Abstract
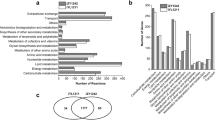
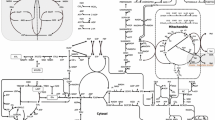
The metabolic flux in microbial fuel cells (MFCs) is significantly different from conventional fermentation because the electrode in MFCs acts as a terminal electron acceptor. In this study, the difference in the carbon metabolism of Klebsiella pnuemoniae L17 (Kp L17) during growth in MFCs and conventional bioreactors was studied using glucose as the sole carbon and energy source. For metabolic flux analysis (MFA), the in silico metabolic flux model of Kp L17 was also constructed. The MFC bioreactor operated in oxidative mode, where electrons are removed by the anode electrode, generated a smaller quantity of reductive metabolites (e.g., lactate, 2,3-butanediol and ethanol) compared to the conventional fermentative bioreactor (non-MFC). Stoichiometric analysis indicated that the cellular metabolism in MFC had partially (or significantly) shifted to anaerobic respiration from fermentation, the former of which was similar to that often observed under micro-aerobic conditions. Electron balance analysis suggested that 30% of the electrons generated from glucose oxidation were extracted from the microbe and transferred to the electrode. These results highlight the potential use of MFCs in regulating the carbon metabolic flux in a bioprocess.
Similar content being viewed by others
References
Lovley, D. R. and K. P. Nevin (2011) A shift in the current: New applications and concepts for microbe-electrode electron exchange. Curr. Opin. Biotechnol. 22: 441–448.
Lovley, D. R. (2011) Powering microbes with electricity: Direct electron transfer from electrodes to microbes. Environ. Microbiol. Rep. 3: 27–35.
Gonzalez, P. J., C. Correia, I. Moura, C. D. Brondino and J. J. Moura (2006) Bacterial nitrate reductases: Molecular and biological aspects of nitrate reduction. J. Inorg. Biochem. 100: 1015–1023.
Ashok, S., S. Mohan Raj, Y. Ko, M. Sankaranarayanan, S. Zhou, V. Kumar, and S. Park (2013) Effect of puuC overexpression and nitrate addition on glycerol metabolism and anaerobic 3-hydroxypropionic acid production in recombinant Klebsiella pneumoniae ?glpK?dhaT. Metab. Eng. 15: 10–24.
Pinchuk, G. E., O. V. Geydebrekht, E. A. Hill, J. L. Reed, A. E. Konopka, A. S. Beliaev, and J. K. Fredrickson (2011) Pyruvate and lactate metabolism by Shewanella oneidensis MR-1 under fermentation, oxygen limitation, and fumarate respiration conditions. Appl. Environ. Microbiol. 77: 8234–8240.
Mordkovich, N., T. Voeikova, L. Novikova, I. Smirnov, V. Il’in, P. Soldatov, A. Y. Tyurin-Kuz’min, T. Smolenskaya, V. Veiko, and R. Shakulov (2013) Effect of NAD+-dependent formate dehydrogenase on anaerobic respiration of Shewanella oneidensis MR-1. Microbiol. 82: 404–409.
Glass, C. and J. Silverstein (1998) Denitrification kinetics of high nitrate concentration water: pH effect on inhibition and nitrite accumulation. Water Res. 32: 831–839.
Gerritse, J., O. Drzyzga, G. Kloetstra, M. Keijmel, L.P. Wiersum, R. Hutson, M. D. Collins, and J. C. Gottschal (1999) Influence of different electron donors and acceptors on dehalorespiration of tetrachloroethene by Desulfitobacterium frappieri TCE1. Appl. Environ. Microbiol. 65: 5212–5221.
Riondet, C., R. Cachon, Y. Wache, G. Alcaraz, and C. Divies (2000) Extracellular oxidoreduction potential modifies carbon and electron flow in Escherichia coli. J. Bacteriol. 182: 620–626.
Du, C., H. Yan, Y. Zhang, Y. Li, and Z. Cao (2006) Use of oxidoreduction potential as an indicator to regulate 1,3-propanediol fermentation by Klebsiella pneumoniae. Appl. Microbiol. Biotechnol. 69: 554–563.
Call, D. F. and B. E. Logan (2011) A method for high throughput bioelectrochemical research based on small scale microbial electrolysis cells. Biosens. Bioelectron. 26: 4526–4531.
Liu, C. G., C. Xue, Y. H. Lin, and F. W. Bai (2013) Redox potential control and applications in microaerobic and anaerobic fermentations. Biotechnol. Adv. 31: 257–265.
Rosenbaum, M. A. and A. E. Franks (2014) Microbial catalysis in bioelectrochemical technologies: status quo, challenges and perspectives. Appl. Microbiol. Biotechnol. 98: 509–518.
Varma, A. and B. O. Palsson (1994) Metabolic flux balancing: Basic concepts, scientific and practical use. Biotechnol. 12.
Jones, J. A., O. D. Toparlak, and M. A. Koffas (2015) Metabolic pathway balancing and its role in the production of biofuels and chemicals. Curr. Opin. Biotechnol. 33: 52–59.
Lee, D. Y., H. Yun, S. Park, and S. Y. Lee (2003) MetaFluxNet: The management of metabolic reaction information and quantitative metabolic flux analysis. Bioinformat. 19: 2144–2146.
Tang, Y. J., H. G. Martin, P. S. Dehal, A. Deutschbauer, X. Llora, A. Meadows, A. Arkin, and J. D. Keasling (2009) Metabolic flux analysis of Shewanella spp. reveals evolutionary robustness in central carbon metabolism. Biotechnol. Bioeng. 102: 1161–1169.
Chen, X.-C., H. Song, T. Fang, J.-M. Cao, H.-J. Ren, J.-X. Bai, J. Xiong, P.-K. Ouyang, and H.-J. Ying (2010) Enhanced cyclic adenosine monophosphate production by Arthrobacter A302 through rational redistribution of metabolic flux. Bioresour. Technol. 101: 3159–3163.
Niu, K., X. Zhang, W. S. Tan, and M. L. Zhu (2011) Effect of culture conditions on producing and uptake hydrogen flux of biohydrogen fermentation by metabolic flux analysis method. Bioresour. Technol. 102: 7294–7300.
Jung, M. Y., S. Mazumdar, S. H. Shin, K.S. Yang, J. Lee, and M. K. Oh (2014) Improvement of 2,3-butanediol yield in Klebsiella pneumoniae by deletion of the pyruvate formate-lyase gene. Appl. Environ. Microbiol. 80: 6195–6203.
Seol, E., S. K. Ainala, B. S. Sekar, and S. Park (2014) Metabolic engineering of Escherichia coli strains for co-production of hydrogen and ethanol from glucose. Int. J. Hydrogen Energy 39: 19323–19330.
Heidelberg, J. F., I. T. Paulsen, K. E. Nelson, E. J. Gaidos, W. C. Nelson, T. D. Read, J. A. Eisen, R. Seshadri, N. Ward, B. Methe, R. A. Clayton, T. Meyer, A. Tsapin, J. Scott, M. Beanan, L. Brinkac, S. Daugherty, R. T. DeBoy, R. J. Dodson, A. S. Durkin, D. H. Haft, J. F. Kolonay, R. Madupu, J. D. Peterson, L.A. Umayam, O. White, A. M. Wolf, J. Vamathevan, J. Weidman, M. Impraim, K. Lee, K. Berry, C. Lee, J. Mueller, H. Khouri, J. Gill, T. R. Utterback, L. A. McDonald, T. V. Feldblyum, H. O. Smith, J. C. Venter, K. H. Nealson and C. M. Fraser (2002) Genome sequence of the dissimilatory metal ion-reducing bacterium Shewanella oneidensis. Nat. Biotechnol. 20: 1118–1123.
Rodionov, D. A., C. Yang, X. Li, I. A. Rodionova, Y. Wang, A. Y. Obraztsova, O. P. Zagnitko, R. Overbeek, M. F. Romine, S. Reed, J. K. Fredrickson, K. H. Nealson, and A. L. Osterman (2010) Genomic encyclopedia of sugar utilization pathways in the Shewanella genus. BMC Genom. 11: 494.
Mao, L. and W. S. Verwoerd (2013) Genome-scale stoichiometry analysis to elucidate the innate capability of the cyanobacterium Synechocystis for electricity generation. J. Ind. Microbiol. Biotechnol. 40: 1161–1180.
Tang, Y. J., A. L. Meadows, J. Kirby, and J. D. Keasling (2007) Anaerobic central metabolic pathways in Shewanella oneidensis MR-1 reinterpreted in the light of isotopic metabolite labeling. J. Bacteriol. 189: 894–901.
Feng, X., Y. Xu, Y. Chen, and Y. J. Tang (2012) Integrating flux balance analysis into kinetic models to decipher the dynamic metabolism of Shewanella oneidensis MR-1. PLoS Comput. Biol. 8: e1002376.
Finch, A. S., T. D. Mackie, C. J. Sund, and J. J. Sumner (2011) Metabolite analysis of Clostridium acetobutylicum: fermentation in a microbial fuel cell. Bioresour. Technol. 102: 312–315.
Zhang, L., S. Zhou, L. Zhuang, W. Li, J. Zhang, N. Lu, and L. Deng (2008) Microbial fuel cell based on Klebsiella pneumoniae biofilm. Electrochem. Commun. 10: 1641–1643.
Bond, D. R. and D. R. Lovley (2003) Electricity production by Geobacter sulfurreducens attached to electrodes. Appl. Environ. Microbiol. 69: 1548–1555.
Lanthier, M., K. B. Gregory, and D. R. Lovley (2008) Growth with high planktonic biomass in Shewanella oneidensis fuel cells. FEMS Microbiol. Lett. 278: 29–35.
Zeng, A. P., K. Menzel, and W. D. Deckwer (1996) Kinetic, dynamic, and pathway studies of glycerol metabolism by Klebsiella pneumoniae in anaerobic continuous culture: II. Analysis of metabolic rates and pathways under oscillation and steadystate conditions. Biotechnol. Bioeng. 52: 561–571.
Arasu, M. V., V. Kumar, S. Ashok, H. Song, C. Rathnasingh, H. J. Lee, D. Seung, and S. Park (2011) Isolation and characterization of the new Klebsiella pneumoniae J2B strain showing improved growth characteristics with reduced lipopolysaccharide formation. Biotechnol. Bioproc. Eng. 16: 1134–1143.
Kim, J. R., G. C. Premier, F. R. Hawkes, J. Rodriguez, R. M. Dinsdale, and A. J. Guwy (2010) Modular tubular microbial fuel cells for energy recovery during sucrose wastewater treatment at low organic loading rate. Bioresour. Technol. 101: 1190–1198.
Li, X., L. Liu, T. Liu, T. Yuan, W. Zhang, F. Li, S. Zhou, and Y. Li (2013) Electron transfer capacity dependence of quinone-mediated Fe(III) reduction and current generation by Klebsiella pneumoniae L17. Chemosphere 92: 218–224.
Kim, J. R., B. Min, and B. E. Logan (2005) Evaluation of procedures to acclimate a microbial fuel cell for electricity production. Appl. Microbiol. Biotechnol. 68: 23–30.
Schilling, C. H., J. S. Edwards, D. Letscher, and B. O. Palsson (2000) Combining pathway analysis with flux balance analysis for the comprehensive study of metabolic systems. Biotechnol. Bioeng. 71: 286–306.
Lu, M., C. Park, S. Lee, B. Kim, M.-K. Oh, Y. Um, J. Kim, and J. Lee (2014) The regulation of 2, 3-butanediol synthesis in Klebsiella pneumoniae as revealed by gene over-expressions and metabolic flux analysis. Bioproc. Biosyst. Eng. 37: 343–353.
Liu, T., X. Li, W. Zhang, M. Hu, and F. Li (2014) Fe(III) oxides accelerate microbial nitrate reduction and electricity generation by Klebsiella pneumoniae L17. J. Colloid Interface Sci. 423: 25–32.
Kim, J. R., S. H. Jung, J. M. Regan, and B. E. Logan (2007) Electricity generation and microbial community analysis of alcohol powered microbial fuel cells. Bioresour. Technol. 98: 2568–2577.
Menzel, K., K. Ahrens, A. Zeng, and W. Deckwer (1998) Kinetic, dynamic, and pathway studies of glycerol metabolism by Klebsiella pneumoniae in anaerobic continuous culture: IV. Enzymes and fluxes of pyruvate metabolism. Biotechnol. Bioeng. 60: 617–626.
Durgapal, M., V. Kumar, T. H. Yang, H. J. Lee, D. Seung, and S. Park (2014) Production of 1, 3-propanediol from glycerol using the newly isolated Klebsiella pneumoniae J2B. Bioresour. Technol. 159: 223–231.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Kim, C., Ainala, S.K., Oh, YK. et al. Metabolic flux change in Klebsiella pneumoniae L17 by anaerobic respiration in microbial fuel cell. Biotechnol Bioproc E 21, 250–260 (2016). https://doi.org/10.1007/s12257-015-0777-6
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12257-015-0777-6