tetracycline (50 µg/ml) and grown overnight at 37°C. After centrifugation, an identical amount of supernatants (50 µl per well per plate) containing phage particles were transferred into two V-bottom plates, and incubated with LNCaP and C4-2B cells, respectively, at RT for 1 h. Following washing with PBS, bound phage were detected by biotin-labeled anti-fd antibodies followed by streptavidin-phycoerythrin (SA-PE), using FACS LSRII (BD Biosciences, San Jose, CA, USA) as described [17, 18].
Recombinant human IgG1
The development of recombinant IgG1 from scFv has been described previously [23]. We have previously constructed a mammalian expression vector that produces recombinant fully human M10A12 IgG1 using genetic information of the M10A12 scFv [23]. CHO DG44 cells stably transfected with the M10A12 IgG1-expressing plasmid were grown in CHO-S-SFM II containing G418 in 1 L spinner flasks on CellgroTMStirrers as described [23]. Supernatants were collected and purified on a HiTrap Protein A column using an AKTAprime (GE Healthcare).
Identification of the target antigen as ICAM-1/CD54/rhinovirus receptor
We used the M10A12 IgG1 to immunoprecipitate the target antigen from C4-2B lysates, and analyzed the product by tandem mass spectrometry. Ten million tumor cells were harvested, washed, and pelleted in an Eppendorf microcentrifuge tube. Cell pellets were resuspended in cell lysis buffer (1% NP-40 in PBS, pH 7.2, with mammalian protease inhibitor cocktails added at v/v 1:200 per manufacturer’s instruction) and incubated on ice for 1 h. Following centrifugation in an Eppendorf 5417R at 10,000×g for 15 min, the supernatants were collected, pre-cleared by incubating with protein A beads on ice for 4 h, centrifuged at 10,000×g at 4°C for 15 min to remove the beads, divided into two parts, and further incubated with the M10A12 IgG1 and a control non-binding human IgG1 respectively on ice for 4 h. Immunoprecipitation products of both the M10A12 IgG1 and the control IgG1 were analyzed on a gradient SDS-PAGE gel (4–20%, Invitrogen), stained with coomassie blue and bands unique to the M10A12 IgG immunoprecipitation products were excised, digested with trypsin, and analyzed by liquid chromatography–tandem mass spectrometry (LC-MS/MS). Peptides were separated by reverse phase chromatography using an Ultimate HPLC (Dionex) and then analyzed on-line using a QSTAR Pulsar Mass Spectrometer (MDS Sciex/Applied Biosystems). Raw data was converted to peaklists using the Mascot dll in Analyst (version 1.6b16), then searched using Batch-Tag in Protein Prospector (version 5.0) [24] against mammalian proteins in the SwissProt Database (downloaded June 2008: 52,897 entries searched), allowing a precursor mass accuracy tolerance of 50 ppm and a fragment mass tolerance of 0.1 Da. Acceptance criteria was a peptide expectation value of less than 0.05. To confirm the identification, CHO (control) and CHO cells stably transfected with the full-length human ICAM-1 gene were incubated with the M10A12 IgG1 at RT for 30 min, washed three times with PBS/0.5% FBS, further incubated with PE-conjugated anti-human Fc antibodies, and then analyzed by FACS. As a further control for background staining, the experiment was repeated using a recombinant anti-botulinum toxin human IgG1, CR-2, which does not bind to prostate cancer cells.
Cell invasion assay
MatriGel basement membrane was used as the matrix for the cell invasion assay. About 2.5 × 105 cells were mixed with 50 µg/ml IgGs at 37°C for 1 h. In the meantime, MatriGel was dissolved in RPMI media at 4°C and placed on the top chamber (insert) at 37°C to solidify. The cells were placed on top of the MatriGel layer and incubated for 48 h. Cells remaining in the top layer of the chamber were removed. After Diff-Quik staining, viable cells that migrated to the lower layer of the chamber were counted under an inverted microscope (Nikon, Japan). The experiments were performed in triplicate and the data were analyzed using a student t test. A p value of less than 0.05 was used as indication of a significant difference.
Results
Selection of scFvs that bind specifically to C4-2B but not the parental LNCaP line
We first counter-selected a 500 million-member naïve phage antibody library on the parental LNCaP cells and then incubated the counter-selected library with the C4-2B cells. After three rounds of counter-selection and selection, we arrayed the output phage antibodies into 96-well plates, collected the supernatants containing monoclonal phage antibodies, and screened for phage antibodies that bind to C4-2B (target) but not LNCaP (control) cells. An example of specific binding phage is shown in Fig. 1. An anti-CD26 mAb was used as a positive control, as CD26 has been shown to be preferentially expressed by C4-2B cells by microarray analysis [8]. We identified a large number of phage antibodies that bound preferentially to C4-2B but not LNCaP (Fig. 2). Phage antibodies that showed specific binding to C4-2B cells were sequenced. Thirty-two unique phage antibodies were identified after screening about 400 clones from the output of the third round of selection. One of the scFvs was identical in sequence to the M10A12 scFv that we previously identified as binding to an unknown antigen expressed by prostate cancer lines PC3 and Du-145 [23]. For uniformity, in subsequent studies we will use the original name M10A12 to refer this scFv.
Identification of ICAM-1 as a C4-2B-associated cell surface antigen
We sought to identify the target antigen bound by the M10A12 antibody. We have previously converted the M10A12 scFv into a full-length human IgG1 [23]. We used the recombinant M10A12 IgG1 to immunoprecipitate prostate cancer cell lysates and analyzed the immunoprecipitation products by mass spectrometry. LC-MS/MS analysis identified two peptides that matched perfectly with human ICAM-1, also known as CD54 or human rhinovirus receptor, and an example of one of these matches is shown in Fig. 3. To confirm this identification, we studied binding patterns of the M10A12 IgG1 to CHO (control) and CHO cells stably transfected with a full-length human ICAM-1 cDNA. We found that the M10A12 antibody bound specifically to CHO-ICAM-1 but not CHO (Figs. 4a, b), further confirming that ICAM-1 is the target of the M10A12 antibody.
The M10A12 IgG1 binds to a neutralizing epitope of ICAM-1
To further study the ICAM-1 epitope bound by the M10A12 antibody, we performed a competition assay using a murine neutralizing anti-ICAM-1 mAb (clone BBIG-I1). This murine mAb targets an ICAM-1 epitope that is involved in ligand binding [25–27]. We found that this neutralizing murine mAb competed with the M10A12 IgG1 for binding to C4-2B cells (Figs. 5a, b), suggesting that the M10A12 IgG1 and the murine mAb bind to the same neutralizing epitope of ICAM-1.
The M10A12 IgG1 binds to a neutralizing epitope of ICAM-1. a Histograms of M10A12 IgG1 binding to C4-2B in the presence or absence of the competing murine neutralizing anti-ICAM-1 mAb (clone BBIG-l1). b MFI values are plotted for the competition experiment. The experiment was done in triplicate. Error bars indicate standard deviations. Control secondary antibodies only
The M10A12 IgG1 blocks C4-2B cell invasion through extracellular matrix components
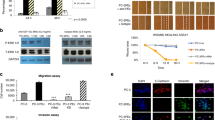
To study the functional consequences of antibody binding to this neutralizing ICAM-1 epitope, we performed an in vitro invasion assay using the C4-2B cells and studied if binding by the M10A12 IgG1 effectively blocks invasion of C4-2B cells through extracellular matrix components using the MatriGel-based in vitro invasion assay. C4-2B and LNCaP cells showed markedly different invasion ability through MatriGel (Fig. 6a). Interestingly, we found that the M10A12 IgG1 caused a significant inhibition of C4-2B invasion through MatriGel while the control IgG1 did not (Fig. 6b). The M10A12 IgG1 did not significantly affect the residual level of invasion activity of LNCaP (Fig. 6b), consistent with the lack of binding of M10A12 IgG1 by LNCaP cells.
Inhibition of C4-2B invasion through MatriGel in vitro by the M10A12 neutralizing anti-ICAM-1 human IgG1. a C4-2B has higher intrinsic invasion ability than LNCaP. The experiment was done in triplicate. P < 0.05 indicates significant difference between C4-2B and LNCaP. b Inhibition of C4-2B invasion by M10A12 IgG1. A non-binding human IgG1 (CR-2) was used as control. The experiment was done in triplicates and the P < 0.05 indicates a significant difference between M10A12- and control CR-2-treated C4-2B cells. There are no significant differences between M10A12- and CR-2-treated LNCaP cells, consistent with a lack of ICAM-1 expression by LNCaP cells
Discussion
To search for molecules that are selectively expressed during transition of prostate cancer cells from being hormone-sensitive to hormone-refractory with enhanced metastatic potential, we have taken an antibody library approach to identify cell surface differences between two isotype-matched prostate cancer cell lines, the parental hormone sensitive LNCaP line and its hormone refractory bone-metastasizing derivative C4-2B. This model system has been widely used by investigators to study prostate cancer development, affording an opportunity to identify meaningful cell surface changes accompanying prostate tumor progression. We have identified 32 unique human scFvs that bind selectively to C4-2B but not LNCaP cells. We have found that one of the C4-2B-targeting scFvs is identical in sequence to a previously identified scFv, M10A12. We used a full-length human IgG1 developed from the M10A12 scFv to immunoprecipitate tumor antigens from prostate cancer cell lysates and used mass spectrometric analysis to identify the target antigen as ICAM-1, also known as CD54 or human rhinovirus receptor. Furthermore, through a competition study with a known neutralizing murine mAb targeting human ICAM-1, we found that the M10A12 IgG1 binds to the same neutralizing epitope and inhibits invasion of the C4-2B cells through extracellular matrix. Taken together, these studies demonstrate that ICAM-1 is differentially expressed during the transition from the parental LNCaP cells to hormone refractory bone-metastasizing C4-2B cells and may play important roles in mediating prostate cancer invasion and metastasis.
Human ICAM gene family members have been implicated in prostate development and progression by genetic studies. A large genome-wide association study identified a loci on chromosome 19 that is associated with predisposition to breast and prostate cancer [28]. By high-density single nucleotide polymorphism mapping, the extent of association was found to span 20 kb and include the ICAM-1 gene and two other genes in the ICAM family [28]. A genome-wide characterization of gene expression variations and DNA copy number changes in prostate cancer cell lines also found that ICAM-1 is associated with an enhanced malignant potential [29]. Despite this genetic evidence and gene expression studies, exactly how ICAM-1 mediates prostate cancer progression remains undetermined. Our study suggests that ICAM-1 mediates invasive behavior of prostate cancer cells, and thus is likely to play an important role in tumor metastasis to distal sites.
We have developed a full-length human neutralizing mAb (M10A12 IgG1) that blocks C4-2B invasion through extracellular matrix components, making this antibody a potential candidate for therapeutic development. However, further studies are needed to evaluate if this neutralizing anti-ICAM-1 human IgG1 can block prostate cancer cell metastasis in vivo.
While we have demonstrated that ICAM-1 is differentially expressed by C4-2B cells and plays a key role in mediating invasion through extracellular matrix by these cells, we have not directly studied if ICAM-1 expression is sufficient to allow the hormone sensitive LNCaP cells to achieve hormone independence and metastasize to the bone. It also remains to be determined if ICAM-1 expression is necessary for C4-2B cells to maintain hormone independence and propensity to localize to the bone. Further studies with appropriate prostate cancer animal models are needed to address these issues.
ICAM-1 is also known as the receptor for human rhinovirus, a causative agent for the common cold. In addition, ICAM-1 is a part of the receptor complex for other Picornavirus family members such as the Coxsackievirus A21. Recently, researchers have exploited ICAM-1 expression on prostate cancer cell lines to achieve efficient infection by oncolytic viruses and observed potent tumor cell killing in vitro [30]. It would be interesting to see if oncolytic viruses can be developed into therapeutic agents to effectively kill in vivo metastatic hormone-refractory prostate tumors that express ICAM-1.
To summarize, our antibody library-based approach is effective in identifying antibodies targeting cell surface molecules that are functionally important for prostate cancer progression. These tumor-targeting scFvs and their derivatives such as IgGs are human in sequence, providing a direct route to the clinic. ICAM-1 has been identified as a cell surface antigen that is differentially expressed in the hormone-refractory metastatic prostate cancer cell line C4-2B, and a fully human neutralizing IgG1 blocks C4-2B invasion through extracellular matrix. In addition to ICAM-1, other antigens defined by this panel of scFvs remain to be identified. Using mass spectrometric analysis described in this report and other novel expression cloning strategies described elsewhere [31, 32], we are identifying additional antigens that play important roles in mediating various functions in advanced prostate tumors including hormone-independence and the propensity to metastasize to the bone and other distal sites.
Abbreviations
- FACS:
-
fluorescence activated cell sorting
- ICAM-1:
-
intercellular adhesion molecule 1
- mAb:
-
monoclonal antibody
- LC-MS/MS:
-
liquid chromatography–tandem mass spectrometry
- CHO:
-
Chinese hamster ovary
- SA-PE:
-
streptavidin-phycoerythrin
- mAb:
-
monoclonal antibody
- FBS:
-
fetal bovine serum
- MFI:
-
mean fluorescence intensity
References
Msaouel P, Pissimissis N, Halapas A, Koutsilieris M (2008) Mechanisms of bone metastasis in prostate cancer: clinical implications. Best Pract Res Clin Endocrinol Metab 22:341–355
Bussard KM, Gay CV, Mastro AM (2008) The bone microenvironment in metastasis; what is special about bone? Cancer Metastasis Rev 27:41–55
Dimitroff CJ, Lechpammer M, Long-Woodward D, Kutok JL (2004) Rolling of human bone-metastatic prostate tumor cells on human bone marrow endothelium under shear flow is mediated by E-selectin. Cancer Res 64:5261–5269
Fu Z, Dozmorov IM, Keller ET (2002) Osteoblasts produce soluble factors that induce a gene expression pattern in non-metastatic prostate cancer cells, similar to that found in bone metastatic prostate cancer cells. Prostate 51:10–20
Edlund M, Sung SY, Chung LW (2004) Modulation of prostate cancer growth in bone microenvironments. J Cell Biochem 91:686–705
Bonfil RD, Chinni S, Fridman R, Kim HR, Cher ML (2007) Proteases, growth factors, chemokines, and the microenvironment in prostate cancer bone metastasis. Urol Oncol 25:407–411
Taichman RS, Cooper C, Keller ET, Pienta KJ, Taichman NS, McCauley LK (2002) Use of the stromal cell-derived factor-1/CXCR4 pathway in prostate cancer metastasis to bone. Cancer Res 62:1832–1837
Liu AY, Brubaker KD, Goo YA, Quinn JE, Kral S, Sorensen CM, Vessella RL, Belldegrun AS, Hood LE (2004) Lineage relationship between LNCaP and LNCaP-derived prostate cancer cell lines. Prostate 60:98–108
Chen ME, Lin SH, Chung LW, Sikes RA (1998) Isolation and characterization of PAGE-1 and GAGE-7. New genes expressed in the LNCaP prostate cancer progression model that share homology with melanoma-associated antigens. J Biol Chem 273:17618–17625
Rubin J, Chung LW, Fan X, Zhu L, Murphy TC, Nanes MS, Rosen CJ (2004) Prostate carcinoma cells that have resided in bone have an upregulated IGF-I axis. Prostate 58:41–49
Lin DL, Tarnowski CP, Zhang J, Dai J, Rohn E, Patel AH, Morris MD, Keller ET (2001) Bone metastatic LNCaP-derivative C4-2B prostate cancer cell line mineralizes in vitro. Prostate 47:212–221
Romanov VI, Durand DB, Petrenko VA (2001) Phage display selection of peptides that affect prostate carcinoma cells attachment and invasion. Prostate 47:239–251
O’Connell D, Becerril B, Roy-Burman A, Daws M, Marks JD (2002) Phage versus phagemid libraries for generation of human monoclonal antibodies. J Mol Biol 321:49–56
Sheets MD, Amersdorfer P, Finnern R, Sargent P, Lindquist E, Schier R, Hemingsen G, Wong C, Gerhart JC, Marks JD (1998) Efficient construction of a large nonimmune phage antibody library: the production of high-affinity human single-chain antibodies to protein antigens. Proc Natl Acad Sci USA 95:6157–6162
Becerril B, Poul MA, Marks JD (1999) Toward selection of internalizing antibodies from phage libraries. Biochem Biophys Res Commun 255:386–393
Liu B, Conrad F, Cooperberg MR, Kirpotin DB, Marks JD (2004) Mapping tumor epitope space by direct selection of single-chain Fv antibody libraries on prostate cancer cells. Cancer Res 64:704–710
Ruan W, Sassoon A, An F, Simko JP, Liu B (2006) Identification of clinically significant tumor antigens by selecting phage antibody library on tumor cells in situ using laser capture microdissection. Mol Cell Proteomics 5:2364–2373
An F, Drummond DC, Wilson S, Kirpotin DB, Nishimura SL, Broaddus VC, Liu B (2008) Targeted drug delivery to mesothelioma cells using functionally selected internalizing human single-chain antibodies. Mol Cancer Ther 7:569–578
Goenaga AL, Zhou Y, Legay C, Bougherara H, Huang L, Liu B, Drummond DC, Kirpotin DB, Auclair C, Marks JD, Poul MA (2007) Identification and characterization of tumor antigens by using antibody phage display and intrabody strategies. Mol Immunol 44:3777–3788
Roth A, Drummond DC, Conrad F, Hayes ME, Kirpotin DB, Benz CC, Marks JD, Liu B (2007) Anti-CD166 single chain antibody-mediated intracellular delivery of liposomal drugs to prostate cancer cells. Mol Cancer Ther 6:2737–2746
Poul MA, Becerril B, Nielsen UB, Morisson P, Marks JD (2000) Selection of tumor-specific internalizing human antibodies from phage libraries. J Mol Biol 301:1149–1161
Hayward SW, Dahiya R, Cunha GR, Bartek J, Deshpande N, Narayan P (1995) Establishment and characterization of an immortalized but non-transformed human prostate epithelial cell line: BPH-1. In Vitro Cell Dev Biol Anim 31:14–24
Liu B, Conrad F, Roth A, Drummond DC, Simko JP, Marks JD (2007) Recombinant full-length human IgG1s targeting hormone-refractory prostate cancer. J Mol Med 85:1113–1123
Chalkley RJ, Baker PR, Medzihradszky KF, Lynn AJ, Burlingame AL (2008) In-depth analysis of tandem mass spectrometry data from disparate instrument types. Mol Cell Proteomics 7:2386–2398
Burke-Gaffney A, Hellewell PG (1998) A CD18/ICAM-1-dependent pathway mediates eosinophil adhesion to human bronchial epithelial cells. Am J Respir Cell Mol Biol 19:408–418
Zhang X, Wojcikiewicz EP, Moy VT (2006) Dynamic adhesion of T lymphocytes to endothelial cells revealed by atomic force microscopy. Exp Biol Med (Maywood) 231:1306–1312
Rosette C, Roth RB, Oeth P, Braun A, Kammerer S, Ekblom J, Denissenko MF (2005) Role of ICAM1 in invasion of human breast cancer cells. Carcinogenesis 26:943–950
Kammerer S, Roth RB, Reneland R, Marnellos G, Hoyal CR, Markward NJ, Ebner F, Kiechle M, Schwarz-Boeger U, Griffiths LR, Ulbrich C, Chrobok K, Forster G, Praetorius GM, Meyer P, Rehbock J, Cantor CR, Nelson MR, Braun A (2004) Large-scale association study identifies ICAM gene region as breast and prostate cancer susceptibility locus. Cancer Res 64:8906–8910
Zhao H, Kim Y, Wang P, Lapointe J, Tibshirani R, Pollack JR, Brooks JD (2005) Genome-wide characterization of gene expression variations and DNA copy number changes in prostate cancer cell lines. Prostate 63:187–197
Berry LJ, Au GG, Barry RD, Shafren DR (2008) Potent oncolytic activity of human enteroviruses against human prostate cancer. Prostate 68:577–587
Bidlingmaier S, Liu B (2006) Construction and application of a yeast surface-displayed human cDNA library to identify post-translational modification-dependent protein-protein interactions. Mol Cell Proteomics 5:533–540
Bidlingmaier S, Liu B (2007) Interrogating yeast surface-displayed human proteome to identify small molecule-binding proteins. Mol Cell Proteomics 6:2012–2020
Acknowledgement
The work is supported in part by grants from the National Institute of Health (R01 CA118919, P50 CA89520, P41 RR01614, and RR012961). We thank Dr. Scott Bidlingmaier for critical reading of the manuscript.
Open Access
This article is distributed under the terms of the Creative Commons Attribution Noncommercial License which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and source are credited.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
Open Access This is an open access article distributed under the terms of the Creative Commons Attribution Noncommercial License (https://creativecommons.org/licenses/by-nc/2.0), which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and source are credited.
About this article
Cite this article
Conrad, F., Zhu, X., Zhang, X. et al. Human antibodies targeting cell surface antigens overexpressed by the hormone refractory metastatic prostate cancer cells: ICAM-1 is a tumor antigen that mediates prostate cancer cell invasion. J Mol Med 87, 507–514 (2009). https://doi.org/10.1007/s00109-009-0446-3
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00109-009-0446-3