Abstract
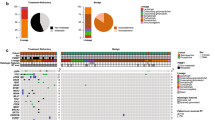
TP53 mutations play a significant role in glioma tumorigenesis. When located in in the DNA binding domain, these mutations can perturb p53 protein conformation and its function, often culminating in altered downstream signaling. Here we describe prevalent pattern of TP53 point mutations in a cohort of 40 glioma patients and show their relevance to gliomagenesis. Point mutations in exon 5–9 of TP53 gene were detected by DNA sequencing. Possible influence of identified mutations at the function of p53 was studied computationally and correlated with the survival. Point mutations in TP53 were detected in 10 glioma samples (25%), out of which 70% were from high grade glioma. A total of 19 TP53 point mutations were identified, out of which 42% were found to be in the DNA binding region of p53. Computational analysis predicted 87.5% of these mutations to be “probably damaging”. In three patients with tumors possessing point mutations R273H, R248Q, Y163H and R175H and poor survival times, structural analysis revealed the nature of these mutations to be disruptive and associated with high risk for cancer progression. In high grade glioma, recurrent TP53 point mutations may be the key to tumor progression, thus, emphasizing their significance in gliomagenesis.
Similar content being viewed by others
References
Jiang Y., Uhrbom L 2012. On the origin of glioma. Ups. J. Med. Sci. 117, 113–121.
Zhang J., Wu G., Miller C.P., Tatevossian R.G., Dalton J.D., Tang B., Orisme W., Punchihewa C., Parker M., Qaddoumi I., Boop F.A., Lu C., Kandoth C., Ding L., Lee R., et al 2013. Whole-genome sequencing identifies genetic alterations in pediatric low grade gliomas. Nat. Genetics. 45, 602–612.
Brennan C.W., Verhaak R.G., McKenna A., Campos B., Noushmehr H., Salama S.R 2013. The somatic genomic landscape of glioblatoma. Cell. 155, 462–477.
Ohgaki H., Kleihues P 2007. Genetic pathways to primary and secondary glioblastomas. Am. J. Pathol. 170, 1445–1453.
Schwartzbaum J.A., Fisher J.L., Aldape K.D., Wrensch M 2006. Epidemiology and molecular pathology of glioma. Nat. Clin. Pract. Neurol. 2, 494–503.
Rasheed B.K., McLendon R.E., Herndon J.E., Friedman H.S., Friedman A.H., Bigner D.D., Bigner S.H 1994. Alterations of the TP53 gene in human gliomas. Cancer Res. 54, 1324–1330.
Hede S.M., Nazarenko I., Nister M., Lindstrom M.S 2011. Novel perspective of p53 function in Neural stem cells and brain tumors. J. Oncol. 11, 1–11. doi 10.1155/2011/852970
Nozaki M., Tada M., Kobayashi H., Zhang C.L., Sawamura Y., Abe H., Ishii N., van Meir E.G 1999. Roles of functional loss of p53 and other genes in astrocytoma tumorigenesis and progression. Neuro Oncol. 1, 124–137.
Nagpal J., Jamoona A., Gulati N.D., Mohan A., Braun A., Murali R., Jhanwar-Uniyal M 2006. Revisiting the role of p53 in primary and secondary glioblastomas. Anticancer Res. 26, 4633–4639.
Ohgaki H., Dessen P., Jourde B., Horstmann S., Nishikawa T., Di Patre P.L., Burkhard C., Schüler D., Probst-Hensch N.M., Maiorka P.C., Baeza N., Pisani P., Yonekawa Y., Yasargil M.G., Lütolf U.M., Kleihues P 2004. Genetic pathways to glioblastoma: A populationbased study. Cancer Res. 64, 6892–6899.
Cho Y., Gorina S., Jeffrey P.D., Pavletich N.P 1994. Crystal structure of p53 tumor suppressor-DNA complex: Understanding tumorigenic mutations. Science. 265, 346–355.
Petitjean A., Mathe E., Kato S., Ishioka C., Tavtigian S.V., Hainaut P., Olivier M 2007. Impact on mutant p53 functional properties on TP53 mutation patterns and tumour phenotype: Lessons from recent developments in the IARC TP53 database. Hum. Mutat. 28, 622–629.
Xu J., Wang J., Hu Y., Qian J., Xu B., Chen H., Zou W., Fang J.Y 2014. Unequal prognostic potentials of p53 gain-of-function mutations in human cancers associate with drug-metabolizing activity. Cell Death Dis. 5, 1–9.
Phatak P., Selvi S.K., Divya T., Hegde A.S., Hegde S., Somasundaram K 2002. Alterations in tumour suppressor gene p53 in human gliomas from Indian patients. J. Biosci. 27, 673–678.
Roy A., Kucukural A., Zhang Y 2010. I-TASSER: A unified platform for automated protein structure and function prediction. Nat. Protoc. 5, 725–738.
DeLano W.L 2014. The Pymol Molecular Graphics Syste 2002. Delano Scientific, San Carlos, CA. http://www.pymol.org.
Jadersten M., Saft L., Pellagatti A., Göhring G., Wainscoat J.S., Boultwood J., Porwit A., Schlegelberger B., Hellström-Lindberg E 2009. Clonal heterogeneity in the 5q-syndrome: p53 expressing progenitors prevail during lenalidomide treatment and expand at disease progression. Haematologica. 94, 1762–1766.
Peraud A., Kreth F.W., Wiestler O.D., Kleihues P., Reulen H.J 2002. Prognostic impact of TP53 mutations and P53 overexpression in supratentorial WHO grade II astrocytomas and oligoastrocytomas. Clin. Cancer Res. 8, 1117–1124.
Olivier M., Hollstein M., Hainaut P 2010. TP53 mutations in human cancers: Origins, consequences, and clinical use. Cold Spring Harb. Perspect. Biol. 2, a001008, 1–17.
Wong K.B., DeDecker B.S., Freund S.M., Proctor M.R., Bycroft M., Fersht A.R 1999. Hot-spot mutants of p53 core domain evince characteristic local structural changes. Proc. Natl. Acad. Sci. U. S. A. 96, 8438–8442.
2BIM, Human p53 Core Domain Mutant M133LV203A-N239Y-N268D-R273 2014. The Protein Data Bank. http://www.rcsb.org/pdb/explore.do? structureId=2BIM.
Brazdova M., Quante T., Togel L., Walter K., Loscher C., Tichy V., Cincarova L., Deppert W., Tolstonog G.V 2009. Modulation of gene expression in U251 glioblastoma cells by binding of mutant p53 R273H to intronic and intergenic sequences. Nucleic Acids Res. 37, 1486–1500.
Freed-Pastor W.A., Prives C 2012. Mutant p53: One name, many proteins. Genes Dev. 26, 1268–1286.
Bullock A.N., Henckel J., Fersht A.R 2000. Quantitative analysis of residual folding and DNA binding in mutant p53 core domain: Definition of mutant states for rescue in cancer therapy. Oncogene. 19, 1245–1256.
Friedlander P., Legros Y., Soussi T., Prives C 1996. Regulation of mutant p53 temperature-sensitive DNA binding. J. Biol. Chem. 271, 25468–25478.
Author information
Authors and Affiliations
Corresponding author
Additional information
Published in Russian in Molekulyarnaya Biologiya, 2017, Vol. 51, No. 2, pp. 334–341.
The article is published in the original.
Rights and permissions
About this article
Cite this article
Sarma, P.P., Dutta, D., Mirza, Z. et al. Point mutations in the DNA binding domain of p53 contribute to glioma progression and poor prognosis. Mol Biol 51, 293–299 (2017). https://doi.org/10.1134/S0026893317020182
Received:
Revised:
Published:
Issue Date:
DOI: https://doi.org/10.1134/S0026893317020182