Abstract
Background
Liver ischemia/reperfusion (I/R) injury is usually caused by hepatic inflow occlusion during liver surgery, and is frequently observed during war wounds and trauma. Hepatocyte ferroptosis plays a critical role in liver I/R injury, however, it remains unclear whether this process is controlled or regulated by members of the DEAD/DExH-box helicase (DDX/DHX) family.
Methods
The expression of DDX/DHX family members during liver I/R injury was screened using transcriptome analysis. Hepatocyte-specific Dhx58 knockout mice were constructed, and a partial liver I/R operation was performed. Single-cell RNA sequencing (scRNA-seq) in the liver post I/R suggested enhanced ferroptosis by Dhx58hep−/−. The mRNAs and proteins associated with DExH-box helicase 58 (DHX58) were screened using RNA immunoprecipitation-sequencing (RIP-seq) and IP-mass spectrometry (IP-MS).
Results
Excessive production of reactive oxygen species (ROS) decreased the expression of the IFN-stimulated gene Dhx58 in hepatocytes and promoted hepatic ferroptosis, while treatment using IFN-α increased DHX58 expression and prevented ferroptosis during liver I/R injury. Mechanistically, DHX58 with RNA-binding activity constitutively associates with the mRNA of glutathione peroxidase 4 (GPX4), a central ferroptosis suppressor, and recruits the m6A reader YT521-B homology domain containing 2 (YTHDC2) to promote the translation of Gpx4 mRNA in an m6A-dependent manner, thus enhancing GPX4 protein levels and preventing hepatic ferroptosis.
Conclusions
This study provides mechanistic evidence that IFN-α stimulates DHX58 to promote the translation of m6A-modified Gpx4 mRNA, suggesting the potential clinical application of IFN-α in the prevention of hepatic ferroptosis during liver I/R injury.
Similar content being viewed by others
Background
Ferroptosis is a form of regulated cell death characterized by iron-dependent lipid peroxidation to lethal levels [1,2,3]. It has been shown to play critical roles in a series of physiological and pathological processes, especially in ischemia/reperfusion (I/R) injury when excessive reactive oxygen species (ROS) are produced [4, 5]. The biochemical mechanism underlying ferroptosis is the accumulation of lethal ROS and iron Fenton reaction-induced lipid peroxides (LPOs) combined with the depletion of glutathione (GSH) and inactivation of the enzyme glutathione peroxidase 4 (GPX4), which is the central suppressor of ferroptosis by catalyzing the conversion of GSH to oxidized GSH (GSSG) and eliminating LPOs [6]. However, as liver I/R injury is the leading cause of surgery-related liver injury, it also commonly occurs during war wounds or trauma [7], the lack of effective and safe clinical precautionary or therapeutic measures is still the main problem in preventing hepatic ferroptosis [8], especially for the potential approach of enhancing GPX4 expression or activity.
DEAD/DExH-box helicase (DDX/DHX) members constitute the largest family of RNA helicases [9, 10]. These members confer RNA binding and unwinding properties and are critical for RNA metabolism, including RNA recognition, modification, splicing, transport, degradation, and translation [11, 12]. Some also belong to the interferon (IFN)-stimulated gene (ISG) family, whose expression can be induced by IFN treatment. Additionally, certain genes within this family have the ability to stimulate IFN production, such as DDX58 [also known as retinoic acid-inducible gene-I (RIG-I)] [13, 14], and DHX58 [also known as laboratory of genetics and physiology 2 (LGP2)] [15, 16]. DDX58 (RIG-I) serves as an intracellular sensor for pathogen-associated molecular patterns present in viral RNA. It possesses a DExD/H-box RNA helicase domain that exhibits ATP hydrolysis activity, along with a C-terminal repressor domain (RD) embedded within the C-terminal domain (CTD) [17]. Both the helicase and RD domains are necessary for recognizing dsRNA and 5’-triphosphate RNA in a synergistic manner [18]. Upon recognizing viral RNA, the two N-terminal tandem caspase-recruiting domains (CARDs) of DDX58 can activate downstream type I IFN production. Compared with DDX58, DHX58 (LGP2) lacks the N-terminal CARDs responsible for recruiting and interacting with downstream antiviral components. DHX58 is regarded as a non-canonical RNA-binding protein (RBP) due to its ability to associate with RNAs [19]. We previously focused on the roles of DDX/DHX family members, including DDX58 and DDX46, in regulating liver physiopathology and inflammation [20,21,22]. However, the potential roles of DDX/DHX family members in the development of hepatic ferroptosis remain unknown up to now.
The regulation of gene expression in eukaryotic cells relies heavily on mRNA metabolism, which is strictly controlled by post-transcriptional modifications, including the highly prevalent N6-methyladenosine (m6A) modification within RNA [23]. The installation of m6A modification is mediated by m6A “writers”, such as the methyltransferase complex methyltransferase-like 3 (METTL3) and METTL14, while their removal is facilitated by m6A “erasers”, including the fat mass and obesity-related gene (FTO) and alkB homolog 5 (ALKBH5). The fate and function of m6A-modified RNAs are primarily administered by m6A “readers”, such as members of the YT521-B homology (YTH) domain family (including YTHDF1, YTHDF2, YTHDF3, YTHDC1, and YTHDC2). These readers regulate mRNA splicing, stability, transport, and translation by recognizing the m6A sites within the mRNAs [24, 25]. Notably, YTHDC2 exerts control over both mRNA translation efficiency and stability in an m6A-dependent manner [26]. However, the specific mechanism by which these m6A enzymes modify or read their corresponding mRNAs remains unclear.
In this study, we screened the expression of DDX/DHX family members during liver I/R injury, and identified DHX58 as the most significantly downregulated member in ISG. Therefore, our focus was on elucidating the potential roles of hepatic DHX58 in the development of I/R injury, including its involvement in regulating hepatocyte death and RNA-binding activity. This study offers insights into understanding its underlying mechanism and provides a basis for preventing liver I/R injury.
Methods
Reagents
Antibodies against cyclooxygenase-2 (COX2; 12282), solute carrier family 7 member 11 (SLC7A11; 98051), V5-tag (13202), and horseradish peroxidase-conjugated secondary antibodies (7074 and 7076) were purchased from Cell Signaling Technology (Danvers, MA, USA). Antibodies specific to 4-hydroxynonenal (4-HNE; ab46545), malondialdehyde (MDA; ab243066), lymphocyte antigen 6 complex locus G (Ly6G; ab238132), F4/80 (ab300421), GPX4 (ab125066), METTL3 (ab195352), acyl-CoA synthetase long-chain family member 4 (ACSL4; ab155282), and YTHDC2 (ab220160) were purchased from Abcam (Cambridge, MA, USA). Antibodies specific to β-actin (A5441) and Flag-tag (F1804) were purchased from Sigma-Aldrich (St. Louis, MO, USA). The antibody specific to DHX58 (11355-1-AP) was purchased from Proteintech (Wuhan, China). Antibody specific to m6A (202003) was obtained from Synaptic Systems (Germany). Antibodies specific to CD45-Bv605 (63-0451-82) and CD11b-Percp.cy5.5 (45-0112-82) were obtained from Invitrogen (Carlsbad, CA, USA). Antibodies specific to F4/80-PE (123110) and Ly6G-FITC (127605) were purchased from BioLegend (San Diego, CA, USA). Protein G Agarose (P4691) and Anti-Flag M2 Affinity Gel (A2220) were from Sigma-Aldrich (St. Louis, MO, USA). Liproxstatin-1 (S7699), Z-VAD-FMK (Zvad; S7023), and necrostatin-1 (Nec-1; S8037) were purchased from SelleckChem (Houston, TX, USA). DMEM (11965092), fetal bovine serum (FBS; 10099141 C), and RPMI 1640 (11875093) were from Gibco (Shanghai, China).
Animals
One hundred C57BL/6 mice were purchased from Joint Ventures Sipper BK Experimental Animal Co. (Shanghai, China). To generate hepatocyte-specific DHX58 deficiency, Dhx58f/f mice (n = 4) were designed and constructed by ViewSolid Biotech (Beijing, China) using clustered regularly interspaced short palindromic repeats/CRISPR-associated 9 (CRISPR/Cas9) techniques, as previously described [27], and then hybridized with Alb-Cre transgenic mice (003574, n = 4) purchased from The Jackson Laboratory (Bar Harbor, ME, USA). Mouse genotyping was performed by PCR analysis of genomic DNA extracted from the tails, as previously described [28]. For the knockdown and overexpression of hepatic Dhx58, the adeno-associated virus serotype 8 (AAV8) constructs were established by OBiO Technology (Shanghai, China) as we previously described [21]. The shRNA target sequence of Dhx58 was 5’-CCTGACTTGAAGCAACAATTT-3’. A single tail vein injection of 2 × 1011 AAV8 was administered, and the mice underwent I/R two weeks post injection. All animal experiments were performed in accordance with the National Institute of Health Guide for the Care and Use of Laboratory Animals, with the approval of the Scientific Investigation Board of Naval Medical University, Shanghai, China.
Mouse models
For the liver I/R mouse model, male mice aged 8–10 weeks were anesthetized with pentobarbital sodium (100 mg/kg). After opening the abdominal cavity, an atraumatic vascular clamp was placed across the hepatic artery, portal vein, and bile duct to interrupt the left lateral and median lobes (70%) of the liver. After 60 min of inducing hepatic ischemia, the clamp was removed to initiate reperfusion. Sham mice underwent the same surgical treatment without vascular occlusion. The mice were sacrificed at the indicated reperfusion time points, and the serum and liver were collected immediately. To reduce ROS, N-acetylcysteine (NAC; A9165, Sigma, USA, 15 mg/kg in sterile phosphate buffered saline, intraperitoneal injection) or butylated hydroxyanisole (BHA; B1253, Sigma, USA, 50 mg/kg in corn oil, gavage) was administered twice daily for 2 d starting 48 h before liver I/R. To activate nuclear factor erythroid 2-related factor 2 (Nrf2) in vivo, 50 mg/kg dimethyl fumarate (DMF, s2586, SelleckChem, USA) was treated daily by gavage for 7 d before liver I/R. For the pharmacological inhibition of cell death, liproxstatin-1 (10 mg/kg), Zvad (10 mg/kg), or Nec-1 (1.65 mg/kg) were administered by intraperitoneal injection 30 min before surgery to inhibit ferroptosis, apoptosis, and necroptosis. Regarding the induction of hepatic ferroptosis, erastin (25 mg/kg) was injected intraperitoneally for 2 d at 12-h intervals, and ferric nitrilotriacetate (Fe-NTA; 22 mg/kg) was injected peritoneally for 3 h. The IFN-α treatment group was intraperitoneally injected with a single dose of IFN-α (1 × 106 U/kg, 752804, BioLegend, USA) at 12 h before liver I/R.
Single-cell RNA sequencing (scRNA-seq)
As described previously, mouse livers were digested to obtain single-cell suspensions A [20]. Dissociated single cells were stained with AO/PI for viability assessment using a Countstar Fluorescence Cell Analyzer. The scRNA-seq libraries were generated using the 10× Genomics Chromium Controller Instrument and Chromium Single Cell 3’V3.1 Reagent Kits (10× Genomics, Pleasanton, CA, USA) according to the recommendations of the manufacturer. scRNA-seq data analysis, including cell communication analysis, quantitative set analysis of gene expression (QuSAGE) analysis, and pathway analysis were performed by NovelBio Bio-Pharm Technology Co., Ltd., using the NovelBrain Cloud Analysis Platform as previous procedures [29,30,31,32]. To characterize the relative activation of a given gene set, such as pathway activation, we performed QuSAGE analysis, in which gene sets, including ferroptosis, described previously, were added. Pathway analysis was used to identify the significant pathways of the marker genes and differentially expressed genes according to the kyoto encyclopedia of genes and genomes (KEGG) database. Fisher’s exact test was used to select significant pathways, and the threshold of significance was defined by the P-value and false discovery rate (FDR).
Isolation of primary hepatocytes and non-parenchymal cells (NPCs)
Primary hepatocytes and NPCs were isolated according to the protocol [33]. Briefly, the mouse liver was perfused in the inferior vena cava above the kidney with 25 ml perfusion buffer and then with 10 ml digestion buffer. Subsequently, the liver sack was ruptured with fine-tip forceps along the liver surface, and the cells were released using a cell lifter. The cells were filtered through a 70 μm cell strainer and then centrifuged at 50 g for 2 min. Only the hepatocytes were pelleted, whereas the other cells remained in the supernatant. Primary hepatocytes and NPCs were collected and purified for further experiments. The viability of primary hepatocytes was evaluated by trypan blue exclusion (> 90%).
Human liver samples
Human normal liver tissues, obtained from distal normal liver tissues of liver hemangioma patients, and human liver tissues post I/R, obtained from biopsy approximately 3 h post reperfusion in donors during liver transplantation, were collected at Huashan Hospital. The donor livers for transplantation underwent 5 to 10 min of warm ischemia followed by 5 to 7 h of cold ischemia. The human subject study was approved by the human ethics committee of Huashan Hospital, Fudan University (KY2021-449).
4D label free proteomics analysis
The proteomics analysis was performed using PTM BIO (Hangzhou, China). Protein lysates from Dhx58f/f and Dhx58hep−/− livers were diluted and digested with trypsin. Tryptic peptides were dissolved in solvent A (0.1% formic acid and 2% acetonitrile/water) and directly loaded onto a home-made reversed-phase analytical column. Peptides were separated with a gradient from 6 to 24% solvent B (0.1% formic acid in acetonitrile) over 70 min, 24 to 35% in 14 min, climbing to 80% in 3 min, and then held at 80% for the last 3 min, all at a constant flow rate of 450 nl/min on a nanoElute UHPLC system (Bruker Daltonics, USA). The peptides were subjected to a capillary source, followed by the timsTOF Pro (Bruker Daltonics, USA) mass spectrometry (MS). Precursors and fragments were analyzed using a TOF detector with an MS/MS scan range of 100–1700 m/z. The resulting MS/MS data were processed using the MaxQuant search engine (v.1.6.15.0). Tandem mass spectra were searched against the human SwissProt database (20422 entries) and concatenated with a reverse-decoy database.
Co-immunoprecipitation (Co-IP) and Western blotting
Cells and livers were collected and lysed using cell lysis buffer (9803, Cell Signaling Technology, MA, USA) supplemented with a protease inhibitor cocktail (539134, Calbiochem, Germany). The protein concentrations in the lysates were measured using a bicinchoninic acid (BCA) protein assay kit (23225, Pierce, USA) and equalized with cell lysis buffer. An equal amount of the extracts was used for immunoprecipitation, which was incubated with the antibodies and protein beads overnight, and the immune complex beads were washed and boiled with sample buffer, or the extracts were directly loaded and subjected to SDS-PAGE, transferred onto nitrocellulose membranes, and then blotted as we described previously [34]. β-actin was used as a loading control.
RNA immunoprecipitation-sequencing (RIP-seq) and RIP-qRT-PCR
Primary hepatocytes seeded in a 15 cm dish at 85% confluence were cross-linked by ultraviolet light and harvested. RIP was performed using a Magna RIP™ RNA-Binding Protein Immunoprecipitation Kit (Millipore, USA) according to the instructions of the manufacturer. Cell lysates were subjected to RIP using rabbit IgG (Millipore, USA), which served as a negative control, or anti-DHX58 antibody (11355-1-AP, Proteintech, China). Input and co-immunoprecipitated RNAs were isolated using TRIzol reagent for sequencing and qRT-PCR.
Study of m6A
m6A RIP was performed according to the instructions of the Magna RIP™ RNA-Binding Protein Immunoprecipitation Kit. Cell lysates were subjected to RIP using rabbit IgG (Millipore, USA), a negative control, or anti-m6A (202003, Synaptic Systems, Germany). Input and co-immunoprecipitated RNAs were isolated using TRIzol reagent for sequencing and qRT-PCR. Global m6A mRNA levels were quantified using an m6A RNA methylation assay kit (ab185912, Abcam, USA) following the protocol by the manufacturer. MeRIP-seq was performed by the Biotechnology Corporation (Shanghai, China).
Polysome profiling
The procedures were performed according to a previously described protocol [35]. The hepatocytes were collected with polysome extraction buffer [20 mmol/L Tris-HCl pH 7.5, 100 mmol/L KCl, 5 mmol/L MgCl2, 0.5% Nonidet P-40, protease inhibitors, RNase inhibitors, and 100 µg/ml cycloheximide (CHX)], and the sample was loaded onto a 10–50% w/v sucrose gradient. The gradients were centrifuged at 4 ℃ for 90 min in a SW41Ti swinging bucket rotor at 39,000 rpm. The gradients were fractioned using a Gradient Station (BioComp, Canada) equipped with an ECONO UV monitor (BioRad, USA) and collected using a fraction collector (FC203B, Gilson, USA). Total RNA from the indicated fractions was extracted using TRIzol reagent for qRT-PCR analysis. For more details, please see Additional file 1: Materials and methods.
Statistical analysis
Data are presented as the mean ± SD of one representative of at least three independent experiments. Statistical comparisons between experimental groups were analyzed by the Student’s t-test in SPSS 17.0 (Chicago, IL, USA), and a two-tailed P < 0.05 was taken to indicate statistical significance.
Results
Excessive ROS during liver I/R injury decreases hepatic DHX58 expression
To comprehensively elucidate the potential roles of DDX/DHX family members in the development of liver I/R injury, we conducted an analysis on their expression and found that Dhx58 was one of the five family members (Ddx4, Ddx11, Ddx25, Ddx43, and Dhx58) with the most markedly decrease in expression after I/R in the liver (Fig. 1a; Additional file 1: Fig. S1a). The basal expression of the first 4 genes was too low to be analyzed (Additional file 1: Fig.S1b), thus our focus shifted to Dhx58. DHX58 expression in the liver was mainly located in parenchymal hepatocytes (Fig. 1b, c). Both protein and mRNA levels of DHX58 were confirmed to be reduced within 6 h in the liver following I/R in vivo (P < 0.01, Fig. 1d, e) and in primary hepatocytes after H/R in vitro (P < 0.01, Fig. 1f, g). Moreover, we collected human liver samples from donors after reperfusion during transplantation and observed a significant decrease in hepatic DHX58 expression (P = 0.0019) during I/R injury, accompanied by an increase of liver COX2 expression. Furthermore, a negative correlation was found between the levels of DHX58 and COX2 (r = -0.8024, P = 0.0017) (Fig. 1h, i). Therefore, the expression of DHX58 in hepatocytes is decreased in response to liver I/R injury.
Hepatic DHX58 expression is markedly decreased during liver I/R injury. a Expression of hepatic DDX/DHX family members following liver I/R injury. b Immunofluorescence staining images of DHX58, hepatocyte and DAPI in liver sections analyzed by confocal microscopy. Scale bar = 50 µm. c DHX58 protein level in the isolated hepatocytes and non-parenchymal cells (NPCs) of the liver was examined by Western blotting. d DHX58 protein level in liver tissues after I/R injury was examined by Western blotting. e Dhx58 mRNA level in liver tissues after I/R injury was examined by qRT-PCR. f DHX58 protein level in primary hepatocytes following H/R injury was examined by Western blotting. g Dhx58 mRNA level in primary hepatocytes following H/R injury was examined by qRT-PCR. h DHX58 and COX2 in human normal liver tissues, obtained from distal normal liver tissues of liver hemangioma patients, and human liver tissues post I/R, obtained from biopsy approximately 3 h post reperfusion in donors during liver transplantation, were examined by Western blotting. The donor livers for transplantation underwent 5 to 10 min of warm ischemia followed by 5 to 7 h of cold ischemia. Quantified DHX58 expression was shown. i The correlation between DHX58 and COX2 in (h) was analyzed by Pearson’s correlation coefficient assay. Data are shown as mean ± SD (n = 3 or indicated) or photographs from one representative of three independent experiments. **P < 0.01 compared with the sham/control group by Student’s two-tailed t-test. DHX58 DExH-box helicase 58, I/R ischemia/reperfusion, DAPI 4’,6-diamidino-2-phenylindole, H/R hypoxia/re-oxygenation, COX2 cyclooxygenase-2, SD standard deviation
The excessive production of ROS is critical for initiating I/R injury [36]. Therefore, we found that H2O2 was responsible for the decrease in DHX58 levels in primary hepatocytes (P < 0.05 or P < 0.01, Fig. 2a, b), as well as in the human hepatocyte HHL5 cell line (P < 0.05 or P < 0.01, Additional file 1: Fig. S2a, b). The expression of DHX58 was also decreased by another ROS-generating compound, menadione (P < 0.01, Additional file 1: Fig. S2c, d). Using NAC or BHA to scavenge ROS, the decreased DHX58 levels were alleviated in the liver after I/R treatment in vivo (P < 0.01, Fig. 2c, d) and in primary hepatocytes treated with H/R or H2O2 in vitro (P < 0.01, Fig. 2e-h). Moreover, these treatments resulted in decreased serum ALT and AST levels (P < 0.05 or P < 0.01), indicating a reduction in liver damage (Additional file 1: Fig. S2e). Additionally, DMF-induced activation of the antioxidant Nrf2 mitigated the reduction of DHX58 in the liver after I/R and in primary hepatocytes following H/R (P < 0.01, Additional file 1: Fig. S2f-i). Taken together, excessive ROS production during I/R injury leads to a reduction in hepatic DHX58 expression, which may contribute to the development of liver I/R injury.
Excessive ROS production during liver I/R injury decreases DHX58 expression. a DHX58 protein level in primary hepatocytes treated with H2O2 was examined by Western blotting. b Dhx58 mRNA level in primary hepatocytes treated with H2O2 was examined by qRT-PCR. c DHX58 protein level in liver tissues with NAC or BHA pre-treatment and then I/R was examined by Western blotting. d Dhx58 mRNA level in liver tissues with NAC or BHA pre-treatment and then I/R was examined by qRT-PCR. e DHX58 protein level in primary hepatocytes with NAC or BHA pre-treatment and then H/R was examined by Western blotting. f Dhx58 mRNA level in primary hepatocytes with NAC or BHA pre-treatment and then H/R was examined by qRT-PCR. g DHX58 protein level in primary hepatocytes with NAC or BHA pre-treatment and then H2O2 (2 mmol/L) administration for 3 h was examined by Western blotting. h Dhx58 mRNA level in primary hepatocytes with NAC or BHA pre-treatment and then H2O2 (2 mmol/L) administration for 3 h was examined by qRT-PCR. Data are shown as mean ± SD (n = 3) or photographs from one representative of three independent experiments. *P < 0.05, **P < 0.01. ROS reactive oxygen species, I/R ischemia/reperfusion, DHX58 DExH-box helicase 58, NAC N-acetylcysteine, BHA butylated hydroxyanisole, H/R hypoxia/re-oxygenation, SD standard deviation
Decreased DHX58 promotes hepatic ferroptosis and I/R injury
To investigate the potential role of decreased DHX58 levels in developing liver I/R injury, we constructed hepatocyte-specific Dhx58 knockout mice Dhx58hep−/− (Fig. 3a; Additional file 1: Fig. S3a, b). The Dhx58hep−/− livers seemed normal but exhibited significantly more severe liver damage (P < 0.01, Fig. 3b-d), higher serum ALT and AST (P < 0.01, Fig. 3e), elevated mRNAs of inflammatory cytokines [interleukin-6 (IL-6), IL-1β, monocyte chemoattractant protein 1 (MCP1)] (P < 0.05 or P < 0.01, Fig. 3f), and enhanced infiltration of inflammatory cells (neutrophils and macrophages; Fig. 3g; Additional file 1: Fig. S3c, d) during liver I/R injury compared with those in Dhx58f/f livers. Moreover, primary hepatocytes isolated from Dhx58hep−/− mice showed lower cell viability when subjected to H/R in vitro (P < 0.01), whereas DHX58 overexpression in primary hepatocytes improved cell viability under H/R compared to their respective controls (P < 0.01) (Additional file 1: Fig. S3e). Thus, the decreased expression of DHX58 during I/R may potentially contribute to liver injury.
Decreased DHX58 aggravates liver I/R injury. a DHX58 protein level in liver tissues and isolated hepatocytes from Dhx58f/f and Dhx58hep−/− mice was confirmed by Western blotting. Liver I/R injury was administrated in Dhx58f/f and Dhx58hep/− mice, liver injury was shown by gross appearances of representative livers, and white arrow indicated injured area (b), liver pathology was analyzed by HE staining (c), necrotic area and Suzuki’s score were analyzed (n = 4) (d), serum ALT and AST were examined (n = 4) (e), IL-6, IL-1β, and MCP1 mRNA levels in liver tissues were examined by qRT-PCR (n = 4) (f), infiltration of neutrophils and macrophages were analyzed by flow cytometry (g). Scale bar = 20 μm. Data are shown as mean ± SD or photographs from one representative of three independent experiments. *P < 0.05, **P < 0.01. DHX58 DExH-box helicase 58, I/R ischemia/reperfusion, HE hematoxylin-eosin, ALT alanine aminotransferase, AST aspartate aminotransferase, IL-6 interleukin-6, IL-1β interleukin-1β, MCP1 monocyte chemoattractant protein 1
To clarify the underlying mechanism of Dhx58hep−/−-induced liver I/R injury, we performed scRNA-seq on the livers of Dhx58f/f and Dhx58hep−/− mice treated with sham or I/R. Intrahepatic cells were divided into 10 clusters, with the largest number of hepatocytes (Fig. 4a, b). Additionally, the scRNA-seq data of Dhx58f/f and Dhx58hep−/− livers after I/R were analyzed, and the cells were clustered into 17 populations (Additional file 1: Fig. S4a), with QuSAGE analysis showing ferroptosis enrichment in some of these cell clusters post I/R, including hepatocyte_c01, hepatocyte_c06, hepatocyte_c09, and hepatocyte_c13 (Additional file 1: Fig. S4b). Therefore, we re-clustered the hepatocytes of Dhx58f/f and Dhx58hep−/− livers under I/R, and divided them into 10 populations (Fig. 4c). QuSAGE analysis was specifically focused on the cell death pathways. The hepatocyte_c00 and c01, which exhibited significant enhancement in Dhx58hep−/− livers (Fig. 4d), showed enrichment for ferroptosis as indicated by QuSAGE analysis (Fig. 4e). Moreover, pathway analysis revealed that ferroptosis was significantly enriched in hepatocyte_c00 and c01 (Additional file 1: Fig. S4c). Furthermore, cell phone analysis suggested potential interaction between hepatocyte_c00, c01, and other clusters, including neutrophils and monocytes (Additional file 1: Fig. S4d, e). Hence, the scRNA-seq data suggest that Dhx58hep−/− may enhance ferroptosis in hepatocyte during I/R injury.
The downregulation of DHX58 expression following I/R injury promotes ferroptosis in hepatocyte. a UMAP visualization of cells in the livers of Dhx58f/f and Dhx58hep−/− mice underwent sham or I/R, and each dot corresponded to one single cell colored according to the cell cluster. b The bubble diagram of signature genes for each cell type is displayed. UMAP visualization of hepatocytes in I/R-treated Dhx58f/f and Dhx58hep−/− mice was shown as indicated (c), cell counts of Dhx58f/f and Dhx58hep−/− mice in each hepatocyte cluster were shown (d), and functional gene enrichment in each hepatocyte cluster by QuSAGE analysis was shown (e). Liver I/R injury was administrated in Dhx58f/f and Dhx58hep/− mice, hepatic iron (f) and LPO (g) levels were analyzed accordingly (n = 4), representative transmission electron microscope images show the morphology of ferroptosis in hepatocyte with black arrows indicated mitochondrion and lipid droplets (h), GSH and GSSG were analyzed and the ratio was calculated (n = 4) (i). Scale bar = 20 μm (f) and 2 μm (h). j Serum ALT and AST of Dhx58f/f and Dhx58hep−/− mice treated with ferroptosis inducer erastin (n = 3). Data are shown as mean ± SD or photographs from one representative of three independent experiments. *P < 0.05, **P < 0.01. DHX58 DExH-box helicase 58, I/R ischemia/reperfusion, UMAP uniform manifold approximation and projection, QuSAGE quantitative set analysis of gene expression, LPO lipid peroxide, GSH glutathione, GSSG oxidized glutathione, ALT alanine aminotransferase, AST aspartate aminotransferase, SD standard deviation, HSEC hepatic sinusoid endothelium cell, GMP granulocyte-monocyte progenitor, NK natural killer, AIM2 absent in melanoma 2, NLRs nucleotide-binding leucine-rich repeat receptors, NLRP1 nucleotide-binding oligomerization domain-like receptor protein 1, IPAF interleukin-1β-converting enzyme-protease activating factor
To confirm the promotion of ferroptosis in hepatocytes by Dhx58hep−/−, livers from Dhx58hep−/− mice were examined after I/R injury. The results revealed an increase in ROS levels (Additional file 1: Fig. S5a), heightened 4-HNE levels (Additional file 1: Fig. S5b, c), elevated hepatic iron accumulation (P < 0.01, Fig. 4f), strengthened lipid peroxidation (P < 0.01, Fig. 4g), raised MDA levels (Additional file 1: Fig. S5d, e), augmented mitochondrial damage (Fig. 4h), increased TUNEL staining intensity (Additional file 1: Fig. S5f), and enhanced GSH/GSSG ratio (P < 0.05, Fig. 4i) compared to those in Dhx58f/f livers. In addition, using the respective inhibitors of ferroptosis, apoptosis, or necroptosis in vivo, only inhibition of ferroptosis abolished the differences in liver I/R injury between Dhx58f/f and Dhx58hep−/− mice (Additional file 1: Fig. S5g-j), confirming Dhx58hep−/− promotes ferroptosis in hepatocyte. We also knocked down or overexpressed hepatic Dhx58 through AAV8-mediated gene delivery. The levels of serum ALT (P < 0.01) and AST (P < 0.05), hepatic iron (P < 0.05), LPO (P < 0.05), and the GSH/GSSG ratio (P < 0.01) were significantly altered in response to Dhx58 knockdown, promoting I/R-induced liver injury and ferroptosis in hepatocyte, whereas DHX58 overexpression alleviated I/R-induced liver injury and ferroptosis in hepatocyte [serum ALT (P < 0.01) and AST (P < 0.05 or P < 0.01), hepatic iron (P < 0.05), LPO (P < 0.05), GSH/GSSG ratio (P < 0.01)] (Additional file 1: Fig. S6a-h). Furthermore, serum ALT and AST (P < 0.01), hepatic iron (P < 0.01), LPO (control: P < 0.05, erastin: P < 0.01), and the GSH/GSSG ratio (P < 0.01) were enhanced by Dhx58hep−/− following ferroptosis inducer erastin treatment (Fig. 4j; Additional file 1: Fig. S6i-k). Following another ferroptosis inducer Fe-NTA treatment, serum ALT and AST (P < 0.01), hepatic iron (P < 0.05), LPO (control: P < 0.01, Fe-NTA: P < 0.05), and the GSH/GSSG ratio (P < 0.01) were also enhanced by Dhx58hep−/− (Additional file 1: Fig. S6l-o). Moreover, cell viability was inhibited by DHX58 deficiency (P < 0.01) while enhanced by DHX58 overexpression (P < 0.05) in primary hepatocytes in vitro (Additional file 1: Fig. S6p, q). Taken together, our findings suggest that downregulation of DHX58 expression contributes to the induction of ferroptosis in hepatocyte during liver I/R injury.
DHX58 associates and promotes the translation of Gpx4 mRNA
The mechanism by which DHX58 inhibits ferroptosis was investigated. The proteomics screening of Dhx58f/f and Dhx58hep−/− livers revealed a total of 213 differentially expressed proteins. DHX58, belonging to the DDX/DHX family with RNA-binding activity, was subjected to RIP-seq using the DHX58 antibody, resulting in the identification of 4617 RNAs. The intersection of these two sets was 124 genes, and together with the literature search of ferroptosis, Gpx4 and Hspb1 were suggested to be differentially expressed between Dhx58f/f and Dhx58hep−/− livers, with their mRNAs levels being associated with DHX58 (Fig. 5a). Using RIP-qRT-PCR, we found that only Gpx4 mRNA was stably associated with DHX58 (P < 0.05, Fig. 5b), with a sequence motif within Gpx4 mRNA enriched in the RIP-seq peaks (Fig. 5c). It was determined that the CTD of DHX58 is responsible for this specific interaction. The binding capacity of the CTD to Gpx4 mRNA closely resembles that of full-length DHX58 (P < 0.01, Fig. 5d). The presented data provide evidence for the association between DHX58 and Gpx4 mRNA.
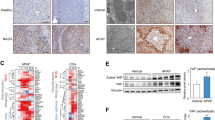
DHX58 associates Gpx4 mRNA and promotes its translation. a Schematic workflow of DHX58 downstream targets analysis. b The association between DHX58 and Gpx4 mRNA in primary hepatocytes were determined by RIP-qRT-PCR. *P < 0.05. c The top motif identified by HOMER of DHX58-bound peaks in Gpx4 mRNA. d The association between the CTD domain of DHX58 and endogenous Gpx4 mRNA in primary hepatocytes was determined by RIP-qRT-PCR. *P < 0.05, **P < 0.01, ns non-significant. e DHX58, ACSL4, COX2, SLC7A11, and GPX4 protein levels in liver tissues of Dhx58f/f and Dhx58hep−/− mice after I/R injury were examined by Western blotting. f DHX58, ACSL4, COX2, SLC7A11, and GPX4 protein levels in primary hepatocytes from Dhx58f/f and Dhx58hep−/− mice following H/R injury were examined by Western blotting. g GPX4 protein level in DHX58-overexpressed primary hepatocytes was examined by Western blotting. h DHX58, ACSL4, COX2, SLC7A11, and GPX4 protein levels in primary hepatocytes from Dhx58f/f or Dhx58hep−/− mice with GPX4 overexpression and treatment of H/R or erastin were examined by Western blotting. i In primary hepatocytes from Dhx58f/f and Dhx58hep−/− mice, relative Gpx4 mRNA distribution in each ribosome fractions was analyzed by qRT-PCR. Data are shown as mean ± SD (n = 3) or photographs from one representative of three independent experiments. #P < 0.05, ##P < 0.01 vs. Dhx58f/f. ▲CTD C-terminal domain deleted, DHX58 DExH-box helicase 58, GPX4 glutathione peroxidase 4, RIP RNA immunoprecipitation, HOMER hypergeometric optimization of motif enrichment, CTD C-terminal domain, ACSL4 acyl-CoA synthetase long chain family member 4, COX2 cyclooxygenase-2, SLC7A11 solute carrier family 7 member 11, I/R ischemia/reperfusion, H/R hypoxia/re-oxygenation, LMW low molecular weight, HMW high molecular weight, SD standard deviation
The ferroptosis suppressor Gpx4 was then examined in the liver in vivo and in primary hepatocytes in vitro. Dhx58hep−/− resulted in a reduction of GPX4 protein levels, while its mRNA level did not show significant change (Fig. 5e, f; Additional file 1: Fig. S7a, b). For other ferroptosis markers ACSL4, COX2, and SLC7A11, both protein and mRNA levels were induced by I/R in vivo and H/R in vitro, and their expression was further increased by Dhx58hep−/− (Fig. 5e, f; Additional file 1: Fig. S7a, b), which is in accordance with the promoted ferroptosis in Dhx58hep−/− liver. The DHX58-mediated increase in GPX4 and decrease in ACSL4, COX2, and SLC7A11 were also validated by knockdown or overexpression of Dhx58, both in the liver in vivo and in primary hepatocytes in vitro (Additional file 1: Fig. S7c-f). DHX58 overexpression in hepatocytes increased the protein level of GPX4 in a dose-dependent manner (Fig. 5g), confirming the binding of DHX58 to Gpx4 mRNA to increase its protein level. Furthermore, in GPX4 overexpressed hepatocytes, the induced ferroptosis markers upon H/R or erastin administration were suppressed, and the differences between Dhx58f/f and Dhx58hep−/− were abolished (Fig. 5h), suggesting that Dhx58hep−/− failed to promote ferroptosis under GPX4 overexpression, and the promotion of ferroptosis by Dhx58hep−/− is dependent on GPX4 reduction. Thus, DHX58 can inhibit liver ferroptosis by binding to Gpx4 mRNA and increasing its protein level.
To elucidate how DHX58 increases GPX4 protein levels, we first tested the degradation of GPX4 protein. Primary hepatocytes were treated with proteasome inhibitor MG132 and lysosome inhibitor chloroquine (CQ) to assess GPX4 expression. However, neither proteasome inhibitor nor lysosome inhibitor resulted in increased expression of GPX4 in Dhx58hep−/− hepatocytes (Additional file 1: Fig. S7g), and these inhibitors also failed to alleviate differences in GPX4 protein levels mediated by DHX58 overexpression (Additional file 1: Fig. S7h), so we excluded that the proteasomal and lysosomal protein degradation of GPX4 was suppressed by DHX58. Next, Gpx4 mRNA levels were not affected by DHX58 overexpression (Additional file 1: Fig. S7i), and its half-life was also unchanged by Dhx58hep−/− (Additional file 1: Fig. S7j), suggesting that the stability of Gpx4 mRNA was not influenced by DHX58. We then performed ribosome profiling of Gpx4 mRNA, and found that its translation was suppressed by Dhx58hep−/− (P < 0.05) (Fig. 5i). Taken together, we conclude that DHX58 associates with Gpx4 mRNA and promotes its translation, thereby increasing GPX4 protein levels and preventing hepatic ferroptosis.
DHX58 recruits YTHDC2 to enhance m6A-dependent translation of Gpx4 mRNA
To elucidate the mechanism underlying DHX58-mediated promotion of Gpx4 mRNA translation, we performed immunoprecipitation of Flag-tagged DHX58 from the lysates of transfected hepatocytes, employed MS to identify proteins associated with DHX58, and selected YTHDC2, an m6A reader with the highest protein score, as a potential candidate (Additional file 1: Fig. S8a-c). The constitutive endogenous association between DHX58 and YTHDC2 was validated (Fig. 6a), with the CTD domain of DHX58 and the R3H domain of YTHDC2 identified as responsible for their interaction (Additional file 1: Fig. S8d, e). Moreover, the expression of GPX4 protein was enhanced by YTHDC2 overexpression, similar to DHX58. Conversely, Ythdc2 knockdown inhibited the GPX4 protein expression (Fig. 6b). However, the mRNA levels remained unchanged (Additional file 1: Fig. S8f). The translation of Gpx4 mRNA was then examined, and it was enhanced by YTHDC2 overexpression, while suppressed by Ythdc2 knockdown (P < 0.01, Fig. 6c). DHX58-promoted GPX4 protein levels were also dependent on YTHDC2 (Fig. 6d). Thus, DHX58 cooperatively enhances Gpx4 mRNA translation by recruiting YTHDC2.
DHX58 recruits YTHDC2 to promote the translation of m6A-modified Gpx4 mRNA. a The endogenous association between DHX58 and YTHDC2 in liver tissues was examined by Co-IP. b GPX4 protein level in primary hepatocytes with Ythdc2 overexpression or knockdown was examined by Western blotting. c In primary hepatocytes with Ythdc2 overexpression or knockdown, the relative distribution of Gpx4 mRNA in each ribosome fraction was analyzed by qRT-PCR. ##P < 0.01 vs. Empty vector. d In primary hepatocytes from Dhx58f/f and Dhx58hep−/− mice with knockdown of Ythdc2, DHX58, YTHDC2, and GPX4 protein levels were examined by Western blotting. e Sequencing read clusters from MeRIP-seq analysis of Gpx4 mRNA in primary hepatocytes and top consensus motif identified by HOMER with MeRIP-seq peaks. f m6A modification of Gpx4 and Acsl4 mRNAs in primary hepatocytes were examined by RIP-qRT-PCR. g GPX4 protein level in primary hepatocytes with knockdown of Mettl3 was examined by Western blotting. h In primary hepatocytes transfected with Flag-tagged DHX58 or YTHDC2, together with Mettl3 knockdown, Flag-tag, METTL3, and GPX4 protein levels were examined by Western blotting. Data are shown as mean ± SD (n = 3) or photographs from one representative of three independent experiments. **P < 0.01. ns non-significant, Si-1 No.1 siRNA targeting YTHDC2, Si-2 No.2 siRNA targeting YTHDC2, DHX58 DExH-box helicase 58, YTHDC2 YT521-B homology domain containing 2, m6A N6-methyladenosine, Gpx4 glutathione peroxidase 4, Co-IP co-immunoprecipitation, MeRIP-seq methylated RNA immunoprecipitation sequencing, HOMER hypergeometric optimization of motif enrichment, Acsl4 acyl-CoA synthetase long chain family member 4, RIP RNA immunoprecipitation, METTL3 methyltransferase complex methyltransferase-like 3, CTD C-terminal domain, SD standard deviation, LMW low molecular weight, HMW high molecular weight
Since YTHDC2 is a typical m6A reader that enhances the translation of m6A-modified mRNAs [26], we examined whether Gpx4 mRNA was m6A-modified and read by YTHDC2. Using m6A immunoprecipitation and sequencing (MeRIP-seq), the m6A peaks present in Gpx4 transcript were determined and the sequence motifs modified at the top were identified (Fig. 6e). The m6A modification of Gpx4 mRNA (P < 0.01) was also confirmed using MeRIP-qRT-PCR (Fig. 6f). To investigate whether m6A-modified Gpx4 mRNA promotes its translation, we knocked down the m6A writer Mettl3 to suppress m6A modification (P < 0.01, Additional file 1: Fig. S8g), and Gpx4 protein levels decreased (Fig. 6g), whereas its mRNA levels remained unchanged (Additional file 1: Fig. S8h), which was similar to repression observed with DHX58 or YTHDC2. Furthermore, the overexpression of DHX58 or YTHDC2 increased GPX4 protein levels in an m6A-dependent manner, and this promotion was abolished when Mettl3 was knocked down (Fig. 6h). Additionally, m6A modification of Gpx4 mRNA was not influenced by DHX58 (Additional file 1: Fig. S8i), and its association with DHX58 was not influenced by the m6A modification (Additional file 1: Fig. S8j). Collectively, our findings suggest that DHX58 exerts a protective role against ferroptosis by recruiting YTHDC2 to read and promote m6A-dependent translation of Gpx4 mRNA, thus enhancing GPX4 protein levels.
IFN-α treatment stimulates DHX58 to prevent hepatic ferroptosis
As a typical ISG, DHX58, is upregulated by IFN-α treatment [37]. We hypothesize that the induction of DHX58 expression by IFN-α treatment may potentially prevent ferroptosis during liver I/R injury. Intraperitoneal injection of IFN-α 12 h before surgery significantly promoted the expression of DHX58 and its downstream GPX4 (Fig. 7a), and liver damage was suppressed (P < 0.01, Fig. 7b-d), especially hepatic ferroptosis was prevented (P < 0.05, Fig. 7e-g; Additional file 1: Fig. S9a, b), accompanied by ameliorated expression of transferrin, hepcidin, and ferroportin after liver I/R injury (Additional file 1: Fig. S9c). Moreover, the preventive effect of IFN-α was abolished in Dhx58hep−/− mice (Fig. 7b-g; Additional file 1: Fig. S9a, b), determining that IFN-α treatment stimulates DHX58 expression, thereby preventing hepatic ferroptosis. Moreover, pretreatment with IFN-α also exhibited inhibitory effects on liver damage (P < 0.01) and ferroptosis (P < 0.05) in erastin-induced hepatic ferroptosis, which was abolished by Dhx58hep−/− (Additional file 1: Fig. S9d-f), thereby confirming the prevention role of IFN-α-stimulated DHX58 in ferroptosis. In conclusion, we propose that excessive production of ROS during liver I/R injury leads to a decrease in DHX58 expression, thereby inhibiting downstream GPX4 and promoting ferroptosis. However, IFN-α can stimulate the expression of DHX58, leading to the recruitment of YTHDC2 to read and promote the translation of m6A-modified Gpx4 mRNA, thus preventing hepatic ferroptosis (Fig. 8).
Pretreatment with IFN-α can inhibit hepatic ferroptosis by stimulating DHX58. a DHX58 and GPX4 protein levels in the liver with IFN-α pretreatment and then I/R. Wild-type (WT) and Dhx58hep/− mice were pretreated with IFN-α, and then underwent I/R, liver damage was examined by serum ALT and AST (b), liver pathology was analyzed by HE staining (c), necrotic area and Suzuki’s score were examined (d), ROS production was analyzed by DHE staining (e), hepatic iron (f) and LPO (g) levels were analyzed accordingly. Scale bar = 20 μm. Data are shown as mean ± SD (n = 4) or photographs from one representative of three independent experiments. *P < 0.05, **P < 0.01. ns non-significant, IFN-α interferon-α, DHX58 DExH-box helicase 58, GPX4 glutathione peroxidase 4, I/R ischemia/reperfusion, ALT alanine aminotransferase, AST aspartate aminotransferase, HE hematoxylin-eosin, ROS reactive oxygen species, DHE dihydroethidium, LPO lipid peroxide, SD standard deviation
The pretreatment with IFN-α induces the activation of DHX58, which recruits YTHDC2 to recognize and enhance the translation of m6A-modified Gpx4 mRNA, thus preventing hepatic ferroptosis. IFN-α interferon-α, DHX58 DExH-box helicase 58, GPX4 glutathione peroxidase 4, ROS reactive oxygen species, YTHDC2 YT521-B homology domain containing 2, m6A N6-methyladenosine
Discussion
Liver I/R injury is an unavoidable leading cause of surgery-related liver injury and commonly occurs during war wounds and trauma. Ferroptosis in hepatocyte plays a critical role in this process, characterized by excessive ROS production and iron-dependent lipid peroxidation [38]. Here, IFN-α pretreatment exerts a protective effect against liver I/R injury, potentially through the upregulation of DHX58 expression and promotion of m6A-modified Gpx4 mRNA translation, thus attenuating ferroptosis during liver I/R injury. Since the lack of effective and safe clinical precautionary or therapeutic measures remains a significant challenge in the prevention of liver I/R injury, we suggest that the treatment of IFN-α may have considerable clinical potential for preventing hepatic ferroptosis and inhibiting liver I/R injury. However, it is important to note that these findings are based on mouse models and still need validation in human subjects.
The roles of IFN in the progression of ferroptosis have been extensively studied, and multiple reports have demonstrated that IFN-γ, specially produced by T cells, can effectively promote ferroptosis in cancer cells [39,40,41,42,43,44,45,46], which appears to contradict the preventive role of IFN-α on ferroptosis in hepatocytes observed in this study. This discrepancy may be due to the different functions between IFN-α and IFN-γ, as well as the disparate responses exhibited by hepatocytes and cancer cells towards the IFN system. Although DHX58 is a typical ISG downstream of IFN, its induction in response to IFN-α and IFN-γ may exhibit variations, which could also differ between hepatocytes and cancer cells. Additionally, the precise role of DHX58 in regulating ferroptosis may vary between hepatocytes and cancer cells. Thus, additional research is necessary to explore the distinctions between IFN-α and IFN-γ in their regulation of ferroptosis, including the downstream ISGs, particularly within the context of cancer. Moreover, a previous report has also mentioned that IFN-α derived from liver plasmacytoid dendritic cells (pDCs) can promote liver I/R injury by enhancing apoptosis through the promotion of IFN regulatory factor 1 (IRF-1) [47]. Regarding the potential contradiction of IFN-α in the prevention of ferroptosis in hepatocyte determined in this study, we propose that the discrepancy may be attributed to the quantity of IFN-α in the liver. It is evident that pDCs account for only a small portion of cells in the liver, and IFN-α derived from them might have a limited impact on the overall significant elevation of IFN-α levels in the liver. Consequently, the pDCs-derived IFN-α may not be adequate to comprehensively induce the expression of the IFN-stimulated gene Dhx58 in the liver and subsequently prevent ferroptosis in hepatocyte. In our study, we administered recombinant IFN-α through intraperitoneal injection, which effectively increased its concentration systematically in the liver and significantly enhanced protein levels of hepatic DHX58 as well as downstream GPX4, which successfully prevented ferroptosis in hepatocyte after liver I/R injury. However, further investigation is required to validate these hypotheses, especially involving human participants.
IFN-α has been used clinically to treat chronic viral infectious diseases, and its safety profile has been validated. Although influenza-like symptoms such as fever are sometimes inevitable, these symptoms often vanish within 12 h post IFN-α injection. In this study, the pretreatment of IFN-α is performed 12 h before liver I/R surgery, with a single dose of approximately 2 × 104 U per mouse. This dosage (converted into human about 4 × 106 U) is lower than the conventional therapeutic dose used for human viral infection or cancer, which is (0.5–1.5) × 107 U per day [48, 49]. Importantly, no abnormalities were observed in liver histology and serum markers associated with liver injury after IFN-α pretreatment, suggesting that the pretreatment of IFN-α is safe for preventing ferroptosis in hepatocyte during liver I/R injury. Nonetheless, further validation in human participants is necessary to evaluate the clinical efficacy and safety of IFN-α pretreatment in the prevention of ferroptosis and I/R injury during liver operation.
During liver I/R injury, damaged mitochondria generate excessive ROS, leading to oxidative stress in hepatocytes, which is one of the main causes of hepatocyte injury and cell death. The present study discovered that ROS produced during liver I/R injury suppresses the expression of DHX58, and this reduction of DHX58 was shown to participate in the progression of ferroptosis in hepatocyte following I/R injury. Both mRNA and protein levels of DHX58 were observed to decrease after liver I/R injury or exposure to ROS, indicating that elevated ROS levels hinder DHX58 expression at the transcriptional level. We investigated the upstream promoter region of the Dhx58 gene, as well as the possible transcription factors responsible for its basal expression in hepatocytes, and their potential interaction with oxidative stress and host antioxidant response. However, no conclusive evidence was found in this regard. Perhaps the alteration of epigenetic factors contributes to the decreased DHX58 following stimulation with excessive ROS, such as modifications at the DNA or histone levels. In particular, intracellular metabolic reprogramming induced by I/R injury and oxidative stress in hepatocytes, metabolic disorders including lactic acid, acetyl coenzyme A, and crotonyl coenzyme A, have been shown to participate in the epigenetic modifications of histones and modulate downstream gene expression under cellular stress conditions [50]. Further metabolic and epigenetic studies are required to validate this hypothesis.
DHX58, also known as LGP2, belongs to the RIG-I (DDX58)-like receptor family and serves as a well-established intracellular sensor for host recognition of invading RNA viruses in innate immune cells. In the present study, DHX58 was also observed to be expressed in parenchymal cells of organs such as hepatocytes, which may imply a specific role associated with organ function. Combined with our previous findings demonstrating predominant expression of the innate sensor RIG-I (DDX58) in parenchymal hepatocytes of the liver and its involvement in regulating hepatic lipid metabolism and inflammation [20, 21], we propose that other immune sensors or immune molecules may also exhibit expression in the parenchymal cells of various organs or systems. Moreover, their biological functions might extend beyond immune activation or regulation. These sensors possess the capability to recognize and bind nucleic acids from invading pathogens, as well as host DNA or RNA [51], thereby modulating their biological processes and functions, consequently governing the corresponding physiological or pathological processes in specific organs. This assumption hypothesis holds promise for future research.
Post-transcriptional modifications of mRNAs, such as the m6A modification investigated in this study, play a crucial role in regulating various biological processes, including splicing, transport, stability, and translation. These modifications are mediated by a group of enzymes known as “writers”, “erasers”, and “readers” [24]. However, the specific mechanisms through which these enzymes modify or recognize their target mRNAs remain an important scientific question due to the widespread occurrence of post-transcriptional modifications in host mRNAs. RBPs possess sequence-specific or selective RNA binding activity and it is speculated that they interact with the corresponding enzymes of post-transcriptional modifications and confer mRNA specificity for their enzymatic activities and modification. In this study, DHX58, an RBP, was found to specifically bind to Gpx4 mRNA and recruit YTHDC2 in an m6A-dependent manner, thereby synergistically enhancing its translation. This finding provides a representative working model supporting the aforementioned hypothesis. Nevertheless, considering that there exists a diverse range of proteins or protein families capable of binding RNAs with specificity towards RNA-binding with specific sequences or structures, it is plausible that these proteins may also associate with post-transcriptionally modified enzymes and provide these enzymes with RNA specificity. Exploring this hypothesis could lead to intriguing future research directions within the fields of RNA modification and protein-RNA interactions.
Conclusions
In summary, this study highlights a promising strategy for preventing hepatic ferroptosis and I/R injury by inhibiting ferroptosis in hepatocytes through upregulating the protein levels of DHX58 expression and its downstream GPX4. Mechanistically, DHX58, possessing RNA-binding activity, constitutively associates with the mRNA of Gpx4 and recruits the m6A reader YTHDC2 to promote the translation of Gpx4 mRNA in an m6A-dependent manner, thus enhancing GPX4 protein levels and effectively preventing hepatic ferroptosis.
Availability of data and materials
The materials of this study are available from the corresponding author (Jin Hou) upon reasonable request and through collaborative investigations. The accession numbers of the RNA-seq data: PRJNA807799, scRNA-seq data: GSE231934, MeRIP-seq data: GSE231842, RIP-seq data: GSE231843, and 4D label free proteomics analysis data in ProteomeXchange dataset: PXD042083. All the unprocessed gels and images, and the original source data for all figures are available at Mendeley Data Reserved https://data.mendeley.com/datasets/g2tk2262sz/1.
Abbreviations
- ACSL4:
-
Acyl-CoA synthetase long-chain family member 4
- ALKBH5:
-
AlkB homolog 5
- ALT:
-
Alanine aminotransferase
- AST:
-
Aspartate aminotransferase
- ActD:
-
Actinomycin D
- BCA:
-
Bicinchoninic acid
- BHA:
-
Butylated hydroxyanisole
- CARD:
-
Caspase-recruiting domain
- CQ:
-
Chloroquine
- Co-IP:
-
Co-immunoprecipitation
- COX2:
-
Cyclooxygenase-2
- CTD:
-
C-terminal domain
- DAPI:
-
4’,6-diamidino-2-phenylindole
- DDX/DHX:
-
DEAD/DExH-box helicase
- DHE:
-
Dihydroethidium
- DHX58:
-
DExH-box helicase 58
- FTO:
-
Fat mass and obesity associated gene
- GPX4:
-
GSH peroxidase 4
- GSH:
-
Glutathione
- GSSG:
-
Oxidized GSH
- HE:
-
Hematoxylin and eosin
- H/R:
-
Hypoxia/re-oxygenation
- 4-HNE:
-
4-hydroxynonenal
- IFN:
-
Interferon
- IL-1β:
-
Interleukin 1β
- IL-6:
-
Interleukin 6
- IP:
-
Immunoprecipitation
- IP-MS:
-
IP-mass spectrometry
- I/R:
-
Ischemia/reperfusion
- ISG:
-
IFN-stimulated gene
- KEGG:
-
Kyoto encyclopedia of genes and genomes
- LGP2:
-
Laboratory of genetics and physiology 2
- LPO:
-
Lipid peroxide
- Ly6G:
-
Lymphocyte antigen 6 complex locus G
- MCP1:
-
Monocyte chemoattractant protein 1
- MDA:
-
Malondialdehyde
- MeRIP-seq:
-
m6A immunoprecipitation and sequencing
- METTL3:
-
Methyltransferase-like 3
- m6A:
-
N6-methyladenosine
- NAC:
-
N-acetylcysteine
- NPC:
-
Non-parenchymal cell
- Nec-1:
-
Necrostatin-1
- OD:
-
Optical density
- pDCs:
-
Plasmacytoid dendritic cells
- qRT-PCR:
-
Quantitative reverse-transcription polymerase chain reaction
- QuSAGE:
-
Quantitative set analysis of gene expression
- RBP:
-
RNA-binding protein
- RD:
-
Repressor domain
- RIG-I:
-
Retinoic acid-inducible gene-I
- RIP-seq:
-
RNA immunoprecipitation-sequencing
- RIP-qRT-PCR:
-
RNA immunoprecipitation-qRT-PCR
- ROS:
-
Reactive oxygen species
- scRNA-seq:
-
Single-cell RNA sequencing
- SD:
-
Standard deviation
- SLC7A11:
-
Solute carrier family 7 member 11
- UAMP:
-
Uniform manifold approximation and projection
- YTHDF1:
-
YTH N6-methyladenosine RNA binding protein F1
- YTHDC2:
-
YT521-B homology domain containing 2
References
Stockwell BR. Ferroptosis turns 10: emerging mechanisms, physiological functions, and therapeutic applications. Cell. 2022;185(14):2401–21.
Jiang X, Stockwell BR, Conrad M. Ferroptosis: mechanisms, biology and role in disease. Nat Rev Mol Cell Biol. 2021;22(4):266–82.
Tang R, Xu J, Zhang B, Liu J, Liang C, Hua J, et al. Ferroptosis, necroptosis, and pyroptosis in anticancer immunity. J Hematol Oncol. 2020;13(1):110.
Stockwell BR, Friedmann Angeli JP, Bayir H, Bush AI, Conrad M, Dixon SJ, et al. Ferroptosis: a regulated cell death nexus linking metabolism, redox biology, and disease. Cell. 2017;171(2):273–85.
Fang X, Zhang J, Li Y, Song Y, Yu Y, Cai Z, et al. Malic enzyme 1 as a novel anti-ferroptotic regulator in hepatic ischemia/reperfusion injury. Adv Sci. 2023;10(13):e2205436.
Tang D, Chen X, Kang R, Kroemer G. Ferroptosis: molecular mechanisms and health implications. Cell Res. 2021;31(2):107–25.
Wu Y, Jiao H, Yue Y, He K, Jin Y, Zhang J, et al. Ubiquitin ligase E3 HUWE1/MULE targets transferrin receptor for degradation and suppresses ferroptosis in acute liver injury. Cell Death Differ. 2022;29(9):1705–18.
Pan Y, Wang X, Liu X, Shen L, Chen Q, Shu Q. Targeting ferroptosis as a promising therapeutic strategy for ischemia-reperfusion injury. Antioxidants. 2022;11(11):2196.
Paine I, Posey JE, Grochowski CM, Jhangiani SN, Rosenheck S, Kleyner R, et al. Paralog studies augment gene discovery: DDX and DHX genes. Am J Hum Genet. 2019;105(2):302–16.
Heaton SM, Gorry PR, Borg NA. DExD/H-box helicases in HIV-1 replication and their inhibition. Trends Microbiol. 2023;31(4):393–404.
Shen L, Pelletier J. General and target-specific DExD/H RNA helicases in eukaryotic translation initiation. Int J Mol Sci. 2020;21(12):4402.
Overwijn D, Hondele M. DEAD-box ATPases as regulators of biomolecular condensates and membrane-less organelles. Trends Biochem Sci. 2023;48(3):244–58.
Xu L, Wang W, Li Y, Zhou X, Yin Y, Wang Y, et al. RIG-I is a key antiviral interferon-stimulated gene against hepatitis E virus regardless of interferon production. Hepatology. 2017;65(6):1823–39.
Jiang Z, Wei F, Zhang Y, Wang T, Gao W, Yu S, et al. IFI16 directly senses viral RNA and enhances RIG-I transcription and activation to restrict influenza virus infection. Nat Microbiol. 2021;6(7):932–45.
Rehwinkel J, Gack MU. RIG-I-like receptors: their regulation and roles in RNA sensing. Nat Rev Immunol. 2020;20(9):537–51.
Gillich N, Zhang Z, Binder M, Urban S, Bartenschlager R. Effect of variants in LGP2 on MDA5-mediated activation of interferon response and suppression of hepatitis D virus replication. J Hepatol. 2023;78(1):78–89.
Loo YM, Gale M Jr. Immune Signaling by RIG-I-like receptors. Immunity. 2011;34(5):680–92.
Kim N, Now H, Nguyen NTH, Yoo JY. Multilayered regulations of RIG-I in the anti-viral signaling pathway. J Microbiol. 2016;54(9):583–7.
Zhu Z, Zhang X, Wang G, Zheng H. The laboratory of genetics and physiology 2: emerging insights into the controversial functions of this RIG-I-like receptor. Biomed Res Int. 2014;2014:960190.
Li Z, Zhou Y, Jia K, Yang Y, Zhang L, Wang S, et al. JMJD4-demethylated RIG-I prevents hepatic steatosis and carcinogenesis. J Hematol Oncol. 2022;15(1):161.
Hou J, Zhou Y, Zheng Y, Fan J, Zhou W, Ng IO, et al. Hepatic RIG-I predicts survival and interferon-α therapeutic response in hepatocellular carcinoma. Cancer Cell. 2014;25(1):49–63.
Zheng Q, Hou J, Zhou Y, Li Z, Cao X. The RNA helicase DDX46 inhibits innate immunity by entrapping m6A-demethylated antiviral transcripts in the nucleus. Nat Immunol. 2017;18(10):1094–103.
Lin X, Wang F, Chen J, Liu J, Lin YB, Li L, et al. N6-methyladenosine modification of CENPK mRNA by ZC3H13 promotes cervical cancer stemness and chemoresistance. Mil Med Res. 2022;9(1):19.
Boulias K, Greer EL. Biological roles of adenine methylation in RNA. Nat Rev Genet. 2023;24(3):143–60.
Zeng C, Huang W, Li Y, Weng H. Roles of METTL3 in cancer: mechanisms and therapeutic targeting. J Hematol Oncol. 2020;13(1):117.
Mao Y, Dong L, Liu XM, Guo J, Ma H, Shen B, et al. m6A in mRNA coding regions promotes translation via the RNA helicase-containing YTHDC2. Nat Commun. 2019;10(1):5332.
Zhou Y, Jia K, Wang S, Li Z, Li Y, Lu S, et al. Malignant progression of liver cancer progenitors requires lysine acetyltransferase 7-acetylated and cytoplasm-translocated G protein GαS. Hepatology. 2023;77(4):1106–21.
Li Z, Zhou Y, Zhang L, Jia K, Wang S, Wang M, et al. microRNA-199a-3p inhibits hepatic apoptosis and hepatocarcinogenesis by targeting PDCD4. Oncogenesis. 2020;9(10):95.
Su M, Pan T, Chen QZ, Zhou WW, Gong Y, Xu G, et al. Data analysis guidelines for single-cell RNA-seq in biomedical studies and clinical applications. Mil Med Res. 2022;9(1):68.
Yao RQ, Zhao PY, Li ZX, Liu YY, Zheng LY, Duan Y, et al. Single-cell transcriptome profiling of sepsis identifies HLA-DRlowS100Ahigh monocytes with immunosuppressive function. Mil Med Res. 2023;10(1):27.
Li PH, Kong XY, He YZ, Liu Y, Peng X, Li ZH, et al. Recent developments in application of single-cell RNA sequencing in the tumour immune microenvironment and cancer therapy. Mil Med Res. 2022;9(1):52.
Bai YM, Yang F, Luo P, Xie LL, Chen JH, Guan YD, et al. Single-cell transcriptomic dissection of the cellular and molecular events underlying the triclosan-induced liver fibrosis in mice. Mil Med Res. 2023;10(1):7.
Charni-Natan M, Goldstein I. Protocol for primary mouse hepatocyte isolation. STAR Protoc. 2020;1(2):100086.
Hou J, Lin L, Zhou W, Wang Z, Ding G, Dong Q, et al. Identification of miRNomes in human liver and hepatocellular carcinoma reveals miR-199a/b-3p as therapeutic target for hepatocellular carcinoma. Cancer Cell. 2011;19(2):232–43.
Panda AC, Martindale JL, Gorospe M. Polysome fractionation to analyze mRNA distribution profiles. Bio Protoc. 2017;7(3):e2126.
Chouchani ET, Pell VR, James AM, Work LM, Saeb-Parsy K, Frezza C, et al. A unifying mechanism for mitochondrial superoxide production during ischemia-reperfusion injury. Cell Metab. 2016;23(2):254–63.
van der Veen AG, Maillard PV, Schmidt JM, Lee SA, Deddouche-Grass S, Borg A, et al. The RIG-I-like receptor LGP2 inhibits dicer-dependent processing of long double-stranded RNA and blocks RNA interference in mammalian cells. EMBO J. 2018;37(4):e97479.
Yamada N, Karasawa T, Wakiya T, Sadatomo A, Ito H, Kamata R, et al. Iron overload as a risk factor for hepatic ischemia-reperfusion injury in liver transplantation: potential role of ferroptosis. Am J Transpl. 2020;20(6):1606–18.
Liao P, Wang W, Wang W, Kryczek I, Li X, Bian Y, et al. CD8+ T cells and fatty acids orchestrate tumor ferroptosis and immunity via ACSL4. Cancer Cell. 2022;40(4):365–78.e6.
Liggett JR, Kang J, Ranjit S, Rodriguez O, Loh K, Patil D, et al. Oral N-acetylcysteine decreases IFN-γ production and ameliorates ischemia-reperfusion injury in steatotic livers. Front Immunol. 2022;13:898799.
Zhang H, Jiao W, Cui H, Sun Q, Fan H. Combined exposure of alumina nanoparticles and chronic stress exacerbates hippocampal neuronal ferroptosis via activating IFN-γ/ASK1/JNK signaling pathway in rats. J Hazard Mater. 2021;411:125179.
Wei TT, Zhang MY, Zheng XH, Xie TH, Wang W, Zou J, et al. Interferon-γ induces retinal pigment epithelial cell ferroptosis by a JAK1-2/STAT1/SLC7A11 signaling pathway in age-related macular degeneration. FEBS J. 2022;289(7):1968–83.
Yang Y, Ma Y, Yu S, Lin Z, Yan C, Wang Y, et al. TIPE2 knockout reduces myocardial cell damage by inhibiting IFN-γ-mediated ferroptosis. Biochim Biophys Acta Mol Basis Dis. 2023;1869(1):166566.
Lv YW, Du Y, Ma SS, Shi YC, Xu HC, Deng L, et al. Proanthocyanidins attenuates ferroptosis against influenza-induced acute lung injury in mice by reducing IFN-γ. Life Sci. 2023;314:121279.
Klune JR, Bartels C, Luo J, Yokota S, Du Q, Geller DA. IL-23 mediates murine liver transplantation ischemia-reperfusion injury via IFN-γ/IRF-1 pathway. Am J Physiol Gastrointest Liver Physiol. 2018;315(6):G991–1002.
Zhao G, Wang S, Wang Z, Sun A, Yang X, Qiu Z, et al. CXCR6 deficiency ameliorated myocardial ischemia/reperfusion injury by inhibiting infiltration of monocytes and IFN-γ-dependent autophagy. Int J Cardiol. 2013;168(2):853–62.
Castellaneta A, Yoshida O, Kimura S, Yokota S, Geller DA, Murase N, et al. Plasmacytoid dendritic cell-derived IFN-α promotes murine liver ischemia/reperfusion injury by induction of hepatocyte IRF-1. Hepatology. 2014;60(1):267–77.
Lo CM, Liu CL, Chan SC, Lam CM, Poon RT, Ng IO, et al. A randomized, controlled trial of postoperative adjuvant interferon therapy after resection of hepatocellular carcinoma. Ann Surg. 2007;245(6):831–42.
Sun HC, Tang ZY, Wang L, Qin LX, Ma ZC, Ye QH, et al. Postoperative interferon alpha treatment postponed recurrence and improved overall survival in patients after curative resection of HBV-related hepatocellular carcinoma: a randomized clinical trial. J Cancer Res Clin Oncol. 2006;132(7):458–65.
Li LY, Han J, Wu L, Fang C, Li WG, Gu JM, et al. Alterations of gut microbiota diversity, composition and metabonomics in testosterone-induced benign prostatic hyperplasia rats. Mil Med Res. 2022;9(1):12.
Liu H, Zhang H, Wu X, Ma D, Wu J, Wang L, et al. Nuclear cGAS suppresses DNA repair and promotes tumorigenesis. Nature. 2018;563(7729):131–6.
Acknowledgements
We thank Prof. Zhengxin Wang and Dr. Jianhua Li from Huashan Hospital, Fudan University for kindly providing the human liver samples.
Funding
This work was supported by grants from the National Key Research and Development Program of China (2023YFC2505900), the National Natural Science Foundation of China (92269204, 82171755, 92369106, 82171749, 82171811, 82073184), the Military Outstanding Youth Program (2020QN06119, 01-SWKJYCJJ07, 23SWAQ53), and the Program of Leading Talents in Shanghai, and Shanghai Shuguang Program (20SG39).
Author information
Authors and Affiliations
Contributions
KWJ, RQY, YWF, DJZ, YZ and MJW performed the experiments and contributed equally to the whole study. LYZ, YD, ZXL, SYW, MW, YHL, LXZ, TL, LCG, SL, YYY, SXW and YZY provided reagents, performed experiments and analyzed the data. JH and YMY analyzed the data and wrote the paper. JH designed the study. All authors have read and approved the final manuscript.
Corresponding authors
Ethics declarations
Ethics approval and consent to participate
All animal experiments were performed in accordance with the National Institute of Health Guide for the Care and Use of Laboratory Animals, with the approval of the Scientific Investigation Board of Naval Medical University, Shanghai, China. The human subject study was approved by the human ethics committee of Huashan Hospital, Fudan University (KY2021-449).
Consent for publication
Not applicable.
Competing interests
The authors declared no competing interests.
Supplementary Information
Additional file 1: Table S1
The primers used in this study. Fig. S1 DHX58 expression is markedly decreased post I/R in the liver. Fig. S2 ROS decreases DHX58 expression in hepatocytes. Fig. S3 Hepatocyte-specific DHX58 deficiency promotes I/R‑induced liver injury and inflammation. Fig. S4 scRNA-seq analysis of Dhx58f/f and Dhx58hep‑/- livers post I/R. Fig. S5 Dhx58hep‑/- promotes ferroptosis in hepatocyte during liver I/R injury. Fig. S6 DHX58 inhibits ferroptosis in hepatocyte during liver I/R injury Fig. S7 DHX58 enhances GPX4 protein level to suppress ferroptosis. Fig. S8 DHX58 associates YTHDC2 to read and promote the translation of Gpx4 mRNA in an m6A-dependent manner. Fig. S9 Pretreatment with IFN-α can inhibit hepatic ferroptosis by stimulating DHX58.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/. The Creative Commons Public Domain Dedication waiver (http://creativecommons.org/publicdomain/zero/1.0/) applies to the data made available in this article, unless otherwise stated in a credit line to the data.
About this article
Cite this article
Jia, KW., Yao, RQ., Fan, YW. et al. Interferon-α stimulates DExH-box helicase 58 to prevent hepatocyte ferroptosis. Military Med Res 11, 22 (2024). https://doi.org/10.1186/s40779-024-00524-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/s40779-024-00524-9