Abstract
In bread wheat (Triticum aestivum L.), initial studies using deletion lines indicated that crossover (CO) events occur mainly in the telomeric regions of the chromosomes with a possible correlation with the presence of genes. However, little is known about the distribution of COs at the sequence level. To investigate this, we studied in detail the pattern of COs along a contig of 3.110 Mb using two F2 segregating populations (Chinese Spring × Renan (F2-CsRe) and Chinese Spring × Courtot (F2-CsCt)) each containing ~2,000 individuals. The availability of the sequence of the contig from Cs enabled the development of 318 markers among which 23 co-dominant polymorphic markers (11 SSRs and 12 SNPs) were selected for CO distribution analyses. The distribution of CO events was not homogeneous throughout the contig, ranging from 0.05 to 2.77 cM/Mb, but was conserved between the two populations despite very different contig recombination rate averages (0.82 cM/Mb in F2-CsRe vs 0.35 cM/Mb in F2-CsCt). The CO frequency was correlated with the percentage of coding sequence in Cs and with the polymorphism rate between Cs and Re or Ct in both populations, indicating an impact of these two factors on CO distribution. At a finer scale, COs were found in a region covering 2.38 kb, spanning a gene coding for a glycosyl transferase (Hga3), suggesting the presence of a CO hotspot. A non-crossover event covering at least 453 bp was also identified in the same interval. From these results, we can conclude that gene content could be one of the factors driving recombination in bread wheat.






Similar content being viewed by others
References
Akhunov ED, Akhunova AR, Linkiewicz AM, Dubcovsky J, Hummel D et al (2003a) Synteny perturbations between wheat homoeologous chromosomes caused by locus duplications and deletions correlate with recombination rates. Proc Natl Acad Sci USA 100:10836–10841
Akhunov ED, Goodyear AW, Geng S, Qi LL, Echalier B et al (2003b) The organization and rate of evolution of wheat genomes are correlated with recombination rates along chromosome arms. Genome Res 13:753–763
Anderson LK, Doyle GG, Brigham B, Carter J, Hooker KD et al (2003) High-resolution crossover maps for each bivalent of Zea mays using recombination nodules. Genetics 165:849–865
Arnheim N, Calabrese P, Tiemann-Boege I (2007) Mammalian meiotic recombination hot spots. Annu Rev Genet 41:369–399
Balfourier F, Roussel V, Strelchenko P, Exbrayat-Vinson F, Sourdille P et al (2007) A worldwide bread wheat core collection arrayed in a 384-well plate. Theor Appl Genet 114:1265–1275
Boeuf C, Prodanovic S, Gay G, Bernard M (2003) Structural organization of the group-1 chromosomes of two bread wheat sister lines. Theor Appl Genet 106:938–946
Borde V, Robine N, Lin W, Bonfils S, Geli V, Nicolas A (2009) Histone H3 lysine 4 trimethylation marks meiotic recombination initiation sites. EMBO J 28:99–111
Buard J, de Massy B (2007) Playing hide and seek with mammalian meiotic crossover hotspots. Trends Genet 23:301–309
Buard J, Barthès P, Grey C, de Massy B (2009) Distinct histone modifications define initiation and repair of meiotic recombination in the mouse. EMBO J 28:2616–2624
Buckler ES, Gaut BS, McMullen MD (2006) Molecular and functional diversity of maize. Curr Opin Plant Biol 9:172–176
Bulgarelli D, Collins NC, Tacconi G, Dellaglio E, Brueggeman R et al (2004) High-resolution genetic mapping of the leaf stripe resistance gene Rdg2a in barley. Theor Appl Genet 108:1401–1408
Chao S, Sharp PJ, Worland AJ, Warham EJ, Koebner RMD, Gale MD (1989) RFLP-based genetic maps of wheat homoeologous group 7 chromosomes. Theor Appl Genet 78:495–504
Choulet F, Wicker T, Rustenholz C, Paux E, Salse J et al (2010) Contrasted patterns of organization and evolution of the wheat gene and transposable element spaces revealed by megabase level genome sequencing. Plant Cell 22:1686–1701
Colas I, Shaw P, Prieto P, Wanous M, Spielmeyer W et al (2008) Effective chromosome pairing requires chromatin remodeling at the onset of meiosis. Proc Natl Acad Sci USA 105:6075–6080
De Massy B (2003) Distribution of meiotic recombination sites. Trends Genet 19:514–522
Dooner HK (2002) Extensive interallelic polymorphisms drive meiotic recombination into a crossover pathway. Plant Cell 14:1173–1183
Dooner HK, Martinez-Ferez IM (1997) Recombination occurs uniformly within the bronze gene, a meiotic recombination hotspot in the maize genome. Plant Cell 9:1633–1646
Dooner HK, Weck E, Adams S, Ralston E, Favreau M, English J (1985) A molecular genetic analysis of insertions in the bronze locus in maize. Mol Gen Genet 200:240–246
Drouaud J, Camilleri C, Bourguignon PY, Canaguier A, Berard A et al (2006) Variation in crossing-over rates across chromosome 4 of Arabidopsis thaliana reveals the presence of meiotic recombination “hotspots”. Genome Res 16:106–114
Drouaud J, Mercier R, Chelysheva L, Berard A, Falque M et al (2007) Sex-specific crossover distributions and variations in interference level along Arabidopsis thaliana chromosome 4. PLoS Genet 3:e106
Endo TR, Gill BS (1996) The deletion Stocks of Common Wheat. J Heredity 87:295–307
Erayman M, Sandhu D, Sidhu D, Dilbirligi M, Baenziger PS et al (2004) Demarcating the gene-rich regions of the wheat genome. Nucleic Acids Res 32:3546–3565
Fengler K, Allen SM, Li B, Rafalski A (2007) Distribution of genes, recombination and repetitive elements in the maize genome. Plant Genome 2:83–95
Flint-Garcia SA, Thornsberry JM, Buckler ES (2003) Structure of linkage disequilibrium in plants. Annu Rev Plant Biol 54:357–374
Fu Y, Springer NM, Gerhardt DJ, Ying K, Yeh CT et al (2010) Repeat subtraction-mediated sequence capture from a complex genome. Plant J 62:898–909
Fung JC, Rockmill B, Odell M, Roeder GS (2004) Imposition of crossover interference through the nonrandom distribution of synapsis initiation complexes. Cell 116:795–802
Gaut BS, Long AD (2003) The lowdown on linkage disequilibrium. Plant Cell 15:1502–1506
Gervais L, Dedryver F, Morlais JY, Bodusseau V, Nègre S et al (2003) Mapping of quantitative trait loci for field resistance to Fusarium head blight in an European winter wheat. Theor Appl Genet 106:961–970
Gill KS, Gill BS, Endo TR, Boyko EV (1996a) Identification and high-density mapping of gene-rich regions in chromosome group 5 of wheat. Genetics 143:1001–1012
Gill KS, Gill BS, Endo TR, Taylor T (1996b) Identification and high-density mapping of gene-rich regions in chromosome group 1 of wheat. Genetics 144:1883–1891
Guillon H, de Massy B (2002) An initiation site for meiotic crossing-over and gene conversion in the mouse. Nat Genet 32:296–299
Guyomarc’h H, Sourdille P, Charmet G, Edwards KJ, Bernard M (2002) Characterisation of polymorphic markers from T. tauschii and transferability to the D-genome of bread wheat. Theor Appl Genet 104:1164–1172
Han L, Su B, Li WH, Zhao Z (2008) CpG island density and its correlations with genomic features in mammalian genomes. Genome Biol 9:R79
Haseneyer G, Ravel C, Dardevet M, Balfourier F, Sourdille P et al (2008) High level of conservation between genes coding for the GAMYB transcription factor in barley (Hordeum vulgare L.) and bread wheat (Triticum aestivum L.) collections. Theor Appl Genet 117:321–331
Horwath A, Didier A, Koenig J, Exbrayat F, Charmet G, Balfourier F (2009) Analysis of diversity and linkage disequilibrium along chromosome 3B of bread wheat (Triticum aestivum L.). Theor Appl Genet 119:1523–1537
Inukai T, Sako A, Hirano HY, Sano Y (2000) Analysis of intragenic recombination at wx in rice: correlation between the molecular and genetic maps within the locus. Genome 43:589–596
Isobe T, Yoshino M, Mizuno KI, Fischer-Lindahl K, Koide T et al (2002) Molecular characterization of the Pb recombination hotspot in the mouse major histocompatibility complex class II region. Genomics 80:229–235
Jeffreys AJ, May CA (2004) Intense and highly localized gene conversion activity in human meiotic crossover hotspots. Nat Genet 36:151–156
Jeffreys AJ, Neumann R (2009) The rise and fall of a human recombination hotspot. Nat Genet 41:625–629
Jeffreys AJ, Murray J, Neumann R (1998) High-resolution mapping of crossovers in human sperm defines a minisatellite-associated recombination hotspot. Mol Cell 2:267–273
Jeffreys AJ, Ritchie A, Neumann R (2000) High resolution analysis of haplotype diversity and meiotic crossover in human TAP2 recombination hotspot. Hum Mol Genet 9:725–733
Jensen-Seaman MI, Furey TS, Payseur BA, Lu Y, Roskin KM et al (2004) Comparative recombination rates in the rat, mouse, and human genomes. Genome Res 14:528–538
Jones GH (1984) The control of chiasma distribution. Symp Soc Exp Biol 38:293–320
Jones GH (1987) Chiasmata. In: Moens PB (ed) Meiosis. Academic, London, pp 213–244
Kauppi L, Jeffreys AJ, Keeney S (2004) Where the crossover are: recombination distribution in mammals. Nat Rev Genet 5:413–424
Kim JS, Islam-Faridi MN, Klein PE, Stelly DM, Price HJ et al (2005) Comprehensive molecular cytogenetic analysis of sorghum genome architecture: distribution of euchromatin, heterochromatin, genes and recombination in comparison to rice. Genetics 171:1963–1976
Kim S, Plagnol V, Hu TT, Toomajian C, Clark RM et al (2007) Recombination and linkage disequilibrium in Arabidopsis thaliana. Nat Genet 39:1151–1155
Kong A, Gudbjartsson DF, Sainz J, Jonsdottir GM, Gudjonsson SA et al (2002) A high-resolution recombination map of the human genome. Nat Genet 31:241–247
Kosambi DD (1944) The estimation of map distances from recombination values. Ann Eugen 12:172–175
Kota R, Spielmeyer W, McIntosh RA, Lagudah ES (2006) Fine genetic mapping fails to dissociate durable stem rust resistance gene Sr2 from pseudo-black chaff in common wheat (Triticum aestivum L.). Theor Appl Genet 112:492
Lewontin RC (1964) The interaction of selection and linkage. I—General considerations, heterotic models. Genetics 49:49–67
Liu S, Zhang X, Pumphrey MO, Stack RW, Gill BS, Anderson JA (2006) Complex microcolinearity among wheat, rice, and barley revealed by fine mapping of the genomic region harboring a major QTL for resistance to Fusarium head blight in wheat. Funct Integr Genomics 6:83–89
Liu S, Yeh CT, Ji T, Ying K, Wu H et al (2009) Mu transposon insertion sites and meiotic recombination events co-localize with epigenetic landmarks for open chromatin across the maize genome. PLoS Genet 5:e1000733
Lukaszewski A, Curtis CA (1993) Physical distribution of recombination in B-genome chromosome of tetraploid wheat. Theor Appl Genet 86:121–127
Mancera E, Bourgon R, Brozzi A, Huber W, Steinmetz LM (2008) High-resolution mapping of meiotic crossovers and non-crossovers in yeast. Nature 454:479–485
McVean GAT, Myers SR, Hunt S, Deloukas P, Bentley DR, Donnelly P (2004) The fine-scale structure of recombination rate variation in the human genome. Science 304:581–584
Mézard C (2006) Meiotic recombination hotspots in plants. Biochem Soc Trans 34:531–534
Myers S, Freeman C, Auton A, Donnelly P, McVean G (2008) A common sequence motif associated with recombination hotspots and genome instability in humans. Nat Genet 40:1124–1129
Nachman MW (2002) Variation in recombination rate across the genome: evidence and implications. Curr Opin Genet Dev 12:657–663
Okagaki RJ, Weil CF (1997) Analysis of recombination sites within the maize waxy locus. Genetics 147:815–821
Paigen K, Szatkiewicz JP, Sawyer K, Leahy N, Parvanov ED et al (2008) The recombinational anatomy of a mouse chromosome. PLoS Genet 4:e1000119
Patterson AH, Bowers JE, Bruggmann R, Dubchak I, Grimwood J et al (2009) The sorghum bicolor genome and the diversification of grasses. Nature 457:551–556
Paux E, Roger D, Badaeva E, Gay G, Bernard M et al (2006) Characterizing the composition and evolution of homoeologous genomes in hexaploid wheat through BAC-end sequencing on chromosome 3B. Plant J 48:463–474
Paux E, Sourdille P, Salse J, Saintenac C, Choulet F et al (2008) A physical map of the 1-gigabase bread wheat chromosome 3B. Science 322:101–104
Petes TD (2001) Meiotic recombination hotspots and cold spots. Nat Rev Genet 2:360–369
Ravel C, Praud S, Murigneux A, Canaguier A, Sapet F et al (2006) Single-nucleotide polymorphisms (SNPs) frequency in a set of selected lines of bread wheat (Triticum aestivum L.). Genome 49:1131–1139
Ravel C, Martre P, Romeuf I, Dardevet M, El-Malki R et al (2009) Nucleotide polymorphism in the wheat transcriptional activator Spa influences its pattern of expression and has pleiotropic effects on grain protein composition, dough viscoelasticity, and grain hardness. Plant Physiol 151:2133–2144
Remington DL, Thornsberry JM, Matsuoka Y, Wilson LM, Whitt SR et al (2001) Structure of linkage disequilibrium and phenotypic associations in the maize genome. Proc Natl Acad Sci USA 98:11479–11484
Röder MS, Korsun V, Wendehake K, Plaschke P, Tixier MH et al (1998) A microsatellite map of the wheat genome. Genetics 149:2007–2023
Safar J, Bartos J, Janda J, Bellec A, Kubalakova M et al (2004) Dissecting large and complex genomes: flow sorting and BAC cloning of individual chromosomes from bread wheat. Plant J 39:960–968
Saintenac C, Falque M, Martin OC, Paux E, Feuillet C, Sourdille P (2009) Detailed recombination studies along chromosome 3B provide new insights on crossover distribution in wheat (Triticum aestivum L.). Genetics 181:393–403
Schnable PS, Ware D, Fulton RS, Stein JC, Wei F et al (2009) The B73 maize genome: complexity, diversity and dynamics. Science 326:1112–1115
See DR, Brooks S, Nelson JC, Brown-Guedira G, Friebe B et al (2006) Gene evolution at the ends of wheat chromosomes. Proc Natl Acad Sci USA 103:4162–4167
Sherman JD, Stack SM (1995) Two-dimensional spreads of synaptonemal complexes from solanaceous plants. VI. High-resolution recombination nodule map for tomato (Lycopersicon esculentum). Genetics 141:683–708
Sidhu D, Gill KS (2004) Distribution of genes and recombination in wheat and other eukaryotes. Plant Cell Tissue Organ Cult 79:257–270
Stein N, Feuillet C, Wicker T, Schlagenhauf E, Keller B (2000) Subgenome chromosome walking in wheat: a 450-kb physical contig in Triticum monococcum L. spans the Lr10 resistance locus in hexaploid wheat (Triticum aestivum L.). Proc Natl Acad Sci USA 97:13436–13441
Stumpf MPH, Goldstein DB (2003) Demography, recombination hotspot intensity and the block structure of linkage disequilibrium. Curr Biol 13:1–8
Tenaillon MI, Sawkins MC, Anderson LK, Stack SM, Doebley J, Gaut BS (2002) Patterns of diversity and recombination along chromosome 1 of maize (Zea mays ssp. mays L.). Genetics 162:1401–1413
Tian Z, Rizzon C, Du J, Zhu L, Bennetzen JL et al (2009) Do genetic recombination and gene density shape the pattern of DNA elimination in rice long terminal repeat retrotransposons? Genome Res 19:2221–2230
Toomajian C, Hu TH, Bomblies K, Laitinen R, Salome P et al (2009) Variation in recombination rates in Arabidopsis: computational inference and direct measurement. Plant & Animal Genome Meeting, San Diego, CA, USA, January 10–14, 2009, W430. http://www.intl-pag.org/17/abstracts/W62_PAGXVII_430.html
Wei F, Wing RA, Wise RP (2002) Genome dynamics and evolution of the Mla (powdery mildew) resistance locus in barley. Plant Cell 14:1903–1917
Werner JE, Endo TR, Gill BS (1992) Toward a cytogenetically based physical map of the wheat genome. Proc Natl Acad Sci USA 89:11307–11311
Wu J, Mizuno H, Hayashi-Tsugane M, Ito Y, Chiden Y et al (2003) Physical maps and recombination frequency of six rice chromosomes. Plant J 36:720–730
Yao H, Zhou Q, Li J, Smith H, Yandeau M et al (2002) Molecular characterization of meiotic recombination across the 140-kb multigenic a1-sh2 interval of maize Proc. Natl Acad Sci U S A 99:6157–6162
Zavattari P, Deidda E, Whalen M, Lampis R, Mulargia A et al (2000) Major factors influencing linkage disequilibrium by analysis of different chromosome regions in distinct populations: demography, chromosome recombination frequency and selection. Hum Mol Genet 9:2947–2957
Zhang X (2008) The epigenetic landscape of plants. Science 320:489–492
Zhou S, Wei F, Nguyen J, Bechner M, Potamousis K et al (2009) A single molecule scaffold for the maize genome. PLoS Genet 5:e1000711
Acknowledgements
We would like to thank J. Philippon, M. Chicard, and D. Boyer for their excellent technical assistance. We also thank the members of the GENTYANE genotyping platform of the UMR GDEC 1095 for their technical support during genotyping. The authors are also grateful to the editor and the referees for their critical comments of the manuscript and to K. Eversole for the editing and English corrections. This work has been supported by grants from the Agence Nationale de la Recherche (ANR-05-BLANC-0258-01 EXEGESE and ANR-GPLA06001G SMART) and from the INRA. CS is funded by a grant from the French Ministry of Research. The sequence of ctg0954b is available under the following accession number FN564434.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by S. Keeney
Cyrille Saintenac and Sébastien Faure contributed equally to this work.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Supplementary Figure 1
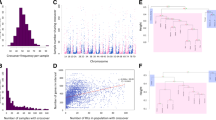
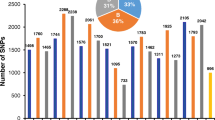
Haplotype network of Hga3 gene. Each pie represents a specific haplotype. Its size is proportional to the number of accessions constitutive of this haplotype. The number of sequence changes is indicated along the branches. The colors correspond to the different groups that structure the wheat core collection as explained in Horwath et al. (2009). Blue, North-West European accessions; red, East European, South-East European, and North American accessions; yellow, Asian accessions (Caucasian, Middle-East, Central-Asian, and South-Eastern accessions); green, accessions from International Maize and Wheat Improvement Centre (CIMMYT) and the International Centre for Agricultural Research in the Dry Areas (ICARDA); and pink, Nepal (PPT 76 kb)
Rights and permissions
About this article
Cite this article
Saintenac, C., Faure, S., Remay, A. et al. Variation in crossover rates across a 3-Mb contig of bread wheat (Triticum aestivum) reveals the presence of a meiotic recombination hotspot. Chromosoma 120, 185–198 (2011). https://doi.org/10.1007/s00412-010-0302-9
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00412-010-0302-9