Abstract
The goal was to elucidate the importance of the waste properties for effective exploitation in biogas production and for soil application, respectively, based on the physicochemical and microbial characterization of biowaste and the corresponding biogas residues. The following waste media were chosen: fruit waste, dairy sewage sludge, corn silage, grass silage, and grain brew to prepare three co-substrates for anaerobic bioconversion. The most satisfactory biogas yield was obtained from biowaste with the following composition: 25% fruit wastes, 25% dairy sewage sludge, 12% corn silage, and 38% grain brew. The study included functional and genetic diversity assessment through the characterization of the catabolic potential and structure of the microbial communities inhabiting the examined organic wastes and their relative biogas residues. The metabolome was based on the use of a Biolog® plate. The elucidation of the metagenome employed the genetic structure of prokaryotes and involved denaturing gradient gel electrophoresis and next-generation sequencing analyses. The usefulness of metagenomics was emphasized by ecotoxicological evaluation of biowaste and in determining the accurate start-up community composition for biogas production, highlighting the pivotal role of anammox and hydrolytic bacteria as marker groups. The high importance of the great diversity of fungi was also revealed based on a functional approach.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The constantly increasing amount of organic wastes produced worldwide is a growing concern (Oleszek et al. 2016; Oszust et al. 2017a). On the other hand, the interest in biowaste has risen recently; it is regarded as a source of renewable energy (Bruni et al. 2010; Lim and Wang 2013; Wartell et al. 2012; Oszust et al. 2017b). In general, large amounts of different organic wastes originating from the production of food and agro-industrial products pose a significant risk to the environmental pollution; however, it also represents a great opportunity to combine waste treatment and energy production (Esposito et al. 2012).
To date, there has been significant research demonstrating the benefits of co-digestion of different substrates, as described by, for example, Rajagopal et al. (2013) or Westerholm et al. (2012). The conceptual conditions necessary for effective biogas production are also related to certain subsequent microbial changes that occur during biowaste decomposition. The initial diversity of microbial communities in co-substrate biowastes and their chemical properties play a very important role in biogas yield enhancement. These factors may be regarded as a prospective useful tool for the determination and subsequent manipulation of appropriate species in anaerobic digesters in order to produce a more stable and satisfactory process.
The biogas residue is regarded as a nutrient-rich end product of anaerobic digestion (Coban et al. 2016). More specifically, the utilization of the biogas residue as exogenous organic matter (EOMs) in field applications is considered to act as a soil conditioner and to enhance crop yield. Therefore, it represents a strategy that may counteract the depletion of organic matter in soils. The application of exogenous organic matter to agricultural land to enrich the soil requires careful monitoring as well as enriched soil (Oszust et al. 2015).
The microbial state of the residue before the application may therefore lead to possible perturbations in soil microbial quality after the application thereof in the soil. Consequently, the evaluation of the structure, function, and activity of the microbial community, either as a whole or in terms of specific phylogenetic or functional groups, presents the optimal method to ascertain the most probable events occurring in soil (Arthurson 2009; Curieses et al. 2016).
Community genomic and bolomic analyses used to characterize biowaste co-substrates for biogas production and its residues are adequate approaches to gain further insight into microbial properties. Schlüter et al. (2008) affirm that the metagenome sequence data from the microbial community provides a sound on the basis of a rational approach to improving the biotechnological process. In essence, environmental metabolomics is the application of metabolomics to the characterization of the interactions of living microorganisms with their environment. The evaluation of the functionality of the co-substrate microbiome may be regarded as the first tool required to identify the phenotype of environmental samples. The assessment of the community genomics and bolomics in biowaste or residues may serve as a sustainable, collateral (to physicochemical) strategy for better characterization of feedstock co-substrates (Bundy et al. 2009).
In this paper, it was proposed to gain further insight into the substrate characteristics of substrates and their combinations, especially in order to devise co-substrate mixtures that will be effective in biogas production. The other objective of this work was to elucidate the land utilization of biogas residues, including the ecotoxicological evaluation of the waste. The advantage of this work is that our research links the standard physicochemical properties with the use of genomic and metabolomic techniques. This allowed for investigations of the composition and functionality of microbial communities directly in the environment of the proposed materials and provided an explanation for the possible relationship with physicochemical conditions.
Materials and methods
Biowaste and biogas residues
The following organic components: dairy sewage sludge, fruit waste, corn silage, grass silage, and grain brew, were used in various proportions to prepare three biowaste mixtures:
-
BW1: 25% fruit wastes + 25% dairy sewage sludge + 12% corn silage + 38% grain brew;
-
BW2: 30% fruit wastes + 35% dairy sewage sludge + 8% corn silage + 27% grain brew;
-
BW3: 25% fruit wastes + 25% dairy sewage sludge + 15% grass silage + 35% grain brew.
The three proposed biowaste mixtures (BW1–BW3) were subjected to the batch anaerobic digestion process, from which biogas residues (BR1–BR3) corresponding to BW1–BW3 were obtained. The anaerobic digestion was conducted under mesophilic conditions at 37 °C in glass chambers with a working volume of 0.5 dm3, with stirring. The operating anaerobic digestion temperature was maintained using a thermostat connected to the fermenter water jacket. The biomass amount introduced into the reactor was properly calculated in order to achieve similar load values of dry organic matter for each of the experimental variants; this amounted to 4.95 g of dry matter for each dm3 in the chamber. The hydration value and organic matter content in each type of biowaste were similar: 11–16% and 88–92%, respectively. The pH of the biowaste was adjusted to 7.0 using Na2CO3 prior to anaerobic digestion. Anaerobic granular sludge taken from an agricultural biogas plant was used as inoculum (20% of the fermenter volume) for anaerobic digestion after concentration by sedimentation.
Biowaste and biogas residues and the particular components of biowaste (dairy sewage sludge, fruit waste, corn silage, grass silage, and grain brew) were subjected to the quantification of total solids (TS), volatile solids (VS), ash content, chemical oxygen demand (COD), total Kjeldahl nitrogen (TKN), and pH using the following standard methods: PN-C-04616-01:1975 (TS, VS, ash), PN-74/C04578.03 (COD), PN-EN 13342:2002 (TKN) and CPI-505 pH meter. Phosphorus and heavy metal contents were evaluated by the inductively coupled plasma with mass spectrometry (ICP-MS) method after microwave digestion (Gałązka and Gembal 2015).
Community-level physiological profiling (CLPP) analysis
Interpretation of the CLPPs was based on metabolic profiles obtained using Biolog® ECO plates designed by Insam (1997) for the bacterial community, Biolog® FF plates for aerobic fungal community, and Biolog® AN plates for the anaerobic microbial community. The recent study of Borowik et al. (2017) or Wolińska et al. (2017) shows that the Biolog® system is an useful tool to explore the shifts in microbial community functional diversity of different environments. One gram portions of biowaste/biogas residues were shaken in 99 ml of sterile peptone water for 20 min at 20 °C and followed by incubation at 4 °C for 30 min. Next, 120 µl of each sample was inoculated into each well of the ECO plates, whereas 100 µl of the suspension was inoculated into the FF and AN plates; this was followed by incubation at 27 °C. The AN plates were incubated in anaerobic jars with a gas mixture (H2: 5%, CO2: 10%, N2: 85%) injected using an Anoxomat® instrument (MART Microbiology B.V., USA). Data were recorded with a plate reader at 590 nm every 24 h until readings that exhibited approximately the same average well colour development (AWCD). The microbial response in each microplate was expressed by the Shannon index (H) and was assessed as described by Oszust et al. (2014).
Genomic DNA extraction
Genomic DNA was extracted from 0.5 g of soil from every sample, using a FastDNA® SPIN Kit for Faeces (MP Biomedicals, Solon, OH, USA), following the protocol of the manufacturer.
Denaturing gradient gel electrophoresis (DGGE) analysis
The PCR was performed in a total volume of 30 μl containing approximately 20 ng of DNA template, 15 μl RedTaq® ReadyMix™ PCR Reaction Mix (Sigma-Aldrich, St. Louis, MO, USA). The volumes of the primers used for PCR amplification were 0.2 μl (10 μM). The primers were as follows: primer R: UNI1401r 5′GCGTGTGTACAAGACCC3′ and primer F: GC-968f5′CGCCCGGGGCGCGCCCCGGGCGGGGCGGG GGCACGGGGGGAACGCGAAGAA3′ (Nübel et al. 1996). Thermal cycling was carried out by an initial denaturation step at 94 °C for 1.5 min, followed by 56 °C for 0.5 min and 72 °C, 45 s. The major cycling program for each primer set was optimized and included (95 °C, 20 s; 56 °C, 30 s; 72 °C, 45 s) × 33; 72 °C, 7 min for biowaste probes and (95 °C, 20 s; 56 °C, 30 s; 72 °C, 45 s) × 27; 72 °C, 7 min for biogas residues..
The 16S rDNA-DGGE was performed using the Dcode System (Universal Mutation Detection System, Bio-Rad). 80 ng of amplicons were loaded in triplicate (top filling method) on 6% polyacrylamide gel containing a denaturant gradient of 46–56% parallel to the electrophoresis direction made of urea and formamide (100% denaturant contains 7 M urea and 40% formamide). The gels were electrophoresed at a constant temperature (60 °C) and electric voltage (70 V) for 16 h, followed by 1-h colouration using SybrGreen I (Sigma-Aldrich, St. Louis, MO, USA), scanned, and analysed with Quantity One software (Bio-Rad, Richmond, California).
Next-generation sequencing (NGS)
The PCR was performed using primers specific for the sequence of the V4 region of the 16S rRNA: 515F and 806R, NEBNext® High-Fidelity PCR Master Mix 2X reaction conditions as recommended by the manufacturer. Libraries were indexed in TruSeq technology, Illumina. The libraries were prepared in an analogous way to the attached Illumina protocol. Sequencing of PE 2 × 250 bp was performed using a v2 Illumina kit. All of the bioinformatic analyses were based on a database of reference sequences Greengenes. The analysis was performed on the available camera MiSeq and QIIME softwares.
Sanitary analysis of the co-substrates and mixtures of organic waste
Sanitary analysis consisted of the detection of the presence of Salmonella spp. with the conventional culture method and the presence of microorganisms belonging to the Enterobacteriaceae family and parasitological analysis. The isolation of Salmonella spp. involved nonselective pre-enrichment of a defined weight of the sample, followed by a selective enrichment step, plating onto selective agars, and biochemical and serological confirmation of suspect colonies. The presence of Enterobacteriaceae was evaluated by the next-generation sequencing analyses. The prevalence analyses of Ascaris sp., Trichuris sp., and Toxocara sp. ova were carried out according to the Wasilkowa method described by Olańczuk-Neyman et al. (2003). A viability assay was performed by incubating the eggs in Petri dishes at 28 °C for 10–14 days.
Statistical analyses
A cluster analysis, including the grouping of treatments and features and H-index calculation, based on Biolog® ECO and FF plates, was performed on the standardized data of the absorbance average values for readings at 144 h and 72 h for AN plates. A dendrogram representing the similarities of the carbon utilization patterns from substrates located on the Biolog ECO, FF, AN plates® between the biowaste samples was set on scaled axis bond distances (Ward’s method, within Euclidean distance), with marked boundaries of Sneath’s criteria (restrictive and less restrictive, 33% and 66%, respectively). The cluster analyses that followed the evaluation of the Biolog plates were performed with Statistica 10.0 software (StatSoft Inc., Tulsa, OK, USA, 2011). In the same mode, a cluster analysis of the DGGE results was performed on Quantity One software (Bio-Rad Laboratories Inc., CA, USA, 2003).
Results and discussion
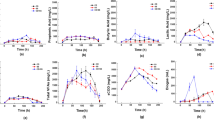
The total solids (TS) amount was 11–16% in the biowaste and approx. 1% in the biogas residues. 88–92.7% volatile solids were detected in the biowaste, while in the residues they reached 55%. The C/N ratio was 27.1, 26.36, and 28.17 in BW1, BW2, and BW3, respectively, and 7.94, 5.65, and 8.96, respectively, in BR1, BR2, and BR3. The C/N ratio varying from about 15 to 30 indicates an optimal carbon and nitrogen content in the tested biowaste for optimal methane fermentation (Wang et al. 2014). The pH value in BW 1–3 was approx. 4.36, 4.52, and 4.69, respectively. In the corresponding biogas residues, it was 7.98, 8.00, and 7.95. The phosphorus content was the highest in BW2 and reached 9.15 g kg−1 TS, whereas 6.87 and 5.77 g kg−1 TS were noted in BW1 and BW3, respectively. The contents of heavy metals such as Cr, Ni, Cu, Cd, Pb, and Hg in the biowaste, biogas residues, and in the particular components of biowaste were relatively low. However, the Zn content was 78.5 ppm, especially in BR1. The growth of methanogens that play a crucial role is dependent on a few cations (Zn, Na, Ni, Co, Fe, Mg, Ca, and K) and molybdate or tungstate and phosphate anions. Among others, they are required for methanogenesis, for the synthesis of enzymes, prosthetic groups, and coenzymes. The work of Zhang et al. (2008) established the restrictive Zn amount at 1 mg l−1. The relatively high concentrations of Zn and other cations obtained in our work might have had a limiting effect on the functionality of the methanogens. Significant differences in volatile solids, nitrogen, and pH in the particular biowaste components were observed. The difference in COD (1.2 and 2.03 g O2 kg−1 TS in corn silage and dairy sewage sludge, respectively) was evidenced, and they might supervene the influence on biogas yield odds (Table 3, Fig. 7). The tested biowaste differed significantly in nitrogen content. The highest values were determined in dairy sewage sludge (7.9% TS) and in grain brew (7.65% TS) (Table 1).
There were differences between the metabolic profiles (CLPPs) of the analysed materials in aerobic fungal (FF), bacterial (ECO) and anaerobic (AN) communities demonstrated by Biolog Plates® as well as the metagenomic approach based on 16S rRNA by next-generation sequencing (NGS) and denaturing gel gradient electrophoresis (DGGE). The results of the Shannon index (H) presented in Table 1 showed the lowest diversity in the bacterial community structure based on DGGE. The functional biodiversity calculated on the FF and AN plates was both relatively high and at the same level (> 4), whereas the ECO plates revealed lower biodiversity (< 3.21) than that shown by the other metabolic approaches. The H-index value based on the AN plates approach revealed a greater degree of diversity for biogas residues compared to that of the biowaste. These results were consistent with the findings of the H-index calculated on the basis of methanogen OTUs (operational taxonomic units) after filtering the NGS data results, which also suggests no considerable presence of methanogens in the biowaste and their differentiated composition in biogas residues. The data of the diversity of methanogens are presented in Table 2. The following genera were revealed in the analysed probes: Methanobacterium, Mathanobrevibacter, Methanocorpusculum, Methanoculleus, Methanoseaeta, Methanosarcina, Methanospharea, Methanospiryllum, Syntrophobacter, and Syntrophomonas. Only Methanospiryllum and Syntrophomonas were found in BW1 and Methanoseaeta in BW2. In BW3, no methanogens were identified. In contrast to the biowaste, biogas residues contained a greater diversity of methanogens. Among all the genera revealed, 34 readings of methanogens, without the presence of Mathanobrevibacter and Syntrophomonas, were noted in BR1. None were detected in BR2, whereas BR3 was characterized by the presence of Mathanobrevibacter and Methanospharea, with 34 and 33 readings, respectively. The differences found in methanogens biodiversity between biowaste and biogas residues result from the stage of anaerobic digestion. The composition of methanogens is in consistent with the results of Gryta et al. (2017). At the beginning of the process, methanogens inoculum is added to the substrates in order to initialize methanogenesis and thus biogas production. Depending on the physicochemical properties of biowaste development, the various groups were quite distinct. Methanosaetaceae were the most abundant organisms in all of the bioresidues tested, which means that primarily acetoclastic conversion to methane was noted (Karakashev et al. 2006). Also, Methanobacteriales, which are hydrogenotrophic methanogens, have been observed. This indicates the occurrence of a second mechanism of methanogenesis. The established hydrogenotrophic methanogenesis encompasses a two-step process. Acetate is first oxidized to H2 and CO2 and subsequently converted to methane. The occurrence of Methanobrevibacter genus members is typical for fruit waste anaerobic digestion (Gryta et al. 2017), as found in the biowaste examined.
Furthermore, the allocation of the assembled contig sequences to Prokaryote genome sequences is presented in Fig. 1. Six phyla among the Bacteria (Proteobacteria, Firmicutes, Planctomycetes, Actinobacteria, Bacteroidetes, and an unidentified one denoted as “x1”) and 1 Archaea (Euryarchaeota) were identified. Among the Proteobacteria, the families Xanthomonadaceae, Rhodobacteraceae, and Rhizobiaceae were identified. Among the Firmicutes, we identified Enterococcaceae, Streptococcaceae, Lactobacillaceae, Peptostreptococcaceae, and Clostridiaceae; Pirellulaceae families represented Planctomycetes. There were also Propionibacteriaceae, Nocardiaceae, Corynebacteriaceae, and Intrasporangiaceae within Actinobacteria. Bacteroidetes were varied and were represented by the families of Marinilabiaceae and Porphyromonadaceae. There were no Archaea revealed in biowaste samples. However, Archaea constituted more than 20% of the biogas residues. An increasing proportion of Firmicutes and Bacteroidetes were noted in biogas residues, compared to the biowaste. In biogas residues, an over 40% decrease was noted in the share of Proteobacteria. Consistently, we noted an increase in Actinobacteria, Proteobacteria, and Planctomycetes. The share of the particular groups differed depending on the residue type.
Fungal functional diversity following Biolog FF plates®a Dendrogram of carbon utilization patterns, b percentage of total carbon source utilization fungal response. Explanations: biowaste; BW1: 25% fruit waste + 25% dairy sewage sludge + 12% corn silage + 38% grain brew, BW2: 30% fruit waste + 35% dairy sewage sludge + 8% corn silage + 27% grain brew; BW3: 25% fruit waste + 25% dairy sewage sludge + 15% grass silage + 35% grain brew; and corresponding biogas residues, after their anaerobic processing. Grouping according to the stringent Sneath’s criterion (33%) and less restrictive criterion (66%), n = 3 (A–C)
To gain more insight into the similarities or differences between the microbial communities inhabiting the tested biowaste and residues, clustering analyses were performed. When the anaerobic fungal community was taken into consideration (Fig. 2a), the bond distance dendrogram revealed that the metabolome of biowaste and biogas residue clustered separately, even based on the strict Sneath’s criterion (33%). This means that the fungal communities showed different modes of the C-substrate utilization. Taking into account the less restrictive Sneath’s criterion (66%), we noted that BR1 and BR2 grouped together and differed from BR3. Figure 2b presents shifts in the utilization pattern of the C-substrate groups located on the FF plate. We showed that the fungal community in the biowaste more readily utilized carbohydrates than in the residues. It may be noted that the community of BR2 more readily utilized amino acids than of BW2. The opposite results in the level of amino acids utilization were met in BR3 and BW3, and simultaneously C-substrates belonging to other groups in BR3 were more preferably metabolized than in BW3. The bond distance in the utilization of C-substrates located on the ECO plate (Fig. 3a) showed clustering results corresponding to the FF plate, i.e. a group composed of BW1, BW2, and BR1 and a separate group comprising only BR2. However, the results shown in Fig. 3b revealed that bacterial communities from the corresponding materials before and after anaerobic digestion utilized carbohydrates in the same manner. The most intense shifts were found in the increasing level of utilization of amines and amides in BR1 compared to BW1 (from 0.2 to 3.3%) and reduction in utilization of polymers from 24% in BW3 to 19% in the corresponding residue (BR3).
Bacterial functional diversity following Biolog ECO plates®a Dendrogram of carbon utilization patterns, b percentage of total carbon source utilization bacterial response. For explanations, please see Fig. 1
Anaerobes functional diversity following Biolog AN plates®a Dendrogram of carbon utilization patterns, b percentage of total carbon source utilization anaerobic community response. For explanations, please see Fig. 1
The bond distance tree of the anaerobic communities (AN plates) (Fig. 4a) revealed clustering into just two groups (regardless of Sneath’s criteria). All residue metabolic profiles grouped together, whereas BW2 and BW3 formed a separate cluster. Surprisingly, BW1 was closer to the residue groups. We noted an increase in the percentage of amino acid utilization in residues BR2 and BR3 (20% and 13%, respectively). Figure 4b presents shifts in the pattern of C-substrate group utilization located on the AN plate. We revealed a reduced utilization level of carbohydrates in biogas residues, compared to biowaste in favour of the increased share of amino acid utilization.
Clustering groups of 16S rDNA based on denaturing gradient gel electrophoresis. For explanations, please see Fig. 1
In the metagenomic approach, the cluster analysis performed on 16S rDNA-DGGE demonstrated two or three clustering groups depending on Sneath’s criteria. Based on the less restrictive criterion, we revealed that BW1 and BW2 clustered together and differed from BW3 (Fig. 5). The NGS neighbour-joining tree showed similarities to the results of the DGGE cluster analysis in revealing a dichotomy in the clustering groups: one group comprising all tested biowaste and the other tested biogas residues (Fig. 6).
16S next-generation sequencing neighbour-joining tree. For explanations, please see Fig. 1
Allocation of assembled contig sequences to microbial genome sequences based on next-generation sequencing. For explanations, please see Fig. 1
The results obtained from the co-digestion analyses of biowaste are presented in Table 3 and Fig. 7. The highest biogas efficiency reached 629.4 dm3 kg−1 TS for BW1, which meets the optimum efficacy of co-digestion, being comparable to the recent findings of other authors (Neshat et al. 2017; Pavi et al. 2017; Rahman et al. 2017). For this substrate, the highest methane yields were obtained as well (402.81 dm3 kg−1 TS, 64%). In contrast, lower biogas and methane yields were obtained from BW3 and BW2 (545.2 and 316.21 dm3 kg−1 TS, respectively). The daily changes in the biogas yield were the highest on the third day of the digestion process. Afterwards, the biogas productivity of all biowaste decreased steadily until the end of the process. Biogas yield effectiveness of bioresidue mixtures depended on their composition and reached relatively high levels compared to substrates analysed separately. For example, the yield of biogas and methane from fruit waste was reported to reach 480 dm3 kg−1 TS and 250 dm3 kg−1 TS, respectively (Zieliński et al. 2016). To the best of our knowledge, there are no reports on the biogas yield from dairy sewage sludge by itself, separate from other waste; however, our unpublished data indicate that this level does not exceed 400 dm3 kg−1 TS, which is not sufficient for our requirements. Among the various types of feedstock substrates available for the production of biogas, corn silage is currently the most advantageous, with biogas production of up to 700 dm3 kg−1 TS, (Oleszek et al. 2016). This is due to the fact that corn silage is a source of proteins, fats, starch, and cellulose, which makes it the preferred starting material for biodegradation in the anaerobic digestion process. Thus, adding some energy-rich substrates, such as corn silage to the feedstock substrate mixture and regulating its composition, should be one of the best options available to resolve adequate biogas effectiveness limitations.
Biogas yield in anaerobic digestion a BW1; b BW2; c BW3. For explanations, please see Fig. 1
Many studies have revealed that anaerobic digestion may be a valuable tool for the management of different wastes. However, it is more probable that higher biogas production will be obtained if combined (co-digestion) (Kamali et al. 2016; Zhang et al. 2014). This is in agreement with our study. We propose waste materials (dairy sewage sludge, fruit waste, corn silage, grass silage, and grain brew) requiring reasonable development, since most of them are produced recently on a large scale (Sawatdeenarunat et al. 2015). However, the data of the physicochemical properties of waste (Table 1) indicate that most of them are not proper for anaerobic bioconversion separately. This particularly applies to dairy sewage sludge which revealed a very low ratio of C/N, amounting only to 3.84. If the ratio of C/N is too low as in dairy sewage sludge, nitrogen is released in the form of ammonia, up to a concentration that is toxic to the methanogenic Archaea (De Vrieze et al. 2015). Consequently, there was a need to combine dairy sewage sludge with other materials, e.g. corn silage or fruit waste (which demonstrated more preferable properties); this solution was proposed for the multicomponent biowaste arrangement. The results demonstrate that a significant improvement was achieved in C/N ratio in the tested combined biowaste, which was relatively high: 26 in BW1. Nonetheless, as suggested by Oleszek et al. (2016) the optimum C/N is 20–30. In BW3, the ratio of C/N was too high (> 38). It is probable that nitrogen was rapidly consumed by methanogens to meet their protein needs and was not available for the actual methane fermentation process. Therefore, we noted the lowest biogas yield from BW3. Surprisingly, the substitution of one ingredient, i.e. barley 12–15% (grass silage in BW3 to corn silage in BW1), led to a better (12%) biogas yield (Table 3). The increased amount of biodegradable organic substances in BW1 introduced with corn silage caused an increase in the amount of biogas produced in comparison with BW3. Modification of the composition of the substrate, compared to a single waste, improved hydration (Table 1) by weight and the balance of the fermentation of biogenic elements. This result is also consistent with the separate cluster of BR3 based on community metabolomics (Figs. 2a, 3a). This shows the differences in the bioconversion efficiency but also the consistency in the characteristics of the physicochemical and microbial properties. Overall, the impact of the relationship between these properties of the process that takes place in a biogas reactor plays a pivotal role in the biogas yield. This is in agreement with the finding reported by Oleszek et al. (2014). What is more, attention should be paid to the composition of BW1 and BW2. Despite the greater differences in the proportions of ingredients between BW1 and BW2 than between BW1 and BW3, the difference in biogas yields was significant and lower biogas yields were noted in BW3. This was evidently caused by the use of grass silage in BW3 and corn silage in BW1 and BW2, since corn silage is one of the most energy-rich substrates for anaerobic digestion (Jankowski et al. 2016).
Since biowaste processing is controlled by microorganisms, mixed wastes ensure better environmental start-up conditions before anaerobic digestion and probably provide a unique microbial community that together cause differences in the biogas yield. The findings described by Satpathy et al. (2016) also indicate that the initial diversity of biomass co-substrates plays a very important role in the methanogenesis process and eventually the biogas yield. The inherent ecological factors, which were found to trigger a prompt start-up for biogas production, focus on the initial relative abundance and concentration of anaerobic ammonium-oxidizing (anammox) bacteria that are capable of ammonium oxidization (De Vrieze et al. 2015; Tao et al. 2013). A considerable share of anammox bacteria belonging to Planctomycetes was identified in biowaste (Fig. 6). The process of assembling of co-substrates might first of all influence the hydrolysis step, since it was identified as the rate-limiting step of anaerobic digestion (Park et al. 2005). The allocation of the assembled contig sequences to the microbial genome sequences in our study produced molecular evidence for the genus-level diversity of Bacteria and Archaea. Bacteria belonging to Firmicutes, Bacteroides, and Spirochaetes seem to be involved in the hydrolysis of cellulose and proteins, especially in BW1 and BW2 (Fig. 6). These groups may be regarded in the genomics approach as a prospective useful tool for the determination and subsequent manipulation of appropriate species and an additional marker in models for the prediction of potential biogas yields. Achinas and Euverink (2016) or Demirel and Scherer (2008) also suggest that these microorganisms are important marker groups in methanogenesis.
On the other hand, the bolomic study based on the Biolog® plates (Table 1) demonstrated higher biodiversity indices of aerobic Fungi and anaerobic microbial communities than those of aerobic Bacteria and Archaea. These findings may reflect the fact that many different types of aerobic Fungi may be engaged equally in organic matter decomposition, followed by anaerobic transformations in the bioreactor. These are probably hydrolytic aerobic Fungi, which are useful in biological pre-treatment for the next step, i.e. anaerobic digestion, as indicated by Vasco-Correa et al. (2016). However, the relatively high-diversity index of the total anaerobic community may also suggest that anaerobic Fungi inhabiting biowaste are involved in biomass decomposition. The presence and role of anaerobic hydrolytic microbes in the conversion of lignocellulosic biomass for biogas production were noted by Azman et al. (2015). The results of our investigations indicating a separate cluster of anaerobic microbial communities in BW1 (Fig. 3a) may impose the significant role of fungi in the effective biodegradation of wastes, as was previously stated by Oszust et al. (2017a). A characteristic trait of the BW anaerobic communities is the high level of carbohydrate utilization, which was not observed in biogas residues (an increased share of microorganisms utilizing amino acids and carboxylic acids was observed instead) (Fig. 3b). The role of anaerobic Fungi in the first step of the hydrolytic degradation of substrates with higher biogas efficiency than anaerobic Bacteria was highlighted by Procházka et al. (2012).
On the other hand, these properties play a crucial role when introducing biogas residues or other exogenous organic matter into the soil as organic fertilizers, because they may change soil properties in a positive or negative manner (Frąc et al. 2012; Malý 2015). Residues incorporated into the soil may impose an additional load of microorganisms and affect the microbial community of soil (Hupfauf et al. 2016). The microbial genetic composition and metabolic capabilities may affect the course of the microbial process and the efficiency of organic material turnover in soil (Oszust et al. 2015). The role of the great diversity of Fungi should also be taken into consideration (Table 1).
Residue characterization is important to provide ecotoxicological and sanitary protection against the introduction of undesirable microbial factors into the soil that could pose a threat, as specified by Regulation (EC) No. 1069/2009 of The European Parliament (Mitter et al. 2016). The presence of Salmonella and Enterobacteriaceae is regarded to be an adverse agent. Besides, the presence of gastrointestinal parasite eggs: Ascaris spp., Trichuris spp., and Toxocara spp., is similarly disadvantageous in residues designed for field application as exogenous organic matter. In our study, no such factors were revealed (Table 1), which indicate that the proposed residues may be allowed for field application as organic fertilizers. The heavy metals content in biowaste and postfermentation biofertilizers was low. The Zn content was relatively high (78.5 mg kg−1 TS), which could have a positive aspect, taking into consideration the role of micronutrients in plant production. Therefore, the potential of biogas residue as a crop fertilizer and soil conditioner appears likely and promising. However, the application of the residue to cropland requires rigorous monitoring to detect early perturbations in soil quality, which may result in reduced crop yield (Abubaker et al. 2012). The plant holobiont comprises the plant and its associated microbiota, which interact with each other and determine holobiont functioning and plant performance.
The investigations demonstrate that biogas residues contain numerous typical representatives of hydrogenotrophic or acetotrophic methanogens (Table 2), whereas no methanogenic microorganisms are present in biowaste. Therefore, biogas residues may successfully be used as seeding sludge (inoculum) for further effective biogas production and may serve as exogenous organic matter.
Conclusion
The main research objective presented in this work involved the microbiological and physicochemical characterization of biowaste and bioresidues to elucidate the significance of their properties for the beneficial exploitation thereof in biogas production or soil application, respectively. Among the waste-based substrates, fruit waste, dairy sewage sludge, corn silage, grass silage, and grain brew were proposed as combined co-substrate loads for anaerobic bioconversion that result in simultaneously relatively high biogas yields; BW1 composed of 25% fruit wastes, 25% dairy sewage sludge, 12% corn silage, and 38% grain brew was the most satisfactory mixture.
The community genomic and bolomic approaches were selected as tools facilitating the characterization of microbial communities in biogas residues, which constitutes a rapid and reliable source of information on the ecotoxicity and quality of wastes in the context of the agricultural application of biogas residues. Most particularly, the usefulness of metagenomics was emphasized in determining the accurate start-up community composition for biogas production, highlighting the pivotal role of anammox and hydrolytic Bacteria as marker groups. The importance of the great diversity of Fungi was also revealed based on the bolomic approach.
References
Abubaker J, Risberg K, Pell M (2012) Biogas residues as fertilisers—effects on wheat growth and soil microbial activities. Appl Energy 99:126–134. https://doi.org/10.1016/j.apenergy.2012.04.050
Achinas S, Euverink GJW (2016) Theoretical analysis of biogas potential prediction from agricultural waste. Resour Eff Technol 2:143–147. https://doi.org/10.1016/j.reffit.2016.08.001
Arthurson V (2009) Closing the global energy and nutrient cycles through application of biogas residue to agricultural land—potential benefits and drawback. Energies 2:226–242. https://doi.org/10.3390/en20200226
Azman S, Khadem AF, Van Lier JB, Zeeman G, Plugge CM (2015) Presence and role of anaerobic hydrolytic microbes in conversion of lignocellulosic biomass for biogas production. Crit Rev Environ Sci 45:2523–2564. https://doi.org/10.1080/10643389.2015.1053727
Borowik A, Wyszkowska J, Oszust K (2017) Functional diversity of fungal communities in soil contaminated with diesel oil. Front Microbiol 8:1862. https://doi.org/10.3389/fmicb.2017.01862
Bruni E, Jensen AP, Angelidaki I (2010) Comparative study of mechanical, hydrothermal, chemical and enzymatic treatments of digested biofibers to improve biogas production. Bioresour Technol 101:8713–8717. https://doi.org/10.1016/j.biortech.2010.06.108
Bundy JG, Davey MP, Viant MR (2009) Environmental metabolomics: a critical review and future perspectives. Metabolomics 5:3. https://doi.org/10.1007/s11306-008-0152-0
Coban H, Miltner A, Centler F, Kästner M (2016) Effects of compost, biochar and manure on carbon mineralization of biogas residues applied to soil. Eur J Soil Sci 67:217–225. https://doi.org/10.1111/ejss.12326
Curieses SP, Sáenz ME, Larramendy M, Di Marzio W (2016) Ecotoxicological evaluation of foundry sands and cosmetic sludges using new earthworm biomarkers. Ecotoxicology 25:914–923. https://doi.org/10.1007/s10646-016-1649-3
De Vrieze J, Saunders AM, He Y, Fang J, Nielsen PH, Verstraete W, Boon N (2015) Ammonia and temperature determine potential clustering in the anaerobic digestion microbiome. Water Res 75:312–323. https://doi.org/10.1016/j.watres.2015.02.025
Demirel B, Scherer P (2008) The roles of acetotrophic and hydrogenotrophic methanogens during anaerobic conversion of biomass to methane: a review. Rev Environ Sci Biotechnol 7:173–190. https://doi.org/10.1007/s11157-008-9131-1
Esposito G, Frunzo L, Giordano A, Liotta F, Panico A, Pirozzi F (2012) Anaerobic co-digestion of organic wastes. Rev Environ Sci Biotechnol 11:325–341. https://doi.org/10.1007/s11157-012-9277-8
Frąc M, Oszust K, Lipiec J (2012) Community level physiological profiles (CLPP), characterization and microbial activity of soil amended with dairy sewage sludge. Sensors 12:3253–3268. https://doi.org/10.1007/s13762-015-0807-0
Gałązka R, Gembal A (2015) Determination of different forms of trace elements and macro elements. In: Bieganowski A, Malý S, Frąc M, Tuf IH, Váňa M, Brzezińska M, Siebielec G, Lipiec J, Šarapatka B (eds) Laboratory manual. Central Institute for Supervising and Testing in Agriculture, Brno, pp 91–96
Gryta A, Oszust K, Brzezińska M, Ziemiński K, Bilińska-Wielgus N, Frąc M (2017) Methanogenic community composition in an organic waste mixture in an anaerobic bioreactor. Int Agrophys 31:327–338
Hupfauf S, Bachmann S, Juárez MF-D, Insam H, Eichler-Löbermann B (2016) Biogas digestates affect crop P uptake and soil microbial community composition. Sci Total Environ 542:1144–1154. https://doi.org/10.1016/j.scitotenv.2015.09.025
Insam H (1997) A new set of substrates proposed for community characterization in environmental samples. Springer, Berlin
Jankowski KJ, Dubis B, Budzyński WS, Bórawski P, Bułkowska K (2016) Energy efficiency of crops grown for biogas production in a large-scale farm in Poland. Energy 109:277–286
Kamali M, Gameiro T, Costa MEV, Capela I (2016) Anaerobic digestion of pulp and paper mill wastes—an overview of the developments and improvement opportunities. Chem Eng J 298:162–182. https://doi.org/10.1111/ejss.12326
Karakashev D, Batstone DJ, Trably E, Angelidaki I (2006) Acetate oxidation is the dominant methanogenic pathway from acetate in the absence of methanosaetaceae. Appl Environ Microb 72:5138–5141. https://doi.org/10.1128/AEM.00489-06
Lim JW, Wang J-Y (2013) Enhanced hydrolysis and methane yield by applying microaeration pretreatment to the anaerobic co-digestion of brown water and food waste. Waste Manag 33:813–819. https://doi.org/10.1016/j.wasman.2012.11.013
Malý S (2015) Egzogenna materia organiczna w relacji do glebowej materii organicznej i funkcji ekosystemu. In: Malý S, Siebielec G (eds) Badania egzogennej materii organicznej w celu bezpiecznego stosowania do gleby. Central Institute for Supervising and Testing in Agriculture, Brno, pp 5–13
Mitter B, Pfaffenbichler N, Sessitsch A (2016) Plant–microbe partnerships in 2020. Microb Biotechnol 9:635–640. https://doi.org/10.1111/1751-7915.12382
Neshat SA, Mohammadi M, Najafpour GD, Lahijani P (2017) Anaerobic co-digestion of animal manures and lignocellulosic residues as a potent approach for sustainable biogas production. Renew Sustain Energy Rev 79:308–322. https://doi.org/10.1016/j.rser.2017.05.137
Nübel U, Engelen B, Felske A, Snaidr J, Wieshuber A, Amann RI et al (1996) Sequence heterogeneities of genes encoding 16S rRNAs in Paenibacillus polymyxa detected by temperature gradient gel electrophoresis. J Bacteriol 178:5636–5643. https://doi.org/10.1128/jb.178.19.5636-5643.1996
Olańczuk-Neyman K, Geneja M, Quant B, Dembińska M, Kruczalak K, Kulbat E et al (2003) Microbiological and biological aspects of the wastewater treatment plant “Wschód” in Gdańsk. Pol Environ Stud J 12:747–757
Oleszek M, Król A, Tys J, Matyka M, Kulik M (2014) Comparison of biogas production from wild and cultivated varieties of reed canary grass. Bioresour Technol 156:303–330. https://doi.org/10.1016/j.biortech.2014.01.055
Oleszek M, Tys J, Wiącek D, Król A, Kuna J (2016) The possibility of meeting greenhouse energy and CO2. BioEnergy Res 9:624–632. https://doi.org/10.1007/s12155-015-9705-z
Oszust K, Frąc M, Gryta A, Bilińska N (2014) The influence of ecological and conventional plant production systems on soil microbial quality under hops (Humulus lupulus). Int J Mol Sci 15:9907–9923. https://doi.org/10.3390/ijms15069907
Oszust K, Frąc M, Lipiec J (2015) Soil microbial functionality in response to dairy sewage sludge and mineral fertilisers application under winter rape. Int J Environ Sci Technol 12:3675–3684. https://doi.org/10.1007/s13762-015-0807-0
Oszust K, Pawlik A, Siczek A, Janusz G, Gryta A, Bilińska-Wielgus N, Frąc M (2017a) Efficient cellulases production by Trichoderma atroviride G79/11 in submerged culture based on soy flour-cellulose-lactose. BioResources 12:8468–8489
Oszust K, Pawlik A, Janusz G, Ziemiński K, Cyran M, Siczek A, Gryta A, Bilińska-Wielgus N, Frąc M (2017b) Characterization and influence of a multi-enzymatic biopreparation for biogas yield enhancement. BioResources 12:6187–6206
Park C, Lee C, Kim S, Chen Y, Chase HA (2005) Upgrading of anaerobic digestion by incorporating two different hydrolysis processes. Biosci Bioeng 100:164–167. https://doi.org/10.1263/jbb.100.164
Pavi S, Kramer LE, Gomes LP, Miranda LAS (2017) Biogas production from co-digestion of organic fraction of municipal solid waste and fruit and vegetable waste. Bioresour Technol 228:362–367. https://doi.org/10.1016/j.biortech.2017.01.003
Procházka J, Dolejš P, Máca J, Dohányos M (2012) Stability and inhibition of anaerobic processes caused by insufficiency or excess of ammonia nitrogen. Appl Microb Biotechnol 93:439–447. https://doi.org/10.1007/s00253-011-3625-4
Rahman MA, Møller HB, Saha CK, Alam MM, Wahid R, Feng L (2017) Optimal ratio for anaerobic co-digestion of poultry droppings and lignocellulosic-rich substrates for enhanced biogas production. Energy Sustain Dev 39:59–66. https://doi.org/10.1016/j.esd.2017.04.004
Rajagopal R, Massé DI, Singh G (2013) A critical review on inhibition of anaerobic digestion process by excess ammonia. Bioresour Technol 143:632–641. https://doi.org/10.1016/j.biortech.2013.06.030
Satpathy P, Steinigeweg S, Cypionka H, Engelen B (2016) Different substrates and starter inocula govern microbial community structures in biogas reactors. Environ Technol 37:1441–1450. https://doi.org/10.1080/09593330.2015.1118559
Sawatdeenarunat C, Surendra K, Takara D, Oechsner H, Khanal SK (2015) Anaerobic digestion of lignocellulosic biomass: challenges and opportunities. Environ Technol 178:178–186. https://doi.org/10.1016/j.biortech.2014.09.103
Schlüter A, Bekel T, Diaz NN, Dondrup M, Eichenlaub R, Gartemann K-H et al (2008) The metagenome of a biogas-producing microbial community of a production-scale biogas plant fermenter analysed by the 454-pyrosequencing technology. Biotechnol J 136:77–90. https://doi.org/10.1016/j.jbiotec.2008.05.008
Tao Y, Gao D-W, Wang H-Y, de Kreuk M, Ren N-Q (2013) Ecological characteristics of seeding sludge triggering a prompt start-up of anammox. Bioresour Technol 133:475–481. https://doi.org/10.1016/j.biortech.2013.01.147
Vasco-Correa J, Ge X, Li Y (2016) Fungal pretreatment of non-sterile miscanthus for enhanced enzymatic hydrolysis. Bioresour Technol 203:118–123. https://doi.org/10.1016/j.biortech.2015.12.018
Wang X, Lu X, Li F, Yang G (2014) Effects of temperature and carbon-nitrogen (C/N) ratio on the performance of anaerobic co-digestion of dairy manure, chicken manure and rice straw, focusing on ammonia inhibition. PLoS ONE 9:e97265. https://doi.org/10.1371/journal.pone.0097265
Wartell BA, Krumins V, Alt J, Kang K, Schwab BJ, Fennell DE (2012) Methane production from horse manure and stall waste with softwood bedding. Bioresour Technol 112:42–50. https://doi.org/10.1016/j.biortech.2012.02.012
Westerholm M, Hansson M, Schnürer A (2012) Improved biogas production from whole stillage by co-digestion with cattle manure. Bioresour Technol 114:314–319. https://doi.org/10.1016/j.biortech.2012.03.005
Wolińska A, Frąc M, Oszust K, Szafranek-Nakonieczna A, Zielenkiewicz U, Stępniewska Z (2017) Microbial biodiversity of meadows under different modes of land use: catabolic and genetic fingerprinting. World J Microb Biotechnol 33:154. https://doi.org/10.1007/s11274-017-2318-2
Zhang P, Zeng G, Zhang G, Li Y, Zhang B, Fan M (2008) Anaerobic co-digestion of biosolids and organic fraction of municipal solid waste by sequencing batch process. Fuel Process Technol 89:485–489. https://doi.org/10.1016/j.fuproc.2007.11.013
Zhang C, Su H, Baeyens J, Tan T (2014) Reviewing the anaerobic digestion of food waste for biogas production. Renew Sustain Energy Rev 38:383–392. https://doi.org/10.1016/j.rser.2014.05.038
Zieliński M, Dębowski M, Krzemieniewski M (2016) Oczyszczanie ścieków mleczarskich w reaktorze beztlenowym z wypełnieniem aktywnym. Eco Eng 47:114–122
Acknowledgements
Scientific work was funded from the budget for science by The National Centre for Research and Development in Poland. LIDER 048/L-2/10. Programme 2011–2014.
Author information
Authors and Affiliations
Corresponding author
Additional information
Editorial responsibility: Xu Han.
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Oszust, K., Gryta, A., Ziemiński, K. et al. Characterization of microbial functional and genetic diversity as a novel strategy of biowaste ecotoxicological evaluation. Int. J. Environ. Sci. Technol. 16, 4261–4274 (2019). https://doi.org/10.1007/s13762-018-2066-3
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13762-018-2066-3