Abstract
We present a model for cell growth, division and packing under soft constraints that arise from the deformability of the cells as well as of a membrane that encloses them. Our treatment falls within the framework of diffuse interface methods, under which each cell is represented by a scalar phase field and the zero level set of the phase field represents the cell membrane. One crucial element in the treatment is the definition of a free energy density function that penalizes cell overlap, thus giving rise to a simple model of cell–cell contact. In order to properly represent cell packing and the associated free energy, we include a simplified representation of the anisotropic mechanical response of the underlying cytoskeleton and cell membrane through penalization of the cell shape change. Numerical examples demonstrate the evolution of multi-cell clusters and of the total free energy of the clusters as a consequence of growth, division and packing.







Similar content being viewed by others
Notes
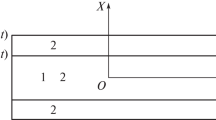
In some simulations, a buffer zone, \(\Omega ^{'}\), is needed around the simulation domain to inhibit unrealistic cell shapes resulting from the enforcement of \(\kappa {\varvec{\nabla }} c_k \cdot {\varvec{n}}= 0\) on \(\partial \Omega \). In addition, this buffer zone also acts as a membrane around the cell cluster. In the buffer zone, an additional term of the form \(\sum _{k=1}^{N}\lambda c_k^2\) is added to the free energy density to penalize the movement of any cells from the active simulation domain (\(\Omega \)) to the buffer zone (\(\Omega ^{'}\)).
In this initial boundary value problem, \(\partial \Omega \) is composed of only a Neumann (flux) boundary (\(\partial \Omega ^{h} = \partial \Omega \)) and hence the Dirichlet boundary is a null set (\(\partial \Omega ^{g} = \varnothing \)).
References
Allen SM, Cahn JW (1979) A microscopic theory for antiphase boundary motion and its application to antiphase domain coarsening. Acta Metall 27(6):1085–1095
Alt S, Ganguly P, Salbreux G (2017) Vertex models: from cell mechanics to tissue morphogenesis. Philos Trans R Soc Lond B Biol Sci 372(1720):20150520
Bangerth W, Hartmann R, Kanschat G (2007) deal. II—a general purpose object oriented finite element library. ACM Trans Math Softw 33(4):24/1–24/27
Brezzi F, Fortin M (1991) Mixed and hybrid finite element methods. Springer, Berlin
Brodland GW (2004) Computational modeling of cell sorting, tissue engulfment, and related phenomena: a review. Appl Mech Rev 57:47–76
Cahn JW, Hilliard JE (1958) Free energy of a nonuniform system. I. Interfacial free energy. J Chem Phys 28(2):258–267
Fletcher A, Osterfield M, Baker RE, Shvartsman SY (2014) Vertex models of epithelial morphogenesis. Biophys J 106(11):2291–2304
Gilbert SF (2000) Developmental biology, 6th edn. Sinauer Associates, Sunderland
Glazier JA, Graner F (1993) Simulation of the differential adhesion driven rearrangement of biological cells. Phys Rev E 47:2128–2154
Goel NS, Rogers G (1978) Computer simulation of engulfment and other movements of embryonic tissues. J Theor Biol 71(1):103–140
Goel N, Campbell RD, Gordon R, Rosen R, Martinez H, Yaas M (1970) Self-sorting of isotropic cells. J Theor Biol 28(3):423–468
Graner F (1993) Can surface adhesion drive cell-rearrangement? Part I: biological cell-sorting. J Theor Biol 164(4):455–476
Graner F, Glazier JA (1992) Simulation of biological cell sorting using a two-dimensional extended Potts model. Phys Rev Lett 69:2013–2016
Heroux M, Bartlett R, Hoekstra VHR, Hu J, Kolda T, Lehoucq R, Long K, Pawlowski R, Phipps E, Salinger A, Thornquist H, Tuminaro R, Willenbring J, Williams A (2003) An overview of Trilinos. Technical report SAND2003-2927, Sandia National Laboratories
Honda H (1978) Description of cellular patterns by Dirichlet domains: the two-dimensional case. J Theor Biol 72(3):523–543
Honda H (1983) Geometrical models for cells in tissues. Int Rev Cytol 81:191–248
Honda H, Yamanaka H, Eguchi G (1986) Transformation of a polygonal cellular pattern during sexual maturation of the avian oviduct epithelium: computer simulation. Development 98(1):1–19
Itskovitz-Eldor J, Schuldiner M, Karsenti D, Eden A, Yanuka O, Amit M, Soreq H, Benvenisty N (2000) Differentiation of human embryonic stem cells into embryoid bodies compromising the three embryonic germ layers. Mol Med 6(2):88
Kamrin K, Rycroft CH, Nave JC (2012) Reference map technique for finite-strain elasticity and fluid–solid interaction. J Mech Phys Solids 60(11):1952–1969
Li XS (2005) An overview of SuperLU: algorithms, implementation, and user interface. ACM Trans Math Softw 31(3):302–325
Mills KL, Kemkemer R, Rudraraju S, Garikipati K (2014) Elastic free energy drives the shape of prevascular solid tumors. PloS ONE 9(7):e103245
Mirams GR, Arthurs CJ, Bernabeu MO, Bordas R, Cooper J, Corrias A, Davit Y, Dunn S-J, Fletcher AG, Harvey DG et al (2013) Chaste: an open source C++ library for computational physiology and biology. PLoS Comput Biol 9(3):e1002970
Mochizuki A, Wada N, Ide H, Iwasa Y (1998) Cell-cell adhesion in limb formation, estimated from photographs of cell sorting experiments based on a spatial stochastic model. Dev Dyn 211(3):204–214
Mosaffa P, Asadipour N, Millán D, Rodríguez-Ferran A, Muñoz JJ (2015) Cell-centred model for the simulation of curved cellular monolayers. Comput Part Mech 2(4):359–370
Mosaffa P, Rodríguez-Ferran A, Muñoz JJ (2017) Hybrid cell-centred/vertex model for multicellular systems with equilibrium preserving remodelling. Int J Numer Methods Biomed Eng 34(3):e2928
Nonomura M (2012) Study on multicellular systems using a phase field model. PLOS ONE 7(4):1–904
Reya T, Morrison SJ, Clarke MF, Weissman IL (2001) Stem cells, cancer, and cancer stem cells. Nature 414(6859):105
Rudraraju S, Mills KL, Kemkemer R, Garikipati K (2013) Multiphysics modeling of reactions, mass transport and mechanics of tumor growth. In: Holzapfel G, Kuhl E (eds) Computer models in biomechanics. Springer, Dordrecht, pp 293–303
Schierenberg E (2006) Embryological variation during nematode development (2 Jan 2006), WormBook, ed. The C. elegans Research Community, WormBook
Swat MH, Thomas GL, Belmonte JM, Shirinifard A, Hmeljak D, Glazier JA (2012) Multi-scale modeling of tissues using compucell3d. In: Asthagiri AR, Arkin AP (eds) Methods in cell biology, vol 110. Elsevier, Amsterdam, pp 325–366
Vedantam S, Patnaik BSV (2006) Efficient numerical algorithm for multiphase field simulations. Phys Rev E 73(1):016703
Verner SN, Garikipati K (2018) A computational study of the mechanisms growth-driven folding patterns on shells, with application to the developing brain. Extreme Mech Lett 18:58–69
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Jiang, J., Garikipati, K. & Rudraraju, S. A Diffuse Interface Framework for Modeling the Evolution of Multi-cell Aggregates as a Soft Packing Problem Driven by the Growth and Division of Cells. Bull Math Biol 81, 3282–3300 (2019). https://doi.org/10.1007/s11538-019-00577-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11538-019-00577-1