Abstract
Oxidative stress can lead to plant growth retardation, yield loss, and death. The atr7 mutant of Arabidopsis thaliana exhibits pronounced tolerance to oxidative stress. Using positional cloning, confirmed by knockout and RNA interference (RNAi) lines, we identified the atr7 mutation and revealed that ATR7 is a previously uncharacterized gene with orthologs in other seed plants but with no homology to genes in lower plants, fungi or animals. Expression of ATR7-GFP fusion shows that ATR7 is a nuclear-localized protein. RNA-seq analysis reveals that transcript levels of genes encoding abiotic- and oxidative stress-related transcription factors (DREB19, HSFA2, ZAT10), chromatin remodelers (CHR34), and unknown or uncharacterized proteins (AT5G59390, AT1G30170, AT1G21520) are elevated in atr7. This indicates that atr7 is primed for an upcoming oxidative stress via pathways involving genes of unknown functions. Collectively, the data reveal ATR7 as a novel seed plants-specific nuclear regulator of oxidative stress response.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Increased levels of reactive oxygen species (ROS) in plants are a consequence of various adverse abiotic conditions such as drought, salinity, extreme temperatures, and pollutants, as well as biotic interactions that trigger the hypersensitive response (HR) to pathogens or programmed cell death (PCD) [1, 2]. An elaborate antioxidant system protects plants from ROS toxicity. In addition to their toxic nature, ROS are important signals that modulate plant growth, developmental programs, and responses to the environment [2]. ROS-induced PCD is essential for processes, such as embryo development, maturation of tracheal elements, formation of leaf shape, and leaf senescence.
Transcription factors (TFs) and regulators are induced under various stresses [3,4,5]. Some of them, such as the heat-inducible HSFA2 or the dehydration-responsive element binding protein (DREB)/C-repeat binding factor (CBF), activate other stress-responsive genes to confer tolerance to single or multiple stresses such as heat, drought, salt, cold, or oxidative stress [3,4,5,6,7,8]. However, our knowledge concerning the intricate ROS network that modulates stress responses, development, and cell death remains limited. Isolation and characterization of mutants with enhanced tolerance to ROS-induced PCD provide a direct way to identify components of the ROS network.
The oxidative stress-tolerant mutant atr7, previously obtained by chemical mutagenesis from its genetic background loh2 (Arabidopsis thaliana ecotype Wassilewskija), displays high tolerance to several ROS-inducing agents such as paraquat (PQ), the catalase inhibitor aminotriazole (AT), and the fungal AAL toxin [9]. PQ is mainly active in chloroplasts, where it generates superoxide radicals by transferring electrons from photosystem I/ferredoxin to molecular oxygen; the superoxide radicals are then quickly converted to hydrogen peroxide (H2O2) by the action of superoxide dismutases [10]. AT is a potent inhibitor of catalases, the main H2O2-detoxifying enzymes in plants, and inhibiting catalase activity leads to PCD similar to the cell death observed in catalase RNAi plants [11, 12]. The loh2 mutant, which is the genetic background of atr7, was obtained by knocking out a gene involved in ceramide synthesis [13] and was used to obtain the atr7 mutant by chemical mutagenesis [9]. Loh2 has the same phenotype as the wild-type, A. thaliana ecotype Wassilewskija, under normal conditions and displays the same sensitivity to PQ- and AT-induced oxidative stress as the A. thaliana ecotype Wassilewskija. For our study on atr7, we chose PQ as the ROS inducing agent. Here, we identify ATR7 by map-based cloning and show that it encodes a novel nuclear-localized protein with a previously unreported function. The gene is specific to seed plants; there are no homologs in lower plants (algae, ferns, lycopods, and mosses), fungi, and animals. Molecular analyses of the atr7 transcriptome (RNA-seq) and metabolome (GC–MS) identified genes and pathways that are highly up- and downregulated in atr7. Their potential role in the oxidative stress response is discussed.
Results
The atr7 mutant tolerates PQ-induced oxidative stress
When plants were grown on Murashige and Skoog (MS) media containing PQ, loh2 seedlings bleached and died while atr7 seedlings stayed green and alive (Fig. 1a), as previously reported [9]. When grown in soil and sprayed with PQ at the rosette stage, loh2 plants developed massive necrotic lesions while atr7 lacked cell death symptoms (Fig. 1b). Trypan blue staining confirmed massive necrosis in loh2 and the absence of cell death in atr7 (Fig. 1c).
Atr7 exhibits enhanced tolerance to oxidative stress. a 1-week-old loh2 and atr7 seedlings on Murashige and Skoog (MS) plant growth media (left) or MS media supplemented with 1 μM paraquat (PQ) (right). b Mature loh2 and atr7 plants grown in soil, sprayed with 15 μM PQ. c Rosette leaves from loh2 and atr7 plants sprayed with 15 μM PQ and stained with trypan blue to detect cell death
Treatment with PQ elevated the endogenous ROS levels, as assessed by staining with diaminobenzidine (detecting hydrogen peroxide) and gene expression analysis of two ROS marker genes (Supplementary Fig. 1). The increase in ROS was much more prominent in loh2 but also evident in atr7. Furthermore, atr7 had higher basal levels in the absence of stress, as compared with unstressed loh2 plants (Supplementary Fig. 1).
Molecular cloning of ATR7 by map-based approach
To identify the mutation responsible for the tolerance to oxidative stress, atr7 plants were crossed with A. thaliana ecotype Columbia-0. All F1 seedlings examined showed sensitivity to 1.5 µM PQ, similar to wild-type (WT) plants. The F2 population of 2909 individuals segregated in a 3 (susceptible): 1 (tolerant) Mendelian fashion, indicating that atr7 is a recessive mutation at a single nuclear locus. Coarse mapping with 50 PQ-tolerant plants located atr7 between the SSLP markers CA72 and NGA139 on chromosome 5 (Fig. 2a). Further fine mapping with a larger population of 604 individuals delimited the atr7 locus within a region of approximately 100 kb (Fig. 2a). Sequencing of the candidate genes in this region using the Illumina technology revealed a point mutation (C/G to T/A transition) in the first exon of gene AT5G21280, resulting in a premature stop codon (Fig. 2b, Supplementary Fig. 2a). Screening of the TAIR database (http://www.arabidopsis.org/) identified a knockout line (KO, SALK_006796) with a T-DNA insertion in the first exon of AT5G21280 (Fig. 2b). Homozygous KO plants were tolerant to PQ, like the originally isolated atr7 mutant (Fig. 2c). The ATR7 KO mutant does not have any obvious phenotype in the absence of stress with plant growth and fertility being normal. End-point RT-PCR with primers recognizing the ends of the ATR7 coding sequence confirmed the absence of full-length transcript in both atr7 and ATR7 KO (Supplementary Fig. 3). The lack of ATR7 expression in both the atr7 point mutant and the ATR7 KO mutant was verified by qRT-PCR with primers upstream of the T-DNA insertion. Additionally, we inhibited ATR7 expression by generating RNAi lines in both loh2 and the Wassilewskija background (Supplementary Fig. 2b). The resulting RNAi lines were as tolerant to PQ-induced oxidative stress as the atr7 mutant (Supplementary Fig. 2b). Finally, a complementation line expressing the wild-type AT5G21280 gene in the atr7 background showed sensitivity to PQ similar to that of wild-type plants (Supplementary Fig. 2c). Taken together, these results demonstrate that ATR7 is AT5G21280.
Identification of ATR7 by molecular cloning. a Genetic mapping of the atr7 mutation. Markers used for the fine mapping and their distance on chromosome 5 are presented above. The relative position of ATR7 between SNP4 and SNP5 is shown below. The red arrow indicates the direction of ATR7 transcription. b Genomic structure of ATR7 (AT5G21280) with the position of the atr7 point mutation (G → A) and the location of the T-DNA insertion in ATR7 in the knockout line SALK_006796 indicated (RB and LB are the right and left borders, respectively, of the T-DNA). c 1-week-old seedlings of loh2, atr7, atr7 knockout (SALK_006796), and wild-type Columbia-0 (Col) grown on Murashige-Skoog (MS) medium (left) or MS medium supplemented with 1 μM paraquat (MS + PQ, right). The loh2 and the wild-type Columbia-0 plants die on media with PQ (100% mortality), whereas all atr7 mutant and atr7 knockout (SALK_006796) plants survive on PQ (100% survival)
ATR7 encodes a novel nuclear-localized protein with unknown function
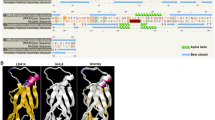
ATR7 encodes a previously uncharacterized protein of unknown function. The ATR7 protein has a length of 302 amino acids (Supplementary Fig. 4), a calculated molecular weight of 33,608.61 Da, and an estimated isoelectric point of 5.14. ATR7 has some similarity with hydroxyproline-rich glycoproteins (HRGPs), but unlike typical HRGPs exported to the cell wall via a signal peptide, ATR7 protein lacks a signal peptide and its hydropathy plot predicts the absence of trans-membrane domains. A multiple sequence alignment shows few stretches of conserved sequences, the longest one is shown in Supplementary Fig. 4. A Pfam search for that sequence did not identify any known motif/domain. Interestingly, however, a significant portion of the ATR7 protein is predicted to be intrinsically disordered in the Database of Disordered Protein Predictions (D2P2; http://d2p2.pro/).
The subcellular localization database for Arabidopsis proteins (SUBA3, http://suba.plantenergy.uwa.edu.au/) predicts nuclear localization of ATR7, which we confirmed by expressing an ATR7-GFP fusion from the Cauliflower Mosaic Virus (CaMV) 35S promoter in stably transformed transgenic plants (Fig. 3). The overexpression of the ATR7-GFP protein or the ATR7 protein alone did not result in any visible phenotype under optimal growth conditions or under PQ-induced oxidative stress.
Nuclear localization of ATR7. GFP signal is detected in nuclei of cells from leaves and roots of Arabidopsis plants stably transformed with the ATR7-GFP construct. Nuclei are counterstained with 4′,6-diamidino-2-phenylindole (DAPI). a DAPI-stained nucleus in the leaf. b GFP signal in the nucleus of the leaf. c Merged image. d DAPI-stained nucleus in the root. e GFP signal in the nucleus of the root. f Merged image
According to the PLAZA 4.0 database for comparative genomics (http://bioinformatics.psb.ugent.be/plaza/) [14], ATR7 homologs are present in most higher plant genomes. We extended the homology study further to cover additional plant genomes available to date and prepared a phylogenetic tree (Fig. 4). A comprehensive list of all ATR7 homologs is presented in Supplementary Table 1. Homologs are present in monocot and dicot crops such as rice, cucumber, cabbage, strawberry, and grapevine. Interestingly, however, no homologs of ATR7 were found in animals or fungi, and no genes with significant sequence similarities were found in lower plants (algae, mosses, ferns, and lycopods). However, there is a single match with Amborella trichopoda, a species believed to be one of the earliest angiosperm plants. A multiple sequence alignment (Supplementary Fig. 4) shows that ATR7 is more closely related to its homologs from the Brassicaceae family than to species from other families. Collectively, our observations indicate recent evolution of the ATR7 gene in flowering plants.
Phylogenetic tree of ATR7 and 80 orthologs from plants with sequenced genomes, available at Phytozome. Multiple sequence alignment of protein sequences was done using MUSCLE. The tree was inferred employing the neighbor-joining method using simple phylogeny after distance correction. ATR7 is shown in red. Description of the sequences is provided in Supplementary Table 1
ATR7 gene expression is induced under oxidative and abiotic stresses
Despite the fact that a mutation of ATR7 has such a striking effect on oxidative stress tolerance and cell death responses in A. thaliana, little is known about its molecular mode of action and its environmentally affected pattern of expression. To fill this gap, we analyzed RNA-seq data to determine how the ATR7 gene is expressed during development, in different organs/tissues, or during abiotic and oxidative stresses. According to the RNA-seq data available through Araport (https://www.araport.org/), ATR7 is expressed in all plant organs. However, the RNA-seq data were scarce, different experiments were not comparable with one another, and crucial information about ATR7 expression under stress was missing. Thus, we generated our own data by performing qRT-PCR expression profiling with plants subjected to oxidative (hydrogen peroxide) and abiotic stresses (heat, cold, CdCl2, and osmotic stress induced by mannitol). According to our data, ATR7 was notably induced by hydrogen peroxide, salt stress, CdCl2, and mannitol but not by sub-optimal temperatures such as cold or heat stress (Supplementary Fig. 5).
Transcriptome analysis reveals activation of both known stress protection genes and a set of novel genes in atr7 that collectively contribute to oxidative stress tolerance
To better understand ATR7´s mode of action, atr7 and its genetic background loh2 were grown under standard condition or subjected to PQ-induced oxidative stress, and RNA was isolated for transcriptome analysis by RNA sequencing (RNA-seq). PQ-induced oxidative stress caused significant transcriptional re-programming in both loh2 and atr7, with more prominent changes in loh2. Gene expression levels are given in Supplementary Table 2. The RNA-seq data were verified by qRT-PCR analysis of eight randomly selected genes. In the absence of stress, expression of 1054, 156, and 72 genes was at least 2-, 5- and 10-fold, respectively, altered (up or down) in atr7 compared to loh2. Importantly, while oxidative stress altered the expression of 7623, 2391, and 1029 genes by at least 2-, 5- and 10-fold in loh2, only 1942, 402, and 157 genes, respectively, were affected in atr7. The 100 genes that were most regulated (up or down) by PQ in loh2 and in atr7, as well as the 100 most regulated (up or down) genes in the atr7 mutant versus loh2 in the absence of stress, were used to generate Fig. 5a. Figure 5b presents an overlap of the genes regulated by more than 2- or 5-fold in the three comparisons (loh2 PQ vs. loh2, atr7 PQ vs. atr7, and atr7 vs. loh2).
Transcriptome re-programming due to paraquat-induced oxidative stress in loh2 and atr7. a Hierarchical linkage clustering of 263 genes representing the 100 most up- or downregulated genes after paraquat (PQ) treatment in the two genotypes, as well as the 100 most regulated genes (induced or repressed) between loh2 and atr7 grown under normal conditions (without stress). Atr7 and its genetic background loh2 were grown on Murashige and Skoog (MS) medium without PQ (unstressed controls) or with 1 μM PQ (oxidative stress), and gene expression was determined by RNA-seq. Each row represents the expression profile of an individual gene, given as mean subtracted average log2 gene expression values. Red color indicates upregulation while green indicates downregulation. Data are log2 normalized mean-centered TMM values. b Venn diagram of genes altered in expression in atr7 in the absence of stress (atr7 vs. loh2, green circles), as well as genes regulated by paraquat in loh2 (loh2 PQ vs. loh2, blue circle) and atr7 (atr7 PQ vs. atr7, red circle). Genes induced by at least 2- or 5-fold are given in red numbers, while genes repressed by at least 2- or 5-fold are in green numbers
Analysis of the differentially expressed genes with the largest fold change (up- and downregulated) represented in Fig. 5a revealed two distinct clusters. Cluster 1 contains genes with a similar expression level in loh2 and atr7 in the absence of stress (Fig. 5a). The genes in cluster 1A are repressed in both genetic backgrounds. It is enriched in genes encoding proteins related to cell wall expansion, growth, and redox processes, such as arabinogalactans, xyloglucan endotransglycosylases, various mono- and dioxygenases, oxidases, and cytochrome P450 proteins (Fig. 5a, Supplementary Table 3). Cluster 1B is enriched in genes encoding proteins involved in photosynthesis and carbon assimilation, such as RuBisCO small subunit, chlorophyll a/b binding protein 2, LHCII subunit B, and carbonic anhydrase (Fig. 5a). This cluster contains genes mainly downregulated in loh2, but not in atr7. This clearly shows that photosynthesis is inhibited in stress-sensitive loh2 but not in stress-tolerant atr7.
Cluster 2 contains genes with expression patterns that are rather different between loh2 and atr7, even in the absence of stress. A small group of genes was highly upregulated by PQ treatment in loh2, but not in atr7 (Fig. 5a, cluster 2A). Indeed, some of these genes were downregulated in atr7 under normal growth conditions and further repressed in the mutant exposed to PQ (Fig. 5a, cluster 2A). Representatives of this group are genes encoding dihydroneopterin aldolase, UDP-glucosyl transferase 76E12, and a glycosyl hydrolase. Sub-clusters 2B and 2C contain genes whose expression levels in unstressed plants were higher in atr7 than in loh2 (Fig. 5a). These clusters are enriched in plant-specific genes, several of which encode nuclear-localized proteins (enzymes, TFs, chromatin modifiers, etc.). Notably, a significant percentage (16%) of those genes has not been functionally characterized so far. The genes selected for clustering contained NAC transcription factors and heat shock proteins all of which are included in cluster 2C (Fig. 4a). Supplementary Table 3 contains the genes most regulated in atr7 in the absence of stress. Gene ontology (GO) enrichment analysis indicated that the two categories with the most upregulated genes in the atr7 mutant are “response to heat” and “rRNA processing”, while the category with the most downregulated genes is “photosynthesis” (Supplementary Fig. 6). Genes encoding key abiotic and oxidative stress-related transcription factors (DREB19, HSFA2, ZAT10, ZAT12) [9], chromatin remodelers (CHR34), and many genes encoding uncharacterized proteins (ANAC085, AT5G59390, AT1G30170, AT1G21520), were constantly upregulated in atr7 in the absence of stress (Table 1, Supplementary Table 3). Among the proteins encoded by the 100 most upregulated genes, 32% are predicted to be localized in the nucleus based on GO annotations. Combined with the nuclear localization of ATR7 protein, the two observations collectively suggest a functional role of ATR7 in the nucleus. It is likely that these key abiotic stress genes upregulated in atr7 jointly contribute to its abiotic and oxidative stress tolerance. DREB19, HSFA2, ZAT10, and ZAT12 have well-established roles in abiotic and oxidative stress responses, playing important roles in tolerance to drought, salinity, and high temperature, as well as for “thermomemory” [4,5,6, 15].
Pathway analysis of the gene expression data using MapMan revealed an upregulation of many abiotic stress-related genes in atr7, as well as of genes from the ubiquitin- and autophagy-dependent degradation pathways (Supplementary Fig. 7). Genes from the mitochondrial electron transport chain were also upregulated in atr7 (Supplementary Fig. 7). On the other hand, genes related to biotic stress, jasmonic acid signalling, chlorophyll biosynthesis, photosynthesis (light reactions, Calvin cycle), and photorespiration were downregulated in atr7 (Supplementary Fig. 7).
To identify genes co-expressed with ATR7, we used the CoNekT platform which uses > 900 A. thaliana samples to build a co-expression network for all genes. The cluster corresponding to ATR7 contains 60 genes (Supplementary Table 4). GO enrichment analysis of these genes showed enrichment of terms such as “response to hypoxia”, “signal transduction”, “regulation of ROS metabolic process”, “seed germination”, “lipid storage”, and “response to abiotic stimulus”. Among these genes, 33 were also induced (at least twofold) in loh2 upon PQ stress (Supplementary Table 4).
Metabolome reconfigurations contribute to the oxidative stress tolerance in atr7
To extend our knowledge about the molecular processes occurring during oxidative stress, we conducted metabolite profiling of loh2 and atr7 plants during PQ-induced oxidative stress. Relative metabolite levels in loh2 and atr7 plants grown in non-stress conditions and under PQ-induced oxidative stress are presented in Fig. 6 and Supplementary Table 5. The global picture reveals that, with a few exceptions, the metabolite profiles of loh2 and atr7 are very similar in the absence of stress. Principal component analysis confirms this (Supplementary Fig. 8). Upon PQ-induced oxidative stress, however, there are a number of changes in atr7, but the most dramatic changes occur in loh2. More specifically, the levels of many amino acids, such as alanine, valine, tyrosine, isoleucine, and lysine, were significantly elevated in loh2, compared with unstressed controls and with PQ-treated atr7. A similar pattern was observed for a number of other metabolites, including the organic acids pyruvic, citric, and benzoic acid. At the same time, other metabolites such as aspartic acid, fumaric acid, malic acid, and putrescine, significantly decreased in loh2 upon PQ treatment, but not in atr7. Moreover, putrescine and proline, two metabolites with prominent stress-protective roles, showed even higher levels in PQ-treated atr7 than in both loh2 and unstressed controls.
Relative levels of primary metabolites in loh2 and atr7 grown under normal conditions or under paraquat-induced oxidative stress (PQ). Red and blue depict increases and decreases, respectively, in content. Asterisks indicate values that are statistically different from the loh2 levels (Student’s t test, p < 0.05). The data are means of six biological replicates
The atr7 mutant is less sensitive to drought stress
The induction of many stress-related TFs in the atr7 mutant, as well as the induction of the ATR7 gene itself by oxidative and abiotic stresses, suggested that the atr7 mutant may be altered in abiotic stress responses. We evaluated the response of atr7 and loh2 to drought stress. Under the conditions used, loh2 showed cessation of growth, visible leaf damage, significant wilting (loss of water, as recorded by the decreased relative water content), and increased electrolyte leakage (Supplementary Fig. 9). Compared with loh2, atr7 showed much less visible damage of the leaves, less growth retardation, preserved relative water content, and showed little increase in ion conductivity (Supplementary Fig. 9).
Discussion
Previously, we showed that the atr7 mutant is more tolerant to oxidative stress induced by PQ and AT than loh2 [9]. Under normal growth conditions, atr7 looks a bit smaller than loh2 and its chlorophyll level is 90% of that of loh2 [9]. Treatment with PQ resulted in a complete loss of chlorophyll in loh2 and 100% plant mortality, while the chlorophyll level of atr7 was fully retained and all atr7 plants remained viable. Furthermore, using a home-made platform for qRT-PCR analysis, we demonstrated that several ROS marker genes and transcription factors are upregulated in loh2 under oxidative stress and these genes have higher basal expression in atr7 compared with loh2 in the absence of stress [9]. These preliminary results suggested a possible link with abiotic stress, as some of the transcription factor genes were known to be involved in abiotic stress responses.
Here, we show that the atr7 mutant, which has a striking oxidative stress-tolerant phenotype, carries a mutation in locus AT5G21280. We also show that the ATR7 gene encodes a novel nuclear-localized protein of unknown function. We identified ATR7 homologs in many flowering plants, including important crops, but no homologs were identified in lower plants such as algae, mosses, ferns, and lycopods. No studies on the biological role of the ATR7 gene or its protein have been published to date and none of the ATR7 homologs in other species have been functionally characterized. Furthermore, there are no known protein domains present that can be used to infer a putative function of the proteins.
While the atr7 mutant is more tolerant to both PQ and AT than loh2, other oxidative stress-tolerant mutants, such as par1, par2, pdr11, or oxt1, are more specific to either PQ or AT, respectively [10, 16,17,18]. Par2 enhances PQ tolerance by elevating nitric oxide (NO) levels, while pdr11 and par1 are blocked in transport of PQ into the cell or chloroplasts, respectively. PAR2 encodes an S-nitrozoglutathione reductase (GSNOR) involved in the metabolism of S-nitrosoglutathione (GSNO), a major bioactive donor of NO [10]. The increased NO levels in par2 upregulate the defense responses against PQ [10]. PDR11 (AT1G66950) encodes an ABC transporter localized in the plasmalemma, while PAR1 encodes an L-type amino acid transporter or amino acid permease localized in the Golgi apparatus [16, 17]. Thus, the two mutants show high levels of tolerance to PQ due to the inability to transport it into the cytoplasm and chloroplasts, but unlike atr7 they are not tolerant to other ROS inducing agents. Another PQ-tolerant mutant, pqt3, is defective in an E3 ubiquitin-protein ligase, which is a negative regulator of histone methylase PRMT4b [19]. PRMT4b itself can activate gene expression of the antioxidant genes ASCORBATE PEROXIDASE 1 (APX1) and GLUTATHIONE PEROXIDASE 1 (GPX1), thus alleviating oxidative stress. Two other mutants, oxt1 and oxt6, have been isolated as being more tolerant to AT. The oxt1 mutant is disrupted in a gene encoding ADENINE PHOSPHORIBOSYLTRANSFERASE (APT1), an enzyme that converts adenine to adenosine monophosphate (AMP), and accordingly oxt1 plants have elevated levels of adenine as well as elevated APX enzyme activities [18]. The oxt6 mutation results in the inactivation of a gene that encodes the 30-kD subunit of THE CLEAVAGE AND POLYADENYLATION SPECIFICITY FACTOR (CPSF30), which eventually results in upregulation of stress-protective genes in the oxt6 mutant [20].
Our RNA-seq analysis revealed clear differences between loh2 and atr7. Of note, many more genes were induced by PQ in loh2 than in atr7. This correlated with a higher level of oxidative stress in PQ-treated loh2. Besides the higher number of genes affected by PQ in loh2, it is obvious that the number of genes repressed by PQ in both loh2 and atr7 is higher than the number of PQ-induced genes. Another notable observation is the large number of genes regulated in the atr7 mutant compared with loh2 in the absence of stress (atr7 vs. loh2, unstressed controls). While the reason for this difference in gene expression is unclear, enhanced basal levels of hydrogen peroxide in atr7 may have contributed to the differential gene expression between atr7 and loh2 in the unstressed plants. The upregulated genes are much more than those downregulated (620 up-/434 downregulated by at least twofold; 116 up-/40 downregulated by at least fivefold).
The differentially expressed gene with the largest upregulation in the atr7 mutant encodes MBS1 (METHYLENE BLUE SENSITIVITY 1), a small zinc finger protein that mediates chloroplast-to-nucleus singlet oxygen (1O2) signaling and is essential for tolerance to photo-oxidative stress [21]. MBS1 is virtually not expressed in loh2, while it is well expressed both in stressed and unstressed atr7. The Arabidopsis loss-of-function mutant mbs1 is hypersensitive to photo-oxidative stress, whereas overexpression of MBS1 leads to enhanced stress tolerance [21]. MBS1 is also essential for acclimation to 1O2-induced oxidative stress and acts downstream of β-cyclocitral, a second messenger that mediates 1O2 responses [22]. Thus, upregulation of MBS1 in atr7 likely contributes to the primed condition against oxidative stress. The second most upregulated gene (AT1G23410) in atr7 encodes ribosomal protein S27a, whose function in Arabidopsis is unknown. In addition, several other ribosomal genes were highly upregulated in atr7 (Supplementary Table 3). GO analysis of the genes upregulated in atr7 indicated enrichment of ribosomal RNA-processing genes, genes involved in 60S ribosome biogenesis, and genes encoding ribosomal proteins, indicating a link between ATR7 and the ribosome (Supplementary Fig. 6).
A number of the genes with high expression in atr7 encode proteins that have not been functionally characterized or have no known functions. Among the top 20 most upregulated genes in atr7 in unstressed conditions, AT5G59390, AT1G30170, and AT1G21520 encode proteins with unknown functions and unknown sub-cellular locations. Incidentally, in a study on 50 proteins with unknown functions, overexpression of AT1G21520 in Arabidopsis was found to alleviate PQ-induced oxidative stress [23]. Thus, it is likely that AT1G21520 together with the other known and unknown proteins mentioned above collectively contribute to the enhanced oxidative stress tolerance of atr7.
The ATR7 gene is induced by a number of oxidative and abiotic stresses, including H2O2, PQ, CdCl2, salinity, and osmotic stress, but not by chilling and heat stress. This, together with the induction of many stress-related genes in the atr7 mutant, suggested a possible altered response of atr7 to abiotic stresses. Indeed, atr7 exhibited reduced sensitivity to drought stress, further substantiating the link between oxidative stress and drought. More studies are needed in the future to determine the response of atr7 in other abiotic stresses such as salt, heavy metals (CdCl2), and extreme temperatures.
Metabolome reconfigurations play important roles in the adaptation of plants to abiotic and oxidative stresses, as readjustments are needed both to maintain cellular homeostasis and to produce metabolites that can protect from stress [24]. The metabolome of PQ-stressed loh2 is strikingly different from the metabolome of unstressed loh2, while the metabolome of PQ-stressed atr7 is not so much different from that ofunstressed atr7. Particularly notable in this respect are the levels of stress metabolites including β-alanine, myo-inositol and proline as well as the amino acids that best reflect enhanced protein degradation, namely lysine and the branched chain and aromatic amino acids [25]. Putrescine and proline show the highest levels in PQ-treated atr7. Proline accumulation in particular is known to ameliorate several stresses [26]. Putrescine has a well-documented role in oxidative and abiotic stress tolerance, including drought stress [27]. With regard to PQ-induced oxidative stress, it has been suggested that putrescine competes with PQ for the polyamine transport system, thereby reducing the uptake of PQ [28]. This notion is supported by the discovery that natural variation in the polyamine transporter RMV1 (resistant to methyl viologen 1) determines PQ tolerance in Arabidopsis [29]. Further substantiating this link, the PQ-resistant mutant par1/pqr2 encodes a defective polyamine transporter [27]. All this collectively supports the conclusion that atr7 is tolerant to oxidative stress.
In conclusion, we identify ATR7 as a novel regulator of oxidative stress tolerance in Arabidopsis thaliana. The fact that oxidative- and abiotic stress-responsive transcription factors are upregulated in the atr7 mutant already in the absence of stress indicates that atr7 is primed for an upcoming oxidative stress. ATR7 is a nuclear protein that represses expression of key oxidative- and abiotic stress-related genes. Many of the genes with altered expression in atr7 encode proteins with unknown functions, suggesting that a previously unidentified molecular mechanism contributes to the oxidative stress tolerance in the atr7 mutant. ATR7 itself is a recently evolved gene, specific for seed plants and with no homologs in lower plants, fungi or animals. The enhanced tolerance of atr7 to drought stress and the presence of ATR7 homologs in agriculturally important species raise the possibility for crop improvement through modulation of ATR7 levels.
Materials and methods
Plant material, growth conditions, stress treatments, and stress assessment
The following plant material was used in this study: Arabidopsis thaliana ecotypes Columbia (Col-0) or Wassilewskija (Ws), Arabidopsis thaliana loh2 and atr7 mutants, described earlier [9, 29] and the atr7 knockout line SALK_006796 obtained from the Nottingham Arabidopsis Stock Center (http://arabidopsis.info/).
Plants were grown either in vitro on Murashige and Skoog (MS) plant media in Percival plant growth chambers (14 h light/10 h dark period, photosynthetic photon flux density 80 μmol m−2 s−1, 22 °C), or on soil under standard greenhouse conditions (14 h light/10 h dark period, photosynthetic photon flux density 400 μmol m−2 s−1, 22 °C and relative humidity 70%). Before sowing, seeds were surface sterilized for 2 h with gaseous chlorine derived from sodium hypochlorite and hydrochloric acid in a closed glass vessel. For most of the in vitro experiments, one-week-old plants were used for different measurements (RNA isolation, extraction of metabolites), while mature plants at rosette leaves stage were used for experiments on soil.
Paraquat (PQ) was either included in MS media at a concentration of 1 or 1.5 μM, or applied by spraying at concentrations of 15 or 25 μM. The presence of dead cells was shown by lacto-phenol trypan blue staining. In brief, PQ-treated and control, loh2 and atr7 rosette leaves were boiled in ethanol-diluted trypan blue solution (10 mL of phenol, 20 mL of 50% glycerol, 10 mL of lactic acid, 10 mL of distilled water, and 0.02 g of trypan blue) for 2 min, followed by 1 h incubation. De-staining was performed by several washings in saturated chloral hydrate solution (1 kg of chloral hydrate dissolved in 400 mL of distilled water, pH 1.2). Four decolorized leaves per plant were examined and the presence of cell death was observed visually.
For the evaluation of drought stress sensitivity, loh2 and atr7 plants were germinated on soil in standard greenhouse conditions. Water supply was stopped 2 weeks after germination and the results were recorded after 2 more weeks. Relative water content (RWC) was measured using the formula RWC (%) = [(FM − DM)/(TM − DM)] × 100, where FM, DM, and TM are the fresh, dry, and turgid masses of the leaves, respectively. DM was determined after drying the leaves at 80 °C for 48 h and TM was measured after immersing the leaves in H2O for 24 h. Electrolyte leakage, which is an indicator of cell damage, was evaluated by measuring the increase in conductivity with an HI 873 conductivity meter (Hanna Instruments, Woonsocket, RI, USA). Leaves from loh2 and atr7 plants grown at optimal conditions and under drought stress were briefly washed with ultrapure water (conductivity of 1 µS). The leaves were then incubated in ultrapure water for 10 min. The conductivity of the resultant solution was measured and compared with the total conductivity obtained after boiling the leaves.
DAB staining to detect hydrogen peroxide
The accumulation of H2O2 in plant tissues was visualized by histochemical detection, using DAB staining. To summarize, PQ-treated and control, loh2 and atr7 leaves were submerged in staining solution (1 mg/ml DAB (3, 3′-diaminobenzidine) in 0.05 M Tris acetate (pH 5)) in aluminium foil-wrapped tubes, followed by vacuum infiltration—two times for 10 min each at 25–100 mbar. Infiltrated samples were incubated in the dark overnight at room temperature. De-staining was performed by boiling in 96% ethanol for 5 min, discarding and replacing with fresh ethanol until no more color was leaching (3–4 times). One decolorized leaf per plant (× 4 plants) was examined and the presence of DAB staining was observed visually and photographed.
Genetic mapping and cloning of ATR7
As atr7 and its genetic background loh2 are in A. thaliana ecotype Wassilewskija (Ws), atr7 plants were crossed with A. thaliana ecotype Col-0 to generate a mapping population. F1 seedlings were also grown on MS media supplemented with 1.5 µM PQ to check if atr7 is dominant or recessive. Thereafter, an F2 population generated from a cross between Col-0 and atr7 was germinated on MS media supplemented with 1.5 µM PQ for 7–12 days to select PQ-tolerant individuals for genetic mapping. The selected plants were transferred to MS media without PQ for a few days before transferring them to pots containing soil. After DNA extraction, atr7 was mapped roughly on chromosome 5 using SSLP (Simple Sequence Length Polymorphism) markers from the TAIR database (The Arabidopsis Information Resource; www.arabidopsis.org) (Supplementary Table 6). Later, a larger F2 population of 604 PQ-tolerant mutant plants was used for fine mapping with SSLP, InDel (Insertion/Deletion) and SNP (Single-Nucleotide Polymorphism) markers (Supplementary Table 7). Potential SNPs were selected by randomly sequencing a 1-kb region of loh2 containing the two mutations. The SNP markers were designed using the Web SNAPER program.
After fine mapping of the atr7 mutation in a region of approximately 100 kb on chromosome 5, the atr7 and loh2 plants were sequenced to find SNPs in that region. The FLORACLEAN™ Plant DNA Isolation kit (MP Biomedicals, CA, USA) was used to obtain nuclear DNA with minimal chloroplast or mitochondrial DNA contamination. Genomic DNA from each of the mutants was sequenced using Illumina HiSeq 2000 (Illumina Inc., CA, USA) at the University Medical Center Groningen (UMCG), according to the manufacturer’s protocol. Sequence contigs of ~ 200 bp obtained from atr7 and loh2 were then separately aligned to the Col-0 ecotype reference genome sequence (GenBank accessions: Chromosome 1, NC_003070, Chromosome 2, NC_003071, Chromosome 3, NC_003074, Chromosome 4 NC_003075, and chromosome 5, NC_003076). SNP list was generated with the help of CLC-Bio software (Qiagen, Hilden, Germany).
Isolation of homozygous atr7 knockout plants, construction of plasmids for complementation analysis, RNAi and overexpression lines, and plant transformation
T-DNA knockouts of the ATR7 gene (locus AT5G21280) were identified from the available lines in the TAIR database (http://www.arabidopsis.org/). One line (KO, SALK_006796) with a T-DNA insertion in the first exon of AT5G21280, close to the nonsense mutation of atr7, was selected. Isolation of homozygous plants was done by genotyping using primers specific for the ATR7 gene flanking the position of the insert and a primer recognizing the left border of the T-DNA (Supplementary Table 8).
To complement the atr7 mutation with a functional ATR7 gene and restore the PQ-sensitive phenotype, the full-length ATR7 gene including both the promoter and 5´-UTR region was amplified from loh2 genomic DNA using TaKaRa Taq™ Polymerase and primers PrRuG3477 and PrRuG3479 (Supplementary Table 8). The amplified product was first cloned into pGEM-T Easy vector (Promega, WI, USA) and then subsequently cloned into the plant expression vector pGreen II 0229 [30]. The integrity of the construct was confirmed by restriction digestion and sequencing. Subsequently, Agrobacterium-mediated gene transfer into atr7 was performed and transgenics were selected using the herbicide Basta [31].
To generate ATR7 RNAi lines, a partial coding region of AT5G21280 was amplified using primers PrRuG3485 and PrRuG3486 (Supplementary Table 8) and cloned into the RNAi vector pFGC5941 (GenBank Accession No. AY310901; Arabidopsis Biological Resource Center stock number CD3-447). The amplified product was cloned in forward orientation using “inner” restriction enzyme sites AscI/SwaI and in reverse orientation using “outer” restriction enzyme sites BamHI/XbaI into plasmid pFGC5941. Thus, two reverse sequences of partial coding region of AT5G21280 are separated by a CHSA intron spacer in the pFGC5941 construct. The integrity of the construct was confirmed by restriction digestion and sequencing. The RNAi construct was transformed into both loh2 and A. thaliana ecotype Wassilewskija via Agrobacterium-mediated transfer. Transgenic plants were selected using the herbicide Basta [31].
Analysis of ATR7 subcellular localization using GFP fusion
The full-length ATR7 coding sequence was amplified without its stop codon from Arabidopsis Col-0 cDNA and the PCR product was cloned into vector pENTR/D–TOPO using pENTR Directional TOPO Cloning kit (Invitrogen, CA, USA). The sequence-verified entry clone was then transferred to vector pK7FWG2.0 using GATEWAY cloning (LR recombination reaction, Invitrogen, CA, USA). The resulting Prom35S:ATR7-GFP construct was verified by sequencing and introduced into Arabidopsis thaliana ecotype Col-0 plants either by transfecting mesophyll cell protoplasts or by Agrobacterium-mediated transformation using the floral dip method. For the transfection of mesophyll protoplasts, 40 µg plasmid DNA (Prom35S:ATR7-GFP) was added to 200 µL of the protoplasts suspension and incubated with the same volume of PEG solution (40% PEG 3500, 0.2 M mannitol, 0.1 M CaCl2). After transfection, the samples were diluted with 3 mL of W5 solution (154 mM NaCl, 125 mM CaCl2, 5 mM KCl, 2 mM MES, pH 5.7), collected by centrifugation at 100 g for 1 min, re-suspended in 1 mL W5 solution, and incubated in dark for 8–12 h in growth chamber before visualization. Agrobacterium-transformed transgenic plants were selected on MS media supplemented with 50 mg L−1 kanamycin. The presence of GFP signal was observed using a Leica TCS SP5 confocal laser scanning microscope. Nuclei were visualized with 4′,6-diamidino-2-phenylindole (DAPI) staining. Briefly, leaves were vacuum infiltrated with 1 µgmL−1 DAPI for 30 min. After rinsing in water, they were immediately used for microscopic analysis using a Leica TCS SP5 confocal microscope. DAPI was excited using the 405 nm laser and emission was collected between 440 and 470 nm.
RNA isolation and qRT-PCR analysis of ATR7 gene expression under environmental stresses
Seedlings from 1-week-old loh2 plants were subjected to different abiotic stresses (heat at 45 °C for 1 h, cold at 1 °C for 2 h, growth on 150 mM NaCl, 250 μM CdCl2, 200 mM mannitol, and 10 mM H2O2) and RNA was extracted with Trizol reagent (Invitrogen) according to the manufacturer’s recommendations. Ten micrograms of total RNA was treated with DNA-free™ Kit (Ambion) to remove eventual DNA contamination. RNA integrity was checked on 1% (w/v) agarose gel and concentration measured with a Nanodrop ND-2000 spectrophotometer before and after DNase I digestion. Additionally, the quality and integrity of the RNA samples were analyzed on an RNA 6000 Lab-on-a-Chip using the Bioanalyzer 2100 (Agilent Technologies, Santa Clara, CA, USA). cDNA was synthesized from 2 µg of total RNA using RevertAid™ First Strand cDNA Synthesis Kit (Fermentas) with oligo-dT primers, according to the manufacturer’s instructions.
Quantitative real-time PCR (qRT-PCR) analysis was performed using an ABI PRISM 7900 HT PCR instrument (Applied Biosystems, Darmstadt, Germany). The following primers, designed using Primer3 software, were used for the qRT-PCR analysis of ATR7 gene expression under abiotic and oxidative stresses: GTGGTGACGTCAGCTTGG (ATR7 forward) and AAGGAAATTCCATGACGTCAC (ATR7 reverse). For the analysis of ATR7 gene expression in the ATR7 KO mutant, the following primers were used: CAAGCAAGAACCACGCGTCT (forward) and GCGACGAGCTTCAGCCATGT (reverse). All reactions contained 10 μL of SYBR Green Master Mix (Applied Biosystems), 25 ng of cDNA, and 200 nM of each gene-specific primer in a final volume of 20 μL. The qRT-PCRs were executed using the following program: 50 °C for 2 min, 95 °C for 10 min, followed by 40 cycles of 95 °C for 15 s and 60 °C for 1 min. Relative mRNA abundance was calculated using the comparative 2−ΔΔCt method and normalized to the corresponding reference gene levels [32].
Transcriptional profiling by RNA sequencing
For RNA-seq transcriptional profiling, total RNA was isolated from loh2 and atr7 seedlings grown on MS media or MS media supplemented with PQ using NucleoSpin® RNA Plant kit of Machery-Nagel (Germany) according to the manufacturer’s protocol (http://www.mn-net.com/tabid/1327/default.aspx). DNase treatment to eliminate DNA contamination is included in the protocol. The concentration of the samples was analyzed with a NanoDrop ND-2000 spectrophotometer (Thermo Fisher Scientific, MA, USA). The quality and integrity of the RNA samples were analyzed on an RNA 6000 Lab-on-a-Chip using the Bioanalyzer 2100 (Agilent Technologies, CA, USA). Sample quality met the requirements for sample preparation. Illumina mRNA-Seq Sample Prep Kits were used to process the samples according to the Illumina protocol ´Preparing Samples for Sequencing of mRNA´ (1,004,898 Rev. D). Briefly, mRNA was isolated from the total RNA using poly-dT-oligo-attached magnetic beads. After fragmentation of the mRNA, cDNA synthesis was performed. The cDNA was used for ligation of the sequencing adapters and subsequent PCR amplification. The quality and yield after sample preparation were measured with a DNA 1000 Lab-on-a-Chip. The size of the resulting products was consistent with the expected size distribution (a broad peak of approximately 200-500 bp).
Sequencing using the Illumina HiSeq 2000 was performed (51 bp single end reads) according to manufacturer’s protocols at ServiceXS (Leiden, The Netherlands). A total of 4.5 pmol of DNA was used. Image analysis, base calling, and quality check were performed with the Illumina data analysis pipeline RTA v1.13.48 and/or OLB v1.9 and CASAVA v1.8.2.
Bioinformatics analysis
Quality of obtained read sequences was tested with FastQC (http://www.bioinformatics.babraham.ac.uk/projects/fastqc). As no overrepresented sequences were detected, an additional 3′-trimming step was skipped.
Quantification was done using kallisto (v.0.43.0; bootstraps: 100) [33] against cDNA sequences (Arabidopsis thaliana: Araport11) [34]. Differential expression analysis was carried out using EdgeR package in R/Bioconductor [35]. FDR cutoff of ≤ 0.05 and log2 fold change ≥ 1 were used to identify significantly differentially expressed genes. Heatmaps and clustering for selected groups of genes were made using pheatmap R-package [36]. Significantly enriched GO terms were identified using GOSeq package [37] in R/Bioconductor with FDR cutoff of ≤ 0.05. Pathway analysis was done using MapMan (v 3.5.1R2) [38]. Co-expressed genes were identified using the CoNekT platform [39]. The multiple sequence alignment of protein sequences, obtained from Phytozome, was established using MUSCLE and the phylogenetic tree was constructed using Simple Phylogeny [40].
Metabolome analysis of primary and secondary metabolites
Primary metabolites were determined by a previously established GC–MS protocol [41], using seedlings as plant material and six biological replicates. Chromatograms and mass spectra were evaluated by Chroma TOF® 4.2 (Leco, MI, USA) and TagFinder 4.0 for the quantification and annotation of the peaks using the MPI Golm Metabolome Database (GMD, http://gmd.mpimp-golm.mpg.de/) [42]. The parameters used for the identification of the metabolites [43] are summarized in Supplementary Table 5. The amounts of metabolites were analyzed as relative metabolite abundances calculated by normalization of signal intensity to that of ribitol, which was added as an internal standard, and then by the fresh weight of the material. The whole dataset is provided in Supplementary Table 5. Data analysis was done using MetaboAnalyst 2.0 (www.metaboanalyst.ca) [44, 45].
Data and materials availability
Sequencing data is available at NCBI-SRA under BioProject ID: PRJNA475098.
References
Apel K, Hirt H (2004) Reactive oxygen species: metabolism, oxidative stress, and signal transduction. Annu Rev Plant Biol 55:373–399
Gechev T, Van Breusegem F, Stone JM, Denev I, Laloi C (2006) Reactive oxygen species as signals that modulate plant stress responses and programmed cell death. Bioessays 28:1091–1101
Jung HS, Crisp PA, Estavillo GM, Cole B, Hong F, Mockler TC et al (2013) Subset of heat-shock transcription factors required for the early response of Arabidopsis to excess light. Proc Natl Acad Sci USA 110:14474–14479
Krishnaswamy S, Verma S, Rahman MH, Kav NN (2011) Functional characterization of four APETALA2-family genes (RAP2. 6, RAP2. 6L, DREB19 and DREB26) in Arabidopsis. Plant Mol Biol 75:107–127
Lämke J, Brzezinka K, Altmann S, Bäurle I (2016) A hit-and-run heat shock factor governs sustained histone methylation and transcriptional stress memory. EMBO J 35:162–175
Li C, Chen Q, Gao X, Qi B, Chen N, Xu S et al (2005) AtHsfA2 modulates expression of stress responsive genes and enhances tolerance to heat and oxidative stress in Arabidopsis. Sci China Ser C: Life Sci 48:540–550
Liu Q, Kasuga M, Sakuma Y, Abe H, Miura S, Yamaguchi-Shinozaki K, Shinozaki K (1998) Two transcription factors, DREB1 and DREB2, with an EREBP/AP2 DNA binding domain separate two cellular signal transduction pathways in drought-and low-temperature-responsive gene expression, respectively, in Arabidopsis. Plant Cell 10:1391–1406
Ogawa D, Yamaguchi K, Nishiuchi T (2007) High-level overexpression of the Arabidopsis HsfA2 gene confers not only increased thermotolerance but also salt/osmotic stress tolerance and enhanced callus growth. J Exp Bot 58:3373–3383
Mehterov N, Balazadeh S, Hille J, Toneva V, Mueller-Roeber B, Gechev T (2012) Oxidative stress provokes distinct transcriptional responses in the stress-tolerant atr7 and stress-sensitive loh2 Arabidopsis thaliana mutants as revealed by multi-parallel quantitative real-time PCR analysis of ROS marker and antioxidant genes. Plant Physiol Biochem 59:20–29
Chen R, Sun S, Wang C, Li Y, Liang Y, An F et al (2009) The Arabidopsis PARAQUAT RESISTANT2 gene encodes an S-nitrosoglutathione reductase that is a key regulator of cell death. Cell Res 19:1377–1387
Gechev T, Minkov I, Hille J (2005) Hydrogen peroxide-induced cell death in Arabidopsis: transcriptional and mutant analysis reveals a role of an oxoglutaratedependent dioxygenase gene in the cell death process. IUBMB Life 57:181–188
Vanderauwera S, Zimmermann P, Rombauts S, Vandenabeele S, Langebartels C, Gruissem W, Van Breusegem F (2005) Genome-wide analysis of hydrogen peroxide-regulated gene expression in Arabidopsis reveals a high light-induced transcriptional cluster involved in anthocyanin biosynthesis. Plant Physiol 139:806–821
Gechev TS, Gadjev IZ, Hille J (2004) An extensive microarray analysis of AAL-toxin-induced cell death in Arabidopsis thaliana brings new insights into the complexity of programmed cell death in plants. Cell Mol Life Sci 61:1185–1197
Van Bel M, Diels T, Vancaester E, Kreft L, Botzki A, Van de Peer Y, Coppens F, Vandepoele K (2017) PLAZA 4.0: an integrative resource for functional, evolutionary and comparative plant genomics. Nucl Acids Res 46(D1):D1190–D1196
Mittler R, Kim Y, Song L, Coutu J, Coutu A, Ciftci-Yilmaz S et al (2006) Gain-and loss-of-function mutations in Zat10 enhance the tolerance of plants to abiotic stress. FEBS Lett 580:6537–6542
Xi J, Xu P, Xiang CB (2012) Loss of AtPDR11, a plasma membrane-localized ABC transporter, confers paraquat tolerance in Arabidopsis thaliana. Plant J 69:782–791
Li J, Mu J, Bai J, Fu F, Zou T, An F, Yang S (2013) Paraquat Resistant1, a Golgi-localized putative transporter protein, is involved in intracellular transport of paraquat. Plant Physiol 162:470–483
Sukrong S, Yun KY, Stadler P, Kumar C, Facciuolo T, Moffatt BA, Falcone DL (2012) Improved growth and stress tolerance in the Arabidopsis oxt1 mutant triggered by altered adenine metabolism. Mol Plant 5:1310–1332
Luo C, Cai XT, Du J, Zhao TL, Wang PF, Zhao PX, Liu R, Xie Q, Cao XF, Xiang CB (2016) PARAQUAT TOLERANCE3 is an E3 ligase that switches off activated oxidative response by targeting histone-modifying PROTEIN METHYLTRANSFERASE4b. PLoS Genet 12(9):e1006332
Zhang J, Addepalli B, Yun KY, Hunt AG, Xu R, Rao S et al (2008) A polyadenylation factor subunit implicated in regulating oxidative signaling in Arabidopsis thaliana. PLoS ONE 3(6):e2410
Shao N, Duan GY, Bock R (2013) A mediator of singlet oxygen responses in Chlamydomonas reinhardtii and Arabidopsis identified by a luciferase-based genetic screen in algal cells. Plant Cell 25:4209–4226
Shumbe L, D’Alessandro S, Shao N, Chevalier A, Ksas B, Bock R, Havaux M (2017) METHYLENE BLUE SENSITIVITY 1 (MBS1) is required for acclimation of Arabidopsis to singlet oxygen and acts downstream of β-cyclocitral. Plant, Cell Environ 40:216–226
Luhua S, Ciftci-Yilmaz S, Harper J, Cushman J, Mittler R (2008) Enhanced tolerance to oxidative stress in transgenic Arabidopsis plants expressing proteins of unknown function. Plant Physiol 148:280–292
Obata T, Fernie AR (2012) The use of metabolomics to dissect plant responses to abiotic stresses. Cell Mol Life Sci 69:3225–3243
Araújo WL, Tohge T, Ishizaki K, Leaver CJ, Fernie AR (2011) Protein degradation—an alternative respiratory substrate for stressed plants. Trends Plant Sci 16:489–498
Kavi Kishor PB, Sreenivasulu N (2014) Is proline accumulation per se correlated with stress tolerance or is proline homeostasis a more critical issue? Plant, Cell Environ 37:300–311
Wu H, Fu B, Sun P, Xiao C, Liu JH (2016) A NAC transcription factor represses putrescine biosynthesis and affects drought tolerance. Plant Physiol 172:1532–1547
Dong S, Hu H, Wang Y, Xu Z, Zha Y, Cai X et al (2016) A pqr2 mutant encodes a defective polyamine transporter and is negatively affected by ABA for paraquat resistance in Arabidopsis thaliana. J Plant Res 129:899–907
Fujita M, Fujita Y, Iuchi S, Yamada K, Kobayashi Y, Urano K et al (2012) Natural variation in a polyamine transporter determines paraquat tolerance in Arabidopsis. Proc Natl Acad Sci USA 109:6343–6347
Hellens RP, Edwards EA, Leyland NR, Bean S, Mullineaux PM (2000) pGreen: a versatile and flexible binary Ti vector for Agrobacterium-mediated plant transformation. Plant Mol Biol 42:819–832
Zhang X, Henriques R, Lin SS, Niu QW, Chua NH (2006) Agrobacterium-mediated transformation of Arabidopsis thaliana using the floral dip method. Nat Protoc 1:641–646
Schmittgen TD, Livak KJ (2008) Analyzing real-time PCR data by the comparative C(T) method. Nat Protoc 3:1101–1108
Bray NL, Pimentel H, Melsted P, Pachter L (2016) Near-optimal probabilistic RNA-seq quantification. Nat. Biotech 34:525–527
Cheng CY, Krishnakumar V, Chan AP, Thibaud-Nissen F, Schobel S, Town CD (2017) Araport11: a complete reannotation of the Arabidopsis thaliana reference genome. Plant J 89:789–804
Robinson MD, McCarthy DJ, Smyth GK (2010) edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26:139–140
Raivo K (2015) Pheatmap: Pretty Heatmaps. R package version 1.0.8. https://CRAN.R-project.org/package=pheatmap
Young MD, Wakefield MJ, Smyth GK, Oshlack A (2010) Gene ontology analysis for RNA-seq: accounting for selection bias. Genome Biol 11:R14
Thimm O, Bläsing O, Gibon Y, Nagel A, Meyer S, Krüger P, Selbig J, Müller LA, Rhee SY, Stitt M (2004) MAPMAN: a user-driven tool to display genomics data sets onto diagrams of metabolic pathways and other biological processes. Plant J 37:914–939
Proost S, Mutwil M (2018) CoNekT: an open-source framework for comparative genomic and transcriptomic network analyses. Nucl Acids Res 46(W1):W133–W140
Chojnacki S, Cowley A, Lee J, Foix A, Lopez R (2017) Programmatic access to bioinformatics tools from EMBL-EBI update: 2017. Nucl Acids Res 45(W1):W550–W553
Lisec J, Schauer N, Kopka J, Willmitzer L, Fernie AR (2006) Gas chromatography mass spectrometry-based metabolite profiling in plants. Nat Protoc 1:387–396
Kopka J, Schauer N, Krueger S, Birkemeyer C, Usadel B, Bergmüller E et al (2005) GMD@ CSB. DB: the Golm metabolome database. Bioinformatics 21:1635–1638
Fernie AR, Aharoni A, Willmitzer L, Stitt M, Tohge T, Kopka J et al (2011) Recommendations for reporting metabolite data. Plant Cell 23:2477–2482
Xia J, Sinelnikov I, Han B, Wishart DS (2015) MetaboAnalyst 3.0—making metabolomics more meaningful. Nucl Acids Res 43:W251–W257
Xia J, Wishart DS (2016) Using MetaboAnalyst 3.0 for comprehensive metabolomics data analysis. Curr Prot Bioinf 55:14.10.1–14.10.91
Acknowledgements
The authors thank Dr. Christian Kappel (University of Potsdam, Potsdam, Germany) for providing the infrastructure for bioinformatics analysis, and Dr. Veselin Petrov (Center of Plant Systems Biology and Biotechnology, CPSBB, Plovdiv, Bulgaria) for technical assistance. This work was financially supported by the Swiss Enlargement Contribution in the framework of the Bulgarian-Swiss Research Programme (grant No. IZEBZ0_143003/1), the European Union FP7 Project PlantSurvivor (GA No. 329816), and the European Union H2020 Projects CropStrengthen (GA No. 642901) and PlantaSYST (SGA-CSA No. 739582 under FPA No. 664620). Collaboration between the German and Bulgarian teams was funded by the Federal Ministry of Education and Research (BMBF), Germany, within the PlantINNO Project (FKZ: 01D516023A).
Author information
Authors and Affiliations
Contributions
TG and JH designed the project. TG, JH, BMR, SB, DW, and AFR suggested and supervised the experiments and provided financial support. NS, NM, TG, MKQ, TO, MB, NS, and MAO performed the wet lab experiments. SG, AF, SP, and DW performed the computational analyses. AF, SP, SG, TG, DW, SB, BMR, and ARF analyzed the data. TG, NS, BMR, JH, and ARF wrote the paper. All coauthors have read and approved the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Sujeeth, N., Mehterov, N., Gupta, S. et al. A novel seed plants gene regulates oxidative stress tolerance in Arabidopsis thaliana. Cell. Mol. Life Sci. 77, 705–718 (2020). https://doi.org/10.1007/s00018-019-03202-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00018-019-03202-5