Summary
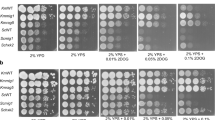
The rag2 mutant of Kluyveromyces lactis cannot grow on glucose when mitochondrial functions are blocked by various mitochondrial inhibitors, suggesting the presence of a defect in the fermentation pathway. The RAG2 gene has been cloned from a K. lactis genomic library by complementation of the rag2 mutation. The amino acid sequence of the RAG2 protein deduced from the nucleotide sequence of the cloned RAG2 gene shows homology to the sequences of known phosphoglucose isomerases (PGI and PHI). In vivo complementation of the pgi1 mutation in Saccharomyces cerevisiae by the cloned RAG2 gene, together with measurements of specific PGI activities and the detection of PGI proteins, confirm that the RAG2 gene of K. lactis codes for the phosphoglucose isomerase enzyme. Complete loss of PGI activity observed when the coding sequence of RAG2 was disrupted leads us to conclude that RAG2 is the only gene that codes for phosphoglucose isomerase in K. lactis. The RAG2 gene of K. lactis is expressed constitutively, independently of the growth substrates (glycolytic or gluconeogenic). Unlike the pgi1 mutants of S. cerevisiae, the K. lactis rag2 mutants can still grow on glucose, however they do not produce ethanol.
Similar content being viewed by others
References
Aguilera A (1986) Deletion of the phosphoglucose isomerase structural gene makes growth and sporulation glucose dependent in Saccharomyces cerevisiae. Mol Gen Genet 204:310–316
Aguilera A, Zimmermann FK (1986) Isolation and molecular analysis of the phosphoglucose isomerase structural gene of Saccharomyces cerevisiae. Mol Gen Genet 202:83–89
Ballou CE (1982) Yeast cell wall and cell surface. In: Strathern NJ, Jones EW, Broach JR (eds) The molecular biology of the yeast Saccharomyces. Metabolism and gene expression. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, New York, pp 335–360
Beggs JD (1978) Transformation of yeast by replicating hybrid plasmid. Nature 275:104–109
Bennetzen JL, Hall BD (1982) Codon selection in yeast. J Biol Chem 257:3026–3031
Bianchi MM, Falcone C, Chen XJ, Wesolowski-Louvel M, Frontali L, Fukuhara H (1987) Transformation of the yeast Kluyveromyces lactis by new vectors derived from the 1.6 μm circular plasmid pkD1. Curr Genet 12:185–192
Casadaban MJ, Martinez-Arias A, Shapira SK, Chou J (1983) β-galactosidase gene fusions for analyzing gene expression in Escherichia coli and yeast. Methods Enzymol 100:293–308
Caubet R, Guérin B, Guerin M (1988) Comparative studies on the glycolytic and hexose monophosphate pathways in Candida parapsilosis and Saccharomyces cerevisiae. Arch Microbiol 149:324–329
Chaput M, Claes V, Portelle D, Cludts I, Cravador A, Burny A, Gras H, Tartar A (1988) The neurotrophic factor neuroleukin is 90% homologous with phosphohexose isomerase. Nature 332:454–455
Chen XJ, Fukuhara H (1988) A gene fusion system using the aminoglycoside 3′-phosphotransferase gene of the kanamycin-reistance transposon Tn903: use in the yeast Kluyveromyces lactis and Saccharomyces cerevisiae. Gene 69:181–192
Ciriacy M, Breitenbach (1979) Physiological effects of seven different blocks in glycolytis in Saccharomyces cerevisiae. J Bacteriol 139:152–160
Clark-Walker GD, Linnane AW (1966) In vivo differentiation of yeast cytoplasmic and mitochondrial protein synthesis with antibiotics. Biochem Biophys Res Commun 25:8–13
De Deken RH (1966a) The Crabtree effect: a regulatory system in yeast. J Gen Microbiol 44:149–156
De Deken RH (1966b) The Crabtree effect and its relation to the peptide mutation. J Gen Microbiol 44:157–165
Eisenberg F (1978) Intermediates in the myo-inositol-1 phosphate synthase reaction. In: Wells W, Eisenberg F (eds) Cyclitols and phopshoinositides. Academic Press, New York, pp 269–275
Ferrero I, Rossi C, Landini MP, Puglisi PP (1978) Role of the mitochondrial protein synthesis in the catabolite repression of the petite negative yeast Kluyveromyces lactis. Biochem Biophys Res Commun 80:340–348
Fraenkel DG (1982) Carbohydrate metabolism. In: Strathern NJ, Jones EW, Broach JR (eds) The molecular biology of the yeast Saccharomyces. Metabolism and gene expression. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, New York, pp 1–37
Goffrini P, Algeri AA, Donnini C, Wesolowski-Louvel M, Ferrero I (1989) RAG1 and RAG2: nuclear genes involved in the dependence-independence on mitochondrial respiratory function for growth on sugars. Yeast 5:99–106
Green JBA, Wright APH, Cheung WY, Lancashire WE, Hartley BS (1988) The structure and regulation of phosphoglucose isomerase in S. cerevisiae. Mol Gen Genet 215:100–106
Hartwell LH, Culotti J, Reid B (1970) Genetic control of the cell-division cycle in yeast. I. Detection of mutants. Proc Natl Acad Sci USA 66:352–359
Henry SA (1982) Membrane lipids of yeast: biochemical and genetic studies. In: Strathern NJ, Jones EW, Broach JR (eds) The molecular biology of the yeast Saccharomyces. Metabolism and gene expression. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, New York, pp 101–158
Lamb AJ, Clark-Walker GD, Linnane AW (1968) The biogenesis of mitochondria. IV. The differentiation of mitochondrial and cytoplasmic protein synthesizing systems in vitro by antibiotics. Biochem Biophys Acta 161:415–427
Maitra PK (1971) Glucose and fructose metabolism in a phosphoglucoisomeraseless mutant of Saccharomyces cerevisiae. J Bacteriol 107:759–769
Maitra PK, Lobo Z (1971) Kinetic study of glycolytic enzyme synthesis in yeast. J Biol Chem 246:475–488
Maitra PK, Lobo Z (1977) Genetic studies with a phosphoglucose isomerase mutant of S. cerevisiae. Mol Gen Genet 156:55–60
Maniatis T, Fritsch EF, Sambrook J (1982) Molecular cloning: a laboratory manual. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, New York
Ohnishi T, Sottocasa G, Ernster L (1966) Current approaches to the mechanism of energy-coupling in the respiratory chain. Studies with yeast mitochondria. Bull Soc Chim Biol 48:1189–1203
Orr-Weaver TL, Szostak JW, Rothstein RJ (1981) Yeast transformation: a model-system for the study of recombination. Proc Natl Acad Sci USA 78:6354–6358
Sherman F, Fink GR, Hicks B (1983) Methods in Yeast Genetics. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, New York
Shuster JR, Moyer D, Irvine B (1987) Sequence of the Kluyveromyces lactis URA3 gene. Nucleic Acids Res 15:8573
Tait RC, Byron EF, Loudencia-Chingcuanco DL, Gottlieb L (1988) Plant phosphoglucose isomerase genes lack introns and are expressed in Escherichia coli. Plant Mol Biol 11:381–388
Tanguy-Rougeau C, Wesolowski-Louvel M, Fukuhara H (1988) The Kluyveromyces lactis KEX1 gene encodes a subtilisin-type serine proteinase. FEBS Lett 234:464–470
Weslowski M, Algeri A, Goffrini P, Fukuhara H (1982) Killer DNA plasmdis of the yeast Kluyveromyces lactis. I. Mutations affecting the killer phenotype. Curr Genet 5:191–197
Wéslowski-Louvel M, Goffrini P, Ferrero I (1988a) The RAG2 gene of the yeast Kluyveromyces lactis codes for a putative phosphoglucose-isomerase. Nucleic Acids Res 16:8714
Wésolowski-Louvel M, Tanguy-Rougeau C, Fukuhara H (1988b) A nuclear gene required for the expression of he linear DNA-associated killer system in the yeast Kluyveromyces lactis. Yeast 4:71–81
Yanisch-Perron C, Vieira J, Messing J (1985) Improved M13 phage cloning vectors and host strains: nucleotide sequence of the M13 mp18 and pUC19 vectors. Gene 33:103–119
Author information
Authors and Affiliations
Additional information
Communicated by C.P. Hollenberg
Rights and permissions
About this article
Cite this article
Goffrini, P., Wésolowski-Louvel, M. & Ferrero, I. A phosphoglucose isomerase gene is involved in the Rag phenotype of the yeast Kluyveromyces lactis . Molec. Gen. Genet. 228, 401–409 (1991). https://doi.org/10.1007/BF00260633
Received:
Issue Date:
DOI: https://doi.org/10.1007/BF00260633