Abstract
Background
Seed dormancy and germination determine wheat resistance to pre-harvest sprouting and thereby affect grain yield and quality. Arabidopsis VQ genes have been shown to influence seed germination; however, the functions of wheat VQ genes have not been characterized.
Results
We identified 65 TaVQ genes in common wheat and named them TaVQ1–65. We identified 48 paralogous pairs, 37 of which had Ka/Ks values greater than 1, suggesting that most TaVQ genes have experienced positive selection. Chromosome locations, gene structures, promoter element analysis, and gene ontology annotations of the TaVQs showed that their structures determined their functions and that structural changes reflected functional diversity. Transcriptome-based expression analysis of 62 TaVQ genes and microarray analysis of 11 TaVQ genes indicated that they played important roles in diverse biological processes. We compared TaVQ gene expression and seed germination index values among wheat varieties with contrasting seed dormancy and germination phenotypes and identified 21 TaVQ genes that may be involved in seed dormancy and germination.
Conclusions
Sixty-five TaVQ proteins were identified for the first time in common wheat, and bioinformatics analyses were used to investigate their phylogenetic relationships and evolutionary divergence. qRT-PCR data showed that 21 TaVQ candidate genes were potentially involved in seed dormancy and germination. These findings provide useful information for further cloning and functional analysis of TaVQ genes and introduce useful candidate genes for the improvement of PHS resistance in wheat.
Similar content being viewed by others
Background
Wheat is a widely cultivated gramineous plant and one of the three most important cereals in the world [1]. It is a heterologous hexaploid derived from three closely related ancestors that have undergone two rounds of natural hybridization. Therefore, the large and complex genome of wheat (17 Gb) poses a significant challenge for wheat genome research [2, 3]. The completion of a whole genome sequence for wheat based on single chromosome sequencing has laid the foundation for wheat genomics research and wheat gene family identification.
Valine-glutamine (VQ) proteins are a class of plant-specific proteins with five highly conserved amino acids in the core FxxxVQxLTG sequence of the VQ motif [4], in which x represents any amino acid (aa) and VQ is a highly conserved pair of aa residues. Research on the VQ proteins has shown that the last three amino acids in almost all species are LTG, although some species have other variants, including FTG, ITG, LTA, and VTG [5]. In some VQ proteins of Gramineae species such as rice, maize, and Moso bamboo, VQ has mutated to VH in the conserved domain [5, 6]. VQ proteins are generally less than 300 aa in length and contain no or few introns [7]. To date, 34, 40, 61, 18, and 74 VQ proteins have been identified in Arabidopsis, rice, maize, grape, and soybean, respectively [6, 8,9,10,11]. According to bioinformatics predictions and experimental verification, some Arabidopsis VQ proteins are located in the nucleus, some in the plastid, and a few partly in the mitochondria [12].
VQ proteins play important roles in the regulation of plant growth and development and the response to abiotic and biotic stress [5,6,7, 13,14,15,16,17]. For instance, AtCaMBP25 (AtVQ15) negatively regulates osmotic stress response during the early stages of seed germination and growth in Arabidopsis [13]. Likewise, AtVQ9 expression responds strongly to NaCl treatment, and its mutation enhances salt stress tolerance in Arabidopsis [16]. VQ54 and VQ19 in maize, as well as VQ2, VQ16, and VQ20 in rice, are highly expressed under drought induction [6, 11]. Soybean VQ6 and VQ53 are highly expressed in roots and stems under low nitrogen conditions [10]. SIB1 (Sigma factor binding protein 1, also known as AtVQ23) was the first VQ motif protein discovered in Arabidopsis and participates in plant disease resistance signaling pathways [15]. AtVQ21 (MSK1) transgenic plants show enhanced resistance to the pathogen Pseudomonas syringae but reduced resistance to Botrytis cinerea [13, 18]. AtVQ22 negatively regulates JA-mediated disease resistance signaling pathways [19], and rice VQ22 shows high expression levels after rice blast infection [5]. AtVQ14 (IKU1) participates in the regulation of endosperm development, thereby affecting the size of Arabidopsis seeds [20]. AtVQ29 is involved in the photomorphogenesis of Arabidopsis seedlings and flowering time regulation [9]. In addition, the growth of VQ17, VQ18, VQ8, and VQ22 transgenic Arabidopsis plants is inhibited, indicating that these genes play crucial roles in plant growth and development [8].
VQ proteins came to the attention of researchers because of their interactions with WRKY transcription factors, which are involved in regulating the plant’s defense response system [13, 15]. WRKY transcription factors belong to a large gene family and are ubiquitous in plants. Studies have shown that WRKY transcription factors are widely involved in plant growth and development and in resistance to adverse conditions [21,22,23,24,25,26,27,28,29,30,31,32,33,34,35,36,37,38]. For example, AtVQ14 and AtWRKY10 interact to form a protein complex that affects seed size in Arabidopsis [20]. AtVQ15 and AtWRKY25 interact and participate in high salt and osmotic stress response [13]. The interaction between AtVQ22 and AtWRKY28 negatively regulates JA-mediated disease resistance signaling pathways [19]. AtVQ23 (SIB1) and AtVQ16 (SIB2) interact with WRKY33 to enhance the binding capacity of WRKY33 to the W-box, thereby regulating plant disease resistance [6, 15]. AtVQ21 can form ternary complexes with AtWRKY33 and AtMPK4 to regulate plant growth and disease resistance [4, 39]. In brief, VQ proteins are transcriptional regulatory cofactors that participate in growth and developmental processes and stress resistance through their interactions with transcription factors. However, until now, the VQ gene family has not been characterized in common wheat.
Pre-harvest sprouting (PHS) refers to the germination of wheat seeds within the spike of the mother plant that occurs in rainy or high moisture conditions before harvest. A series of physiological and biochemical reactions take place in wheat grains when PHS occurs. The activity of hydrolases such as amylase and proteolytic enzymes is enhanced, leading to starch and protein degradation and seriously affecting wheat processing quality and utilization value. In the international wheat market, when the germination rate of commercial wheat reaches 5%, it is regarded as feed wheat and its price is reduced, causing serious economic losses to producers [40, 41]. Seed dormancy and germination traits determine wheat PHS resistance: wheat varieties with higher levels of dormancy or lower germination percentages show higher resistance to PHS. Therefore, the identification of candidate genes that control seed dormancy and germination may help to reduce yield and quality losses caused by PHS. Previously, in Arabidopsis, VQ18 and VQ26 were found to be involved in seed germination via the ABA signaling pathway [42]. However, the functions of VQ genes in common wheat are largely unknown.
The objectives of this study were to identify TaVQ genes and to perform bioinformatics analysis, including phylogenetic tree construction and characterization of gene structures, conserved domains, chromosome positions, expression patterns, and promoter elements. In addition, we measured the expression levels of TaVQ genes in wheat varieties with contrasting seed dormancy and germination phenotypes by qRT-PCR to identify TaVQ gene family members that were potentially involved in seed dormancy and germination.
Results
Identification and attribute analysis of VQ candidate genes in wheat
A total of 65 TaVQ genes were identified, mapped to wheat chromosomes, and named TaVQ1–TaVQ65. The length of the encoded proteins ranged from 127 to 723 aa, with an average length of 220 aa. Their MWs ranged from 13,377.16 Da (TaVQ52) to 61,926.76 Da (TaVQ1). Information on chromosome positions, ORF lengths, and exon numbers is provided in Table 1. The majority of genes included one exon, and only four genes (TaVQ13/-17/-18/-53) contained two exons.
Phylogenetic trees of VQ proteins from wheat, maize, poplar, rice, and Arabidopsis
To explore the evolutionary relationships among VQ genes from wheat, Arabidopsis, rice, poplar, and maize, we downloaded published VQ protein sequences from these species (Table S1) [8,9,10] and constructed a phylogenetic tree (Fig. 1). Based on the original division and naming of VQ subfamilies in Arabidopsis and rice, we divided the 251 VQ genes (34 AtVQ genes, 40 OsVQ genes, 51 PtVQ genes, 61 ZmVQ genes, and 65 TaVQ genes) into seven subfamilies (VQI, VQII, VQIII, VQIV, VQV, VQVI, and VQVII). The VQII subfamily contained the largest number of genes (28) (Fig. 1). Members of the VQ family from rice, maize, and wheat, which belong to the Gramineae, were interspersed, whereas those of Arabidopsis, a model plant from the Cruciferae, formed separate clades, probably due to the relatively distant relationship between monocots and dicots.
Phylogeny of VQ proteins from wheat, rice, poplar, maize, and Arabidopsis. The 65 TaVQ proteins, 40 OsVQ proteins, 51PtVQ proteins, 61 ZmVQ proteins, and 34AtVQ proteins are clustered into seven subfamilies. Details of VQ genes from Arabidopsis, maize, poplar and rice are listed in Table S1. The red circle, black square, yellow triangle, purple diamond and green inverted triangle represent the VQ gene families of wheat, rice, poplar, maize and Arabidopsis respectively. Different colors of inner ring and outer ring represent different subfamilies
Structural analysis of the VQ gene family
We constructed a wheat VQ phylogenetic tree and a gene structure diagram (Fig. 2a). The structure of each VQ gene contained one to three parts: the untranslated region (yellow rectangle), the exon region (green rectangle), and the intron region (solid gray line). Among the 48 TaVQ paralogous pairs (Table 2), only two pairs (TaVQ10/-17 and TaVQ53/-58) differed in intron number, having lost or gained one intron (Fig. 2A and Table S2). Further analysis revealed that 94% of the TaVQ genes had no introns, and only four genes (TaVQ13/-17/-18/-53) contained one intron. This result is consistent with previous studies in other species: 78%, 88%, 89% and 93% of the VQ genes in poplar, Arabidopsis, maize, and rice have no introns, respectively, whereas only 28% of moss VQ genes have no introns (Fig. 2b). Based on comparisons of many species, including angiosperms (rice, poplar, soybean, Chinese cabbage, etc.) and bryophytes (moss), we speculate that most VQ genes tend to lose introns during long-term evolution [6, 8, 9, 43, 44].
Gene structure analysis of TaVQ genes. a Phylogenetic relationships and gene structures of TaVQ genes. Exons, introns, and untranslated regions (UTRs) are indicated by green rectangles, gray lines, and yellow rectangles, respectively. Coloured boxes indicate the subfamily based on the phylogenetic analysis. b The numbers of VQ genes and the numbers of VQ genes without introns in different species. Pp: Moss, At: Arabidopsis, Pt: Poplar, Br: Chinese Cabbage, Gm: Soybean, Os: Rice, Zm: Maize, Vv: Grape, Pe: Moso bamboo, Ta: Wheat
Using published information on characteristics of the VQ domain as a reference, we aligned the protein sequences of wheat and analyzed their VQ domains. The 65 TaVQ proteins all contained conserved VQ domains, but they differed slightly and could be grouped into three types: FxxxVQxLTG (52/65), FxxxVQxFTG (10/65), and FxxxVQxITG (3/65) (Fig. 3a). We further analyzed the VQ domains from multiple species (Fig. 3b) and found that the FxxxVQxLTG sequence was most prevalent and that two additional VQ domain types (LTG/FTG) were also common. There were differences in VQ domain sequence between monocots and dicots. In addition to the more common domain sequences, monocot VQ domains also included ITG, ATG, and LTA, and dicot VQ domains included LTS, LTD, YTG, LTR, and LTV (Table S3).
Multiple sequence alignment of TaVQ proteinsand domain type analysis in different species. a Multiple sequence alignment of VQ proteins in wheat. b VQ domain type in different species. Pp: Moss, At: Arabidopsis, Pt: Poplar, Br: Chinese Cabbage, Gm: Soybean, Os: Rice, Zm: Maize, Vv: Grape, Pe: Moso bamboo, Ta: Wheat. c Schematic representation of 20 conserved motifs in the TaVQ genes. Different colored boxes represent different motifs. Box lengths are not proportional to actual motif size
A total of 20 conserved motifs were identified in the TaVQ gene family (Table S4). All 65 TaVQ proteins shared one conserved motif (core motif 1, Motif 1) (Fig. 3c). The MEME diagram showed that wheat VQ genes from the same subfamily tended to share the same conserved motifs. Only 7 of the 48 TaVQ paralogous pairs (TaVQ12/-15, TaVQ32/-36, TaVQ32/-41, TaVQ44/-49, TaVQ47/-49, TaVQ55/-60, and TaVQ60/-65) differed in their motifs.
Evolution and divergence of the VQ gene family in wheat, rice, and maize
In total, 48 homologous pairs were identified in wheat, 22 in wheat and rice, and 14 in wheat and maize (Table 2). The Ks (number of synonymous substitutions per synonymous site) values of the wheat paralogous pairs ranged from 0.0163 to 1.5197, indicating that duplication events occurred in this species approximately 1.2538 to 116.8985 million years ago (MYA). The Ks values of orthologous pairs from wheat and rice ranged from 0.5821 to 1.6479, indicating that duplication events occurred approximately 44.78 to 126.7631 MYA. The Ks values of orthologous pairs from wheat and maize ranged from 0.6453 to 1.0257, indicating that the duplication events occurred approximately 49.6415 to 78.8962 MYA (Table 3).
To investigate the role of natural selection in the evolution of the VQ gene family in Gramineae, we analyzed the Ka (number of non-synonymous substitutions per non-synonymous site)/Ks ratios of all homologous pairs and generated sliding window graphs (Figure S1 and Table 3). Among the 48 paralogous pairs, 11 had Ka/Ks ratios less than one, and 37 pairs had Ka/Ks ratios greater than one, indicating that wheat VQ genes were mainly under positive selection during the evolutionary process. The Ka/Ks ratio of all orthologous pairs was greater than one, indicating that the VQ gene family in wheat, rice, and maize had primarily undergone positive selection.
Expression pattern analysis of the TaVQ gene family
Transcriptome data (FPKM values) were obtained for all TaVQ genes with the exception of TaVQ13/-18/-45 (Fig. 4a and Table S5). The expression patterns of VQ genes differed among varieties and within time periods in the same variety. Most TaVQ genes were highly expressed in J411, especially at 4 h after seed imbibition, and only four genes (TaVQ4/-7/-8/-20) were expressed at a low level. It is worth noting that TaVQ8 and TaVQ20 were both highly expressed in HMC21 and expressed at a low level in J411. We further analyzed the expression of 48 paralogous gene pairs. Only one pair showed a similar expression pattern, whereas the rest were differentially expressed among varieties and within different time periods of the same variety.
Expression profiles of TaVQ genes in different tissues and at different developmental stages. a Heatmap shows hierarchical clustering of the 62 TaVQ genes based on transcriptome results. b Heatmap shows hierarchical clustering of 11TaVQ genes from different tissues. Abbreviations represent specific developmental stages. GSC, germinating seed coleoptile; GSR, germinating seed root; GSE, germinating seed, embryo; SR, seedling root; SC, seedling crown; SL, seedling leaf; II, immature inflorescence; Fba, floral bracts before anthesis; Pba, pistil before anthesis; Aba, anthers before anthesis; 3–5 DAP C, 3–5 DAP caryopsis; 22 DAP EM, 22 DAP embryo; 22 DAP EN, 22 DAP endosperm
Microarray data were obtained for 11 TaVQ genes to further investigate wheat VQ gene expression (Fig. 4b and Table S6). TaVQ16, TaVQ31, and TaVQ35 were highly expressed in Aba and at 22 DAP EM (22 days after planting—embryo), but they showed little expression elsewhere. Further analysis of paralogous pairs showed that three pairs (TaVQ55/-60, TaVQ55/-65, and TaVQ60/-65) had similar expression patterns in different tissues.
Promoter analysis and gene ontology annotation of the TaVQ gene family
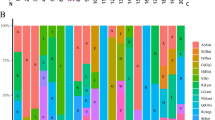
Two categories of response element were analyzed in the promoter regions of the TaVQ genes (Fig. 5a and Table S7). The first category included elements associated with biotic stress, such as ABRE, CGTCA motif, TGACG motif, TGA element, AuxRR core, TCA element, GARE motif, and P-box. The second category included elements associated with abiotic stress, such as MBS, LTR, and TC-rich repeats. The most common biotic stress response elements in the TaVQ promoters were associated with methyl jasmonate (CGTCA motif and TGACG motif) (42.77%) and ABA (ABRE) (41.85%) (Fig. 5b). The drought-associated MBS element (4.31%) was the most common abiotic stress response element (Fig. 5c).
Cis-acting element analysis of the promoter regions of TaVQ genes. Based on functional annotation data, cis-acting elements were classified into two major classes: phytohormone responsive elements (i.e. those responsive to ABA, auxin, GA, MeJA, and/or SA) and abiotic stress responsive elements (e.g. those involved in plant defense, drought stress response, and/or low temperature stress response). a Percentage of total cis-acting elements in the promoter region of the TaVQ gene. b and c The percentage of cis-acting elements in different categories
The 65 TaVQ genes were annotated with 15 GO terms (Fig. 6 and Table S8): three, three, and nine terms in the molecular function, cellular component, and biological process categories, respectively. Among these terms, GO:0,005,634 (cellular component), GO:0,003,674 (molecular function), and GO:0,008,150 (biological process) were most common and were assigned to 28, 21, and 13 genes, respectively.
Chromosome locations and subcellular localization predictions for the TaVQ gene family
The TaVQ genes were unevenly distributed on wheat chromosomes 1–7, and no TaVQ genes were present on chromosomes 1B and 1D (Figure S2). One gene was located on chromosome 1A, five were located on chromosomes 5B, 7A, 7B, and 7D, and two to four were located on each of the other chromosomes. We defined a single gene cluster as a chromosomal region of less than 200 kb that contained two or more TaVQ genes [45]. Two gene clusters containing six genes were identified on chromosomes 4A and 4B (Figure S2).
Subcellular localization prediction indicated that the TaVQ proteins were present in three locations. Most were predicted to be located in the periplasmic region (47, 72.3%), some in the extracellular region (15, 23.1%), and the rest in the cytoplasm (3, 4.6%) (Table S9).
Responses of TaVQ genes to water imbibition
We investigated the responses of 65 TaVQ genes (Table 1 and Table S10) in six wheat varieties with different seed dormancy and germination phenotypes after water imbibition for 0, 6, and 10 h. Seeds from three highly dormant varieties (HMC21, YXM, and SNTT) showed no seed germination, whereas partial seeds from three low-dormancy varieties (J411, ZY9507, and ZM895) germinated after 10 h of imbibition with an average germination index (GI) of 0.33, 0.31, and 0.41, respectively (Table S11). We found that the TaVQ genes were differentially expressed in the six wheat varieties. The expression levels of 13 genes (TaVQ8/-9/-13/-17/-25/-32/-34/-43/-48/-49/-53/-59/-62) were higher in the low-dormancy varieties than in the high-dormancy varieties. Eight genes showed the opposite expression trend (TaVQ4/-16/-20/-35/-38/-42/-51/-56) (Fig. 7).
Discussion
The plant-specific VQ proteins initially attracted attention due to their interactions with WRKY transcription factors [15]. Additional in-depth studies showed that the VQ gene family not only participated in responses to biotic and abiotic stress, but was also involved in the regulation of plant growth and development [5,6,7, 13,14,15,16,17]. Genome-wide surveys of VQ proteins have now been performed in a number of species, although functional research has remained focused on Arabidopsis. VQ genes have not previously been characterized in wheat, and we therefore performed basic bioinformatics analyses to better understand the VQ gene family in wheat.
We identified 65 VQ genes from wheat and classified them into seven subfamilies. The VQ genes of five species (wheat, rice, maize, poplar, and Arabidopsis) were distributed in each subfamily, but the number of subfamily members differed among species, indicating that the VQ genes have developed in multiple directions over the course of evolution. The VQ genes of monocots (rice, maize, and wheat) were interspersed and clustered together, whereas the VQ genes of Arabidopsis and poplar were clustered into separate clades, indicating that proteins encoded by wheat VQ genes were highly similar to those of rice and maize [8, 9]. These results highlight the evolutionary conservation of the VQ gene family.
Phylogenetic trees represent the genetic relationships among gene families from different species and reflect the similarity of protein-coding genes. From structural analysis of the VQ gene family, we found that most VQ genes were intron-free [6, 8,9,10,11, 43, 44]. Based on comparisons of several species, we speculate that this gene family tends to lose introns during evolution. Amino acid sequence alignment and motif analysis indicated that the sequences of most VQ domains from different species were similar, although a small number of variants existed. In general, members of the same subfamily had similar types and numbers of conserved motifs, but there were also cases in which members of the same family had different types and numbers of conserved motifs. In addition, VQ had mutated to VH in the VQ domain of several Gramineae species [5, 6]. Taken together, these results indicate that the VQ gene family is highly conserved and diverse, reflecting the functional diversity of the gene family members.
With the development of next generation sequencing technology, the genomes of Arabidopsis, rice, maize, and wheat have recently been sequenced [2, 3, 8,9,10]. Their genome sizes are 164 Mb, 389 Mb, 2500 Mb, and 17 Gb, respectively. Based on genome size and chromosome number, the number of VQ genes among the four species is expected to be the highest in wheat, followed by maize, rice, and Arabidopsis. The numbers of maize, rice, and Arabidopsis VQ genes are 61, 40, and 34, and the number of wheat VQ genes in this study was 65, consistent with predictions based on genome size. By calculating the Ks value of homologous pairs to estimate the time of duplication events, the time range for whole genome duplication events in wheat was approximately 1.2538 to 116.8985 MYA. The Ka/Ks ratio of most paralogous pairs (37, 77%) was greater than one, indicating that the TaVQ gene family had undergone positive selection. A sliding window graph demonstrated that the Ka/Ks ratio of homologous pairs differed among different coding segments: some had Ka/Ks ratios greater than one, and some had Ka/Ks ratios less than one, indicating that the homologous pairs had undergone different evolutionary selection pressures. These results show that natural selection has played an important role in the evolution and differentiation of the VQ gene family.
TaVQ55/-60/-65 were highly expressed in GSC, GSR, GSE, SR, SL, Fba, and Pba; TaVQ2/-5/-8 were highly expressed in SL, Aba, 3–5 DAP C (22 days after planting—caryopsis), and 22 DAP EM; and TaVQ16/-31/-35 were highly expressed in SL, Aba, and 22 DAP EM. These results indicate that the VQ gene family is active during multiple plant growth and developmental stages. Previous studies on Arabidopsis have shown that IKU1 (AtVQ14, At2g35230) regulates endosperm development and seed size [20]. In this study, TaVQ48, which belongs to the same subfamily as AtVQ14/-29, was also expressed in floral bracts before anthesis, in 22 DAP EN, and in 22 DAP EM. In addition, TaVQ48 was strongly expressed in germinating seeds, roots, seedling roots, and seedling leaves. These results will guide further exploration of the functions of the TaVQ gene family.
GO annotations of 62 TaVQ genes were extracted from transcriptome data. The most common GO terms were from the biological process category (43, 38.1%), especially GO:0,006,952 (13 genes) and GO:0,008,150 (15 genes). GO:0,006,952 is related to defense response, and combined with promoter analysis, we found that 5 TaVQ genes (TaVQ1/-2/-32/-41/-51) had this function in both analyses. GO:0,010,337 is related to the regulation of salicylic acid (SA) metabolism, but only TaVQ14 was assigned this annotation. TaVQ14 also had an SA cis-acting element in the promoter analysis. These results indicate that gene structure determines function, and the diversity of structure reflects the diversity of function.
In the present study, we measured the expression of the 65 TaVQ genes during seed imbibition of six wheat varieties (HMC21, YXM, SNTT, J411, ZY9507, and ZM895). The expression of thirteen TaVQ genes (TaVQ8/-9/-13/-17/-25/-32/-34/-43/-48/-49/-53/-59/-62) was consistently higher in low-dormancy varieties than in high-dormancy varieties. By contrast, the expression levels of 8 TaVQ genes (TaVQ4/-16/-20/-35/-38/-42/-51/-56) were consistently higher in high-dormancy varieties. These 21 TaVQ genes may therefore participate in the regulation of seed dormancy and germination. According to phylogenetic analysis, three of these 21 genes (TaVQ8, TaVQ13, and TaVQ59) are members of the VQV subfamily. Interestingly, Arabidopsis AtVQ18 and AtVQ26 involved in seed germination also belong to the VQV subfamily. These results suggest that TaVQ8/-13/-59 may have similar functions in the regulation of seed dormancy and germination, a hypothesis that requires future validation.
Conclusions
We investigated the phylogeny and diversification of VQ genes in wheat by multiple methods, including phylogenetic tree construction and characterization of gene structures, conserved domains, chromosome positions, expression patterns, and promoter elements. In addition, we measured the expression levels of TaVQ genes in wheat varieties with contrasting seed dormancy and germination phenotypes by qRT-PCR to identify genes that were potentially involved in seed dormancy and germination. Sixty-five TaVQ proteins were identified for the first time in common wheat, and qRT-PCR data showed that 21 were potentially involved in seed dormancy and germination. These findings provide valuable information for further cloning and functional analysis of TaVQ genes, as well as useful candidate genes for improvement of PHS resistance in wheat.
Methods
Plant materials
We measured TaVQ gene expression in six wheat varieties with extreme dormancy levels [46]: J411 (Jing 411, average germination index [GI] = 0.89, average germination rate [GR] = 98.00%), HMC21 (Hongmangchun 21, average GI = 0.04, average GR = 10.00%), SNTT (Suiningtuotuo, average GI = 0.06, average GR = 16.00%), ZM895 (Zhongmai 895, average GI = 0.81, average GR = 96.00%), ZY9507 (Zhongyou 9507, average GI = 0.90, average GR = 98.00%), and YXM (Yangxiaomai, average GI = 0.03, average GR = 9.00%) (Tables S11 and S12). J411 and HMC21 were provided by Shihe Xiao from the Chinese Academy of Agricultural Sciences, and ZM895, ZY9507, YXM, and SNTT were provided by Xianchun Xia from the Chinese Academy of Agricultural Sciences.
Germination index and germination rate assays
Freshly harvested seeds were used to measure the GI as described in our previous study [46]. Fifty seeds from each genotype were placed in Φ 90 Petri dishes on filter paper with 9 ml distilled water, then grown in a 20 °C greenhouse with a 14 h day/10 h night photoperiod at 80% humidity. The number of germinated seeds in each culture dish was counted at the same time every day, and germinated seeds were removed. The GI value was calculated after 3 days as GI = ([3 × n1] + [2 × n2] + [3 × n1])/3 × N. The GR was also calculated after 3 days of seed imbibition as GR = [(n1 + n2 + n3)/N] × 100%. In these equations, n1, n2, and n3 are the numbers of seeds germinated on the first, second, and third days, and N is the total number of seeds. Each genotype was replicated three times, and germination was defined as visible rupture of the pericarp and testa.
Identification of wheat VQ genes
To determine the number of VQ genes in common wheat, we used sequences obtained from the Ensembl database to build a local wheat database [46]. The VQ domain hidden Markov model (PF05678) was used to identify candidate genes by BLAST in the established local wheat database. To ensure the accuracy of the results, all candidate genes were inspected, repetitive sequences were removed, and Pfam, SMART, and NCBI online tools were used to verify the existence of the conserved VQ domain in all candidate genes [46, 47]. The ExPASy online tool was used to predict the isoelectric point (PI), protein molecular weight (MW), open reading frame (ORF), and other attributes of the VQ proteins.
Phylogenetic tree and multiple sequence alignment
FASTA sequence files were opened in ClustalX2.11 software [48,49,50,51] and used to generate a multiple sequence alignment from which a phylogenetic tree was constructed using the neighbor-joining method with 1000 bootstrap replicates in MEGA7.0 [43, 52,53,54]. The same method was used to build a composite phylogenetic tree of VQ protein sequences from maize, rice, poplar, Arabidopsis, and wheat.
Intron/exon structure and conserved motif analysis
The distribution and structure of exons and introns were determined by uploading CDS and genomic sequences to the Gene Structure Display Server (http://gsds.cbi.pku.edu.cn/) for plotting and analysis [7, 54, 55].
To predict structural differences among the TaVQ proteins, all candidate protein sequences were uploaded to the MEME online tool (http://memesuite.org/tools/meme) for conserved motif analysis using standard operating parameters [54, 56].
Identification of homologous pairs and calculation of Ka/Ks values
Using previously reported methods for the identification of homologous gene pairs (paralogs and orthologs), the nucleotide sequences of VQ genes from wheat and other species were compared using BLASTN [57, 58].
Wheat homologous gene pairs were compared and aligned in ClustalX 2.11, and the aligned sequences were analyzed in MEGA7.0 [59]. The results were uploaded to DnaSP v5.10.1 [60] to calculate the values of Ka (non-synonymous nucleotide mutation rate) and Ks (synonymous nucleotide mutation rate) for all homologous pairs. The formula T = (Ks/2λ) × 10−6 was used to estimate the approximate dates of divergence events. To further analyze Ka/Ks values, we used GraphPad Prism 5 software to generate a sliding window graph [7, 61]. A Ka/Ks ratio less than 1 indicates that a DNA mutation is harmful and under purifying selection, whereas a Ka/Ks ratio greater than 1 indicates that a DNA mutation is beneficial and under positive selection. A Ka/Ks ratio of 1 indicates neutral selection [62].
Chromosome location and gene ontology annotation
The chromosome locations of TaVQ genes were downloaded from the Ensembl database, and chromosome maps were built using MapGene2Chromosome v2.0 [63]. Gene ontology (GO) annotations in the biological process, cellular component, and molecular function categories were assigned based on our transcriptome data (http://amigo.geneontology.org/amigo) [54].
Promoter analysis and subcellular location prediction
The 1500-bp sequence upstream of the transcription start site of each VQ gene was downloaded from the Ensembl website, and cis-acting elements in the promoter region were identified using the PlantCARE online tool [64]. WOLF was used to predict the subcellular localization of the TaVQ proteins [65].
Tissue expression pattern analysis
We collected three replicate seed tissue samples from HMC21 and J411 at 4, 6, and 10 h after seed imbibition for transcriptome sequencing. In addition, we obtained microarray data for 13 different tissues (three biological replicates each) from the Gene Expression Omnibus database (accession number GSE12508) [66, 67]. Mapper Plus was used to generate an expression heat map [45, 68].
RNA extraction and RT-qPCR analysis
Total RNA was extracted from seeds using the TaKaRa MiniBEST Universal RNA Extraction Kit. Primer Premier 5.0 was used to design 65 TaVQ gene-specific primers (Table S10), and TaActin was used as the reference gene [69]. The total PCR volume was 10 μl. The reaction process was 94 °C denaturation for 30 s, followed by 40–45 cycles of 94 °C for 5 s, 50–60 °C for 15 s, and 72 °C for 10 s. We performed three biological replicates for each sample. Finally, we processed the data and created the corresponding figure in GraphPad version 5 [70].
Availability of data and materials
The datasets generated and analysed during the current study are available in the corresponding author on reasonable request. The genome sequences of wheat, maize, rice and Arabidopsis were downloaded from the Ensembl database (http://plants.ensembl.org/index.html), PlantTFDB (http://planttfdb.gao-lab.org/), Rice Genome Annotation Project database (http://rice.plantbiology.msu.edu/analyses_search_locus.shtml) and Arabidopsis Information Resource (http://www.arabidopsis.org).
Abbreviations
- VQ:
-
Valine–glutamine
- qRT-PCR:
-
Quantitative real-time PCR
- Ks:
-
Number of synonymous substitutions per synonymous site
- Ka:
-
Number of non-synonymous substitutions per non-synonymous site
- CDS:
-
Coding sequence
- bp:
-
Base pair
- aa:
-
Amino acids
- MW:
-
Molecular weight
- Da:
-
Dalton
- PHS:
-
Pre-harvest sprouting
- GI:
-
Germination index
- pI:
-
Isoelectric point
- GO:
-
Gene ontology
- GSC:
-
Germinating seed coleoptile
- GSR:
-
Germinating seed root
- GSE:
-
Germinating seed, embryo
- SR:
-
Seedling root
- SC:
-
Seedling crown
- SL:
-
Seedling leaf
- II:
-
Immature inflorescence
- Fba:
-
Floral bracts before anthesis
- Pba:
-
Pistil before anthesis
- Aba:
-
Anthers before anthesis
- 3–5 DAP C:
-
3–5 Days after planting caryopsis
- 22 DAP EM:
-
22 Days after planting embryo
- 22 DAP EN:
-
22 Days after planting endosperm
References
Brenchley R, Spannagl M, Pfeifer M, Barker GLA, DAmore R, Allen AM, McKenzie N, Kramer M, Kerhornou A, Dan B. Analysis of the bread wheat genome using whole genome shotgun sequencing. Nature. 2012;491:705–10.
Luo MC, Gu YQ, Puiu D, Wang H, Twardziok SO, Deal KR, Huo N, Zhu T, Wang L, Wang Y, McGuire PE, Liu S, Long H, Ramasamy RK. Genome sequence of the progenitor of the wheat D genome Aegilops tauschii. Nature. 2007;551:498–502.
Ling H, Ma B, Shi X, Liu H, Dong L, Sun H, Cao Y, Gao Q, Zheng S, Li Y, Yu Y, Du H, Qi M, Li Y, Lu H, Yu H, Cui Y, Wang N. Genome sequence of the progenitor of wheat A subgenome Triticum urartu. Nature. 2018;557:424–8.
Pecher P, Eschen-Lippold L, Herklotz S, Kuhle K, Naumann K, Bethke G, Uhrig J, Weyhe M, Scheel D, Lee J. The Arabidopsis thaliana mitogen-activated protein kinases MPK3 and MPK6 target a subclass of ‘VQ-motif’-containing proteins to regulate immune responses. New Phy. 2014;203:592–606.
Kim DY, Kwon SI, Choi C, Lee H, Ahn I, Park SR, Bae SC, Lee SC, Hwang DJ. Expression analysis of rice VQ genes in response to biotic and abiotic stresses. Gene. 2013;529:208–14.
Song W, Zhao H, Zhang X, Lei L, Lai J. Genome-wide identification of VQmotif-containing proteins and their expression profiles under abiotic stresses in Maize. Front Plant Sci. 2016;6:1177.
Wang Y, Liu H, Zhu D, Gao Y, Yan H, Yan X. Genome-wide analysis of VQ motif-containing proteins in Moso bamboo (Phyllostachys edulis). Planta. 2017;246:165.
Cheng Y, Chen Z. Structural and functional analysis of VQ motif-containing proteins in Arabidopsis as interacting proteins of WRKY transcription factors. Plant P. 2012;159:810–25.
Li N, Li X, Xiao J, Wang S. Comprehensive analysis of VQ motif-containing gene expression in rice defense responses to three pathogens. Plant Cell Rep. 2014;33:1493–505.
Wang X, Zhang H, Sun G, Jin Y, Qiu L. Identification of active VQ motif-containing genes and the expression patterns under low nitrogen treatment in soybean. Gene. 2014;543:237–43.
Wang M, Vannozzi A, Wang G, Zhong Y, Corso M, Cavallini E, Cheng ZM. A comprehensive survey of the grapevine VQ gene family and its transcriptional correlation with WRKY proteins. Front Plant Sci. 2015;6:417.
Schnable PS, Ware D, Fulton RS. The B73 maize genome: complexity, diversity, and dynamics. Science. 2009;326:1112–5.
Perruc E, Charpenteau M, Ramirez BC, Jauneau A, Galaud JP, Ranjeva R, Ranty B. A novel calmodulin-binding protein functions as a negative regulator of osmotic stress tolerance in Arabidopsis thaliana seedlings. Plant J. 2004;38:410–20.
Cao J, Huang J, Yang Y, Hu X. Analyses of the oligopeptide transporter gene family in poplar and grape. BMC Genomics. 2011;12:459–64.
Lai Z, Li Y, Wang F, Cheng Y, Fan B, Yu JQ, Chen Z. Arabidopsis sigma factor binding proteins are activators of the WRKY33 transcription factor in plant defense. Plant Cell. 2011;23:3824.
Hu Y, Chen L, Wang H, Zhang L, Wang F, Yu D. Arabidopsis transcription factor WRKY8 functions antagonistically with its interacting partner VQ9 to modulate salinity stress tolerance. Plant J. 2013;74:730–45.
Zhang GY, Wang FD, Li JJ. Genome-wide identification and analysis of the VQmotif-containing protein family in Chinese Cabbage (Brassica rapa L. ssp. Pekinensis). Intj Mol Sci. 2015;16:28683–704.
Fill BK, Petersen M. Constitutive expression of MKS1confers susceptibility to Botrytis cinerea infection independent of PAD3 expression. Plant Signaling Behavior. 2011;6:1425–7.
Hu P, Zhou W, Cheng Z, Fan M, Wang L, Xie D. JAV1controls jasmonate-regulated plant defense. MolCell. 2013;50:504–15.
Wang A, Garcia D, Zhang H, Feng K, Chaudhury A, Berger F, Peacock WJ, Dennis ES, Luo M. The VQ motif protein IKU1 regulates endosperm growth and seed size in Arabidopsis. Plant J. 2010;63:670–9.
Chen CH, Chen ZX. Isolation and characterization of two pathogen- and salicylic acid-induced genes encoding WRKY DNA-binding proteins from tobacco. Plant Mol Biol. 2000;42:387–96.
Du L, Chen Z. Identification of genes encoding receptor-like protein kinases as possible targets of pathogen- and salicylic acid-induced WRKY DNA-binding proteins in Arabidopsis. Plant J. 2000;24:837–47.
Eulgem T, Rushton PJ, Robatzek S, Somssich IE. The WRKY superfamily of plant transcription factors. Trends Plant Sci. 2000;5:199–206.
Hara K, Yagi M, Kusano T, Sano H. Rapid systemic accumulation of transcripts encoding a tobacco WRKY transcription factor upon wounding. Mol Gen Genet. 2000;263:30–7.
Yu D, Chen C, Chen Z. Evidence for an important role of WRKY DNA binding proteins in the regulation of NPR1 gene expression. Plant Cell. 2001;13:1527–40.
Cormack RS, Eulgem T, Rushton PJ, Kochner P. Hahlbrock K and Somssich IE Leucine zipper-containing WRKY proteins widen the spectrum of immediate early elicitor-induced WRKY transcription factors in parsley. Biochim Biophys Acta. 2002;1576:92–100.
Pnueli L, Hallak-Herr E, Rozenberg M, Cohen M, Goloubinoff P, Kaplan A, Mittler R. Molecular and biochemical mechanisms associated with dormancy and drought tolerance in the desert legume Retama raetam. Plant J. 2002;31:319–30.
Seki M, Narusaka M, Ishida J, Nanjo T, Fujita M, Oono Y, Kamiya A, Nakajima M, Enju A, Sakurai T, Satou M, Akiyama K, et al. Monitoring the expression profiles of 7000 Arabidopsis genes under drought, cold and high-salinity stresses using a full-length cDNA microarray. Plant J. 2002;31:279–92.
Izaguirre MM, Scopel AL, Baldwin IT, Ballare CL. Convergent responses to stress. Solar ultraviolet-B radiation and Manduca sexta herbivory elicit overlapping transcriptional responses in field-grown plants of Nicotiana longiflora. Plant Physiol. 2003;132:1755–67.
Guo ZJ, Kan YC, Chen XJ, Li DB, Wang DW. Characterization of a rice WRKY gene whose expression is induced upon pathogen attack and mechanical wounding. Acta Botanica Sinica. 2004;46:955–64.
Li J, Brader G, Palva ET. The WRKY70 transcription factor: a node of convergence for jasmonate-mediated and salicylatemediated signals in plant defense. Plant Cell. 2004;16:319–31.
Xie Z, Zhang ZL, Zou X, Huang J, Ruas P, Thompson D, Shen QJ. Annotations and functional analyses of the rice WRKY gene superfamily reveal positive and negative regulators of abscisic acid signaling in aleurone cells. Plant Physiol. 2005;137:176–89.
Ralph S, Oddy C, Cooper D, Yueh H. Genomics of hybrid poplar (Populus trichocarpa 9 deltoides) interacting with forest tent caterpillars (Malacosoma disstria): normalized and full length cDNA libraries, expressed sequence tags, and a cDNA microarray for the study of insect-induced defences in poplar. Mol Ecol. 2006;15:1275–97.
Cui X, Yan Q, Gan S, Xue D, Wang H, Xing H, Guo N. GmWRKY40, a member of the WRKY transcription factor genes identified from Glycine max L., enhanced the resistance to Phytophthora sojae. BMC Plant Biol. 2019;19:1–15.
Kanofsky K, Riggers J, Staar M, Strauch CJ, Arndt LC, Hehl R. A strong NF-κB p65 responsive cis-regulatory sequence from Arabidopsis thaliana interacts with WRKY40. Plant Cell Rep. 2019;38:1139–50.
Zhao K, Zhang D, Lv K, Zhang X, Cheng Z, Li R, Jiang T. Functional characterization of poplar WRKY75 in salt and osmotic tolerance. Plant Science. 2019;289:110259.
Khuman A, Arora S, Makkar H, Patel A, Chaudhary B. Extensive intragenic divergences amongst ancient WRKY transcription factor gene family is largely associated with their functional diversity in plants. Plant Gene. 2020;22:100222.
Wang H, Zou S, Li Y, Lin F, Tang D. An ankyrin-repeat and WRKY-domain-containing immune receptor confers stripe rust resistance in wheat. Nat Commun. 2020;11:1–11.
Andreasson E, Jenkins T, Brodersen P, Thorgrimsen S. The MAP kinase substrate MKS1 is a regulator of plant defense responses. EMBO J. 2005;24:2579–89.
Clerkx EJM, Vries HBD, Ruys GJ, Groot SPC, Koornneef M. Characterization of green seed, an enhancer of abi3–1 in arabidopsis that affects seed longevity. Plant Physiol. 2003;132:1077–84.
Finkelstein R, Reeves W, Ariizumi T. Molecular aspects of seed dormancy. Annual Rev Plant Biol. 2008;59:387–415.
Pan JJ, Wang HP, Hu YR, Yu DQ. Arabidopsis VQ18 and VQ26 proteins interact with ABI5 transcription factor to negatively modulate ABA response during seed germination. Plant J. 2018;95:529–44.
Chu W, Liu B, Wang Y, Pan F, Chen Z, Yan H, Yan X. Genome-wide analysis of poplar VQ gene family and expression profiling under PEG, NaCl, and SA treatments. Tree Genet Genom. 2016;12:124.
Wang H, Hu Y, Pan J, Yu D. Arabidopsis VQ motif-containing proteins VQ12 and VQ29 negatively modulate basal defense against Botrytis cinerea. Sci Rep. 2015;5:14185.
Sturn A, Quackenbush J, Trajanoski Z. Genesis: cluster analysis of microarray data. Bioinformatics. 2002;18:207–8.
Cheng XR, Wang SX, Xu DM, Liu X, Zhang HP, Chang C, Ma CX. Identification and analysis of the GASR gene family in common wheat (Triticum aestivum L.) and characterization of TaGASR34, a gene associated with seed dormancy and germination. Fron Genet. 2019;4:515.
Chen Z, Chen X, Yan H, Li W, Li YY, Cai R, Xiang Y. The lipoxygenaseGene Family in Poplar: identification, classification, and expression in responseto MeJA treatment. PloS One. 2015;10:e0125526.
Thompson JD, Gibson TJ, Plewniak F, Higgins DG. The CLUSTAL_X windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res. 1995;25:4876–82.
Gao YM, Liu H, Wang Y, Li F, Xiang Y. Genome-wide identification of PHD-finger genes and expression pattern analysis under various treatments in moso bamboo (Phyllostachys edulis). Plant Physiol Bioch. 2017;123:S0981942817304357.
Cao Y, Han Y, Li D, Yi L, Cai Y. MYB Transcription Factors in Chinese Pear (Pyrus bretschneideri Rehd.): Genome-wide identification, classification, and expression profiling during fruit development. Front Plant Sci. 2016;7:557.
Li Q, Li L, Liu Y, LY Q, Zhang H, Zhu J, Li XJ. Influence of TaGW2–6a on seed development in wheat by negatively regulating gibberellin synthesis. Plant Science. 2017;263:226.
Saitou N, Nei M. The neighbor-joining method: a new method for reconstructing phylogenetic trees. Molbiolevol. 1987;4:406–25.
Kumar S, Stecher G, Tamura K. MEGA7: Molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol Biol Evol. 2016;33:1870–4.
Cheng XR, Xiong R, Liu HL, Wu M, Chen F, Yan HW, Xiang Y. Basic helix-loop-helix gene family: Genome wide identification, phylogeny, and expression in Moso bamboo. Plant physiol bioch. 2018;132:104–19.
Guo AY, Zhu QH, Chen X. GSDS: a gene structure display server. Hereditas. 2007;29:1023–6.
Bailey TL, Boden M, Buske FA. MEME SUITE: tools for motif discovery and searching. Nucleic Acids Res. 2009;37:W202–8.
Altschul SF, Madden TL, Schäffer AA, Zhang J, Zhang Z, Miller W, Lipman DJ. Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic acids res. 1997;25:3389–402.
Blanc G, Wolfe KH. Widespread paleopolyploidy in model plant species inferred from age distributions of duplicate genes. Plant Cell. 2004;16:1667–78.
Chen D, Chen Z, Wu M, Wang Y, Wang Y, Yan H, Xiang Y. Genome-wide identification and expression analysis of the HD-Zip gene family in Moso Bamboo (Phyllostachys edulis). J Plant Grow Re. 2017;36:323–37.
Librado P, Rozas J. DnaSP v5: a software for comprehensive analysis of DNA polymorphism data. Bioinformatics. 2009;25:1451–2.
Wang Y, Feng L, Zhu Y, Li Y, Yan H, Xiang Y. Comparative genomic analysis of the WRKY III gene family in populus, grape, arabidopsis and rice. Biol Direct. 2015;10:48.
Zhang Z, Li J, Zhao XQ, Wang J, Wong GK, Yu J. KaKs Calculator: Calculating Ka and Ks through model selection and model averaging. Genomics Proteomics Bioinformatics. 2006;4:259–63.
Voorrips RE. Map Chart: Software for the graphical presentation of linkage maps and QTLs. J Hered. 2002;93:77–8.
Zhao P, Wang D, Wang R, Kong N, Zhang C, Yang C, Wu W, Ma H, Chen Q. Genome-wide analysis of the potato Hsp20 gene family: identification, genomic organization and expression profiles in response to heat stress. BMC Genom. 2018;19:61.
Park KJ, Horton P, Obayashi T, Fujita N, Harada H, Adamscollier CJ, Nakai K. WoLF PSORT: protein localization predictor. Nucleic Acids Res. 2007;35:585–7.
Barrett T, Edgar R. Gene expression omnibus: microarray data storage, submission, retrieval, and analysis. Methods in Enzymol. 2006;411:352–69.
Wilkins O, Nahal H, Foong J, Provart NJ, Campbell MM. Expansion and diversification of the Populus R2R3-MYB family of transcription factors. Plant Physiol. 2009;149:981–93.
Kiana T, Brady SM, Ryan A, Eugene L, Provart NJ. The botany array resource: e-northerns, expression angling, and promoter analyses. Plant J. 2005;43:153–63.
Sun T, Wang Y, Wang M, Li T, Zhou Y, Wang X, Wei S, He G, Yang G. Identification and comprehensive analyses of the CBL and CIPK gene families in wheat (Triticum aestivum L.). BMC Plant Biol. 2015;15:269.
Bryfczynski SP, Pargas RP. GraphPad: a graph creation tool for CS2/CS7. ACM, 2009:389-389.
Acknowledgements
We would like to thank TopEdit (www.topeditsci.com) for English language editing of this manuscript. Thanks to teachers Shihe Xiao and Xianchun Xia from the Chinese Academy of Agricultural Sciences for providing six pairs of parental germplasm.
Funding
This work was supported by grants from the National Natural Science Foundation of China (31871608), The China Agriculture Research System (CARS-03), Jiangsu Collaborative Innovation Center for Modern Crop Production (JCIC-MCP), and The Agriculture Research System of Anhui province (AHCYTX-02). These funding bodies had no role in the design of the study; in the collection, analyses, or interpretation of data; in the writing of the manuscript, and in the decision to publish the results. Publication costs are defrayed by these funding.
Author information
Authors and Affiliations
Contributions
XRC and CG conceived the study, put into effect the main bioinformatics analyses, and drafted the manuscript. DMX and XL took part in the experiments and drafting of the manuscript. XP, WG, SNY, HY, JJC, XYM and JL processed the experimental data and helped to draft the manuscript. HPZ, CC, and CXM conceived and guided the experiments, and helped in coordinating the project and drafting the manuscript. All authors read and accepted the final manuscript.
Corresponding authors
Ethics declarations
Ethics approval and consent to participate
Not applicable.
Consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing interests.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Additional file 1:
Figure S1. Sliding window plots of the VQ genes. Figure S2. Chromosomal locations of TaVQ genes. Chromosome numbers are indicated above each bar. Table S1 Detailed information about the ZmVQ, OsVQ, PtVQ and AtVQ genes. Table S2. Numbers of VQ genes and VQ genes without introns in different species. Table S3. VQ domain types in different species. Table S4. Information on 20 conserved motifs of the TaVQ protein family. Table S5. Transcriptome data for VQ genes. Table S6. Microarray data for VQ genes. Table S7. Promoter analysis of the TaVQ protein family. Table S8. Gene ontology (GO) annotations of TaVQ proteins. Table S9. Subcellular localization of TaVQs predicted by WOLF PSORT. Table S10. qRT-PCR primers for TaVQ genes. Table S11. Data of seed germination index (GI) of six wheat varieties. Table S12. Data of seed germination rate (GR) of six wheat varieties.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/. The Creative Commons Public Domain Dedication waiver (http://creativecommons.org/publicdomain/zero/1.0/) applies to the data made available in this article, unless otherwise stated in a credit line to the data.
About this article
Cite this article
Cheng, X., Gao, C., Liu, X. et al. Characterization of the wheat VQ protein family and expression of candidate genes associated with seed dormancy and germination. BMC Plant Biol 22, 119 (2022). https://doi.org/10.1186/s12870-022-03430-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/s12870-022-03430-1