Abstract
Key message
Fine mapping and sequencing revealed 28 genes in the non-recombining haplotype containing Fhb1 . Of these, only a GDSL lipase gene shows a pathogen-dependent expression pattern.
Abstract
Fhb1 is a prominent Fusarium head blight resistance locus of wheat, which has been successfully introgressed in adapted breeding material, where it confers a significant increase in overall resistance to the causal pathogen Fusarium graminearum and the fungal virulence factor and mycotoxin deoxynivalenol. The Fhb1 region has been resolved for the susceptible wheat reference genotype Chinese Spring, yet the causal gene itself has not been identified in resistant cultivars. Here, we report the establishment of a 1 Mb contig embracing Fhb1 in the donor line CM-82036. Sequencing revealed that the region of Fhb1 deviates from the Chinese Spring reference in DNA size and gene content, which explains the repressed recombination at the locus in the performed fine mapping. Differences in genes expression between near-isogenic lines segregating for Fhb1 challenged with F. graminearum or treated with mock were investigated in a time-course experiment by RNA sequencing. Several candidate genes were identified, including a pathogen-responsive GDSL lipase absent in susceptible lines. The sequence of the Fhb1 region, the resulting list of candidate genes, and near-diagnostic KASP markers for Fhb1 constitute a valuable resource for breeding and further studies aiming to identify the gene(s) responsible for F. graminearum and deoxynivalenol resistance.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
One of the most prevalent pathogens to hexaploid wheat (Triticum aestivum) is the hemibiotrophic fungus Fusarium graminearum. The related disease Fusarium head blight (FHB) leads to severe reduction in grain yield and quality causing globally devastating economic losses. Infected grain may be contaminated with mycotoxins such as deoxynivalenol (DON) and other heat-stable trichothecene type B toxins, which remain in processed food stuffs and feed and constitute a serious threat to food and feet safety (Pestka 2010). The European Union and many other countries have enacted maximum levels for DON in food stuffs (The European Commission 2006) and the United States Food and Drug Administration has issued advisory levels. Conventional agronomical measures to control the disease, such as changed crop rotations, tillage regimes or the use of fungicides, are costly and/or frequently not applicable. Breeding for resistance against FHB in wheat and other small grain cereals is a sustainable and environmentally friendly strategy to achieve durable and cost efficient resistance.
A broad range of resistance sources exist and about 100 quantitative trait loci (QTL) were described to date by genetic mapping in diverse hexaploid wheat germplasm (Buerstmayr et al. 2009). Yet, most are only minor contributors to overall resistance. Among the strongest and most reliable QTL is Fhb1, which has been the focus of several studies aiming to fine-map, identify the causal gene and define its biological mode of action. Fhb1 (syn. Qfhs.ndsu-3BS) was first described as a strong contributor to type 2 resistance (resistance against spreading of the disease) located on wheat chromosome 3BS by Waldron et al. (1999) in a biparental recombinant-inbred population derived from the highly resistant Chinese spring wheat landrace Sumai-3 using RFLP markers. The QTL was confirmed by Anderson et al. (2001) with additional SSR markers and a second population generated from Sumai-3 derivative ND2603 as the resistance donor and in an independent study by Buerstmayr et al. (2002), who employed a doubled haploid (DH) population with CM-82036, a CYMMIT-derived offspring of Sumai-3 as the resistance donor. Fine-mapping Fhb1 (Cuthbert et al. 2006; Liu et al. 2006) narrowed the original confidence interval from SSR markers gwm493 and gwm533 (Anderson et al. 2001) to a 1.2 cM interval between sts3B-189 and sts3B-206 (Liu et al. 2006). BAC sequencing of the syntenic region in the susceptible wheat reference cultivar Chinese Spring yielded UMN10 (Liu et al. 2008), a near-diagnostic marker for Fhb1 that is widely used for marker-assisted selection in wheat. A single recombinant line in the same study further narrowed the interval down to 0.4 cM with sts3B-32 replacing sts3B-206. The sequence of a large genomic contig harboring the susceptible Fhb1 region in Chinese Spring has been reported (Choulet et al. 2010) spurring comparisons with the sequence in resistant cultivars. An association mapping study of FHB-related traits identified several significant marker associations yet it seemed difficult to reconcile the genetic map with the physical sequence of the reference genotype (Hao et al. 2012).
Type 2 resistance conferred by Fhb1 is associated with its ability to inactivate DON. F. graminearum requires DON to spread from the initial infection site and penetrate further into the rachis and adjacent spikelets (Jansen et al. 2005). Lemmens et al. (2005) showed that the QTL co-localizes with the higher ability to transform DON into the non-toxin DON-3-O-glucoside. To date, several UDP-glucosyltransferases (UGT) capable of effectively inactivating DON have been identified in several species of the Gramineae family (Schweiger et al. 2010, 2013). The barley-derived HvUGT13248 when transformed into susceptible wheat is sufficient to confer high level spreading resistance against F. graminearum (Li et al. 2015). Yet, no such gene could be associated with Fhb1. Also the genomic region harboring the susceptible Fhb1 allele from Chinese Spring does not contain a small molecule-accepting UGT gene between markers flanking the QTL.
Several proteomic, metabolomic and transcriptomic studies have sought to pinpoint the mechanism underlying Fhb1 (Walter et al. 2008; Gunnaiah et al. 2012; Kugler et al. 2013; Schweiger et al. 2013; Xiao et al. 2013; Zhuang et al. 2013; Warth et al. 2014; Nussbaumer et al. 2015; Hofstad et al. 2016). A combined proteomic and metabolomic study by Gunnaiah et al. (2012) suggested that the higher accumulation of phenylpropanoids leads to the resistance effect observed in near-isogenic lines (NILs) harboring the QTL. Using RNA-seq data, Xiao et al. (2013) observed changed jasmonic acid signaling in a deletion mutant of resistant cv. Wangshuibai and suggested this to be as a possible mechanism for the resistance mediated by the QTL. The to date only study that associated transcript abundances to genetically mapped positions stems from a small eQTL study by Zhuang et al. (2013), who suggested that a pectin methyl esterase inhibitor gene mapping into the confidence interval, which is down-regulated in susceptible lines, could be the causal gene. Only the most recent large-scale RNA-seq studies (Nussbaumer et al. 2015; Hofstad et al. 2016) include whole genome mapping data based on the recent release of the full wheat gene models of Chinese Spring (Mayer et al. 2014). Both studies using unrelated pairs of NILs presented viable candidate genes mapped to the Fhb1 interval that shows stress-dependent and QTL-associated expression, which do show some overlaps to previous transcriptomics studies. Yet, despite the seemingly completeness of the reference genome, it may still not include the causative gene: in our own co-expression network study (Nussbaumer et al. 2015), an F-box protein showing strong constitutive expression for lines including Fhb1 originates from the QTL-homoeologous region on chromosome 3D. Possibly this and other genes are not present in the susceptible genotypes and thus such transcripts map to homologs or homoeologs elsewhere in the reference gene set. To unambiguously identify genes present in the QTL interval, it is therefore essential to establish the sequence of the respective genomic region in a resistant cultivar.
Here, we report sequencing and analysis of a 1 Mb genomic contig harboring Fhb1 from the Sumai-3 derivative CM-82036. The region has been thoroughly analyzed for gene content and transcriptional activity with dense time-course RNA-seq data derived from a F. graminearum and mock-challenged NIL pair differing in Fhb1. Additional fine mapping identified recombinant lines with cross-over events mapping to the sequenced contig, thus, successfully localizing the Fhb1 gene on a region of about 860 kb harboring 28 genes. These findings provide a relevant resource for work focused on identifying the underlying gene.
Methods
Generation of plant material
NILs for Fhb1 and Qhfs.ifa-5A have been developed from a cross of the highly resistant donor cv. CM-82036 and the highly susceptible cultivar Remus (Sappo/Mex//Famos) using CM-82036 as the recurrent parent. CM-82036 originates from the cross Sumai-3/Thornbird-S and was developed in a shuttle breeding program between CIMMYT Mexico and South America. Remus is a German spring wheat cultivar with well-adapted agronomic characters, developed at the Bavarian State Institute for Agronomy in Freising, Germany (Buerstmayr et al. 2002). In the BC5F2 generation sister lines either homozygous for both resistant alleles at Fhb1 and Qhfs.ifa-5A (CM-NIL38), susceptible alleles for Fhb1 and resistant for Qhfs.ifa-5A (CM-NIL47) or the susceptible alleles at both QTL (CM-NIL51) were selected. The presence of the resistance QTL on 3BS was verified with marker UMN10 (Liu et al. 2006). The genotype of Qfhs.ifa-5A was confirmed with flanking simple sequence repeat (SSR) markers gwm304, barc186 and barc1 (Buerstmayr et al. 2003).
BAC library construction
A bacterial artificial chromosome (BAC) library based on CM-82036 has been constructed from 20 g of deep-frozen fresh leave tissue harvested from 12-day-old seedlings at CNRGV-INRA (Toulouse, France). The extraction of nuclear DNA and preparation of high molecular weight (HMW)-DNA-agarose plugs to limit DNA shearing followed a protocol using sucrose extraction buffer (SEB) as described in Peterson et al. (2000) with slight modifications (given in the Supplementary File methods).
Following a partial HindIII test restriction digestion to establish the conditions yielding the highest fraction of 100–250 kb sized HMW-DNA fragments, a total of six plugs were digested with 0.4–0.7 units/mL HindIII yielding pools A, B and C. The protocols for digestions, fragment sizing and elution followed Peterson et al. (2000) with modifications (Supplementary File methods). Eluted HMW-DNA from all three pools was ligated into a HindIII linearized and dephosphorylated pIndigoBAC5 vector using T4 ligase for 10 h at 16 °C. For transformation by electroporation, 100 μL of Escherichia coli strain DH10B suspension was transformed with 15 μL desalted ligation product and left to recover in 2 mL SOC medium at 37 °C for 1 h. 50 μL of the suspension was plated as a control to test transformation efficiency on LB medium supplemented with X-Gal, IPTG and chloramphenicol (X/I/C). The remaining 1.95 mL was supplemented with 200 μL glycerol and aliquoted to contain about 600–1500 individual clones per pool in deep well plates. Pools were incubated for 16 h at 37 °C. From these stocks 300 μL working stocks were transferred into separate deep well plates. All stocks and aliquots were stored at −80 °C. At least 20 individual clones per pool were picked from the test plate, plasmid DNA was isolated, NotI digested and separated by pulsed-field electrophoresis to estimate yielding estimated average fragment sizes of 99.8, 138.1 and 163.7 kb for pools A, B and C, respectively. After this preliminary size estimation, remaining ligations from fractions B and C were transformed, characterized and pooled. Global isothermal amplification of BAC DNA was performed by amplification of 1 μL of denatured working stock with Phi29 (GenomiPhi V2 DNA Amplification Kit, GE Healthcare, Buckinghamshire, UK) at 30 °C for 2 h followed by heat inactivation of the enzyme. Phi29-amplified DNA was stored at −80 °C and diluted 1:200 for PCR screening of BAC DNA. This resource and dedicated screening tools are available upon request at http://cnrgv.toulouse.inra.fr.
Screening, isolation, sequencing of BACs and assembly of the Fhb1 sequence
Phi29 amplified pools were screened with published and newly generated PCR-markers (Supplementary Table S1) that were tested on genomic DNA of CM-82036 and nullisomic–tetrasomic substitution lines for chromosome 3B of Chinese Spring to ensure chromosome specificity. Novel markers were designed based on genic sequences and TE junctions from initially sequenced and annotated BACs. BAC-clone pools positive for either marker were identified based on distinct melting curve differences and/or PCR product formation on a BIO-RAD CFX386 qPCR. To isolate single BAC clones dilutions of 50 μL of the original working stock bacterial pools were plated on X/I/C qTrays and single colonies were picked and distributed into 386-well plates containing selective medium. Single colonies from overnight-incubated 386 well plates were pooled into one sample and screened for the respective marker. If these were positive the single colonies were pooled by rows and columns which were further screened to identify single candidate BACs. Isolated and validated BACs were fingerprinted by NotI digestion to estimate clone sizes and sequencing of BAC ends was done by Sanger sequencing. Midi-prepped (Nucleobond Xtra midi kit, Macherey–Nagel, Düren, Germany) single BAC clones were subjected to 454 sequencing (clones 235H14, 217J06, 131J06, 90G13, 114J13, 2O7, 28L19, 18G11) or 454 paired-end sequencing (305I1) at CNRGV or sequenced as one indexed sample (238O16) on a PacBio SMRT cell along with other unrelated samples by a commercial sequencing provider (GATC, Konstanz, Germany). The BAC names given here are abbreviations from the complete clone names (e.g., Tae-B-82036-ng-235H14). Raw 454 read data were processed and assembled using Newbler (version 2.7). Resulting contigs were preliminary ordered by mapping BAC end sequences and genetic markers on the contigs to the published homologous region on 3B of the reference cultivar Chinese Spring (GenBank accession: FN564434). The contig order was refined and remaining gaps were closed by sequencing equimolar pooled BACs 217J06, 235H15, 18G11, 114J13 and 90G13 on one PacBio SMRT cell (GATC, Konstanz, Germany) cells using AHA scaffolding (Bashir et al. 2012) for orientating and ordering contigs and PBJelly (English et al. 2012) for filling gaps. Ab initio gene models refined by BLASTx and protein domain prediction algorithms were retrieved using the semi-automated annotations pipeline TriAnnot (Leroy et al. 2012). The finished annotated contig sequence is deposited at NCBI (GenBank accession: KU641029).
Establishing a fine map for the Fhb1 region
In the BC5F2 generation of the NIL development (see generation of plant material), four plants heterozygous for markers spanning the Fhb1 region (gwm493, UMN10, barc133) and homozygous for the resistant alleles at Qhfs.ifa-5A were selected, further multiplied and plants heterozygous at the Fhb1 region were selected and harvested. 3000 of these BC5F2 lines, which are in the F2 generation for the Fhb1 region, comprise the fine mapping population. These BC5F2 lines were grown in the greenhouse and genotyped using flanking markers gwm493 and barc133 in 2013 and 2014. Lines with recombinations between both markers, but still heterozygous for one of them, were brought to the next generation to select homozygous recombinant plants.
The recombinant NILs were further genotyped with a set of seven newly developed SNP markers using KASP-marker assays (LGC-Genomics, Middlesex, UK), based on genic sequences from the established genomic sequence covering Fhb1 and snp3BS-8 (Bernardo et al. 2011). The original UMN10 marker from Liu et al. (2008) was replaced with a novel KASP assay derived from the sequenced UMN10 PCR product of cv. Remus. In addition, dominant gene-specific markers were developed for five of the annotated genes in the Fhb1 region, which were scored on agarose gels. All marker data and primer sequences are collected in Supplementary Table S1.
Greenhouse trials for FHB and DON resistance
The greenhouse experiments were conducted in 2015 as described in Steiner et al. (2009). F. graminearum conidia spores from strain IFA65 required for inoculation were produced on defined SNA medium under UV-light at 25 °C. After two weeks, conidia were harvested and diluted to 50,000 conidia/mL. Aliquots were stored at −80 °C. For every individual recombinant line in the fine-mapping population and the control lines CM-82036, Remus, CM-NIL38, CM-NIL47 and CM-NIL51 two pots were sown with five plants each. The experimental design was a randomized complete block design, with two replications. Temperature in the greenhouse was on average 18/12 °C (day/night) from tillering to heading with 12–14 h daylight. During flowering time, the conditions in the greenhouse were controlled and set at 22 and 17 °C during night with a 16 h photoperiod at 15,000 lux.
FHB resistance evaluations: At anthesis two central adjacent spikelets (four florets) of at least five heads per genotype and replication were inoculated by pipetting 10 µL of conidia spore suspension (500 conidia) between palea and lemma of the two basal florets. Inoculated heads were sprayed with water to provide high humidity and covered with plastic bags for 48 h. The number of diseased spikelets was counted 26 days after inoculation (dai). For each pot, the mean FHB severity as number of diseased spikelets/head was used for further analyses.
Similarly, an additional experiment was carried out to sample tissue for RNA profiling using the exact same protocol but the two NILs CM-NIL38 and CM-NIL51 only. The experimental design was a randomized complete block design with three blocks (=replications) each representing the four possible combinations of two genotypes (CM-NIL38, CM-NIL51) by two treatments (F. graminearum, mock) combinations. Six central spikelets were inoculated to increase the amount of uniformly treated tissue with either mock or F. graminearum spore suspension. Five heads were sampled for each of these conditions at 3, 6, 12, 24, 36 and 48 h after inoculation (hai) in three biological replicates (a total of 360 plants).
DON resistance evaluation: a subset of 35 recombinant NILs representing eight of the nine detected haplotypes was tested for DON resistance. The production and application of the toxin in the heads were conducted as described by Lemmens et al. (2005) with slight modifications: At anthesis, four distal florets of two central spikelets were treated once with 20 μL of a DON solution (12 g/L DON, 0.1 % Tween). Treated heads were sprayed with water and covered with plastic bags for 24 h. The number of DON-bleached spikelets was assessed 26 dai. For each pot, mean DON severity as number of DON-bleached spikelets/head was used for further analyses.
RNA sequencing
For RNA profiling, frozen tissue samples were ground under sterile conditions and pooled to comprise a single sample/data point as described in Kugler et al. (2013). 100 mg frozen tissue was used to extract RNA using the RNeasy Plant Mini Kit (Qiagen, Venlo, Netherlands). Quality and quantity were checked on an automated electrophoresis-system (Experion, #701-7000, Bio-Rad, Hercules, CA, USA). Samples were sequenced on Illumina HiSeq 2000/MiSeq machines (Eurofins MGW, Ebersberg, Germany) yielding at least 20 M 100 bp paired-end reads per sample (Supplementary Table S2). The respective data sets are available in the EBI ArrayExpress (http://www.ebi.ac.uk/arrayexpress/) repository under the accession number E-MTAB-4222. Tophat (Trapnell et al. 2012) was used to assign reads from the susceptible and resistant genotype to the Fhb1 contig (Supplementary Table S2). Reads were kept which showed at maximum one error in the alignment over its entire read length. Next, the contig-mapped reads were compared to the same data set mapped to the high confidence and low confidence gene sets generated from all 21 chromosomes of Chinese Spring (Mayer et al. 2014) to detect falsely mapped reads. Reads were removed from the contig alignment when they found better matches to contigs from 20 Chinese Spring chromosomes excluding chromosome 3B. HTSeq (Anders et al. 2015) was used to extract the reads counts for annotated gene models. Significant differences between treatments and lines were obtained using the exactTest function in the R package EdgeR (Robinson et al. 2010).
SNP in CDS regions were detected by searching for polymorphisms present only in mapped reads from CM-NIL51, which includes the Remus susceptible Fhb1 region compared to reads originating from the CM-82036 Fhb1-region (CM-NIL38). We used the software package Geneious 8.1.7 (Kearse et al. 2012) to call SNP covered by at least five reads.
Results
The resistant locus of Fhb1 includes a 395 kb highly dissimilar sequence compared to the region in the susceptible Chinese Spring reference.
We have established a BAC library based on the Fhb1 donor line CM-82036 yielding 488,390 clones (576 pools) with a mean insert size of clones of 146 kb, corresponding to a total coverage of 3.9 genome equivalents. The library was first screened for BACs harboring flanking markers sts32 and sts189 (Liu et al. 2006, 2008) or the near-diagnostic UMN10 (Liu et al. 2008), which yielded several BACs (18G17, 114J13, 2O7, 90G13) that bridge the distance between sts32 and UMN10, but not between the latter and sts189 for which one BAC was isolated (28L19, Fig. 1a). Additional subgenome-specific markers (Supplementary Table S1) based on transposable element-junctions and genic sequences were generated from BAC end sequences or fully sequenced BACs. Ultimately, ten BACs of which five cover the entire contig were sequenced and assembled to form a contig of 1029 kb (Fig. 1a, GenBank accession: KU641029).
Reconstruction of the Fhb1 region in CM-82036. a Sequenced BACs bridge a distance of 1.027 kb including flanking markers sts32 and sts189 and near-diagnostic marker UMN10. Arrows indicate positions and orientations of annotated genes. Light gray arrows relate to low confidence gene models. b Dot plot analysis comparing the sequenced Fhb1 contig to the colinear segment of the published sequence of Chinese Spring (ctg0954b). c Comparison of gene content between CM-82036 and Chinese Spring. Regions with highly similar genes are shaded in gray. Genes with homology <80 % are indicated by dashed lines
The distance between flanking markers sts32 and sts189 stretches 800.5 kb compared to 752 kb of the same interval in the Chinese Spring. While markers sts32 and UMN10 are evenly spaced in both cultivars, the distance between UMN10 and sts189 is around 49 kb larger in CM-82036. A dot plot analysis using a 100 bp word size to compare collinearity between the Fhb1-contig and the respective region in Chinese Spring showed more extensive rearrangements. Both sequences are highly dissimilar between positions 143 and 517 kb of the Fhb1-contig, which also includes UMN10 at position 426.0 kb, but high similarities exist in the flanking regions (Fig. 1b). Two gaps (positions 860.7 and 888.3 kb) remain unresolved after 454 sequencing of either BAC and additional PacBio sequencing. Both gaps are located in annotated transposable elements and are covered by the same two BACs, 217J6 and 131J6. The missing sequences are most likely not longer than several kb each and remain part of the flanking transposons (Supplementary File Fig. S1). The sequence lengths of the finished BAC sequences of BACs 217J6 (229 kb) and 131J6 (136 kb) are matched well by size estimates of restriction endonuclease-digested BAC clones (235 and 135 kb, respectively, Supplementary File Fig. S2). While the gap at position 860.7 kb is part of a segment not present in Chinese Spring, the distance between flanking sequence of the gap at 888.3 kb spans 4.3 kb in Chinese Spring.
Unique genes in the Fhb1 locus
Thirty-tree genes were identified in the contig by TriAnnot (Leroy et al. 2012) (Table 1; Fig. 1c). Of these, 14 gene models were classified as high confidence, with clear biological evidences for start/stop codons and intron–exon junctions. Two low confidence genes (#13 and #19) located in the 5′-UTR or N-terminal regions of genes #12 and #18, respectively, most likely represent erroneous gene calls. A BLASTn search for the best matches against the genomic contig ctg0954b covering the Fhb1 region in Chinese Spring and against the IWGSC 2.2 high and low confidence gene sets identified the best matches against the Chinese Spring reference (Table 1; Fig. 1c; Supplementary Table S3). Here, we found that in accordance with the direct comparison of genomic sequences also the gene content in the Fhb1 region is different to Chinese Spring. With the exception of four low confidence gene models located on the very distal end of the sequenced region (genes #1–#4 in contig range 25–128 kb), genes in the flanking sections of the sequenced genomic region match genes annotated on ctg0954b well: genes #6–#9 correspond to TAA_ctg0954b.00280.1–TAA_ctg0954b.00310.1 and genes #28–#33 match TAA_ctg0954b.00420.1–TAA_ctg0954b.00490.1. For two additional low confidence genes in Chinese Spring TAA_ctg0954b.00430.1 and TAA_ctg0954b.00480.1, encoding an unknown protein and an F-box domain-containing protein, no evidence was found that either gene is also present in the Fhb1 region.
The central section does—despite the high sequence divergence to Chinese Spring—harbor several genes present in both genotypes. Most of these are located in a segment between sts32 and the central marker UMN10: six genes (#10–#15, between positions 155 and 228 kb) are annotated in both cultivars. Additionally, Chinese Spring harbors TAA_ctg0954b.00350.1, encoding a C-terminally truncated UDP-glucose dehydrogenase (lacking active site residues), which is not present in the Fhb1 region of CM-82036.
A second segment between marker UMN10 and sts189 hosts 12 genes (#16–#27, positions 288–530 kb) of which only few find best matches to the annotated genes on ctg0954b. These share overall poor coverage and similarity to the BLASTn matched genes. None of the six predicted high confidence genes in this segment finds matches to 3B-mapped IWGSC high confidence genes and only one finds a match in the low confidence gene set (Supplementary Table S3). Several, however, find best matches to putative homoeoallelic loci on chromosomes 3D (#26 and #27) and 3A (#17), while others (#22, #24 and #25) share high similarities to genes located on chromosomes 2A, 2B and 4D, respectively. In contrast, the Chinese Spring sequence contains one gene, TAA_ctg0954b.00400.1 (unknown), which is not predicted by TriAnnot on Fhb1. Yet two of the four ab initio pipelines employed by TriAnnot suggest a gene model, which finds no further evidence in database comparisons to expressed genes.
At the distal end of the sequenced contig around marker sts189 genes #28 to #33 find best matches in the Chinese Spring reference sequence. The Fhb1 region seems to lack TAA_ctg0954b.t00480.1, an F-box domain-containing protein located between genes #32 and #33, for which no gene models were predicted. This highly colinear region to Chinese Spring also covers the two remaining small gaps in the sequenced contig with no predicted Chinese Spring genes mapping therein.
Gene expression profiles in the Fhb1 region in response to F. graminearum
We added expression data to the sequenced contig to further substantiate gene predictions and to gain an impression of expression dynamics in this region in the presence or absence of the pathogen. Expression profiles were generated from 72 RNA-seq libraries originating from F. graminearum and mock-inoculated wheat head tissues sampled in a dense time-course series from three to 48 h after inoculation. The sampled tissue derives from a newly developed NIL pair with the resistance donor CM-82036 as the recurring parent containing either Fhb1 and Qfhs.ifa-5A (CM-NIL38) or susceptible alleles originating from the German spring wheat cultivar Remus (CM-NIL51). To account for polymorphisms in CM-NIL51 to the sequenced contig, we allowed one mismatch per read when mapping reads to the Fhb1 region. We controlled falsely mapped reads by mapping reads also against the IWGSC high and low confidence gene sets. About 10 % of the mapped reads found better matches in these data sets excluding genes from chromosome 3B and were excluded from any further analysis.
Despite these considerations, only few genes in the central segment of the sequenced contig were hit by reads originating from the susceptible CM-NIL51 (genes #8–#27), while genes in the flanking non-divergent regions are generally expressed at comparable levels in either NIL (Fig. 2a; Supplementary Tables S4 and S5 including raw read counts and significant changed genes, respectively). Other large constitutive expression differences have only been detected for gene #3, encoding a protein of unknown function, for which we find higher transcript abundances in the susceptible CM-NIL51.
Expression analysis of genes located in the Fhb1 region of CM-82036. a log2-transformed RNA-seq read counts for genes located in the region. Each subpanel comprises time-course-derived data (3–48 h) after inoculation with F. graminearum or mock for either the Fhb1-carrying CM-NIL38 or the susceptible CM-NIL51. Genes with no mapped reads are given in dark blue, highly expressed genes in red. b Differentially expressed genes in contrasts comparing F. graminearum-challenged to mock-treated samples in the resistant CM-NIL38 and the susceptible CM-NIL51 (left panels) and comparing mock-treated and F. graminearum-inoculated samples between the two NILs (right panels). Positive log2-transformed fold-change values indicate significant higher expression in response to the pathogen (left panels) or higher expression for the susceptible CM-NIL51 when comparing similarly treated samples between NILs (right panels). White spaces represent samples with no significant (FDR >0.05) differences (yet differences in read counts as indicated in a might occur) (color figure online)
Genes #10 and #11, both coding for unknown proteins, show a circadian-type expression with expression peaks at 3, 24, and 48 hai (representing morning hours) in the resistant CM-NIL38, while in the susceptible CM-NIL51 only gene 10 shows a similar expression pattern albeit to a much weaker extent (Fig. 2a). None of the genes in the susceptible NIL are significantly different expressed in response to the pathogen and only three show significant changes in expression in the resistant NIL (Fig. 2b): transcripts corresponding to gene #4 encoding a coatamer subunit domain-containing protein (G4) were 3.61-fold less abundant at 48 hai in the F. graminearum-challenged samples compared to mock, while at the same time point transcript abundances for gene #9, encoding an alanyl-tRNA synthase, were 2.25-fold increased in response to the pathogen. Striking differences were observed for gene #24 encoding a GDSL lipase (acylhydrolase): significantly higher transcript abundances in F. graminearum-treated samples were detected at 12, 36 and 48 hai and a 15.76-fold expression increase at 48 hai was observed.
Fine-mapping Fhb1
A large mapping population was developed to genetically narrow down the Fhb1 region and thereby reducing the number of Fhb1 candidates. Polymorphisms for marker design were detected utilizing mapped RNA-seq data from the susceptible Remus to the Fhb1 region of CM-82036. Using only SNPs unique to CM-NIL38-mapped reads with a frequency of >90 %, we detected 30 polymorphisms within predicted coding sequences, which comprise a conservative estimate of SNP in the region (Supplementary Table S6). We used these and SNP identified by mapping reads from a previous RNA-seq project using similar NILs (Kugler et al. 2013) to design six KASP genotyping assays. An additional KASP assay was constructed based on resequencing the PCR product of UMN10 from the susceptible and highly polymorphic locus of cv. Remus (Supplementary File Fig. S3). To design markers for regions not covered by RNA-seq due to the absence of genes in the susceptible parent, we generated four PCR assays that produce a dominant-type PCR amplicon from annotated genes found only in the Fhb1-region of CM-82036. All together 14 markers separated the 100 recombinant NILs in nine haplotypes, whereas nine of the markers co-segregated resulting in a 703.4 kb region, between contig positions 133.5 and 836.9 kb, with no recombination events found within (Fig. 3; Supplementary File Fig. S4, which includes the number of inoculated heads).
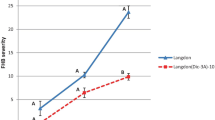
Graphical illustration of the nine haplotypes and control lines in the Fhb1 interval for 16 markers and their phenotypes for FHB and DON resistance. Resistant (CM-82036) alleles for the markers are illustrated by dashed boxes, whereas susceptible (Remus) alleles are shown in white. The FHB/DON resistance levels are measured in number of diseased/DON-bleached spikelets per head 26 days after F. graminearum-inoculation or DON infiltration. The dotted lines define the interval harboring Fhb1. Resistant (R) and susceptible (S) haplotypes significantly different from each other are indicated by the respective letters (FHB severity: mean LSD α = 5 % = 2.0, DON severity: mean LSD α = 5 % = 3.0)
Disease severity separated CM-82036 and the near-isogenic control lines into two phenotypic classes depending on the presence of Fhb1. The control lines, possessing Fhb1, developed disease symptoms on the two inoculated spikelets per spike only; no further spreading of disease symptoms was observed, whereas the NILs with susceptible alleles at Fhb1, CM-NIL47 and CM-NIL51, showed disease symptoms in 4.7 and 6.2 spikelets/head on average. The FHB severity of the highly susceptible cultivar Remus was three times higher as observed for the susceptible NILs.
Statistical analysis of the 100 recombinant NILs representing nine haplotypes detected three phenotypic classes: for the FHB-resistant class (haplotypes 3, 4, 5, 6, 7) just the two point-inoculated spikelets per head exhibited disease symptoms, as also observed for the resistant control lines. These lines have CM-82036 alleles for nine markers in common, from contig position 133.5–836.9 kb. The FHB susceptible haplotypes (1, 8, 9) were grouped in two phenotypic classes, with haplotype 9, being slightly more diseased with on average 7.2 symptomatic spikelets per spike compared to haplotypes 1 and 8 with 4.4 and 5.2 diseased spikelets. Haplotype 2, representing lines with recombinations between snp3BS-8 and IFA-FMG1 and CM-82036 alleles for snp3BS-8, but susceptible alleles at IFA-FMG1 exhibited an intermediate FHB resistance phenotype, being not significantly different from the resistant and the susceptible class. The mean of 3.6 symptomatic spikelets/spike rather points towards the susceptible class.
In addition, a subset of the recombinant NILs and the control lines were tested for DON resistance after application of the toxin DON in the head. The toxin treatment to the flowering heads induced typical FHB symptoms, straw-like color, spreading in both acropetal and basipetal directions, only in DON-sensitive lines (Supplementary File Fig. S5). DON severity (measured as number of DON-bleached spikelets per spike 26 days after treatment) divided all tested lines into two distinct phenotypic classes: the DON resistant class with almost no symptoms and the DON susceptible class with about ten DON-bleached spikelets per spike. The DON resistant class comprises haplotypes 3, 4, 6, and 7. The DON susceptible class with haplotypes 1, 2, 8 and 9 positioned Fhb1 in the same contig interval as obtained for FHB severity, with the flanking markers snp3BS-8 and IFA-FM958 mapping to the contig positions 91.9 and 958.3 kb, respectively. The separation of all lines, including control lines and the highly susceptible cultivar Remus into two distinct classes, identifies Fhb1 as the only gene for toxin resistance in CM-82036.
We could successfully connect the physical map of the Fhb1 region to the genetic map; the two markers, snp3BS-8 and IFA-FM958, place the causal gene(s) behind Fhb1 between the contig positions 91.9 and 958.3 kb, respectively. One or several genes in this region confer FHB and DON resistance.
Discussion
We have determined the genomic sequence of the wheat FHB resistance QTL Fhb1 in the resistant donor cultivar CM-82036 and positioned the gene within a region covering 860 kb using a fine-mapping panel for Fhb1 that has been phenotyped for both FHB and DON resistance. 28 candidate genes including 13 high confidence genes located in this interval have been further characterized in a dense time-course RNA-seq study.
Suppressed recombination in the highly divergent region of Fhb1
Our findings show that the sequence containing Fhb1 in CM-82036 differs significantly from the susceptible Chinese Spring reference in gene content and size. The core region of the sequenced contig is highly dissimilar to Chinese Spring when comparing genomic DNA; also markers designed for this region in CM-82036 failed to amplify in the susceptible region of cv. Remus. A dot plot analysis did not identify structural rearrangements such as genomic inversions or duplications (Fig. 1b). The in part large differences in distance between the genes present in resistant and susceptible cultivars may be attributed to different transposon insertions events (Scherrer et al. 2005). Most genes unique for Fhb1 reside in a compact cluster (genes #18–#27). The observed differences can be explained in part by pseudogenisation of genes #21, #26 and #27 in Chinese Spring where partial overlaps were still identifiable in non-annotated regions. However, multiple genes are unique for either the Fhb1 contig or Chinese Spring.
Haplotype divergences and loss of microcolinearity between cultivars on a comparable scale as detected for Fhb1 have been observed several times before (i.e., for the barley Rph7 locus (Scherrer et al. 2005) and wheat Lr10 (Isidore et al. 2005)) and also in two recent studies: Yeo et al. (2015) have resequenced the resistant and susceptible loci from two barley cultivars differing in the Puccinia hordei resistance gene Rphq2 and identified entirely different haplotypes of which the resistant locus harbors unique candidate genes. Mago et al. (2014) established the genomic region harboring the wheat stem rust resistance gene Sr2, which includes a cluster of germin-like proteins missing in the susceptible reference cultivar Chinese Spring. The authors showed that this cluster is shared between other resistant accessions containing Sr2.
In rice, more than 6 % of the genome is occupied by regions of high divergence (Tang et al. 2006). The higher genome plasticity of polyploid wheat led to a high rate of gene deletions and activity of repetitive elements buffered by the redundancies within the three homoeoallelic subgenomes (Dubcovsky and Dvorak 2007). Most likely the rate of highly divergent regions between cultivars is more frequent than generally assumed. Redundant gene content may also more easily accommodate introgressions of rare resistance haplotypes from landraces under selective pressure by replacing genes with redundant gene activity.
Highly diverging haplotypes generally lead to strongly reduced meiotic recombinations in such regions. In the Fhb1 locus, loss of colinearity has a direct effect on recombination frequency. While flanking recombining regions harbor 4 and 6 recombinations in an interval of 42.6 and 122 kb, respectively (Fig. 3) about 700 kb remain unresolved. On a larger scale, the locus itself resides in a highly recombining telomeric region of chromosome 3B with an average of 0.85 cM/Mb (Saintenac et al. 2009), which would relate to 17 expected recombinations within the Fhb1 non-recombining region. Tracking recombinants from two mapping populations of which neither parent harbored Fhb1 the same authors detected a recombination hotspot that covers the region around UMN10 in one of their populations leading to high recombination rates, while only a below average rate was observed in the second population (Saintenac et al. 2011). Apparently, cross-over hotspots exist in this region, but these need to be met with matching crossing partners.
As CM-82036 is a direct derivative from a cross of the Fhb1 donor Sumai-3, the sequence obtained from this cultivar should be directly comparable to the sequence of Sumai-3, which has been target of several fine-mapping studies (Cuthbert et al. 2006; Liu et al. 2008; Bernardo et al. 2011). Among the markers used in both studies, the widely used UMN10 is the only marker we found mapping within the non-colinear region, while all others mapped to flanking regions. The original UMN10 from Liu et al. (2008) detects a length polymorphism. This marker has been converted into an easily applicable KASP-SNP assay, which has been successfully employed on diverse germplasm conducted by several groups so far and should remain a reliable marker for broader use.
The closest reported recombination event is based on a single recombinant line for marker sts32 reported by Liu et al. (2008) which mapped to position 155.4 kb on the Fhb1 contig. Consideration of this single event would exclude genes #1–#10 from the list of Fhb1 candidates. To further substantiate these findings, we identified 100 recombinant lines in our own fine-mapping panel for the gwm493 and barc133 interval. Yet, we failed to identify additional recombinant lines that would reach as far or further in the ‘core’ region of Fhb1.
Candidate genes in the Fhb1 locus
Fhb1 expresses a dominant phenotype (Xie et al. 2007 and own unpublished data). Consequently, possible explanations for the phenotypic difference could be induced expression of the underlying gene in response to the pathogen or constitutive expression in the resistant genotype, or absence of the respective gene in susceptible lines as Chinese Spring and Remus. Furthermore, gain of function polymorphisms through changes in protein sequence may cause the resistance phenotype. Also susceptibility factors encoded in the interval in lines lacking Fhb1 need to be considered, although such a scenario is more difficult to reconcile with the reported dominance. Our results also show that DON resistance (determined as bleaching resistance after application of high concentrations of pure toxin) is clearly associated with Fhb1. Either DON resistance itself could simultaneously lead to FHB resistance, or the gene causing DON resistance might be tightly linked to the gene conferring FHB resistance by a different mechanism. The fact that increased F. graminearum resistance was achieved by increasing DON resistance [due to overexpression of a barley glucosyltransferase (Li et al. 2015)] suggests that genes with an effect on toxin resistance should be considered as prime candidate Fhb1 genes. With the sequenced region at hand and mapped expression data, these scenarios can now be considered much better; albeit lacking a higher resolved map still many candidate genes remain: We discuss putative functions and expression patterns of the candidate genes and the implications for functional testing.
Gene #17 (ubiquitin-2 like Rad60 SUMO-like protein) is unique for the CM-82036 sequence, deleted in the susceptible cultivar (see Fig. 1c). In yeast, it has been shown that reducing the ubiquitin pool by disruption of the stress responsive polyubiquitin gene leads to reduced DON resistance of ubi4 mutants (Abolmaali et al. 2008). Yet, gene #17 is practically not expressed, neither in the control nor following F. graminearum-inoculation. Similarly, genes #21 (general transcription factor IIE subunit) and #25 (cystatin) are not present in the susceptible Chinese Spring reference, but also not expressed under both conditions. We, therefore, consider them unlikely candidates for Fhb1.
All other genes within the diverging region are expressed in the Fhb1-containing CM-NIL38 but not in the susceptible CM-NIL51. Expression levels range from few reads per sequenced sample to hundreds of reads per sample. While highly expressed genes may present themselves as ‘more-likely’ candidates, comparably lowly expressed genes should not be ruled out as RNA levels may not directly reflect protein expression levels and the abundances to establish a specific function may be vastly different for individual gene products.
Gene #19 (terpene synthase), gene #20 (unknown protein) and gene #23 (E3-Ubiquitin ligase) are also present in Chinese Spring but they are only expressed in Fhb1 containing lines. Terpene synthases act in the biosynthesis of secondary metabolites, which play a role in defense against herbivores or pathogens (Lange 2015). Many terpenoid phytoalexins from Poaceae have been described (Ejike et al. 2013). These include phytoalexins derived from monoterpenes and sesquiterpenes, which have a direct antimicrobial effect (Schmelz et al. 2011; Inoue et al. 2013). Sesquiterpenoid phytoalexins active against F. graminearum have for instance been described in maize (Huffaker et al. 2011). Volatile terpenes also may act as messengers upon pathogen attack (Nagegowda 2010). The Fhb1-associated terpene synthase most likely acts in synthesis of cytosolic sesquiterpenoids from farnesyl diphosphate as a BLASTp result suggests (delta-cadinene synthase isozyme A, e value = 0). However, the overall low expression level of the terpene synthase encoded on Fhb1, with no observable differences between F. graminearum treatment and mock, suggests that this gene does not play an active role in the response to the pathogen.
A secondary annotation for gene #20 (unknown protein) suggests a role in calcium sensing (sarcoplasmic reticulum histidine-rich calcium-binding protein precursor, blastp, e value = 8e−38) and, consequently, may lead to changes in gene expression following external cues such as abiotic and biotic stresses (Reddy et al. 2011). Yet, its strong constitutive expression suggests a different role for this gene. The lack of expression in the susceptible NIL is not due to pseudogenisation of the CS ortholog. Both gene models seem intact and share high amino acid sequence similarity (95.7 %, Supplementary File Fig. S6). In contrast, the promotor regions are highly divergent in the first 1 kb upstream of the short 5’UTR region with multiple sequence deletions in the Fhb1-contig, which most likely are responsible for the observed large expression differences. Despite the lack of clear indications about its potential mode of action, gene #20 should be considered a candidate for Fhb1.
Gene #23 is a predicted E3-ubiquitin ligase of the ‘seven in absentia’ (SINA) type. Such proteins mediate ubiquitination and proteasome-mediated degradation of specific proteins (in response to a stimulus). Some SINA proteins and their client proteins have been implicated in plant–pathogen and plant–symbiont interactions (Kim et al. 2006; Den Herder et al. 2012). They contain an N-terminal RING domain and a C-terminal conserved domain implicated in dimerization and substrate binding. Interestingly, the gene models for this protein differ largely due to an internal deletion of 31 nt in the Fhb1 reading frame compared to the gene model in Chinese Spring. The consequence is a frameshift and premature stop codon, removing the entire SINA domain. The Fhb1-resistant line, therefore, possesses most likely a nonfunctional version of the protein. Yet, the expression is higher in the Fhb1 background, so potentially the truncated protein might act in a dominant negative fashion, so that it may be premature to exclude this gene as a candidate.
The Fhb1 region hosts four clearly expressed genes absent in the susceptible reference
Gene #22 encodes protein with domains encoding agglutinin and ‘pore-forming toxin-like’. This weakly expressed gene might have direct antifungal activity by binding to fungal cell wall carbohydrate structures and permeating membranes. The role of lectins in plant defense is well established (Lannoo and Van Damme 2014). Wheat germ agglutinin has been shown to bind to N-acetyl-d-glucosamine (Levy 1979), a monomer of the fungal cell wall chitin and as such constitutes a pathogen recognition mechanism that elicits further, early defense responses. Wheat germ agglutinin exhibits also a negative effect on hyphal growth of various fungi including F. graminearum (Mirelman et al. 1975; Ciopraga et al. 1999). This proposed mechanism is, however, more consistent with type I than type II resistance against spreading of the disease and resistance to DON. Expression of the gene #22 cDNA in Saccharomyces cerevisiae under control of the inducible GAL1 promoter did not affect the growth of the transformed yeast strain on galactose medium (data not shown).
Gene #24 encoding a GDSL lipase is the only gene in the sequenced contig that exhibits a significant increase in expression in response to the pathogen. GDSL lipase/esterases comprise a structurally diverse gene family in plants. For instance, 114 members exist in the rice genome (Chepyshko et al. 2012). They act in regulation of a variety of physiological functions including defense. A chain of studies (Kwon et al. 2009; Kim et al. 2014) demonstrated the role of an Arabidopsis thaliana GDSL lipase 1 in modulating systemic immunity through the regulation of ethylene signaling in response to necrotrophic pathogens. However, the present GDSL lipase does not share similarity with the A. thaliana lipase 1 gene. The expression pattern and its possible role in defense warrant further investigations.
Gene #26 (F-box protein) is among the strongest constitutively expressed genes on the contig. No similar gene is annotated in this region of Chinese Spring, yet mapping of the coding sequence of this gene onto Chinese Spring identified a likely pseudogene with weak similarity (Table 1). F-box proteins are part of the ubiquitination complex, which form specific interaction with target proteins. Consequently, the gene family is very large with 779 genes in rice (Xu et al. 2009). The F-box protein could be involved in reducing the levels of protein encoding a susceptible factor for FHB. Its target protein, which is most likely not encoded in the Fhb1 region, would need to be genetically fixed and must not segregate, to be in agreement with the absence of epistasis at Fhb1. Potentially, the F-box protein could also directly target an unknown effector protein of the pathogen. In A. thaliana, the F-box protein encoded by COI1 is involved in jasmonate signaling and is the target of the jasmonic acid mimicking bacterial toxin coronatine which increases susceptibility (Geng et al. 2012).
Also gene #27 (hypothetical protein) cannot be excluded as gene candidate. However, only few reads map to the predicted CDS of this low confidence gene for which no annotation could be retrieved.
The genes on the right half of the Fhb1 interval (Fig. 1c) have again counterparts in the susceptible line. The genes #28 and #29 are constitutively expressed and are discussed below. Gene #30 encodes a predicted zinc finger C3H4 type (RING finger) domain-containing protein showing low expression in both NILs, and no response to F. graminearum infection. Zinc finger-containing proteins have functions ranging from transcription, translation, mRNA trafficking, cytoskeleton organization, protein folding, chromatin remodeling and more. Only a domain of unknown function (DUF3675) is additionally recognized. But since the gene model is identical with that of Chinese Spring, this gene showing no significant expression difference between NILs and in response to F. graminearum can be excluded.
Also genes #31 and #32 are unlikely candidates due to lacking expression. A CYP450 gene could encode an enzyme involved in the biosynthesis of an antifungal metabolite, or a detoxification enzyme leading to chemical modification of the toxin structure. A bacterial cytochrome P450 detoxifying DON by hydroxylation of C16 has been described (Ito et al. 2013). The product of gene #32 contains an NB-ARC domain, which is found in plant disease resistance genes (van der Biezen and Jones 1998). Besides the nucleotide binding domain, also leucine-rich repeats can be recognized. A highly similar protein from Aegilops tauschii has been annotated as ‘putative disease resistance RPP13-like protein 1’ (GenBank accession: EMT27135.1). The version of the susceptible Chinese spring gene is identical in 898 of 905 amino-acids, leaving room for functional differences (Supplementary File Fig. S7). Yet, lack of expression is hard to reconcile with the otherwise suggestive role of this candidate disease resistance gene.
How can genes on the Fhb1 contig help explain the higher ability to inactivate DON?
Lemmens et al. (2005) have associated the Fhb1 locus with the higher ability to metabolize DON into the non-toxic DON-3-O-glycoside, which is a product of the activity of toxin-specific UDP-glucosyltransferases (UGT, Poppenberger et al. 2003). No such gene is encoded on the Fhb1 contig, gene #6 annotated as a HGA-like UGT does share similarities to the large super family encoding small molecule conjugating UGTs (Ross et al. 2001), but most likely acts on the formation of homogalacturonan (HGA) as part of the cell wall (Yin et al. 2010). Toxicity of DON is caused by inhibition of protein biosynthesis; therefore, genes involved in translation may counteract the adverse effect of DON by increasing overall translation fidelity or exerting a greater tolerance to DON in other ways. Genes #7 and #9 encode leucyl- and alanyl-tRNA synthases, respectively. While gene #7 shows no significant expression difference, gene #9 is clearly higher expressed in the Fhb1-containing NIL (Fig. 2) and is, therefore, more attractive. In addition, potentially relevant sequence differences exist (Supplementary File Fig. S8). Recently, it has been shown that overexpression of a methionyl-tRNA synthase from wheat when overexpressed in A. thaliana causes increased DON resistance in transformants (Zuo et al. 2016). Yet, as stated above, if the reported single recombinant line at sts32 (Liu et al. 2008) is indeed correct, all genes up to #10 can be ruled out as candidates. Gene #13 (tRNA-modifying methyltransferase) has a ribosome-associated function, likewise #28 (translation initiation factor). A possible role of this gene for methylation-associated resistance of ribosomes against trichothecene toxins has been proposed by Iglesias and Ballesta (1994), who found that in Fusarium oxysporum adaptive toxin resistance of ribosomes can be obtained by enzymatic modification of an unknown ribosomal component upon incubation with S-adenosylmethionin. Despite its low expression in the Fhb1-containing NIL, this gene should not be ruled out as candidate. Also #29 has a predicted methyltransferase domain.
With the sequence of Fhb1 at hand the genes described in this study are a valuable resource for further functional analysis of the QTL. Based on expression profiles and annotations, some genes can be ruled out, but many remain for which further functional assessments are required. The knowledge of which gene is causing FHB resistance is not irrelevant, as breeders unknowingly may deploy proteins with potentially undesired health effects (lectin/pore-forming toxin) or increase the levels of antifungal compounds with unknown toxicological properties (terpenoid synthase). The most promising approach to further characterize Fhb1 is the characterization of EMS-generated stable loss-of-function mutants (Slade and Knauf 2005) for which polyploid wheat is especially well suited due to the high possible mutation rates. RNA-interference methods such as VIGS may not yield clear phenotypes for targeted candidate genes as the silencing is only partial and transient. This residual expression levels bear the risk of providing sufficiently high mRNA levels to produce relevant amounts of protein to confer the resistance phenotype. Generating stable wheat transformants in a type 2 susceptible cultivar is a viable alternative to assess candidates for FHB resistance (Li et al. 2015) and should bring clarity about the gene underlying FHB and DON resistance. While the simplest hypothesis is that only one gene is causing both phenotypes, also the scenario of two different resistance genes cosegregating due to repressed recombination in the region cannot be excluded.
Author contribution statement
Generation of plant material: GS, BS. BAC library construction, screening and sequencing: SV, WS. Contig assembly and annotation: WS. Greenhouse trials: BS, FJ, MZ. Toxin and strain provision: ML. Marker design and genotyping: BS, VG, FJ, WS. RNAseq data acquisition and analysis: MZ, WS, TM. HB (BOKU), KFXM, GA and HB (INRA) conceived this study and obtained funding. Manuscript writing: WS, BS and GA. The manuscript was finally approved by all coauthors.
References
Abolmaali S, Mitterbauer R, Spadiut O et al (2008) Engineered bakers yeast as a sensitive bioassay indicator organism for the trichothecene toxin deoxynivalenol. J Microbiol Methods 72:306–312. doi:10.1016/j.mimet.2007.12.013
Anders S, Pyl PT, Huber W (2015) HTSeq—a Python framework to work with high-throughput sequencing data. Bioinformatics 15:166–169. doi:10.1093/bioinformatics/btu638
Anderson J, Stack R, Liu S, Waldron B (2001) DNA markers for Fusarium head blight resistance QTLs in two wheat populations. Theor Appl Genet 102:1164–1168. doi:10.1007/s001220000509
Bashir A, Klammer AA, Robins WP et al (2012) A hybrid approach for the automated finishing of bacterial genomes. Nat Biotechnol 30:701–707. doi:10.1038/nbt.2288
Bernardo AN, Ma H, Zhang D, Bai G (2011) Single nucleotide polymorphism in wheat chromosome region harboring Fhb1 for Fusarium head blight resistance. Mol Breed 29:477–488. doi:10.1007/s11032-011-9565-y
Buerstmayr H, Lemmens M, Hartl L et al (2002) Molecular mapping of QTLs for Fusarium head blight resistance in spring wheat. I. Resistance to fungal spread (type II resistance). Theor Appl Genet 104:84–91. doi:10.1007/s001220200009
Buerstmayr H, Steiner B, Hartl L et al (2003) Molecular mapping of QTLs for Fusarium head blight resistance in spring wheat. II. Resistance to fungal penetration and spread. Theor Appl Genet 107:503–508. doi:10.1007/s00122-003-1272-6
Buerstmayr H, Ban T, Anderson JA (2009) QTL mapping and marker-assisted selection for Fusarium head blight resistance in wheat: a review. Plant Breed 128:1–26. doi:10.1111/j.1439-0523.2008.01550.x
Chepyshko H, Lai C-P, Huang L-M et al (2012) Multifunctionality and diversity of GDSL esterase/lipase gene family in rice (Oryza sativa L. japonica) genome: new insights from bioinformatics analysis. BMC Genom 13:309. doi:10.1186/1471-2164-13-309
Choulet F, Wicker T, Rustenholz C et al (2010) Megabase level sequencing reveals contrasted organization and evolution patterns of the wheat gene and transposable element spaces. Plant Cell 22:1686–1701. doi:10.1105/tpc.110.074187
Ciopraga J, Gozia O, Tudor R et al (1999) Fusarium sp. growth inhibition by wheat germ agglutinin. Biochim Biophys Acta 1428:424–432. doi:10.1016/S0304-4165(99)00085-9
Cuthbert PA, Somers DJ, Thomas J et al (2006) Fine mapping Fhb1, a major gene controlling fusarium head blight resistance in bread wheat (Triticum aestivum L.). Theor Appl Genet 112:1465–1472. doi:10.1007/s00122-006-0249-7
Den Herder G, Yoshida S, Antolín-Llovera M et al (2012) Lotus japonicus E3 ligase SEVEN IN ABSENTIA4 destabilizes the symbiosis receptor-like kinase SYMRK and negatively regulates rhizobial infection. Plant Cell 24:1691–1707. doi:10.1105/tpc.110.082248
Dubcovsky J, Dvorak J (2007) Genome plasticity a key factor in the success of polyploid wheat under domestication. Science 316:1862–1866. doi:10.1126/science.1143986
Ejike CECC, Gong M, Udenigwe CC (2013) Phytoalexins from the Poaceae: biosynthesis, function and prospects in food preservation. Food Res Int 52:167–177. doi:10.1016/j.foodres.2013.03.012
English AC, Richards S, Han Y et al (2012) Mind the gap: upgrading genomes with Pacific Biosciences RS long-read sequencing technology. PLoS One 7:e47768. doi:10.1371/journal.pone.0047768
Geng X, Cheng J, Gangadharan A, Mackey D (2012) The coronatine toxin of Pseudomonas syringae is a multifunctional suppressor of Arabidopsis defense. Plant Cell 24:4763–4774. doi:10.1105/tpc.112.105312
Gunnaiah R, Kushalappa AC, Duggavathi R et al (2012) Integrated metabolo-proteomic approach to decipher the mechanisms by which wheat QTL (Fhb1) contributes to resistance against Fusarium graminearum. PLoS One 7:e40695. doi:10.1371/journal.pone.0040695
Hao C, Wang Y, Hou J et al (2012) Association mapping and haplotype analysis of a 3.1-Mb genomic region involved in Fusarium head blight resistance on wheat chromosome 3BS. PLoS One 7:e46444. doi:10.1371/journal.pone.0046444
Hofstad AN, Nussbaumer T, Akhunov E et al (2016) Examining the transcriptional response in wheat near-isogenic lines to infection and deoxynivalenol treatment. Plant Genome 9:1–15. doi:10.3835/plantgenome2015.05.0032
Huffaker A, Kaplan F, Vaughan MM et al (2011) Novel acidic sesquiterpenoids constitute a dominant class of pathogen-induced phytoalexins in maize. Plant Physiol 156:2082–2097. doi:10.1104/pp.111.179457
Iglesias M, Ballesta JP (1994) Mechanism of resistance to the antibiotic trichothecin in the producing fungi. Eur J Biochem 223:447–453. doi:10.1111/j.1432-1033.1994.tb19012.x
Inoue H, Hayashi N, Matsushita A et al (2013) Blast resistance of CC-NB-LRR protein Pb1 is mediated by WRKY45 through protein-protein interaction. Proc Natl Acad Sci USA 110:9577–9582. doi:10.1073/pnas.1222155110
Isidore E, Scherrer B, Chalhoub B et al (2005) Ancient haplotypes resulting from extensive molecular rearrangements in the wheat A genome have been maintained in species of three different ploidy levels. Genome Res 15:526–536. doi:10.1101/gr.3131005
Ito M, Sato I, Ishizaka M et al (2013) Bacterial cytochrome P450 system catabolizing the Fusarium toxin deoxynivalenol. Appl Environ Microbiol 79:1619–1628. doi:10.1128/AEM.03227-12
Jansen C, von Wettstein D, Schafer W et al (2005) Infection patterns in barley and wheat spikes inoculated with wild-type and trichodiene synthase gene disrupted Fusarium graminearum. Proc Natl Acad Sci USA 102:16892–16897. doi:10.1073/pnas.0508467102
Kearse M, Moir R, Wilson A et al (2012) Geneious basic: an integrated and extendable desktop software platform for the organization and analysis of sequence data. Bioinformatics 28:1647–1649. doi:10.1093/bioinformatics/bts199
Kim Y-S, Ham B-K, Paek K-H et al (2006) An Arabidopsis homologue of human seven-in-absentia-interacting protein is involved in pathogen resistance. Mol Cells 21:389–394. doi:10.1109/TCAD.2005.853689
Kim HG, Kwon SJ, Jang YJ et al (2014) GDSL lipase 1 regulates ethylene signaling and ethylene-associated systemic immunity in Arabidopsis. FEBS Lett 588:1652–1658. doi:10.1016/j.febslet.2014.02.062
Kugler KG, Siegwart G, Nussbaumer T et al (2013) Quantitative trait loci-dependent analysis of a gene co-expression network associated with Fusarium head blight resistance in bread wheat (Triticum aestivum L.). BMC Genom 14:728. doi:10.1186/1471-2164-14-728
Kwon SJ, Jin HC, Lee S et al (2009) GDSL lipase-like 1 regulates systemic resistance associated with ethylene signaling in Arabidopsis. Plant J 58:235–245. doi:10.1111/j.1365-313X.2008.03772.x
Lange BM (2015) The evolution of plant secretory structures and emergence of terpenoid chemical diversity. Annu Rev Plant Biol 66:139–159. doi:10.1146/annurev-arplant-043014-114639
Lannoo N, Van Damme EJM (2014) Lectin domains at the frontiers of plant defense. Front Plant Sci 5:397. doi:10.3389/fpls.2014.00397
Lemmens M, Scholz U, Berthiller F et al (2005) The ability to detoxify the mycotoxin deoxynivalenol colocalizes with a major quantitative trait locus for Fusarium head blight resistance in wheat. Mol Plant-Microbe Interact 18:1318–1324. doi:10.1094/MPMI-18-1318
Leroy P, Guilhot N, Sakai H et al (2012) TriAnnot: a versatile and high performance pipeline for the automated annotation of plant genomes. Front Plant Sci 3:5. doi:10.3389/fpls.2012.00005
Levy NJ (1979) Wheat germ agglutinin blockage of chlamydial attachment sites: antagonism by N-acetyl-d-glucosamine. Infect Immun 25:946–953
Li X, Shin S, Heinen S et al (2015) Transgenic wheat expressing a barley UDP-glucosyltransferase detoxifies deoxynivalenol and provides high levels of resistance to Fusarium graminearum. Mol Plant Microbe Interact 28:1237–1246. doi:10.1094/MPMI-03-15-0062-R
Liu S, Zhang X, Pumphrey MO et al (2006) Complex microcolinearity among wheat, rice, and barley revealed by fine mapping of the genomic region harboring a major QTL for resistance to Fusarium head blight in wheat. Funct Integr Genomics 6:83–89. doi:10.1007/s10142-005-0007-y
Liu S, Pumphrey MO, Gill BS et al (2008) Toward positional cloning of Fhb1, a major QTL for Fusarium head blight resistance in wheat. Cereal Res Commun 36:195–201. doi:10.1556/CRC.36.2008.Suppl.B.15
Mago R, Tabe L, Vautrin S et al (2014) Major haplotype divergence including multiple germin-like protein genes, at the wheat Sr2 adult plant stem rust resistance locus. BMC Plant Biol 14:1–11. doi:10.1186/s12870-014-0379-z
Mayer KFX, Rogers J, el Dole J et al (2014) A chromosome-based draft sequence of the hexaploid bread wheat (Triticum aestivum) genome. Science 345:1251788-1–1251788-11. doi:10.1126/science.1251788
Mirelman D, Galun E, Sharon N, Lotar R (1975) Inhibition of fungal growth by wheat germ agglutinin. Nature 256:414–416. doi:10.1038/256414a0
Nagegowda DA (2010) Plant volatile terpenoid metabolism: biosynthetic genes, transcriptional regulation and subcellular compartmentation. FEBS Lett 584:2965–2973. doi:10.1016/j.febslet.2010.05.045
Nussbaumer T, Warth B, Sharma S et al (2015) Joint transcriptomic and metabolomic analyses reveal changes in the primary metabolism and imbalances in the subgenome orchestration in the bread wheat molecular response to Fusarium graminearum. G3 (Bethesda) 5:2579–2592. doi:10.1534/g3.115.021550
Pestka JJ (2010) Deoxynivalenol: mechanisms of action, human exposure, and toxicological relevance. Arch Toxicol 84:663–679. doi:10.1007/s00204-010-0579-8
Peterson DG, Tomkins JP, Frisch DA et al (2000) Construction of plant bacterial artificial chromosome (BAC) libraries: an illustrated guide. J Agric Genomics 5:1–91
Poppenberger B, Berthiller F, Lucyshyn D et al (2003) Detoxification of the Fusarium mycotoxin deoxynivalenol by a UDP-glucosyltransferase from Arabidopsis thaliana. J Biol Chem 278:47905–47914. doi:10.1074/jbc.M307552200
Reddy ASN, Ali GS, Celesnik H, Day IS (2011) Coping with stresses: roles of calcium- and calcium/calmodulin-regulated gene expression. Plant Cell 23:2010–2032. doi:10.1105/tpc.111.084988
Robinson MD, McCarthy DJ, Smyth GK (2010) edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26:139–140. doi:10.1093/bioinformatics/btp616
Ross J, Li Y, Lim EK, Bowles DJ (2001) Higher plant glycosyltransferases. Genome Biol 2 2(2): reviews3004.1–reviews3004.6
Saintenac C, Falque M, Martin OC et al (2009) Detailed recombination studies along chromosome 3B provide new insights on crossover distribution in wheat (Triticum aestivum L.). Genetics 181:393–403. doi:10.1534/genetics.108.097469
Saintenac C, Faure S, Remay A et al (2011) Variation in crossover rates across a 3-Mb contig of bread wheat (Triticum aestivum) reveals the presence of a meiotic recombination hotspot. Chromosoma 120:185–198. doi:10.1007/s00412-010-0302-9
Scherrer B, Isidore E, Klein P et al (2005) Large intraspecific haplotype variability at the Rph7 locus results from rapid and recent divergence in the barley genome. Plant Cell 17:361–374. doi:10.1105/tpc.104.028225
Schmelz EA, Kaplan F, Huffaker A et al (2011) Identity, regulation, and activity of inducible diterpenoid phytoalexins in maize. Proc Natl Acad Sci USA 108:5455–5460. doi:10.1073/pnas.1014714108
Schweiger W, Boddu J, Shin S et al (2010) Validation of a candidate deoxynivalenol-inactivating UDP-glucosyltransferase from barley by heterologous expression in yeast. Mol Plant-Microbe Interact 23:977–986. doi:10.1094/MPMI-23-7-0977
Schweiger W, Pasquet JC, Nussbaumer T et al (2013) Functional characterization of two clusters of Brachypodium distachyon UDP-glycosyltransferases encoding putative deoxynivalenol detoxification genes. Mol Plant Microbe Interact. doi:10.1094/MPMI-08-12-0205-R
Slade AJ, Knauf VC (2005) TILLING moves beyond functional genomics into crop improvement. Transgenic Res 14:109–115. doi:10.1007/s11248-005-2770-x
Steiner B, Kurz H, Lemmens M, Buerstmayr H (2009) Differential gene expression of related wheat lines with contrasting levels of head blight resistance after Fusarium graminearum inoculation. Theor Appl Genet 118:753–764. doi:10.1007/s00122-008-0935-8
Tang T, Lu J, Huang J et al (2006) Genomic variation in rice: genesis of highly polymorphic linkage blocks during domestication. PLoS Genet 2:e199. doi:10.1371/journal.pgen.0020199
The European Commission (2006) Commission recommendation (EC) no. 1881/2006 of 19 December 2006 setting maximum levels for certain contaminants in foodstuffs. Off J Eur Union Legis 364:5–24
Trapnell C, Roberts A, Goff L et al (2012) Differential gene and transcript expression analysis of RNA-seq experiments with TopHat and Cufflinks. Nat Protoc 7:562–578. doi:10.1038/nprot.2012.016
van der Biezen EA, Jones JD (1998) The NB-ARC domain: a novel signalling motif shared by plant resistance gene products and regulators of cell death in animals. Curr Biol 26:R226–R227. doi:10.1016/S0960-9822(98)70145-9
Waldron BL, Moreno-Sevilla B, Anderson JA et al (1999) RFLP mapping of QTL for Fusarium head blight resistance in wheat. Crop Sci 39:805–811. doi:10.2135/cropsci1999.0011183X003900030032x
Walter S, Brennan JM, Arunachalam C et al (2008) Components of the gene network associated with genotype-dependent response of wheat to the Fusarium mycotoxin deoxynivalenol. Funct Integr Genomics 8:421–427. doi:10.1007/s10142-008-0089-4
Xiao J, Jin X, Jia X et al (2013) Transcriptome-based discovery of pathways and genes related to resistance against Fusarium head blight in wheat landrace Wangshuibai. BMC Genom 14:197. doi:10.1186/1471-2164-14-197
Xie GQ, Zhang MC, Chakraborty S, Liu CJ (2007) The effect of 3BS locus of Sumai 3 on Fusarium head blight resistance in Australian wheats. Aust J Exp Agric 47:603–607. doi:10.1071/EA05250
Xu G, Ma H, Nei M, Kong H (2009) Evolution of F-box genes in plants: different modes of sequence divergence and their relationships with functional diversification. Proc Natl Acad USA 106:835–840. doi:10.1073/pnas.0812043106
Yeo FKS, Wang Y, Vozabova T et al (2015) Haplotype divergence and multiple candidate genes at Rphq2, a partial resistance QTL of barley to Puccinia hordei. Theor Appl Genet 289–304. doi:10.1007/s00122-015-2627-5
Yin Y, Chen H, Hahn MG et al (2010) Evolution and function of the plant cell wall synthesis-related glycosyltransferase family 8. Plant Physiol 153:1729–1746. doi:10.1104/pp.110.154229
Zhuang Y, Gala A, Yen Y (2013) Identification of functional genic components of major Fusarium head blight resistance quantitative trait loci in wheat cultivar sumai 3. Mol Plant Microbe Interact 26:442–450. doi:10.1094/MPMI-10-12-0235-R
Zuo D, Yi S, Liu R et al (2016) A deoxynivalenol-activated methionyl-tRNA synthetase gene from wheat encodes a nuclear localized protein and protects plants against Fusarium pathogens and mycotoxins. Phytopathology. doi:10.1094/PHYTO-12-15-0327-R. (epub ahead of print)
Acknowledgments
Open access funding provided by University of Natural Resources and Life Sciences Vienna (BOKU). We would like to acknowledge S. Cauet, G. Beydon and N. Rodde for assisting in BAC library screening and sequencing in Toulouse, France. A. Müllner, D. Fink and S. Zimmerl have assisted in marker development and fine mapping in Tulln, Austria. C. Ametz is acknowledged for bioinformatical support. This work was financed by the Austrian Science Fund (FWF) (SFB F3705, F3708, F3711) and the Deutsche Forschungsgemeinschaft (DGF) project LAP3714.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Additional information
Communicated by H.-Q. Ling.
W. Schweiger and B. Steiner share first authorship.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Schweiger, W., Steiner, B., Vautrin, S. et al. Suppressed recombination and unique candidate genes in the divergent haplotype encoding Fhb1, a major Fusarium head blight resistance locus in wheat. Theor Appl Genet 129, 1607–1623 (2016). https://doi.org/10.1007/s00122-016-2727-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00122-016-2727-x