Abstract
Mitochondria play important roles in the regulation of key cellular processes, including energy metabolism, oxidative stress response, and signaling towards cell death or survival, and are distinguished by carrying their own genome (mtDNA). Mitochondrial dysfunction has emerged as a prominent cellular mechanism involved in neurodegeneration, including Parkinson’s disease (PD), a neurodegenerative movement disorder, characterized by progressive loss of dopaminergic neurons and the occurrence of proteinaceous Lewy body inclusions. The contribution of mtDNA variants to PD pathogenesis has long been debated and is still not clearly answered. Cytoplasmic hybrid (cybrid) cell models provided evidence for a contribution of mtDNA variants to the PD phenotype. However, conclusive evidence of mtDNA mutations as genetic cause of PD is still lacking. Several models have shown a role of somatic, rather than inherited mtDNA variants in the impairment of mitochondrial function and neurodegeneration. Accordingly, several nuclear genes driving inherited forms of PD are linked to mtDNA quality control mechanisms, and idiopathic as well as familial PD tissues present increased mtDNA damage. In this review, we highlight the use of cybrids in this PD research field and summarize various aspects of how and to what extent mtDNA variants may contribute to the etiology of PD.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Mitochondria are cellular organelles that carry their own genome, the mitochondrial DNA (mtDNA), and their own protein synthesis machinery [1, 2]. These organelles, often labelled as the ‘power stations’ of the cell, play an important role in the regulation of a plethora of key cellular processes, including energy metabolism, calcium homeostasis, oxidative stress response, and cellular signaling towards cell survival or death. Neurons have special energetic needs to fuel axonal and dendritic transport, the release and re-uptake of neurotransmitters, and vesicle trafficking at synapses. Therefore, it does not come unexpectedly that mitochondrial dysfunction has emerged as one of the most prominent cellular processes involved in neuronal dysfunction and neurodegeneration in Parkinson’s disease (PD) [3].
PD is the most common neurodegenerative movement disorder, with a particularly high and rising prevalence in Europe and North America [4, 5]. The neuropathological hallmarks of PD include the progressive loss of dopaminergic neurons in the substantia nigra pars compacta (SNpc) and the occurrence of proteinaceous inclusions known as Lewy bodies (LBs) in the remaining neurons [6, 7]. The subsequent dopamine deficiency in the basal ganglia triggers cellular and synaptic dysfunction, leading to the classical parkinsonian motor symptoms, which include tremor, rigidity, bradykinesia, and postural instability. In addition, significant non-motor symptoms, such as mental health issues, sleep disorders, pain, or fatigue are associated with PD and can largely precede the motor symptoms [5]. At present, only symptomatic treatments are available, with levodopa (L-Dopa) as the gold standard, and so during treatment, the disease pathology continues to progress.
PD has been widely accepted as a multifactorial disorder, with both genetic and environmental factors playing an important role, and age being the biggest risk factor [8]. The large majority of PD cases is classified as idiopathic (iPD, i.e., with an unknown etiology, ~ 90%), and about 10% of cases represent rare monogenic forms with Mendelian inheritance patterns, with more than 20 genes identified to date [9]. However, several of these genes lack replication and are, therefore, still debated. In addition, about 90 common variants, which exert an increase in heritable PD risk estimated between 16 and 36%, were identified by genome-wide association studies [10]. Several environmental risk factors, including pesticide exposure and traumatic brain injury, are also linked to the risk of developing PD. Conversely, cigarette smoking, caffeine intake and physical activity have been described as protective factors [11]. Interestingly, a recent study has shown that tobacco use is associated also with later age at onset (AAO) in patients carrying the p.G2019S mutation in LRRK2, and smoking intensity and duration was also correlated with AAO [12].
The earliest report linking mitochondria and PD dates back to the 1980s, when two incidents of drug abuse were reported, where patients showed typical signs of L-Dopa-responsive parkinsonism after injecting synthetic heroin contaminated with MPTP (1-methyl-4-phenyl-1,2,3,6-tetrahydropyridine) [13, 14]. As a highly lipophilic compound, MPTP can pass the blood brain barrier and, once in the brain, it is converted into 1-methyl-4-phenylpyridinium (MPP+), which selectively enters dopaminergic neurons through the dopamine transporter [15, 16]. Within neurons, MPP+ concentrates in mitochondria, where it inhibits complex I of the respiratory chain (NADH:ubiquinone oxidoreductase), leading to reactive oxygen species (ROS) production and ATP depletion [17, 18]. The link between mitochondrial malfunction and PD was further corroborated some years later when reduced activities of complexes I, II and IV were found in postmortem SNpc homogenates of iPD patients [19,20,21]. Complex I is dysfunctional also in platelets, lymphocytes and fibroblasts of PD patients [22,23,24]. Moreover, evidence is mounting that exposure to environmental inhibitors of the electron transport chain, like the pesticide rotenone, is associated to an increased risk for PD [25, 26], which further links mitochondria with PD. Ultimately, genetic evidence also implicates mitochondrial function as an integral disease component, since several familial PD genes are associated either directly or indirectly with mitochondrial function [27]. Mutations in PINK1, PRKN (Parkin), DJ-1, and CHCHD2 directly impact mitochondrial function, whereas mutations in SNCA (α-synuclein), LRRK2, ATP13A2, FBXO7, VPS13C, VPS35, and GBA1 genes mostly show indirect effects on mitochondrial function (reviewed recently in [28]). Interestingly, mutations in the mitochondrial Ubiquinol-Cytochrome C Reductase Core Protein 1 (UQCRC1) gene have been found recently to cause autosomal dominant parkinsonism with polyneuropathy, characterized molecularly by mitochondrial complex III dysfunction [29]. However, mutations in this gene were not replicated in other PD cohorts [30,31,32].
Cybrid cell models
The question of how and to what extent mtDNA variants contribute to the pathology in PD has long been debated and is still not clearly answered. Genetic manipulations of the mtDNA are difficult to achieve, which complicates direct molecular studies and the creation of model systems. A sophisticated model to study the contribution of mitochondrially encoded genes to cellular phenotypes is represented by cytoplasmic hybrids (cybrids). After their first description in 1989 by King and Attardi [54], cybrid cells became important models to study mtDNA contributions to pathologies, such as mitochondrial disorders, cancer, and neurodegenerative diseases.
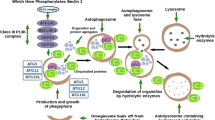
The creation of cybrid cells involves the elimination of mtDNA from recipient cells, which is mostly achieved by long-term treatment with Ethidium Bromide or through enzymatic methods, such as mitochondrially targeted restriction enzymes [55] or base excision repair pathway enzymes [56]. Cells lacking mtDNA are called Rho0 (ρ0) cells and are characterized by respiratory chain deficiency. Rho0 cells, although missing the mitochondrial genome, still contain mitochondrial organelles that can carry out several biochemical functions. The creation of cybrid cells is then achieved by repopulating Rho0 cells with donor mitochondria through cytoplasmic fusion with cytoplasts. Donor mitochondria can be obtained from platelets or from enucleated cytoplasts derived from cells carrying the mitochondria of interest. Studies of patient-derived mitochondria are mostly carried out by fusion of Rho0 cells with platelets that naturally lack the nucleus but contain mitochondria and mtDNA. Because Rho0 cells lack a functional respiratory chain, they are auxotrophic for pyruvate and uridine, a property that can be exploited to negatively select for Rho0 cells or positively select for cybrids repopulated by donor mitochondria. In this context, uridine is thought to supplement de novo pyrimidine synthesis, blocked at the dihydroorotate dehydrogenase step in Rho0 cells, while pyruvate may establish a favorable cellular redox balance [54]. Figure 1 gives an overview of the cybrid model generation process.
Schematic outline of cybrid cell model generation. The creation of cybrid cells involves the collection of cells devoid of nuclear DNA either by enucleation of the mitochondria donor cells (cytoplasts) or by the collection of platelets from blood. Cells providing the nuclear genome for the resulting cybrids are obtained by ridding cells of mtDNA through chemical (e.g., Ethidium Bromide) or genetic (e.g., mitochondria-targeted restriction enzymes or base excision repair pathway enzymes) means. Selection of resulting mtDNA-depleted Rho0 cells can be performed based on their auxotrophy for uridine and pyruvate. Finally, cybrid cells are created by polyethylene glycol (PEG)-mediated cytoplasmic fusion of mitochondria-donor cytoplasts or platelets with nuclear-donor Rho0 cells and positively selected against uridine and pyruvate auxotrophy
Cybrid studies of mitochondrial physiology in PD
Cybrid models were successfully used to prove the influence of mtDNA variants on cell function for many inherited mitochondrial disorders (reviewed in [57]). The recognition of mitochondrial defects, and in particular complex I deficiency in PD, led to the speculation that the mitochondrial genome may contribute to disease etiology. Therefore, cybrids were early on and extensively employed to shed light on several aspects of PD disease biology, such as mitochondrial defects within PD patient cells, genetic alterations in PD, signaling pathways and therapeutic options. While initial cybrid models were generated using osteosarcoma cell lines [54], the first neuronal cybrid models were established in 1996 by creation of respiratory-deficient SH-SY5Y neuroblastoma cell lines and repopulation with exogenous human platelet mitochondria. Such cells could successfully be differentiated into cells with neuron-like phenotypes [58]. These experiments were rapidly followed by repopulation of neuronal Rho0 cells (human neuroblastoma SH-SY5Y and teratocarcinoma NT2 cells) with mitochondria from iPD patients and demonstrated that patient-derived mitochondria are responsible for reduced respiratory chain complex I activity, increased ROS, lack of antioxidant enzyme upregulation, abnormal morphological mitochondrial ultrastructure, alterations in calcium homeostasis and increased susceptibility to cell death under stress [59,60,61,62]. Yet, in-depth and comprehensive genetic analyses of the mtDNA in these models were not reported. More recent biochemical analyses of NT2 cell cybrids showed that basal oxygen consumption was similar in PD and control cybrids, but maximal respiratory capacity was decreased, and proton leakage was increased in PD cybrids [63]. Differentiation of SH-SY5Y cybrids into neuronal cells allowed for controlled genetic conditions and led to the observation that PD cybrid neuronal cells had a lower differentiating potential compared to controls and were characterized by reduced axonal transport of mitochondria and higher dopamine release [64, 65].
However, the application of cybrid technology to a larger number of iPD patients evidenced that such models showed large heterogeneity between clones from patient and control subjects. While several mitochondrial physiological parameters including respiratory chain defects were mostly confirmed in cybrids carrying PD patient-derived mitochondria, substantial variations between clones were observed in histochemical and functional assays. Such assays included respiratory chain function, as measured by oxygen consumption and respiratory chain enzyme activities, genomic properties, as determined by mtDNA copy number assays and a non-sequence-specific heteroplasmy assay (i.e., Surveyor Nuclease®), as well as physiological properties, including mitochondrial movement and depolarization [66,67,68,69,70]. This variability may underlie the fact that some studies could observe only little effect of PD mitochondria on cellular phenotypes or no significant alterations of electron transport chain subunit protein levels [67, 69]. A comprehensive summary of cybrid studies evaluating the role of mtDNA alterations on mitochondrial functions in PD was recently reviewed by Buneeva et al. [71]. Importantly, studies aiming to evaluate physiological changes based on subtle somatic mtDNA alterations warrant cautious interpretation of results and should carefully control the experimental design with age-matched controls, since many studies have shown that aging alone leads to the accumulation of somatic mtDNA mutations in various tissues [72, 73].
Cybrid studies of molecular defects in PD
Nevertheless, cybrid models have proved to be valid and very useful tools to investigate signaling pathways and disease cascades involved in PD etiology. Indeed, a comparison between iPD cybrid cells with brain tissue samples from iPD patients showed a strong correlation between protein expression profiles of nuclear and mtDNA-encoded electron transport chain genes, suggesting that cybrid cells reliably mimic bioenergetic properties of the PD brain [74]. Pathophysiological investigations showed that cybrids with PD patient-derived mitochondria have an increased propensity to undergo apoptosis and have an altered response to cellular stress [59, 75]. Studies of PD cybrids also corroborated the importance of protein aggregates in the pathology of PD. Long-term cultures of cybrid cell lines can develop fibrillar and vesicular inclusions with a typical protein composition of PD-related Lewy bodies [76]. While the expression of the yeast single-subunit NADH dehydrogenase (Ndi1) in SH-SY5Y cybrids increased mitochondrial respiration and expression of mitochondrial genes, it was not sufficient to rescue cybrid Lewy body formation. Also, mitochondrial function remained related to the mtDNA and not the Lewy body status of selected cell clones [70]. Furthermore, cybrid models have allowed to link mitochondrial function with intracellular trafficking and associated homeostasis processes, such as autophagy. Disruption of microtubule-dependent intracellular trafficking in PD cybrids can impair autophagic degradation, as well as mitochondrial dynamics and health [77, 78]. Conversely, ATP depletion in PD cybrids can disrupt microtubule protein motors, leading to axonal transport disruption and microtubule depolymerization, which can trigger α-synuclein oligomerization [79].
Cybrid studies of inherited mtDNA variants in PD
The possible causal contribution of inherited mtDNA variants to PD was exemplified in a few cases and followed up in cybrid studies. Swerdlow et al. provided early evidence for a potential contribution of mtDNA variants to the PD phenotype in a family with maternally inherited parkinsonism [80]. The comparison of several cybrid lines established with mitochondria from maternal and paternal descendants of the family showed a lower complex I activity, increased ROS levels and abnormal mitochondrial morphology in cybrids derived from maternal descendants. The potential influence of pathogenic mtDNA variants on parkinsonism was further illustrated by a young patient with a complex phenotype including features of parkinsonism. This patient carried high levels of an MT-CYB frameshift mutation, which was responsible for respiratory, metabolic, and biochemical deficiencies in cybrid cells [81]. Also another inherited heteroplasmic mtDNA variant, m.T1095C, affecting MT-RNR1 was described in a family with maternally inherited parkinsonism, sensorineural deafness, and neuropathy [82]. Cybrids with mitochondria from an affected member of this family demonstrated depletion of mitochondrial glutathione and decreases in complex II/III enzymatic activity, in addition to increased apoptosis under aminoglycoside antibiotic [83].
Cybrid cell lines carrying known pathogenic mtDNA mutations, such as those found in patients with mtDNA-derived mitochondrial disorders, have been used to study cellular reactions to mitochondrial insults. For instance, by investigating cybrid cells carrying mtDNA mutations leading to respiratory chain complex I or complex IV defects, it was observed that aggregated α-synuclein can induce mitochondrial dysfunction specifically through inhibiting complex I [84]. Conversely, the overexpression of wild-type Parkin, known to affect mitochondrial quality control, can ameliorate certain mitochondrial parameters in long-term cybrid cultures of cells carrying heteroplasmic mtDNA mutations (e.g., COXI mutation [85]). Physiologically, the Parkin–mitophagy pathway seems to be upregulated in cells carrying pathogenic mtDNA mutations, such as the MERRF (Myoclonic Epilepsy with Ragged-Red Fibers) syndrome-causing m.A8344G mutation, perhaps as a protective feedback mechanism [86] (Fig. 2).
Outline of the contribution of inherited and somatic mtDNA variants to idiopathic and genetic forms of PD. mtDNA genome alterations can contribute to PD as inherited or somatic variants. 1) Inherited mtDNA variants are maternally transmitted and can influence mitochondrial function. While mtDNA haplogroup-driving variants can increase the risk or protect against PD, pathogenic mtDNA mutations decrease mitochondrial functions and can lead to PD-related phenotypes in patients. 2) Somatic mtDNA variants can accumulate in post-mitotic tissues, such as the brain. Various types of mtDNA variants, including point mutations and base damage, mtDNA deletions, transcription/replication‐associated 7S DNA molecules, and variations in mtDNA copy numbers have been investigated in PD patient tissues. While no clear genotype–phenotype correlation has been established to date, somatic mtDNA variations accumulate in affected tissues, such as neurons of iPD patients, suggesting an underlying defective mitochondrial quality control mechanism. In genetic forms of PD, nuclear gene mutations can directly affect mitophagy pathways (e.g. PRKN, PINK1). This can exacerbate the accumulation of somatic mtDNA variants, which may add to the mitochondrial dysfunction phenotype and aggravate the pathogenic cascade in both genetic and idiopathic forms of PD
More in general, some population genetic variants in the mtDNA have been shown to contribute to the genetic component of complex disorders, including PD. For example, cybrid studies found that cells carrying mitochondria from haplogroup H displayed differences in several mitochondrial physiology parameters (mtDNA and mtRNA levels, mitochondrial protein synthesis, cytochrome oxidase activity and amount, oxygen consumption, mitochondrial inner membrane potential and growth capacity) when compared with those of the haplogroup Uk [87]. This concept underlines the fact that mtDNA evolution is an active process and not all variants are detrimental. Clinical investigations and cybrid experiments provided data supporting the role of variant mtDNA in protecting against PD. For example, haplogroup B5 with the m.A10398G (MT-ND3 p.T114A) and m.G8584A (MT-ATP6 p.A20T) variants was proposed to protect against PD, and cybrids with these variants were more resistant to rotenone and had higher thresholds to induce autophagy and apoptosis, compared to cybrids carrying mitochondria from B4 haplogroup without these variants [88] (Fig. 2).
In addition to strictly investigating the contribution of mtDNA variants to PD-related phenotypes, cybrid cells were also used to explore mitochondrial dysfunction in familial PD. A distinct mitochondrial phenotype could be observed between cybrids carrying mtDNA from iPD patients as compared to cybrids from patients with a familial autosomal dominant form of PD, caused by SNCA mutations. Cybrids created from PD patients from a Contursi kindred in Southern Italy with an SNCA mutation did not manifest complex I deficiencies, when compared to iPD cybrids. However, in line with most PD-related cellular models, mitochondrial oxidative stress was still observed [89].
Taken together, while mtDNA sequence variants may influence PD pathogenesis, mtDNA mutations as ultimate genetic cause of PD have not been established to date.
Alternatives to cybrid models
Due to technical challenges and the heterogeneous nature of cybrid models, in recent years, they were not extensively applied in PD research. New methodological advances of genome editing strategies, such as the clustered regularly interspaced short palindromic repeats (CRISPR)/Cas9 system, have allowed the development of alternative models that can be adapted to specific research questions. To investigate genetic contributions to the phenotype of known pathogenic PD-driving nuclear gene mutations or PD disease-relevant genes, isogenic cell models can be developed by precise genome editing. Restoration of wild-type alleles in PD patient-derived cells or the insertion of PD-relevant mutations in control cells allow to explore genetic contributions to disease etiology [90]. Meanwhile, genetic engineering of mutations in nuclear-encoded mitochondrial genes can help to examine mitochondrial contributions to the PD phenotypes [91]. In addition, to study the phenotypic contribution of mtDNA variants, big progress has been made over the last years in developing tools for precise genome editing of the mtDNA. Such genome editing technologies include restriction endonucleases, zinc finger nuclease, transcription activator-like effector nucleases (TALEN), including the Double-stranded DNA Deaminase toxin A- derived cytosine base editor (DdCBE), the CRISPR system, and prokaryotic Argonaute proteins (pAgos)-based systems (reviewed in [92]). While such mtDNA genome editing strategies still need to be improved, models combining PD-relevant nuclear gene alterations with engineered mtDNA mutations may further contribute to the dissection of mitochondrial contributions in PD pathogenesis.
mtDNA alterations in Parkinson’s disease
mtDNA alterations in iPD
While several of the genes causing monogenic PD can be shown to directly affect mitochondrial quality, nuclear genome mutations can explain only a small fraction of PD. Alterations of the mtDNA are, therefore, an appealing target to investigate as a contributing factor for PD development. Once acquired, somatic mutations of the mtDNA are expected to accumulate in specific tissues due to clonal expansion or as a consequence of continued replication during life. Such mutations might be more likely to significantly expand in post-mitotic tissues, such as the brain, as compared to peripheral tissues, like blood cells, which are characterized by a quick turnover and potentially by a selection against cells with mitochondrial dysfunction caused by mutated mtDNA [93,94,95]. In general, a threshold of 60–90% heteroplasmic/mutant mtDNA results in phenotypic expression of mitochondrial dysfunction; however, this threshold depends on the type of mutation as well as cell and tissue types [96, 97]. In post-mitotic neurons, a lower level of heteroplasmy might be sufficient to induce phenotypic effects in patients.
In iPD patients, four different types of mtDNA alterations, namely mtDNA point mutations, mtDNA deletions, variations in mtDNA copy number and transcription/replication‐associated 7S DNA molecules have been investigated in postmortem brains or peripheral tissues including blood and skeletal muscle [98,99,100,101,102,103,104], but no specific rare variant and no clear genotype–phenotype correlation have been found so far that would establish a direct relationship between germline mtDNA variants and PD (Table1). However, a high proportion of multiple heteroplasmic mtDNA deletions were found to accumulate in the SNpc and other brain regions with age and were significantly more abundant in iPD patients [105, 106]. Higher levels of somatic mtDNA deletions were also found in another study investigating dopaminergic SN neurons of individuals with iPD as compared to age-matched controls [107]. Furthermore, ultra-deep sequencing revealed a large and heterogeneous pool of low-frequency mtDNA deletions in addition to a few (1–4) abundant species of mtDNA deletions in single dopaminergic neurons of two PD patients [108]. Moreover, in single SN neurons of iPD patients, a reduction of 7S DNA molecules, which bind to the D-Loop region during mtDNA transcription and replication, and mtDNA depletion were observed, which was more pronounced in neurons with severe complex I deficiency. On the contrary, in fibroblasts of iPD patients, accumulation of 7S DNA, low mtDNA replication, high heavy strand transcription and low mtDNA release was detected [109]. Using a histochemical approach able to detect abasic sites in the mtDNA, in which there is a loss of either a purine or a pyrimidine base, postmortem tissue sections of iPD patients showed a selective accumulation of mtDNA damage in nigral dopaminergic neurons [110] (Table1, Fig. 2). Of note, when investigating single cells of three different brain regions (putamen, frontal cortex, SN) of Alzheimer’s disease patients and controls, mtDNA deletion load was higher in SN neurons compared to the other brain regions, possibly pointing to a specific susceptibility of dopaminergic neurons to accumulate mtDNA damage, which was independent of the disease state [111]. Interestingly, the same selectivity of mtDNA damage in midbrain neurons was detected in rats exposed to the pesticide rotenone, where the mtDNA lesions appeared before any signs of neurodegeneration under conditions of mitochondrial impairment [110]. Such mitochondrial phenotypes and genotypes correlated with a decrease in mtDNA transcription factors TFAM and TFB2M [112]. Previous studies have shown that heterozygous loss of Tfam in mouse cells and tissues lead to a reduction in mtDNA copy number, respiratory chain deficiency, and increased oxidative mtDNA damage [113, 114].
Some studies have shown that mtDNA integrity and copy number specifically in SN neurons is correlated also with complex I and cytochrome c oxidase (COX) deficiency, suggesting that mtDNA alterations might be linked to impaired mitochondrial respiration [105, 112]. The particular susceptibility of dopaminergic neurons to mitochondrial dysfunction may be due to multiple factors, including their morphology, such as large axonal arbors and their unmyelinated or only lightly myelinated axons, their high overall pace-making function, and the energy-demanding and oxidatively challenging dopamine-producing metabolism (reviewed in [115]). Increased energy requirements of SN neurons in PD might be impacted by additional mitochondrial stressors, like mtDNA impairments, possibly contributing to the unique selective vulnerability, although more data on mtDNA integrity in SN neurons need to be collected. Compared to nuclear DNA, mtDNA is more vulnerable to ROS because it is physically close to ROS-producing sites, such as respiratory chain complexes I and III, and it lacks a protective chromatin structure. In addition, mitochondria possess fewer DNA repair pathways, and the mitochondrial DNA polymerase (POLG) shows different properties at incorporating modified nucleotides than nucleus DNA polymerases (reviewed in [116]). Accordingly, after exposing cultured cells to oxidative stress, the damage to mtDNA was higher and persisted longer than that to nuclear DNA, indicating that mtDNA is a cellular target for ROS [117]. This pathway may lead to a vicious cycle whereby damaged mitochondria increase ROS production, which exacerbates mtDNA damage. This mechanism was exemplified in a model of acute ischemic kidney injury, where mitochondrial ROS reduced the abundance of TFAM, resulting in reduced mtDNA synthesis and mitochondrial biogenesis, leading to decreased energy metabolism [118].
On top of known PD risk genes, nuclear genetic variants may contribute to mitochondrial defects and mtDNA alterations described in PD. A polygenic enrichment of rare nonsynonymous variants in genes belonging to pathways controlling mtDNA maintenance has been shown to influence the risk of iPD [119]. Similarly, the analysis of mitochondria-specific polygenic risk scores evidenced a cumulative small effect of common genetic variation within genes implicated in mitochondrial function on PD risk and AAO [120].
mtDNA alterations in genetic forms of parkinsonism
mtDNA alterations may also be the driving force of the mitochondrial phenotype in the presence of nuclear gene mutations involved in parkinsonism. Mutations in POLG, which lead to the accumulation of multiple mtDNA deletions over time, have been associated with Alpers-Huttenlocher syndrome (AHS) in children and chronic progressive external ophthalmoplegia (CPEO) in adults [121,122,123,124]. Parkinsonism occurs in a fraction of patients of both disorders. In addition to mtDNA deletions, POLG variants were also correlated with decreased mtDNA copy numbers in blood cells of PD patients [104]. Notably, neuronal cell loss was observed in patients with mutations in POLG, but not in other mitochondrial disorders, suggesting that patients with acquired (somatic) mtDNA mutations are more susceptible to neuronal degeneration compared to individuals with inherited mtDNA mutations [125].
It is becoming clear that PINK1 and Parkin are involved in several aspects for mitochondrial quality control and mtDNA maintenance (see BOX: Mitophagy). 7S DNA levels were found to be reduced in induced pluripotent stem cells (iPSC)-derived neuronal progenitor cells of patients with PRKN mutations as compared to age-matched controls, while the abundance of major arc deletions was unchanged in these individuals [126]. By genome-wide chromatin immunoprecipitation experiments, Parkin was found to interact with the mtDNA, most probably through an indirect interaction with the mitochondrial transcription factor TFAM [127]. Furthermore, in cell lines stably overexpressing Parkin, it was shown to increase mtDNA replication and transcription of mitochondrial genes [127, 128]. Parkin, together with PINK1, also controls mitochondrial biogenesis through Parkin interacting substrate (PARIS) and downstream regulation of Peroxisome proliferator-activated receptor gamma coactivator 1-alpha (PGC-1α) in a mechanism that proved to be critical for the survival of dopaminergic neurons [129,130,131].
Furthermore, also neural cells derived from iPSCs from patients carrying a homozygous or a heterozygous LRRK2 p.G2019S mutation, or asymptomatic individuals carrying a heterozygous LRKK2 p.R1441C mutation, showed significantly increased mitochondrial lesions as compared to control cells. After zinc finger nuclease-mediated repair of the LRRK2 p.G2019S mutation in iPSCs, mtDNA damage was not detected anymore in differentiated neural and neuroprogenitor cells, demonstrating that LRKK2 mutations are linked to mtDNA lesions in PD patients and asymptomatic mutation carriers [110, 132]. Recently, when investigating mtDNA integrity in fibroblasts of manifesting and non-manifesting carriers of the LRRK2 p.G2019S mutation, higher levels of somatic major arc deletions were detected in manifesting mutation carriers [133]. In addition, mtDNA copy number was found to be elevated in fibroblasts of manifesting mutation carriers as compared to non-manifesting carriers, and 7S DNA levels were reduced in LRRK2 p.G2019S mutation carriers independent of disease status [103]. These data suggest the involvement of mtDNA dyshomeostasis in the penetrance of LRRK2 mutations. Extending on these findings, alterations in the mtDNA maintenance pathway, including mtDNA copy number and 7S DNA, were found in PRKN-mutant iPSC-derived neurons, highlighting these mtDNA markers as potential modifiers of disease manifestation [134]. Furthermore, in LRRK2 p.G2019S mutation carriers, cell-free mtDNA concentration in cerebrospinal fluid correlated with α-synuclein levels, which was higher in mutation carriers compared to iPD patients [135] (Table1, Fig. 2).
In addition, PD-associated α-synuclein mutations let to mitochondrial DNA damage, altered mitochondrial transport and morphology, and reduced mitochondrial membrane potential [136,137,138]. Metabolic and cellular bioenergetics impairments were linked to accumulation of pathogenic α-synuclein aggregation at the mitochondrial membranes [139, 140] and a potential functional link was provided by an α-synuclein overexpressing mouse model that presented increased mtDNA deletions and oxidative DNA damage, which was related to the mitochondrial protein transport machinery [141]. mtDNA variation might, therefore, be a risk factor for increased phenotype severity and/or earlier AAO in genetic forms of parkinsonism.
Impact of impaired mitophagy on mtDNA dynamics in PD
As exemplified by the fact that genetic forms of PD can be driven by mutations in central mitophagy genes (e.g. PRKN, PINK1), mitophagy defects resulting in a decline of mitochondrial function have emerged as common feature in PD (reviewed recently by [162]). In general, a decline of mitochondrial fitness is observed during aging, which may be driven by ROS or a drop in the removal of malfunctioning mitochondria. Evidence for this stems from research that showed the accumulation of abnormal mitochondria and a rise in ROS production upon knockout of autophagy genes in immune cell models [163]. Likewise, accumulation of mtDNA double strand breaks were shown to underly mitochondrial dysfunction in mouse cardiomyocytes, in which Mitofusin2 knockout was used to model mitophagy deficiency [164]. On the reverse, increasing mitophagy can increase the selection against defective mitochondria, as evidenced by long-term overexpression of Parkin that decreased the heteroplasmy load of a deleterious COXI mtDNA mutation in cybrid cells [85].
As outlined above, there is evidence that somatic rather than inherited mtDNA mutations might be involved in the pathogenesis of PD and that Parkin and PINK1 might contribute to the removal of mtDNA mutations through mitophagy [165]. However, the exact mechanism of the interplay between these factors is poorly understood.
Mitophagy defects have been connected also to the pathogenesis of various inflammatory diseases, including neuroinflammation, an early event in the pathogenesis cascade of PD (reviewed in [166]). A dysfunctional mitochondrial quality control pathway, as caused by PINK1 or PRKN mutations, has recently been proposed to be linked to increased risk for PD through inflammation. The intra- and extracellular release of damage-associated molecular patterns (DAMPs), including mtDNA fragments, from damaged mitochondria is a potent trigger of the innate immune system. mtDNA can trigger inflammation through intracellular sensors, including nucleotide-binding oligomerization domain (NOD)-like receptor (NLR), Toll-like receptor (TLR), and the cytosolic cyclic GMP-AMP synthase (cGAS)-stimulator of interferon genes (STING) DNA sensing pathway. Inhibition of mitophagy has been shown to cause accumulation of mitochondrial ROS and damaged mitochondria, which in turn can trigger inflammation through the activation of the MAPK pathway [167] or the NLRP3 inflammasome [168]. Activation of these cellular response pathways can trigger an interferon response, followed by secretion of pro-inflammatory cytokines (e.g. IL-6), a condition recently described in PD patients carrying PRKN/PINK1 mutations, and therefore provides a rationale for the connection between mitochondrial dysfunction and inflammation in PD [165, 169].
mtDNA homeostasis in in vivo models of PD
Translational studies of human diseases, including PD, benefit greatly from in vivo animal models. Chemically induced rodent models of PD rely mostly on toxins that affect either mitochondrial function (e.g., MPTP) or dopamine metabolism (e.g., 6-hydroxydopamine, 6-OHDA). Similarly, genetic mouse models have focused on engineering genes known to cause familial PD or to cause mitochondrial dysfunctions.
The MitoPark mice, with disruption of the gene for mitochondrial transcription factor A (TFAM) in dopaminergic neurons, are characterized by a marked depletion of mtDNA, impairment of oxidative phosphorylation, dopaminergic neuron degeneration and motor deficits that mimic human parkinsonism [170]. Similarly, the “TwinkPark” mice, characterized by knockout of Prkn, together with SN-specific mtDNA deletions due to a mutation in the mtDNA helicase Twinkle (C10orf2), presented decreased mitochondrial function in SN cells, accompanied by increased neurobehavioral deficits and parkinsonian phenotypes compared to single mutant control animals [171]. Notably, mutations in Twinkle were described in association with CPEO/parkinsonism patients [172, 173].
In the Mutator mouse model, which possesses a proofreading-deficient form of POLG, resulting in time-dependent accumulation of multiple deletions in the mtDNA, neuroprotective compensatory mechanisms were observed at the mitochondrial level. These data suggested that somatic mtDNA deletions per se are not sufficient to trigger SNpc dopaminergic neuron death in animal models [174]. In contrast, in the mtDNA mutator Drosophila model that showed mitochondrial dysfunction, shortened lifespan, progressive locomotor deficit, and loss of dopaminergic neurons, a positive selection of deleterious mtDNA mutations was observed [175]. After crossing the Mutator mouse with a Prkn knockout mouse, which both by themselves do not show neurodegeneration, mitochondrial dysfunction and PD pathology became apparent. This was linked to a greater predicted pathogenicity score of mtDNA mutations. Surprisingly, however, the overall abundance and type of these mutations (synonymous vs. nonsynonymous) was unchanged [176]. This study highlights a key role of mtDNA mutations in dopaminergic neurons and a protective role of Parkin, since its loss synergizes with mitochondrial dysfunction resulting in neurodegeneration. The Prkn knockout:Mutator mouse model was also used to show a role for mitophagy at mitigating inflammation through the STING pathway [165].
Attempts at modeling PD in vivo based on mitochondrial dysfunction through genetic disruption of nuclear-encoded complex I subunits evidenced a complicated picture of complex I dysfunction in PD. Ndufs4 knockout mouse models with a mild complex I deficiency in vivo displayed an altered dopamine metabolism but did not present neurodegeneration phenotypes [177, 178]. On the contrary, a recently described model with Ndufs2 knockout in dopaminergic neurons (MCI-Park mice) showed that complete complex I disruption can lead to an axonal-first progressive neurodegeneration that produces a human-like parkinsonism in which the loss of nigral dopamine release makes a critical contribution to motor dysfunction. This model allowed the authors to follow the PD disease cascade over time and between different brain regions [179].
PINK1 and Parkin have been shown to play important roles in the mitochondrial quality control through the mitophagy pathway and may, therefore, also be involved in the selection against deleterious mtDNA mutations. However, their role in vivo under physiological conditions is still debated. Comparisons of heteroplasmic mtDNA variants between wild type, Pink1 knockout and Parkin knockout mice showed an increase in early onset single nucleotide variants in Pink1 knockout mice and an increased variant allele frequency of mtDNA mutations in SN of aged Pink1 and Prkn knockout mice [180]. Furthermore, in a transgene-based Drosophila model carrying a heteroplasmic mtDNA deletion in adult muscle tissue, genetic induction of the Pink1-Parkin pathway resulted in a selective decrease in mtDNA deletion molecules that was at least partially dependent on autophagy [181]. These studies indicated that selection against dysfunctional mitochondria can be achieved in vivo in post-mitotic tissues. Another elegant model to study the effect of mtDNA variants and their selection process in vivo was provided by Drosophila carrying a heteroplasmic, temperature-sensitive COXI mutation induced by a mitochondria-targeted restriction enzyme and cytoplasm transfer [182, 183]. While Parkin was not involved in the selection process of deleterious mtDNA in this model, PINK1 did preferentially accumulate on mitochondria enriched for the pathogenic mtDNA mutations and was required for selection of healthy mitochondria. However, rather than inducing the canonical mitophagy pathway, PINK1 was shown to inhibit local protein translation through phosphorylation of Larp, which induced preferential replication of normal mitochondria and hampered the propagation of defective organelles [184, 185]. Collectively, these data provide evidence for an interplay between PD-related nuclear gene functions and mtDNA alterations in the etiology of PD.
Treatment strategies in PD aimed at mitochondrial dysfunction and mtDNA
In recent years, the microbiome–gut–brain axis has gained particular attention as underlying pathological mechanism in PD, linked through metabolic alterations, bacterial metabolites, and inflammatory responses (reviewed in [186]). In this context, pro- and prebiotics may represent promising options to prevent or ameliorate PD symptoms through modulation of inflammatory pathways or metabolic functions [186]. The use of dietary supplements to improve mitochondrial function has long been applied to primary mitochondrial disorders and has been suggested also for PD. A functional interaction between diet and mtDNA has been proposed in fly models, where mtDNA plays a role in modulating diet-dependent fitness [187]. Also in humans, a higher diet quality was associated with a greater whole blood-derived mtDNA copy number [188].
In the context of PD, dietary supplement research focused primarily on the antioxidant pathway as potential therapeutic approach to counteract the detrimental effects of ROS and to preserve mitochondrial function (reviewed in [189]). For example, coenzyme Q10 (ubiquinone) that functions as an electron carrier in the electron transport chain shows potent antioxidant properties, and hence an effect also on mtDNA content and integrity [190]. While several preclinical studies showed beneficial effects of coenzyme Q10 dietary supplementation in PD models, meta-analyses of randomized, placebo-controlled trials in human PD found no evidence that coenzyme Q10 improved motor-related symptoms or delayed the progression of the disease when compared to placebo [191, 192]. Similarly, a mitochondrially targeted quinone, MitoQ (mitoquinone), proved beneficial in several preclinical models, but not in clinical trials in PD patients [193]. Furthermore, polyphenols have been proposed as beneficial molecules against PD due to their antioxidant and anti-inflammatory actions, but conclusive pharmacological benefits are outstanding (reviewed in [194]). A more direct effect on cellular metabolism and influence on mtDNA variations can be expected by metabolic fuels, such as proteins and carbohydrates. In fly models, it was suggested that a diet with a low protein to carbohydrate ratio and rich in certain fatty acids would be beneficial for cells with respiratory chain complex I deficiencies and Parkin mutations [195, 196]. Also, calorie restriction may play a role in mtDNA integrity and prolonging lifespan through the regulation of multiple molecular pathways, with direct implications for NDDs, including PD (reviewed in [197]). However, the effect of dietary supplements on mitochondrial metabolism and mtDNA are mainly based on preclinical models. Translation of such treatment strategies to clinical practice have mostly failed to show significant benefits and will warrant better patient stratification and improved biomarkers.
In addition to dietary adjustments, exercise has been shown in preclinical and clinical studies to provide a health benefit to PD patients, which can at least in part be explained by mitochondrial metabolic improvements and may potentially involve the mitochondrial genome. Notably, in the MitoPark mouse, which is characterized by inactivation of TFAM, exercise was shown to improve behavioral parameters and nigrostriatal dopamine input as well as oxidative phosphorylation [198]. Furthermore, exercise prevented alterations of mitochondrial biogenesis signaling (PGC-1α/NRF-1/TFAM) and respiratory chain activity in a PD rat model [199]. However, exercise-induced motor improvements in PD were associated with metabolic changes but not with muscle mtDNA content [200].
Summary
There is accumulating evidence demonstrating that mitochondrial dysfunction either directly underlies the neurodegenerative disease process in PD or constitutes one part of a complex series of events. The establishment of human cybrid cell lines provided the opportunity to show direct molecular consequences of the expression of mtDNA variants from iPD patients in vitro. Although no rare inherited mtDNA mutation was linked to PD as yet, studies performed in postmortem brains or peripheral tissues including blood and skeletal muscle identified four different types of somatic mtDNA alterations, namely mtDNA point mutations, mtDNA deletions, variations in mtDNA copy number and transcription/replication‐associated 7S DNA molecules. Also, in different tissues of genetic PD patients, higher levels of somatic mtDNA mutations were detected. Genetic mouse models suggest that dysfunctional degradation of mitochondria carrying mtDNA mutations through the process of mitophagy might be associated with the accumulation of somatic mutations at different heteroplasmy levels in different tissues. A schematic overview of contributions of inherited vs. somatic mtDNA variations to the pathogenesis of PD is represented in Table 1 and Fig. 2.
Methodological advances are adding to our understanding of biological and genetic basis of PD. For example, recently established long-read sequencing methodologies (i.e., Nanopore sequencing) allowed for the investigation of mtDNA methylation architecture and provided some evidence for CpG methylation differences between tissues and between PD patients and healthy controls [45]. Application of refined genomic and biochemical methods will be needed to accurately measure mtDNA base modifications and adducts in order to quantify stress-induced mtDNA alterations and better understand mtDNA repair mechanisms. As such, deep sequencing of tissues and cell line models, ideally by long read sequencing technologies to capture structural variants, will help to clarify the picture of mtDNA alterations in PD. Furthermore, transcriptomics and sequencing studies at the single cell level should provide information on the somatic mitochondrial genetic architecture of brain tissues. Performance of such studies in appropriate animal models, where early stages of the disease can be evaluated may allow to place mitochondrial and mtDNA dysfunction into the timeline of the disease cascade of PD.
Mitochondrial and mtDNA alterations result in metabolic changes of the affected cells. To better connect genetic and physiological alterations in PD, but also to properly select and stratify patients for clinical trials, it will be important to pair genetic analyses with studies of metabolic alterations in model systems and brain tissues. Functional imaging techniques that allow the assessment of mitochondrial metabolism and bioenergetic disturbances may proof helpful tools and can be performed longitudinally to compare different stages of disease (e.g., reviewed in [189]). The altered mitochondrial metabolism may also offer a therapeutic handle against PD, and comprehensive metabolomics characterizations of PD models are providing the bases for such approaches [201]. Treatment options could be inspired by recent work in the field of mitochondrial disorders, where it was shown that metabolic restrictions can suppress pathological mtDNA replication. By limiting the energy output of glycolysis with a glucose analog such as 2-Deoxy-D-glucose (2DG) or 5-thioglucose, wild-type mtDNA was selected over mutant mtDNA copies in the context of a pathogenic mtDNA mutation [202]. Furthermore, accelerating the removal of damaged mitochondria by pharmacologically enhancing mitophagy may provide an additional therapeutic approach against PD [162].
Mitochondrial function is also tightly connected with a proper function of the immune system and mitochondrial dysfunctions can trigger inflammation, e.g., through release of mtDNA fragments. The understanding that inflammation plays an important and early role in PD pathogenesis suggests that targeting the immune system with anti-inflammatory drugs may provide another promising therapeutic avenue against PD [165].
An intact mitochondrial genome is central to proper mitochondrial function, but mtDNA is vulnerable to insults. Mitochondrial impairment appears to be centrally involved in the neurodegenerative process of PD, independent of the root cause of the disease in a single patient. Our understanding of the underlying genetic and pathophysiological mechanism is, therefore, providing the basis for therapeutic approaches against PD.
Data availability
Enquiries about data availability should be directed to the authors.
References
Anderson S, Bankier AT, Barrell BG, de Bruijn MH, Coulson AR, Drouin J et al (1981) Sequence and organization of the human mitochondrial genome. Nature 290(5806):457–465. https://doi.org/10.1038/290457a0
Watanabe K (2010) Unique features of animal mitochondrial translation systems. The non-universal genetic code, unusual features of the translational apparatus and their relevance to human mitochondrial diseases. Proc Jpn Acad Ser B Phys Biol Sci 86(1):11–39. https://doi.org/10.2183/pjab.86.11
Perier C, Vila M (2012) Mitochondrial biology and Parkinson’s disease. Cold Spring Harb Perspect Med 2(2):a009332. https://doi.org/10.1101/cshperspect.a009332
Collaborators GBDN (2019) Global, regional, and national burden of neurological disorders, 1990–2016: a systematic analysis for the Global Burden of Disease Study 2016. Lancet Neurol 18(5):459–480. https://doi.org/10.1016/S1474-4422(18)30499-X
Kalia LV, Lang AE (2015) Parkinson’s disease. Lancet 386(9996):896–912. https://doi.org/10.1016/S0140-6736(14)61393-3
Spillantini MG, Schmidt ML, Lee VM, Trojanowski JQ, Jakes R, Goedert M (1997) Alpha-synuclein in Lewy bodies. Nature 388(6645):839–840. https://doi.org/10.1038/42166
Forno LS (1996) Neuropathology of Parkinson’s disease. J Neuropathol Exp Neurol 55(3):259–272. https://doi.org/10.1097/00005072-199603000-00001
Reeve A, Simcox E, Turnbull D (2014) Ageing and Parkinson’s disease: why is advancing age the biggest risk factor? Ageing Res Rev 14:19–30. https://doi.org/10.1016/j.arr.2014.01.004
Blauwendraat C, Nalls MA, Singleton AB (2020) The genetic architecture of Parkinson’s disease. Lancet Neurol 19(2):170–178. https://doi.org/10.1016/S1474-4422(19)30287-X
Nalls MA, Blauwendraat C, Vallerga CL, Heilbron K, Bandres-Ciga S, Chang D et al (2019) Identification of novel risk loci, causal insights, and heritable risk for Parkinson’s disease: a meta-analysis of genome-wide association studies. Lancet Neurol 18(12):1091–1102. https://doi.org/10.1016/S1474-4422(19)30320-5
Ascherio A, Schwarzschild MA (2016) The epidemiology of Parkinson’s disease: risk factors and prevention. Lancet Neurol 15(12):1257–1272. https://doi.org/10.1016/S1474-4422(16)30230-7
Luth T, Konig IR, Grunewald A, Kasten M, Klein C, Hentati F et al (2020) Age at Onset of LRRK2 p.Gly2019Ser is related to environmental and lifestyle factors. Mov Disord 35(10):1854–1858. https://doi.org/10.1002/mds.28238
Langston JW, Ballard P, Tetrud JW, Irwin I (1983) Chronic Parkinsonism in humans due to a product of meperidine-analog synthesis. Science 219(4587):979–980. https://doi.org/10.1126/science.6823561
Langston JW, Ballard P (1984) Parkinsonism induced by 1-methyl-4-phenyl-1,2,3,6-tetrahydropyridine (MPTP): implications for treatment and the pathogenesis of Parkinson’s disease. Can J Neurol Sci 11(1 Suppl):160–165. https://doi.org/10.1017/s0317167100046333
Heikkila RE, Hess A, Duvoisin RC (1984) Dopaminergic neurotoxicity of 1-methyl-4-phenyl-1,2,5,6-tetrahydropyridine in mice. Science 224(4656):1451–1453. https://doi.org/10.1126/science.6610213
Javitch JA, D’Amato RJ, Strittmatter SM, Snyder SH (1985) Parkinsonism-inducing neurotoxin, N-methyl-4-phenyl-1,2,3,6 -tetrahydropyridine: uptake of the metabolite N-methyl-4-phenylpyridine by dopamine neurons explains selective toxicity. Proc Natl Acad Sci U S A 82(7):2173–2177. https://doi.org/10.1073/pnas.82.7.2173
Nicklas WJ, Vyas I, Heikkila RE (1985) Inhibition of NADH-linked oxidation in brain mitochondria by 1-methyl-4-phenyl-pyridine, a metabolite of the neurotoxin, 1-methyl-4-phenyl-1,2,5,6-tetrahydropyridine. Life Sci 36(26):2503–2508. https://doi.org/10.1016/0024-3205(85)90146-8
Ramsay RR, Salach JI, Singer TP (1986) Uptake of the neurotoxin 1-methyl-4-phenylpyridine (MPP+) by mitochondria and its relation to the inhibition of the mitochondrial oxidation of NAD+-linked substrates by MPP+. Biochem Biophys Res Commun 134(2):743–748. https://doi.org/10.1016/s0006-291x(86)80483-1
Schapira AH (2007) Mitochondrial dysfunction in Parkinson’s disease. Cell Death Differ 14(7):1261–1266. https://doi.org/10.1038/sj.cdd.4402160
Schapira AH, Cooper JM, Dexter D, Clark JB, Jenner P, Marsden CD (1990) Mitochondrial complex I deficiency in Parkinson’s disease. J Neurochem 54(3):823–827. https://doi.org/10.1111/j.1471-4159.1990.tb02325.x
Schapira AH, Cooper JM, Dexter D, Jenner P, Clark JB, Marsden CD (1989) Mitochondrial complex I deficiency in Parkinson’s disease. Lancet 1(8649):1269. https://doi.org/10.1016/s0140-6736(89)92366-0
Parker WD Jr, Boyson SJ, Parks JK (1989) Abnormalities of the electron transport chain in idiopathic Parkinson’s disease. Ann Neurol 26(6):719–723. https://doi.org/10.1002/ana.410260606
Muftuoglu M, Elibol B, Dalmizrak O, Ercan A, Kulaksiz G, Ogus H et al (2004) Mitochondrial complex I and IV activities in leukocytes from patients with parkin mutations. Mov Disord 19(5):544–548. https://doi.org/10.1002/mds.10695
Mortiboys H, Thomas KJ, Koopman WJ, Klaffke S, Abou-Sleiman P, Olpin S et al (2008) Mitochondrial function and morphology are impaired in parkin-mutant fibroblasts. Ann Neurol 64(5):555–565. https://doi.org/10.1002/ana.21492
Tanner CM, Kamel F, Ross GW, Hoppin JA, Goldman SM, Korell M et al (2011) Rotenone, paraquat, and Parkinson’s disease. Environ Health Perspect 119(6):866–872. https://doi.org/10.1289/ehp.1002839
Horst CH, Schlemmer F, de Aguiar MN, Domingues ACM, Ferreira GG, da Silva Ribeiro CY et al (2018) Signature of aberrantly expressed microRNAs in the striatum of rotenone-induced Parkinsonian rats. Neurochem Res 43(11):2132–2140. https://doi.org/10.1007/s11064-018-2638-0
Park JS, Davis RL, Sue CM (2018) Mitochondrial dysfunction in Parkinson’s disease: new mechanistic insights and therapeutic perspectives. Curr Neurol Neurosci Rep 18(5):21. https://doi.org/10.1007/s11910-018-0829-3
Trinh D, Israwi AR, Arathoon LR, Gleave JA, Nash JE (2021) The multi-faceted role of mitochondria in the pathology of Parkinson’s disease. J Neurochem 156(6):715–752. https://doi.org/10.1111/jnc.15154
Lin CH, Tsai PI, Lin HY, Hattori N, Funayama M, Jeon B et al (2020) Mitochondrial UQCRC1 mutations cause autosomal dominant parkinsonism with polyneuropathy. Brain 143(11):3352–3373. https://doi.org/10.1093/brain/awaa279
Courtin T, Tesson C, Corvol JC, Lesage S, Brice A (2021) French Parkinson’s disease genetics g. Lack of evidence for association of UQCRC1 with autosomal dominant Parkinson’s disease in Caucasian families. Neurogenetics 22(4):365–366. https://doi.org/10.1007/s10048-021-00647-4
Zhao YW, Pan HX, Wang CY, Zeng Q, Wang Y, Fang ZH et al (2021) UQCRC1 variants in Parkinson’s disease: a large cohort study in Chinese mainland population. Brain 144(6):e54. https://doi.org/10.1093/brain/awab137
Senkevich K, Bandres-Ciga S, Ganor Z, Krohn L, International Parkinson’s Disease Genomics C (2021) Lack of evidence for association of UQCRC1 with Parkinson’s disease in Europeans. Neurobiol Aging 101:297. https://doi.org/10.1016/j.neurobiolaging.2020.10.030
Johnston IG, Williams BP (2016) Evolutionary inference across eukaryotes identifies specific pressures favoring mitochondrial gene retention. Cell Syst 2(2):101–111. https://doi.org/10.1016/j.cels.2016.01.013
DiMauro S, Schon EA (2003) Mitochondrial respiratory-chain diseases. N Engl J Med 348(26):2656–2668. https://doi.org/10.1056/NEJMra022567
Ojala D, Montoya J, Attardi G (1981) tRNA punctuation model of RNA processing in human mitochondria. Nature 290(5806):470–474. https://doi.org/10.1038/290470a0
Mercer TR, Neph S, Dinger ME, Crawford J, Smith MA, Shearwood AM et al (2011) The human mitochondrial transcriptome. Cell 146(4):645–658. https://doi.org/10.1016/j.cell.2011.06.051
Wang G, Yang E, Mandhan I, Brinkmeyer-Langford CL, Cai JJ (2014) Population-level expression variability of mitochondrial DNA-encoded genes in humans. Eur J Hum Genet 22(9):1093–1099. https://doi.org/10.1038/ejhg.2013.293
Ali AT, Boehme L, Carbajosa G, Seitan VC, Small KS, Hodgkinson A (2019) Nuclear genetic regulation of the human mitochondrial transcriptome. Elife. https://doi.org/10.7554/eLife.41927
Patil V, Cuenin C, Chung F, Aguilera JRR, Fernandez-Jimenez N, Romero-Garmendia I et al (2019) Human mitochondrial DNA is extensively methylated in a non-CpG context. Nucleic Acids Res 47(19):10072–10085. https://doi.org/10.1093/nar/gkz762
Shock LS, Thakkar PV, Peterson EJ, Moran RG, Taylor SM (2011) DNA methyltransferase 1, cytosine methylation, and cytosine hydroxymethylation in mammalian mitochondria. Proc Natl Acad Sci U S A 108(9):3630–3635. https://doi.org/10.1073/pnas.1012311108
Saini SK, Mangalhara KC, Prakasam G, Bamezai RNK (2017) DNA Methyltransferase1 (DNMT1) Isoform3 methylates mitochondrial genome and modulates its biology. Sci Rep 7(1):1525. https://doi.org/10.1038/s41598-017-01743-y
Bellizzi D, D’Aquila P, Scafone T, Giordano M, Riso V, Riccio A et al (2013) The control region of mitochondrial DNA shows an unusual CpG and non-CpG methylation pattern. DNA Res 20(6):537–547. https://doi.org/10.1093/dnares/dst029
Wong M, Gertz B, Chestnut BA, Martin LJ (2013) Mitochondrial DNMT3A and DNA methylation in skeletal muscle and CNS of transgenic mouse models of ALS. Front Cell Neurosci 7:279. https://doi.org/10.3389/fncel.2013.00279
Lopes FCA (2020) Mitochondrial metabolism and DNA methylation: a review of the interaction between two genomes. Clin Epigenetics 12(1):182. https://doi.org/10.1186/s13148-020-00976-5
Luth T, Wasner K, Klein C, Schaake S, Tse R, Pereira SL et al (2021) Nanopore single-molecule sequencing for mitochondrial DNA methylation analysis: investigating parkin-associated Parkinsonism as a proof of concept. Front Aging Neurosci 13:713084. https://doi.org/10.3389/fnagi.2021.713084
Bicci I, Calabrese C, Golder ZJ, Gomez-Duran A, Chinnery PF (2021) Single-molecule mitochondrial DNA sequencing shows no evidence of CpG methylation in human cells and tissues. Nucleic Acids Res. https://doi.org/10.1093/nar/gkab1179
Wang YE, Marinov GK, Wold BJ, Chan DC (2013) Genome-wide analysis reveals coating of the mitochondrial genome by TFAM. PLoS ONE 8(8):e74513. https://doi.org/10.1371/journal.pone.0074513
Bonekamp NA, Larsson NG (2018) SnapShot: Mitochondrial Nucleoid. Cell 172(1–2):388-e1. https://doi.org/10.1016/j.cell.2017.12.039
Chen X, Prosser R, Simonetti S, Sadlock J, Jagiello G, Schon EA (1995) Rearranged mitochondrial genomes are present in human oocytes. Am J Hum Genet 57(2):239–47
D’Erchia AM, Atlante A, Gadaleta G, Pavesi G, Chiara M, De Virgilio C et al (2015) Tissue-specific mtDNA abundance from exome data and its correlation with mitochondrial transcription, mass and respiratory activity. Mitochondrion 20:13–21. https://doi.org/10.1016/j.mito.2014.10.005
Luo S, Valencia CA, Zhang J, Lee NC, Slone J, Gui B et al (2018) Biparental inheritance of mitochondrial DNA in humans. Proc Natl Acad Sci U S A 115(51):13039–44. https://doi.org/10.1073/pnas.1810946115
Lutz-Bonengel S, Parson W (2019) No further evidence for paternal leakage of mitochondrial DNA in humans yet. Proc Natl Acad Sci U S A 116(6):1821–2. https://doi.org/10.1073/pnas.1820533116
Wei W, Chinnery PF (2020) Inheritance of mitochondrial DNA in humans: implications for rare and common diseases. J Intern Med 287(6):634–44. https://doi.org/10.1111/joim.13047
King MP, Attardi G (1989) Human cells lacking mtDNA: repopulation with exogenous mitochondria by complementation. Science 246(4929):500–3. https://doi.org/10.1126/science.2814477
Heller S, Schubert S, Krehan M, Schafer I, Seibel M, Latorre D et al (2013) Efficient repopulation of genetically derived rho zero cells with exogenous mitochondria. PLoS ONE 8(9):e73207. https://doi.org/10.1371/journal.pone.0073207
Spadafora D, Kozhukhar N, Chouljenko VN, Kousoulas KG, Alexeyev MF (2016) Methods for efficient elimination of mitochondrial DNA from cultured cells. PLoS ONE 11(5):e0154684. https://doi.org/10.1371/journal.pone.0154684
Wilkins HM, Carl SM, Swerdlow RH (2014) Cytoplasmic hybrid (cybrid) cell lines as a practical model for mitochondriopathies. Redox Biol 2:619–31. https://doi.org/10.1016/j.redox.2014.03.006
Miller SW, Trimmer PA, Parker WD Jr, Davis RE (1996) Creation and characterization of mitochondrial DNA-depleted cell lines with “neuronal-like” properties. J Neurochem 67(5):1897–907. https://doi.org/10.1046/j.1471-4159.1996.67051897.x
Swerdlow RH, Parks JK, Miller SW, Tuttle JB, Trimmer PA, Sheehan JP et al (1996) Origin and functional consequences of the complex I defect in Parkinson’s disease. Ann Neurol 40(4):663–71. https://doi.org/10.1002/ana.410400417
Sheehan JP, Swerdlow RH, Parker WD, Miller SW, Davis RE, Tuttle JB (1997) Altered calcium homeostasis in cells transformed by mitochondria from individuals with Parkinson’s disease. J Neurochem 68(3):1221–33. https://doi.org/10.1046/j.1471-4159.1997.68031221.x
Cassarino DS, Fall CP, Swerdlow RH, Smith TS, Halvorsen EM, Miller SW et al (1997) Elevated reactive oxygen species and antioxidant enzyme activities in animal and cellular models of Parkinson’s disease. Biochim Biophys Acta 1362(1):77–86. https://doi.org/10.1016/s0925-4439(97)00070-7
Trimmer PA, Swerdlow RH, Parks JK, Keeney P, Bennett JP Jr, Miller SW et al (2000) Abnormal mitochondrial morphology in sporadic Parkinson’s and Alzheimer’s disease cybrid cell lines. Exp Neurol 162(1):37–50. https://doi.org/10.1006/exnr.2000.7333
Esteves AR, Lu J, Rodova M, Onyango I, Lezi E, Dubinsky R et al (2010) Mitochondrial respiration and respiration-associated proteins in cell lines created through Parkinson’s subject mitochondrial transfer. J Neurochem 113(3):674–82. https://doi.org/10.1111/j.1471-4159.2010.06631.x
Trimmer PA, Schwartz KM, Borland MK, De Taboada L, Streeter J, Oron U (2009) Reduced axonal transport in Parkinson’s disease cybrid neurites is restored by light therapy. Mol Neurodegener 4:26. https://doi.org/10.1186/1750-1326-4-26
Appukuttan TA, Ali N, Varghese M, Singh A, Tripathy D, Padmakumar M et al (2013) Parkinson’s disease cybrids, differentiated or undifferentiated, maintain morphological and biochemical phenotypes different from those of control cybrids. J Neurosci Res 91(7):963–70. https://doi.org/10.1002/jnr.23241
Gu M, Cooper JM, Taanman JW, Schapira AH (1998) Mitochondrial DNA transmission of the mitochondrial defect in Parkinson’s disease. Ann Neurol 44(2):177–86. https://doi.org/10.1002/ana.410440207
Aomi Y, Chen CS, Nakada K, Ito S, Isobe K, Murakami H et al (2001) Cytoplasmic transfer of platelet mtDNA from elderly patients with Parkinson’s disease to mtDNA-less HeLa cells restores complete mitochondrial respiratory function. Biochem Biophys Res Commun 280(1):265–73. https://doi.org/10.1006/bbrc.2000.4113
Esteves AR, Domingues AF, Ferreira IL, Januario C, Swerdlow RH, Oliveira CR et al (2008) Mitochondrial function in Parkinson’s disease cybrids containing an nt2 neuron-like nuclear background. Mitochondrion 8(3):219–28. https://doi.org/10.1016/j.mito.2008.03.004
Keeney PM, Dunham LD, Quigley CK, Morton SL, Bergquist KE, Bennett JP Jr (2009) Cybrid models of Parkinson’s disease show variable mitochondrial biogenesis and genotype-respiration relationships. Exp Neurol 220(2):374–82. https://doi.org/10.1016/j.expneurol.2009.09.025
Cronin-Furman EN, Borland MK, Bergquist KE, Bennett JP Jr, Trimmer PA (2013) Mitochondrial quality, dynamics and functional capacity in Parkinson’s disease cybrid cell lines selected for Lewy body expression. Mol Neurodegener 8:6. https://doi.org/10.1186/1750-1326-8-6
Buneeva O, Fedchenko V, Kopylov A, Medvedev A (2020) Mitochondrial Dysfunction in Parkinson’s disease: focus on mitochondrial DNA. Biomedicines. https://doi.org/10.3390/biomedicines8120591
Michikawa Y, Mazzucchelli F, Bresolin N, Scarlato G, Attardi G (1999) Aging-dependent large accumulation of point mutations in the human mtDNA control region for replication. Science 286(5440):774–9. https://doi.org/10.1126/science.286.5440.774
Wolf AM (2021) MtDNA mutations and aging-not a closed case after all? Signal Transduct Target Ther 6(1):56. https://doi.org/10.1038/s41392-021-00479-6
Borland MK, Mohanakumar KP, Rubinstein JD, Keeney PM, Xie J, Capaldi R et al (2009) Relationships among molecular genetic and respiratory properties of Parkinson’s disease cybrid cells show similarities to Parkinson’s brain tissues. Biochim Biophys Acta 1792(1):68–74. https://doi.org/10.1016/j.bbadis.2008.09.014
Cassarino DS, Halvorsen EM, Swerdlow RH, Abramova NN, Parker WD Jr, Sturgill TW et al (2000) Interaction among mitochondria, mitogen-activated protein kinases, and nuclear factor-kappaB in cellular models of Parkinson’s disease. J Neurochem 74(4):1384–92. https://doi.org/10.1046/j.1471-4159.2000.0741384.x
Trimmer PA, Borland MK, Keeney PM, Bennett JP Jr, Parker WD Jr (2004) Parkinson’s disease transgenic mitochondrial cybrids generate Lewy inclusion bodies. J Neurochem 88(4):800–12. https://doi.org/10.1046/j.1471-4159.2003.02168.x
Arduino DM, Esteves AR, Cortes L, Silva DF, Patel B, Grazina M et al (2012) Mitochondrial metabolism in Parkinson’s disease impairs quality control autophagy by hampering microtubule-dependent traffic. Hum Mol Genet 21(21):4680–702. https://doi.org/10.1093/hmg/dds309
Esteves AR, Palma AM, Gomes R, Santos D, Silva DF, Cardoso SM (2019) Acetylation as a major determinant to microtubule-dependent autophagy: relevance to Alzheimer’s and Parkinson disease pathology. Biochim Biophys Acta Mol Basis Dis 1865(8):2008–23. https://doi.org/10.1016/j.bbadis.2018.11.014
Esteves AR, Arduino DM, Swerdlow RH, Oliveira CR, Cardoso SM (2009) Oxidative stress involvement in alpha-synuclein oligomerization in Parkinson’s disease cybrids. Antioxid Redox Signal 11(3):439–48. https://doi.org/10.1089/ARS.2008.2247
Swerdlow RH, Parks JK, Davis JN 2nd, Cassarino DS, Trimmer PA, Currie LJ et al (1998) Matrilineal inheritance of complex I dysfunction in a multigenerational Parkinson’s disease family. Ann Neurol 44(6):873–81. https://doi.org/10.1002/ana.410440605
Rana M, de Coo I, Diaz F, Smeets H, Moraes CT (2000) An out-of-frame cytochrome b gene deletion from a patient with parkinsonism is associated with impaired complex III assembly and an increase in free radical production. Ann Neurol 48(5):774–81
Thyagarajan D, Bressman S, Bruno C, Przedborski S, Shanske S, Lynch T et al (2000) A novel mitochondrial 12SrRNA point mutation in parkinsonism, deafness, and neuropathy. Ann Neurol 48(5):730–6
Muyderman H, Sims NR, Tanaka M, Fuku N, Raghupathi R, Thyagarajan D (2012) The mitochondrial T1095C mutation increases gentamicin-mediated apoptosis. Mitochondrion 12(4):465–71. https://doi.org/10.1016/j.mito.2012.06.006
Reeve AK, Ludtmann MH, Angelova PR, Simcox EM, Horrocks MH, Klenerman D et al (2015) Aggregated alpha-synuclein and complex I deficiency: exploration of their relationship in differentiated neurons. Cell Death Dis 6:e1820. https://doi.org/10.1038/cddis.2015.166
Suen DF, Narendra DP, Tanaka A, Manfredi G, Youle RJ (2010) Parkin overexpression selects against a deleterious mtDNA mutation in heteroplasmic cybrid cells. Proc Natl Acad Sci U S A 107(26):11835–40. https://doi.org/10.1073/pnas.0914569107
Villanueva-Paz M, Povea-Cabello S, Villalon-Garcia I, Alvarez-Cordoba M, Suarez-Rivero JM, Talaveron-Rey M et al (2020) Parkin-mediated mitophagy and autophagy flux disruption in cellular models of MERRF syndrome. Biochim Biophys Acta Mol Basis Dis 1866(6):165726. https://doi.org/10.1016/j.bbadis.2020.165726
Gomez-Duran A, Pacheu-Grau D, Lopez-Gallardo E, Diez-Sanchez C, Montoya J, Lopez-Perez MJ et al (2010) Unmasking the causes of multifactorial disorders: OXPHOS differences between mitochondrial haplogroups. Hum Mol Genet 19(17):3343–53. https://doi.org/10.1093/hmg/ddq246
Liou CW, Chuang JH, Chen JB, Tiao MM, Wang PW, Huang ST et al (2016) Mitochondrial DNA variants as genetic risk factors for Parkinson disease. Eur J Neurol 23(8):1289–300. https://doi.org/10.1111/ene.13020
Swerdlow RH, Parks JK, Cassarino DS, Binder DR, Bennett JP Jr, Di Iorio G et al (2001) Biochemical analysis of cybrids expressing mitochondrial DNA from Contursi kindred Parkinson’s subjects. Exp Neurol 169(2):479–85. https://doi.org/10.1006/exnr.2001.7674
Beylina A, Langston RG, Rosen D, Reed X, Cookson MR (2021) Generation of fourteen isogenic cell lines for Parkinson’s disease-associated leucine-rich repeat kinase (LRRK2). Stem Cell Res 53:102354. https://doi.org/10.1016/j.scr.2021.102354
Choi WS, Kim HW, Tronche F, Palmiter RD, Storm DR, Xia Z (2017) Conditional deletion of Ndufs4 in dopaminergic neurons promotes Parkinson’s disease-like non-motor symptoms without loss of dopamine neurons. Sci Rep 7:44989. https://doi.org/10.1038/srep44989
Yang X, Jiang J, Li Z, Liang J, Xiang Y (2021) Strategies for mitochondrial gene editing. Comput Struct Biotechnol J 19:3319–29. https://doi.org/10.1016/j.csbj.2021.06.003
Maeda K, Kawai H, Sanada M, Terashima T, Ogawa N, Idehara R et al (2016) Clinical Phenotype and Segregation of Mitochondrial 3243A>G Mutation in 2 Pairs of Monozygotic Twins. JAMA Neurol 73(8):990–3. https://doi.org/10.1001/jamaneurol.2016.0886
Rajasimha HK, Chinnery PF, Samuels DC (2008) Selection against pathogenic mtDNA mutations in a stem cell population leads to the loss of the 3243A–>G mutation in blood. Am J Hum Genet 82(2):333–43. https://doi.org/10.1016/j.ajhg.2007.10.007
DiMauro S, Schon EA, Carelli V, Hirano M (2013) The clinical maze of mitochondrial neurology. Nat Rev Neurol 9(8):429–44. https://doi.org/10.1038/nrneurol.2013.126
Rossignol R, Faustin B, Rocher C, Malgat M, Mazat JP, Letellier T (2003) Mitochondrial threshold effects. Biochem J 370(Pt 3):751–62. https://doi.org/10.1042/BJ20021594
Holt IJ, Harding AE, Petty RK, Morgan-Hughes JA (1990) A new mitochondrial disease associated with mitochondrial DNA heteroplasmy. Am J Hum Genet 46(3):428–33
Simon DK, Mayeux R, Marder K, Kowall NW, Beal MF, Johns DR (2000) Mitochondrial DNA mutations in complex I and tRNA genes in Parkinson’s disease. Neurology 54(3):703–9. https://doi.org/10.1212/wnl.54.3.703
Ikebe S, Tanaka M, Ozawa T (1995) Point mutations of mitochondrial genome in Parkinson’s disease. Brain Res Mol Brain Res 28(2):281–95. https://doi.org/10.1016/0169-328x(94)00209-w
Brown MD, Shoffner JM, Kim YL, Jun AS, Graham BH, Cabell MF et al (1996) Mitochondrial DNA sequence analysis of four Alzheimer’s and Parkinson’s disease patients. Am J Med Genet 61(3):283–9. https://doi.org/10.1002/(SICI)1096-8628(19960122)61:3%3c283::AID-AJMG15%3e3.0.CO;2-P
Vives-Bauza C, Andreu AL, Manfredi G, Beal MF, Janetzky B, Gruenewald TH et al (2002) Sequence analysis of the entire mitochondrial genome in Parkinson’s disease. Biochem Biophys Res Commun 290(5):1593–601. https://doi.org/10.1006/bbrc.2002.6388
Wei W, Keogh MJ, Wilson I, Coxhead J, Ryan S, Rollinson S et al (2017) Mitochondrial DNA point mutations and relative copy number in 1363 disease and control human brains. Acta Neuropathol Commun 5(1):13. https://doi.org/10.1186/s40478-016-0404-6
Delcambre S, Ghelfi J, Ouzren N, Grandmougin L, Delbrouck C, Seibler P et al (2020) Mitochondrial mechanisms of LRRK2 G2019S penetrance. Front Neurol 11:881. https://doi.org/10.3389/fneur.2020.00881
Gui YX, Xu ZP, Lv W, Zhao JJ, Hu XY (2015) Evidence for polymerase gamma, POLG1 variation in reduced mitochondrial DNA copy number in Parkinson’s disease. Parkinsonism Relat Disord 21(3):282–6. https://doi.org/10.1016/j.parkreldis.2014.12.030
Kraytsberg Y, Kudryavtseva E, McKee AC, Geula C, Kowall NW, Khrapko K (2006) Mitochondrial DNA deletions are abundant and cause functional impairment in aged human substantia nigra neurons. Nat Genet 38(5):518–20. https://doi.org/10.1038/ng1778
Bender A, Krishnan KJ, Morris CM, Taylor GA, Reeve AK, Perry RH et al (2006) High levels of mitochondrial DNA deletions in substantia nigra neurons in aging and Parkinson disease. Nat Genet 38(5):515–7. https://doi.org/10.1038/ng1769
Dolle C, Flones I, Nido GS, Miletic H, Osuagwu N, Kristoffersen S et al (2016) Defective mitochondrial DNA homeostasis in the substantia nigra in Parkinson disease. Nat Commun 7:13548. https://doi.org/10.1038/ncomms13548
Nido GS, Dolle C, Flones I, Tuppen HA, Alves G, Tysnes OB et al (2018) Ultradeep mapping of neuronal mitochondrial deletions in Parkinson’s disease. Neurobiol Aging 63:120–7. https://doi.org/10.1016/j.neurobiolaging.2017.10.024
Podlesniy P, Puigros M, Serra N, Fernandez-Santiago R, Ezquerra M, Tolosa E et al (2019) Accumulation of mitochondrial 7S DNA in idiopathic and LRRK2 associated Parkinson’s disease. EBioMedicine 48:554–67. https://doi.org/10.1016/j.ebiom.2019.09.015
Sanders LH, McCoy J, Hu X, Mastroberardino PG, Dickinson BC, Chang CJ et al (2014) Mitochondrial DNA damage: molecular marker of vulnerable nigral neurons in Parkinson’s disease. Neurobiol Dis 70:214–23. https://doi.org/10.1016/j.nbd.2014.06.014
Bender A, Schwarzkopf RM, McMillan A, Krishnan KJ, Rieder G, Neumann M et al (2008) Dopaminergic midbrain neurons are the prime target for mitochondrial DNA deletions. J Neurol 255(8):1231–5. https://doi.org/10.1007/s00415-008-0892-9
Grunewald A, Rygiel KA, Hepplewhite PD, Morris CM, Picard M, Turnbull DM (2016) Mitochondrial DNA depletion in respiratory chain-deficient Parkinson disease neurons. Ann Neurol 79(3):366–78. https://doi.org/10.1002/ana.24571
Larsson NG, Wang J, Wilhelmsson H, Oldfors A, Rustin P, Lewandoski M et al (1998) Mitochondrial transcription factor A is necessary for mtDNA maintenance and embryogenesis in mice. Nat Genet 18(3):231–6. https://doi.org/10.1038/ng0398-231
Woo DK, Green PD, Santos JH, D’Souza AD, Walther Z, Martin WD et al (2012) Mitochondrial genome instability and ROS enhance intestinal tumorigenesis in APC(Min/+) mice. Am J Pathol 180(1):24–31. https://doi.org/10.1016/j.ajpath.2011.10.003
Haddad D, Nakamura K (2015) Understanding the susceptibility of dopamine neurons to mitochondrial stressors in Parkinson’s disease. FEBS Lett 589(24 Pt A):3702–13. https://doi.org/10.1016/j.febslet.2015.10.021
Kowalska M, Piekut T, Prendecki M, Sodel A, Kozubski W, Dorszewska J (2020) Mitochondrial and nuclear DNA oxidative damage in physiological and pathological aging. DNA Cell Biol 39(8):1410–20. https://doi.org/10.1089/dna.2019.5347
Yakes FM, Van Houten B (1997) Mitochondrial DNA damage is more extensive and persists longer than nuclear DNA damage in human cells following oxidative stress. Proc Natl Acad Sci U S A 94(2):514–9. https://doi.org/10.1073/pnas.94.2.514
Zhao M, Wang Y, Li L, Liu S, Wang C, Yuan Y et al (2021) Mitochondrial ROS promote mitochondrial dysfunction and inflammation in ischemic acute kidney injury by disrupting TFAM-mediated mtDNA maintenance. Theranostics 11(4):1845–63. https://doi.org/10.7150/thno.50905
Gaare JJ, Nido GS, Sztromwasser P, Knappskog PM, Dahl O, Lund-Johansen M et al (2018) Rare genetic variation in mitochondrial pathways influences the risk for Parkinson’s disease. Mov Disord 33(10):1591–600. https://doi.org/10.1002/mds.64
Billingsley KJ, Barbosa IA, Bandres-Ciga S, Quinn JP, Bubb VJ, Deshpande C et al (2019) Mitochondria function associated genes contribute to Parkinson’s Disease risk and later age at onset. NPJ Parkinsons Dis 5:8. https://doi.org/10.1038/s41531-019-0080-x
di Bandettini Poggio M, Nesti C, Bruno C, Meschini MC, Schenone A, Santorelli FM (2013) Dopamine-agonist responsive Parkinsonism in a patient with the SANDO syndrome caused by POLG mutation. BMC Med Genet 14:105. https://doi.org/10.1186/1471-2350-14-105
Balafkan N, Tzoulis C, Muller B, Haugarvoll K, Tysnes OB, Larsen JP et al (2012) Number of CAG repeats in POLG1 and its association with Parkinson disease in the Norwegian population. Mitochondrion 12(6):640–3. https://doi.org/10.1016/j.mito.2012.08.004
Luoma P, Melberg A, Rinne JO, Kaukonen JA, Nupponen NN, Chalmers RM et al (2004) Parkinsonism, premature menopause, and mitochondrial DNA polymerase gamma mutations: clinical and molecular genetic study. Lancet 364(9437):875–82. https://doi.org/10.1016/S0140-6736(04)16983-3
Davidzon G, Greene P, Mancuso M, Klos KJ, Ahlskog JE, Hirano M et al (2006) Early-onset familial parkinsonism due to POLG mutations. Ann Neurol 59(5):859–62. https://doi.org/10.1002/ana.20831
Reeve A, Meagher M, Lax N, Simcox E, Hepplewhite P, Jaros E et al (2013) The impact of pathogenic mitochondrial DNA mutations on substantia nigra neurons. J Neurosci 33(26):10790–801. https://doi.org/10.1523/JNEUROSCI.3525-12.2013
Wasner K, Grunewald A, Klein C (2020) Parkin-linked Parkinson’s disease: From clinical insights to pathogenic mechanisms and novel therapeutic approaches. Neurosci Res 159:34–9. https://doi.org/10.1016/j.neures.2020.09.001
Rothfuss O, Fischer H, Hasegawa T, Maisel M, Leitner P, Miesel F et al (2009) Parkin protects mitochondrial genome integrity and supports mitochondrial DNA repair. Hum Mol Genet 18(20):3832–50. https://doi.org/10.1093/hmg/ddp327
Kuroda Y, Mitsui T, Kunishige M, Shono M, Akaike M, Azuma H et al (2006) Parkin enhances mitochondrial biogenesis in proliferating cells. Hum Mol Genet 15(6):883–95. https://doi.org/10.1093/hmg/ddl006
Stevens DA, Lee Y, Kang HC, Lee BD, Lee YI, Bower A et al (2015) Parkin loss leads to PARIS-dependent declines in mitochondrial mass and respiration. Proc Natl Acad Sci U S A 112(37):11696–701. https://doi.org/10.1073/pnas.1500624112
Pirooznia SK, Yuan C, Khan MR, Karuppagounder SS, Wang L, Xiong Y et al (2020) PARIS induced defects in mitochondrial biogenesis drive dopamine neuron loss under conditions of parkin or PINK1 deficiency. Mol Neurodegener 15(1):17. https://doi.org/10.1186/s13024-020-00363-x
Shin JH, Ko HS, Kang H, Lee Y, Lee YI, Pletinkova O et al (2011) PARIS (ZNF746) repression of PGC-1alpha contributes to neurodegeneration in Parkinson’s disease. Cell 144(5):689–702. https://doi.org/10.1016/j.cell.2011.02.010
Sanders LH, Laganiere J, Cooper O, Mak SK, Vu BJ, Huang YA et al (2014) LRRK2 mutations cause mitochondrial DNA damage in iPSC-derived neural cells from Parkinson’s disease patients: reversal by gene correction. Neurobiol Dis 62:381–6. https://doi.org/10.1016/j.nbd.2013.10.013
Ouzren N, Delcambre S, Ghelfi J, Seibler P, Farrer MJ, Konig IR et al (2019) Mitochondrial DNA deletions discriminate affected from unaffected LRRK2 mutation carriers. Ann Neurol 86(2):324–6. https://doi.org/10.1002/ana.25510
Wasner K, Smajic S, Ghelfi J, Delcambre S, Prada-Medina CA, Knappe E et al (2022) Parkin deficiency impairs mtDNA dynamics and propagates inflammation. Mov Disord. https://doi.org/10.1002/mds.29025
Podlesniy P, Vilas D, Taylor P, Shaw LM, Tolosa E, Trullas R (2016) Mitochondrial DNA in CSF distinguishes LRRK2 from idiopathic Parkinson’s disease. Neurobiol Dis 94:10–7. https://doi.org/10.1016/j.nbd.2016.05.019
Martin LJ, Pan Y, Price AC, Sterling W, Copeland NG, Jenkins NA et al (2006) Parkinson’s disease alpha-synuclein transgenic mice develop neuronal mitochondrial degeneration and cell death. J Neurosci 26(1):41–50. https://doi.org/10.1523/JNEUROSCI.4308-05.2006
Pozo Devoto VM, Dimopoulos N, Alloatti M, Pardi MB, Saez TM, Otero MG et al (2017) alphaSynuclein control of mitochondrial homeostasis in human-derived neurons is disrupted by mutations associated with Parkinson’s disease. Sci Rep 7(1):5042. https://doi.org/10.1038/s41598-017-05334-9
Little D, Luft C, Mosaku O, Lorvellec M, Yao Z, Paillusson S et al (2018) A single cell high content assay detects mitochondrial dysfunction in iPSC-derived neurons with mutations in SNCA. Sci Rep 8(1):9033. https://doi.org/10.1038/s41598-018-27058-0
Wang X, Becker K, Levine N, Zhang M, Lieberman AP, Moore DJ et al (2019) Pathogenic alpha-synuclein aggregates preferentially bind to mitochondria and affect cellular respiration. Acta Neuropathol Commun 7(1):41. https://doi.org/10.1186/s40478-019-0696-4
Zambon F, Cherubini M, Fernandes HJR, Lang C, Ryan BJ, Volpato V et al (2019) Cellular alpha-synuclein pathology is associated with bioenergetic dysfunction in Parkinson’s iPSC-derived dopamine neurons. Hum Mol Genet 28(12):2001–13. https://doi.org/10.1093/hmg/ddz038
Bender A, Desplats P, Spencer B, Rockenstein E, Adame A, Elstner M et al (2013) TOM40 mediates mitochondrial dysfunction induced by alpha-synuclein accumulation in Parkinson’s disease. PLoS ONE 8(4):e62277. https://doi.org/10.1371/journal.pone.0062277
Greene JC, Whitworth AJ, Kuo I, Andrews LA, Feany MB, Pallanck LJ (2003) Mitochondrial pathology and apoptotic muscle degeneration in Drosophila parkin mutants. Proc Natl Acad Sci U S A 100(7):4078–83. https://doi.org/10.1073/pnas.0737556100
Park J, Lee SB, Lee S, Kim Y, Song S, Kim S et al (2006) Mitochondrial dysfunction in Drosophila PINK1 mutants is complemented by parkin. Nature 441(7097):1157–61. https://doi.org/10.1038/nature04788
Clark IE, Dodson MW, Jiang C, Cao JH, Huh JR, Seol JH et al (2006) Drosophila pink1 is required for mitochondrial function and interacts genetically with parkin. Nature 441(7097):1162–6. https://doi.org/10.1038/nature04779
Kim Y, Park J, Kim S, Song S, Kwon SK, Lee SH et al (2008) PINK1 controls mitochondrial localization of Parkin through direct phosphorylation. Biochem Biophys Res Commun 377(3):975–80. https://doi.org/10.1016/j.bbrc.2008.10.104
Narendra D, Tanaka A, Suen DF, Youle RJ (2008) Parkin is recruited selectively to impaired mitochondria and promotes their autophagy. J Cell Biol 183(5):795–803. https://doi.org/10.1083/jcb.200809125
Narendra DP, Jin SM, Tanaka A, Suen DF, Gautier CA, Shen J et al (2010) PINK1 is selectively stabilized on impaired mitochondria to activate Parkin. PLoS Biol 8(1):e1000298. https://doi.org/10.1371/journal.pbio.1000298
Vives-Bauza C, Zhou C, Huang Y, Cui M, de Vries RL, Kim J et al (2010) PINK1-dependent recruitment of Parkin to mitochondria in mitophagy. Proc Natl Acad Sci U S A 107(1):378–83. https://doi.org/10.1073/pnas.0911187107
Matsuda N, Sato S, Shiba K, Okatsu K, Saisho K, Gautier CA et al (2010) PINK1 stabilized by mitochondrial depolarization recruits Parkin to damaged mitochondria and activates latent Parkin for mitophagy. J Cell Biol 189(2):211–21. https://doi.org/10.1083/jcb.200910140
Hasson SA, Kane LA, Yamano K, Huang CH, Sliter DA, Buehler E et al (2013) High-content genome-wide RNAi screens identify regulators of parkin upstream of mitophagy. Nature 504(7479):291–5. https://doi.org/10.1038/nature12748
Deas E, Plun-Favreau H, Gandhi S, Desmond H, Kjaer S, Loh SH et al (2011) PINK1 cleavage at position A103 by the mitochondrial protease PARL. Hum Mol Genet 20(5):867–79. https://doi.org/10.1093/hmg/ddq526
Yamano K, Youle RJ (2013) PINK1 is degraded through the N-end rule pathway. Autophagy 9(11):1758–69. https://doi.org/10.4161/auto.24633
Kane LA, Lazarou M, Fogel AI, Li Y, Yamano K, Sarraf SA et al (2014) PINK1 phosphorylates ubiquitin to activate Parkin E3 ubiquitin ligase activity. J Cell Biol 205(2):143–53. https://doi.org/10.1083/jcb.201402104
Kazlauskaite A, Kondapalli C, Gourlay R, Campbell DG, Ritorto MS, Hofmann K et al (2014) Parkin is activated by PINK1-dependent phosphorylation of ubiquitin at Ser65. Biochem J 460(1):127–39. https://doi.org/10.1042/BJ20140334
Koyano F, Okatsu K, Kosako H, Tamura Y, Go E, Kimura M et al (2014) Ubiquitin is phosphorylated by PINK1 to activate parkin. Nature 510(7503):162–6. https://doi.org/10.1038/nature13392
Lazarou M, Sliter DA, Kane LA, Sarraf SA, Wang C, Burman JL et al (2015) The ubiquitin kinase PINK1 recruits autophagy receptors to induce mitophagy. Nature 524(7565):309–14. https://doi.org/10.1038/nature14893
Palikaras K, Lionaki E, Tavernarakis N (2018) Mechanisms of mitophagy in cellular homeostasis, physiology and pathology. Nat Cell Biol 20(9):1013–22. https://doi.org/10.1038/s41556-018-0176-2
McWilliams TG, Prescott AR, Allen GF, Tamjar J, Munson MJ, Thomson C et al (2016) mito-QC illuminates mitophagy and mitochondrial architecture in vivo. J Cell Biol 214(3):333–45. https://doi.org/10.1083/jcb.201603039
Sun N, Yun J, Liu J, Malide D, Liu C, Rovira II et al (2015) Measuring in vivo mitophagy. Mol Cell 60(4):685–96. https://doi.org/10.1016/j.molcel.2015.10.009
McWilliams TG, Prescott AR, Montava-Garriga L, Ball G, Singh F, Barini E et al (2018) Basal mitophagy occurs independently of PINK1 in mouse tissues of high metabolic demand. Cell Metab 27(2):439–49. https://doi.org/10.1016/j.cmet.2017.12.008
Lee JJ, Sanchez-Martinez A, Martinez Zarate A, Beninca C, Mayor U, Clague MJ et al (2018) Basal mitophagy is widespread in Drosophila but minimally affected by loss of Pink1 or parkin. J Cell Biol 217(5):1613–22. https://doi.org/10.1083/jcb.201801044
Clark EH, de Vazquez Torre A, Hoshikawa T, Briston T (2021) Targeting mitophagy in Parkinson’s disease. J Biol Chem 296:100209. https://doi.org/10.1074/jbc.REV120.014294
Garcia-Prat L, Martinez-Vicente M, Perdiguero E, Ortet L, Rodriguez-Ubreva J, Rebollo E et al (2016) Autophagy maintains stemness by preventing senescence. Nature 529(7584):37–42. https://doi.org/10.1038/nature16187
Chen Y, Sparks M, Bhandari P, Matkovich SJ, Dorn GW 2nd (2014) Mitochondrial genome linearization is a causative factor for cardiomyopathy in mice and Drosophila. Antioxid Redox Signal 21(14):1949–59. https://doi.org/10.1089/ars.2013.5432
Sliter DA, Martinez J, Hao L, Chen X, Sun N, Fischer TD et al (2018) Parkin and PINK1 mitigate STING-induced inflammation. Nature 561(7722):258–62. https://doi.org/10.1038/s41586-018-0448-9
Zhao Y, Huang S, Liu J, Wu X, Zhou S, Dai K et al (2018) Mitophagy contributes to the pathogenesis of inflammatory diseases. Inflammation 41(5):1590–600. https://doi.org/10.1007/s10753-018-0835-2
Kamata H, Honda S, Maeda S, Chang L, Hirata H, Karin M (2005) Reactive oxygen species promote TNFalpha-induced death and sustained JNK activation by inhibiting MAP kinase phosphatases. Cell 120(5):649–61. https://doi.org/10.1016/j.cell.2004.12.041
Zhou R, Yazdi AS, Menu P, Tschopp J (2011) A role for mitochondria in NLRP3 inflammasome activation. Nature 469(7329):221–5. https://doi.org/10.1038/nature09663
Borsche M, Konig IR, Delcambre S, Petrucci S, Balck A, Bruggemann N et al (2020) Mitochondrial damage-associated inflammation highlights biomarkers in PRKN/PINK1 parkinsonism. Brain 143(10):3041–51. https://doi.org/10.1093/brain/awaa246
Ekstrand MI, Terzioglu M, Galter D, Zhu S, Hofstetter C, Lindqvist E et al (2007) Progressive parkinsonism in mice with respiratory-chain-deficient dopamine neurons. Proc Natl Acad Sci U S A 104(4):1325–30. https://doi.org/10.1073/pnas.0605208103
Song L, McMackin M, Nguyen A, Cortopassi G (2017) Parkin deficiency accelerates consequences of mitochondrial DNA deletions and Parkinsonism. Neurobiol Dis 100:30–8. https://doi.org/10.1016/j.nbd.2016.12.024
Baloh RH, Salavaggione E, Milbrandt J, Pestronk A (2007) Familial parkinsonism and ophthalmoplegia from a mutation in the mitochondrial DNA helicase twinkle. Arch Neurol 64(7):998–1000. https://doi.org/10.1001/archneur.64.7.998
Kiferle L, Orsucci D, Mancuso M, Lo Gerfo A, Petrozzi L, Siciliano G et al (2013) Twinkle mutation in an Italian family with external progressive ophthalmoplegia and parkinsonism: a case report and an update on the state of art. Neurosci Lett 556:1–4. https://doi.org/10.1016/j.neulet.2013.09.034
Perier C, Bender A, Garcia-Arumi E, Melia MJ, Bove J, Laub C et al (2013) Accumulation of mitochondrial DNA deletions within dopaminergic neurons triggers neuroprotective mechanisms. Brain 136(Pt 8):2369–78. https://doi.org/10.1093/brain/awt196
Samstag CL, Hoekstra JG, Huang CH, Chaisson MJ, Youle RJ, Kennedy SR et al (2018) Deleterious mitochondrial DNA point mutations are overrepresented in Drosophila expressing a proofreading-defective DNA polymerase gamma. PLoS Genet 14(11):e1007805. https://doi.org/10.1371/journal.pgen.1007805
Pickrell AM, Huang CH, Kennedy SR, Ordureau A, Sideris DP, Hoekstra JG et al (2015) Endogenous parkin preserves dopaminergic substantia nigral neurons following mitochondrial DNA mutagenic stress. Neuron 87(2):371–81. https://doi.org/10.1016/j.neuron.2015.06.034
Sterky FH, Hoffman AF, Milenkovic D, Bao B, Paganelli A, Edgar D et al (2012) Altered dopamine metabolism and increased vulnerability to MPTP in mice with partial deficiency of mitochondrial complex I in dopamine neurons. Hum Mol Genet 21(5):1078–89. https://doi.org/10.1093/hmg/ddr537
Kim HW, Choi WS, Sorscher N, Park HJ, Tronche F, Palmiter RD et al (2015) Genetic reduction of mitochondrial complex I function does not lead to loss of dopamine neurons in vivo. Neurobiol Aging 36(9):2617–27. https://doi.org/10.1016/j.neurobiolaging.2015.05.008
Gonzalez-Rodriguez P, Zampese E, Stout KA, Guzman JN, Ilijic E, Yang B et al (2021) Disruption of mitochondrial complex I induces progressive parkinsonism. Nature 599(7886):650–6. https://doi.org/10.1038/s41586-021-04059-0
Li J, Xue C, Gao Q, Tan J, Wan Z (2020) Mitochondrial DNA heteroplasmy rises in substantial nigra of aged PINK1 KO mice. Biochem Biophys Res Commun 521(4):1024–9. https://doi.org/10.1016/j.bbrc.2019.10.112
Kandul NP, Zhang T, Hay BA, Guo M (2016) Selective removal of deletion-bearing mitochondrial DNA in heteroplasmic Drosophila. Nat Commun 7:13100. https://doi.org/10.1038/ncomms13100
Xu H, DeLuca SZ, O’Farrell PH (2008) Manipulating the metazoan mitochondrial genome with targeted restriction enzymes. Science 321(5888):575–7. https://doi.org/10.1126/science.1160226
Ma H, Xu H, O’Farrell PH (2014) Transmission of mitochondrial mutations and action of purifying selection in Drosophila melanogaster. Nat Genet 46(4):393–7. https://doi.org/10.1038/ng.2919
Hill JH, Chen Z, Xu H (2014) Selective propagation of functional mitochondrial DNA during oogenesis restricts the transmission of a deleterious mitochondrial variant. Nat Genet 46(4):389–92. https://doi.org/10.1038/ng.2920
Zhang Y, Wang ZH, Liu Y, Chen Y, Sun N, Gucek M et al (2019) PINK1 inhibits local protein synthesis to limit transmission of deleterious mitochondrial DNA mutations. Mol Cell 73(6):1127–37. https://doi.org/10.1016/j.molcel.2019.01.013
Klann EM, Dissanayake U, Gurrala A, Farrer M, Shukla AW, Ramirez-Zamora A et al (2021) The Gut-Brain axis and its relation to Parkinson’s disease: a review. Front Aging Neurosci 13:782082. https://doi.org/10.3389/fnagi.2021.782082
Camus MF, Moore J, Reuter M (2020) Nutritional geometry of mitochondrial genetic effects on male fertility. Biol Lett 16(2):20190891. https://doi.org/10.1098/rsbl.2019.0891
Ma J, Liu X, Zhang Y, Cheng H, Gao W, Lai CQ et al (2022) Diet quality scores are positively associated with whole blood-derived mitochondrial DNA copy number in the Framingham heart study. J Nutr 152(3):690–7. https://doi.org/10.1093/jn/nxab418
Prasuhn J, Davis RL, Kumar KR (2020) Targeting mitochondrial impairment in Parkinson’s disease: challenges and opportunities. Front Cell Dev Biol 8:615461. https://doi.org/10.3389/fcell.2020.615461
Lee D, Shim MS, Kim KY, Noh YH, Kim H, Kim SY et al (2014) Coenzyme Q10 inhibits glutamate excitotoxicity and oxidative stress-mediated mitochondrial alteration in a mouse model of glaucoma. Invest Ophthalmol Vis Sci 55(2):993–1005. https://doi.org/10.1167/iovs.13-12564
Negida A, Menshawy A, El Ashal G, Elfouly Y, Hani Y, Hegazy Y et al (2016) Coenzyme Q10 for patients with Parkinson’s disease: a systematic review and meta-analysis. CNS Neurol Disord Drug Targets 15(1):45–53. https://doi.org/10.2174/1871527314666150821103306
Zhu ZG, Sun MX, Zhang WL, Wang WW, Jin YM, Xie CL (2017) The efficacy and safety of coenzyme Q10 in Parkinson’s disease: a meta-analysis of randomized controlled trials. Neurol Sci 38(2):215–24. https://doi.org/10.1007/s10072-016-2757-9
Snow BJ, Rolfe FL, Lockhart MM, Frampton CM, O’Sullivan JD, Fung V et al (2010) A double-blind, placebo-controlled study to assess the mitochondria-targeted antioxidant MitoQ as a disease-modifying therapy in Parkinson’s disease. Mov Disord 25(11):1670–4. https://doi.org/10.1002/mds.23148
Kujawska M, Jodynis-Liebert J (2018) Polyphenols in Parkinson’s disease: a systematic review of in vivo studies. Nutrients. https://doi.org/10.3390/nu10050642
Aw WC, Towarnicki SG, Melvin RG, Youngson NA, Garvin MR, Hu Y et al (2018) Genotype to phenotype: Diet-by-mitochondrial DNA haplotype interactions drive metabolic flexibility and organismal fitness. PLoS Genet 14(11):e1007735. https://doi.org/10.1371/journal.pgen.1007735
Bajracharya R, Youngson NA, Ballard JWO (2019) Dietary macronutrient management to treat mitochondrial dysfunction in Parkinson’s disease. Int J Mol Sci. https://doi.org/10.3390/ijms20081850
Lam J, McKeague M (2019) Dietary modulation of mitochondrial DNA damage: implications in aging and associated diseases. J Nutr Biochem 63:1–10. https://doi.org/10.1016/j.jnutbio.2018.07.003
Lai JH, Chen KY, Wu JC, Olson L, Brene S, Huang CZ et al (2019) Voluntary exercise delays progressive deterioration of markers of metabolism and behavior in a mouse model of Parkinson’s disease. Brain Res 1720:146301. https://doi.org/10.1016/j.brainres.2019.146301
Ferreira AFF, Binda KH, Singulani MP, Pereira CPM, Ferrari GD, Alberici LC et al (2020) Physical exercise protects against mitochondria alterations in the 6-hidroxydopamine rat model of Parkinson’s disease. Behav Brain Res 387:112607. https://doi.org/10.1016/j.bbr.2020.112607
Krumpolec P, Vallova S, Slobodova L, Tirpakova V, Vajda M, Schon M et al (2017) Aerobic-strength exercise improves metabolism and clinical state in Parkinson’s disease patients. Front Neurol 8:698. https://doi.org/10.3389/fneur.2017.00698
Okarmus J, Havelund JF, Ryding M, Schmidt SI, Bogetofte H, Heon-Roberts R et al (2021) Identification of bioactive metabolites in human iPSC-derived dopaminergic neurons with PARK2 mutation: altered mitochondrial and energy metabolism. Stem Cell Reports 16(6):1510–26. https://doi.org/10.1016/j.stemcr.2021.04.022
Pantic B, Ives D, Mennuni M, Perez-Rodriguez D, Fernandez-Pelayo U, de Lopez Arbina A et al (2021) 2-Deoxy-D-glucose couples mitochondrial DNA replication with mitochondrial fitness and promotes the selection of wild-type over mutant mitochondrial DNA. Nat Commun 12(1):6997. https://doi.org/10.1038/s41467-021-26829-0
Acknowledgements
Figures were created using elements from Servier Medical Art (https://smart.servier.com), which is licensed under a Creative Commons Attribution 3.0 Unported Generic License (https://creativecommons.org/licenses/by/3.0/).
Funding
This work was funded by an International Mobility grant of the Autonomous Province of Bolzano to ML under Project number D54I18000150003, Parkinson's disease: Research into mitochondria and autophagy – PRIMA, by the Department of Innovation, Research and University of the Autonomous Province of Bozen/Bolzano, Italy, through a core funding initiative to the Institute for Biomedicine, and by the Deutsche Forschungsgemeinschaft to AG, AAH and PPP (FOR2488).
Author information
Authors and Affiliations
Contributions
ML and IP conceived and wrote the manuscript. All authors commented on previous versions of the manuscript and read and approved the final manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors have no relevant financial or non-financial interests to disclose.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Lang, M., Grünewald, A., Pramstaller, P.P. et al. A genome on shaky ground: exploring the impact of mitochondrial DNA integrity on Parkinson’s disease by highlighting the use of cybrid models. Cell. Mol. Life Sci. 79, 283 (2022). https://doi.org/10.1007/s00018-022-04304-3
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00018-022-04304-3