Abstract
Cancer stem cells (CSCs), their properties and interaction with microenvironment are of interest in modern medicine and biology. There are many studies on the emergence of CSCs and their involvement in tumor pathogenesis. The most important property inherent to CSCs is their stemness. Stemness combines ability of the cell to maintain its pluripotency, give rise to differentiated cells, and interact with environment to maintain a balance between dormancy, proliferation, and regeneration. While adult stem cells exhibit these properties by participating in tissue homeostasis, CSCs behave as their malignant equivalents. High tumor resistance to therapy, ability to differentiate, activate angiogenesis and metastasis arise precisely due to the stemness of CSCs. These cells can be used as a target for therapy of different types of cancer. Laboratory models are needed to study cancer biology and find new therapeutic strategies. A promising direction is three-dimensional tumor models or spheroids. Such models exhibit properties resembling stemness in a natural tumor. By modifying spheroids, it becomes possible to investigate the effect of therapy on CSCs, thus contributing to the development of anti-tumor drug test systems. The review examines the niche of CSCs, the possibility of their study using three-dimensional spheroids, and existing markers for assessing stemness of CSCs.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
INTRODUCTION
Cancer stem cells (CSCs) are a small subpopulation of tumor cells [1-3], which have the properties of stem cells (SCs) [4, 5]. SCs and CSCs have similarities and differences. The similarities include identical cell surface markers as well as ability to differentiate and self-renew using common signaling pathways [6]. One of the differences between SCs and CSCs is the degree of their dependence on the niche in which they are located. Unlike SCs, CSCs can change the niche with the help of signals stimulating proliferation. SCs are stable and contain a normal diploid genome, but CSCs are aneuploid with chromosomal rearrangements. In addition, SCs tend to be dormant and have relatively long telomeres, which is not characteristic of CSCs.
Presence of CSCs in the tumor tissue determines formation of phenotypically different subclones and leads to differentiated cell composition of the tumor [7]. Difference between the tumor growth and tissue renewal with participation of stem cells lies in the further developmental pathway of the transient-amplifying cells: usually they differentiate and die, but in the case of tumor cells they accumulate in the body and accelerate tumor growth [8].
Unlike the differentiated tumor cells, which constitute a significant part of the tumor and use glycolysis as a way of energy production, CSCs possess a distinct metabolic phenotype, which, depending on the cancer type, can have highly glycolytic phenotype or phenotype inherent to normal cells – oxidative phosphorylation [9]. The distinctive feature of CSCs is their oncogenic activity, which allows them to form tumors when transplanted to laboratory animals, which is impossible in the case of transplantation of normal SCs [10].
CSCs can influence metabolism of the neighboring cells, supply nutrients and create favorable conditions for tumor growth. Presence of CSCs in the tumor microenvironment (TME) leads to formation of the heterogeneous cell populations with high plasticity potential [11, 12] and high resistance to stress factors in TME (such as low oxygen or nutrient levels) [13, 14].
Proportion of CSCs in the tumor tissues is very small and usually does not exceed 2% of the total tumor mass. CSCs are characterized by certain markers depending on the type of tissue in which they are located (Table 1). About 73% of the surface markers of CSCs are present on the membranes of embryonic or adult stem cells and are rarely expressed on the membranes of differentiated cells [15]. The leukemia stem cells have been shown to possess the CD34+CD38– phenotype, in which absence of CD38 distinguishes these cells from the normal hematopoietic stem cells (HSCs) [16]. Singh et al. showed the presence of CD133+ HSCs in the glioblastoma tumors [17], which are resistant to the chemotherapeutic agent temozolomide compared to the CD133– cells [18].
In addition, not only surface antigens but also microRNAs (miR-21, miR-210, miR-34a, miR-16, etc.) can be CSCs markers [45].
Stemness refers to the general molecular processes underlying properties of the SC self-renewal and generation of differentiated progeny. Stemness allows maintaining cell population and balance between dormancy, proliferation, and regeneration [46]. It is known that the tumor cell stemness is manifested by the heredity maintenance, ability to survive stress and chemotherapeutic drugs influence, and is a key feature of cancer progression [47, 48]. It was shown that the CD133+ stem cells of colon carcinoma are resistant to fluorouracil and oxaliplatin in contrast to the CD133– cells [49].
Understanding genetic and epigenetic changes in tumor cells that are transformed into stem cells is the key to developing new therapeutic approaches. By expanding the knowledge about pathophysiology of CSCs in tumor it would be possible to create qualitative model systems that provide applied knowledge about molecular properties of the malignant tumors [46]. Xenografts are commonly considered as a model for studying CSCs stability. For example, the xenografts obtained from pancreatic tumor tissue of patients treated with gemcitabine showed resistance to the therapeutic agent due to CSCs [50]. In the T-cell acute lymphoblastic leukemia, xenografts had a stronger genetic similarity to the recurrence samples than to the main tumor at the time of diagnosis, which may imply natural selection of resistant clones during xenografting [51].
However, there are still many questions remaining regarding CSCs. One of the biggest challenges is identifying CSCs. Another challenge is understanding relation between the stemness and tumor progression. Furthermore, it is unclear how stemness contributes to the emergence of metastases and resistance to therapy [52].
Despite these challenges, progress is being made in the study of tumor stemness. Novel technologies and procedures are emerging for the detection and separation of CSCs, and scientists are commencing to elucidate molecular pathways that govern stemness in both normal and malignant cells. As this knowledge base matures, it would probably lead to better therapies and improved treatment results for cancer patients.
TRANSCRIPTION FACTORS AND STEM SIGNALING PATHWAYS
Disruption of the regulation of expression and activity of the tumor transcription factors allows the cancer cell to acquire stemness manifested by the ability to self-renew, differentiate, and form tumors in vivo. Such cancer stem cells have a significant potential for metastasis and resistance to therapy, which requires deep understanding of the molecular mechanisms of changes in the transcription factors associated with stemness. The main transcription factors that play a key role in the regulation of CSCs growth include OCT4, SOX2, NANOG, KLF4, and MYC [53].
OCT4 controls self-renewal and maintains pluripotency of stem cells [54]. OCT4 overexpression has been described in some types of cancers and promotes SC self-renewal and development of drug resistance [55, 56]. OCT4 knockdown reduces tumor cell stemness [57]. Correlation between the OCT4 expression and colorectal cancer metastasis to liver has been established. For example, Fujino et al. investigated clinical samples and the OCT4+ cells of colorectal adenocarcinoma. The level of OCT4 expression correlated with the frequency of liver metastasis in the patients, and OCT4+ cells had the ability to self-renew and differentiate similarly to CSCs [58]. Expression of OCT4 by the lung cancer cells leads to polarization of M2 macrophages, inducing secretion of the macrophage colony-stimulating factor (M-CSF), which promotes tumor growth and metastasis [59].
SOX2 plays a role in the development and maintenance of undifferentiated embryonic stem cells (ESCs), performs key functions in the regulation of proliferation, self-renewal, and other processes in stem cells, and is also involved in the development of tumor cell stemness [60, 61]. SOX2 overexpression has been established in the triple-negative breast cancer. Suppression of the SOX2 expression led to decrease in proliferation of the triple-negative breast cancer cells, repression of cell invasion ability, induction of apoptosis in vitro, and reduction of tumor growth and metastasis in vivo [62]. In the mouse xenograft model with osteosarcoma, the osteoblast-specific conditional knockout of SOX2 has been shown to cause a dramatic decrease in the incidence of tumors. All newly emerged rare tumors found in the SOX2 knockout experimental animals were SOX2-positive, indicating that they originated from the cells that avoided SOX2 deletion [63]. Also, it was discovered that the simultaneous occurrence of CD133 and SOX2 expressed together correlated with the more negative prognosis for the patient survival [64].
NANOG is a key factor for pluripotency and self-renewal of ESCs [65] and is often overexpressed in many types of cancers [66, 67]. In oral squamous cell carcinoma with increasing degree of dysplasia, NANOG expression was found to be increased, which could be an early predictor of the risk of malignant neoplasm development [68]. It was shown that the NANOG expression is associated with the increased aldehyde dehydrogenase (ALDH) activity and cellular radioresistance (through activation of NOTCH1 and AKT signaling pathways), as well as with stimulation of the DNA double-strand break repair [69]. Mutations in the SPOP gene have been shown to stop the SPOP-mediated degradation of NANOG, thereby causing increased synthesis of the stem cell markers by the prostate tumor cells and reducing patient survival [70].
KLF4 plays an important role in the regulation of proliferation, differentiation, apoptosis, and reprogramming of somatic cells [71]. KLF4 triggers expression of the telomerase reverse transcriptase (TERT) and enhances capacity of ESCs to maintain their pluripotency and continuous self-regeneration [72]. KLF4 can play a dual role in carcinogenesis, either stimulating this process or functioning as an oncosuppressor. Overexpression of KLF4 in different types of human melanomas was established. Inhibiting apoptosis through ectopic expression of KLF4 promotes higher division rate of the melanoma cells. In contrast, the KLF4 knockdown decreases melanoma cell proliferation and causes cell death. Moreover, KLF4 depletion reduces growth rate of the melanoma xenograft in vivo [73]. KLF4 plays an important role in regulation of the osteosarcoma cell stemness, and the KLF4-p38 MAPK signaling pathway could be a potential therapeutic target for osteosarcoma treatment [74]. Inhibition of the KLF4 expression has been demonstrated to enhance migratory and invasive properties of the non-small cell lung carcinoma. When the KLF4 expression was restored, decrease in the tumor cell invasion was noted [75]. Moreover, sensitivity of the ovarian cancer cells that were genetically modified and transduced with the lentivirus for KLF4 overexpression to the effects of paclitaxel and cisplatin was enhanced as a result of increased cell responsiveness [76].
MYC proteins (c-Myc, L-Myc, S-Myc and N-Myc) are a family of transcription factors that regulate growth, differentiation, self-renewal, metabolism, and cell cycle of stem cells [77]. Overexpression of MYC is often observed in the human tumors compared to normal tissue. MYC dysregulation plays an important role in maintaining the number of invasive CSCs. It was found that MYC overexpression in the glioblastoma CSCs induces tumor cell proliferation and invasion, and inhibits apoptosis [78]. It was shown that dysregulation of MYC, including increase of this gene expression, leads to the development of invasive ductal carcinoma [79]. Notably, c-MYC activates SOX4 expression through binding to its promoter, thus, c-MYC overexpression may contribute to the prostate cancer progression through the SOX4 activation [80].
There is increasing evidence that the overactive or abnormal signaling pathways may contribute to the survival of CSCs. It was found that the malignant neoplasms are characterized by dysregulation of Wnt, NF-κB, Notch, Hedgehog, JAK/STAT, and PI3K/AKT/mTOR signaling pathways, which play an important role in the CSCs stemness maintenance (Table 2) [53]. Moreover, the proteins of the above signaling pathways are involved in migration, development/maintenance of resistance and tumor formation, as well as in the epithelial-mesenchymal (EMT) and mesenchymal-epithelial (MET) transitions [81]. Genes of different signaling pathways can be expressed at different stages of carcinogenesis in different tumor types as well as have cross interaction in the CSCs regulation [82]. Thus, signaling pathways regulating CSCs proliferation are complex networks of signaling mediators interacting with each other [83]. Also, there is quite a large number of additional molecular agents affecting the CSC stemness (Table 3).
Thus, the study of transcription factors and signaling pathways not only provides scientific knowledge in the field of stem cell biology, but also makes a significant contribution to the application of this knowledge to the creation of new therapeutic strategies (Table 4). A large number of studies are aimed at identifying targeted therapies using components of signaling pathways as targets. For example, understanding of the tumor stem-driven interactions between the breast cancer subtypes is fundamental to identifying possible therapeutic approaches. In the study by Mei et al. difference between the cell subtypes was determined using the CRISPR/dCas9 system targeting genes OCT4, KLF, MYC, SOX2. It has been shown that the cells synthesizing epidermal growth factor receptor 2 (HER2) are precursors of the luminal A cells, hence, similar antitumor drugs can be used for the therapy targeting these tumor cells [120].
STEMNESS AND TUMOR MICROENVIRONMENT
In the absence of specific microenvironmental factors, stem cells are unable to sustain their viability beyond their respective niches [121]. It is known that extrinsic stimuli can promote emergence of the new stem cells, as the cells in general retain the ability to dedifferentiate [122]. Exposure to extracellular microenvironmental factors such as mesenchymal stem cells (MSCs), immune cells, tumor-associated fibroblasts (TAFs), extracellular matrix (ECM), vesicles, and hypoxia promotes acquisition of stemness by the tumor cell or reprogramming into CSCs [53]. Vascular endothelial growth factor (VEGF) contributes to acquisition of the tumor stemness phenotype by the cells in squamous cell skin cancer [123]. In the head and neck squamous cell carcinoma, epidermal growth factor (EGF) secreted by endothelial cells induces EMF with the cells acquiring stem cell characteristics [124]. Transforming growth factor (TGF)-β secreted by TAFs promotes ECM and increases proliferation and emergence of metastases of breast cancer cells [125].
Hypoxia also affects stemness and other properties of tumor cells by increasing synthesis of the hypoxia-inducible factor (HIF)-1α [126, 127].
The effect of ECM proteins on stemness modulation is interesting. For example, Zhang et al. studied a fibrin gel that was used to create a three-dimensional (3D) ECM of colorectal cancer cells. The soft fibrin gel (90 Pascal) was the most effective for enrichment of tumor colonies. In particular, these tumor colonies exhibited carcinogenicity, activated the stem cell markers, and exhibited anti-chemotherapeutic properties. These structures were called tumor repopulating cells (RTCs). It should be noted that NANOG factor was induced in the 3D-cultured colon RTCs [128]. It is also known that the collagen type I (Col-I) induces tumor stemness. The work of Valadão et al. determined dependence of this effect on the Col-I density. The results showed that high Col-I density increased the number of the CD44+CD24 breast cancer stem cells, but did not promote cell self-renewal or clonogenicity. Thus, it was concluded that high Col-I density by itself is not sufficient for full development of the breast cancer stem cell phenotype [129].
Factors associated with inflammation can activate the reprogramming network leading to generation of CSCs. The serum IL-6 levels were found to be higher in the patients with osteosarcoma than in the controls. IL-6 also increased proliferation and spherulation of the osteosarcoma cells as well as their invasive and migratory potential, thus contributing to formation of the cells with an EMT-like phenotype and increased chemoresistance. In a xenograft model of osteosarcoma, it was determined that the tumor size and weight were higher in the mice treated with recombinant IL-6 than in the controls [130]. It was found that IL-1β can induce spherulation of the colorectal cancer cells that were characterized by activation of the stemness factor genes (BMI1 and NESTIN) and increased drug resistance, which is a hallmark of CSCs [131]. Hong et al. showed that the tumor necrosis factor (TNF)-α promotes oral carcinogenesis associated with human papilloma virus by increasing tumor stemness [132].
Extracellular vesicles (EVs) of tumor and stromal cells mediate intercellular communication in the tumor microenvironment (TME) and participate in various stages of carcinogenesis such as proliferation, angiogenesis, metastasis, and resistance to therapeutic drugs. Because EVs carry various substances in them, they can participate in intercellular communication and play a role in the acquisition of stemness by cells. For example, EVs produced by the glioblastoma stem cells promote endothelial tube formation [133]. Similarly, in the colorectal adenocarcinoma, exosomes of TAFs promoted increase of the CSCs percentage in the tumor by activating the Wnt signaling pathway [134]. EVs from MSCs can have two effects: they can enhance tumor growth and metastasis, or they can promote cell apoptosis and cause tumor regression [135, 136].
Thus, extracellular factors released by the tumor microenvironment affect cell stemness and heterogeneity through regulation of the signaling pathways.
SPHEROIDS AS A MODEL SYSTEM FOR STUDYING CANCER STEM CELLS
To date, the use of 3D models in tumor research attracts attention of investigators. One of such models is the tumor spheroid, which can be divided into several types: (i) a multicellular spheroid first described in the early 1970s and generated by subjecting tumor cell lines to non-adherent culture dishes; (ii) oncosphere – a prototype for expansion of CSCs utilizing serum-free medium in combination with added growth factors; (iii) an organotypic multicellular spheroid created by dissociating tumor tissue via mechanical and enzymatic methods [137].
Multicellular spheroids are used as an in vitro model system for studying micrometastases and avascular tumor regions. In addition, this type of spheroids is suitable for studying cell adhesion-dependent resistance.
The terms “sphere”, “tumor sphere” [138], and “oncosphere” [139] are used to denote spherical structures originating from CSCs. Remarkably, the names of spheres can be derived from the type of tissue from which they are derived. For example, spheres obtained from stem cell cultures of the brain and mammary gland have been named “neurospheres” and “mammospheres”, respectively. Studies aiming in vitro evaluation and characterization of CSCs are usually performed using this model [140-142].
Organotypic multicellular spheroids are engineered to mimic TME exactly, which includes tumor cells, non-tumor cells, and stromal components [140].
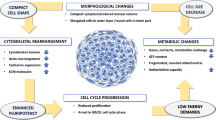
In the literature, these models are placed between the 2D and xenograft models, suggesting that the 3D models can fill the gaps that exist between them [140]. 3D models are characterized by high translatability of biological characteristics of the original tumors, which gives advantages when using them [143]. 3D models have architecture and metabolic responses similar to the native tumors. Histomorphological analysis shows that the 3D model, in particular a tumor spheroid, is divided into several zones, just like a natural tumor. These include the outer zone characterized predominantly by proliferating cells, the middle zone with dormant cells, and the central necrotic zone [144]. This can be explained by insufficient oxygenation, lack of nutrients, and accumulation of metabolites. The outer zone of spheroid is characterized by the large amount of lactate and minimal amount of adenosine triphosphate (ATP), mainly due to the significant consumption by the proliferating cell layer. Glucose distribution occurs uniformly in all areas of the spheroid [145].
The listed tumor spheroid types can contain different cells supporting genetic heterogeneity and interaction between the tumor cells and stroma. 3D models offer a possibility to study intercellular contacts, microenvironment, and interaction between the immune and tumor cells [146]. Experiments with the 3D cell culture could provide more accurate data on the effects of therapeutic agents, metabolic profiling, stem cell research [147], and the processes such as EMT and MET, which are involved in the regulation of stemness. Activation of several transcription factors, including SNAI1 and SNAI2, at the molecular level accounts for the effect of EMT. This effect is associated with the loss of intercellular contact molecules, such as E-CAD, as well as with acquisition of mesenchymal markers, including vimentin. EMT has been particularly studied in various types of tumor spheroids as well as in normal stem cells, including cardiospheres, neurospheres, and retinal spheres. Additionally, EMT activation studies have been performed with the embryonic stem cells (blastulation), embryoid cells, and cells subjected to induced reprogramming [148].
Tumor spheroids can be obtained by aggregation or spontaneous self-assembly. Aggregation can be induced by forcing cell-to-cell contact using various methods, such as hanging drop, cell suspension, microwell (96-well round-bottom plates), or microhydrodynamics (i.e., encapsulation in a gel) [140].
Studies of the spheroid model systems using novel specific agents targeting Notch (MK-0752, R4733, and TR-4) [113], PI3K (CAL-101 and XL-147) [114], AKT (perifosine and arhexin) [149], and Hedgehog (BMS-863923 and IPI-926) signaling pathways are of great interest for the development of anti-tumor therapy [150]. It was found that salinomycin (ionophore polyether antibiotic) is able to selectively kill the mammary CSCs and induce epithelial differentiation of tumor cells contributing to tumor elimination [138]. Namiki et al. showed that epigallocatechin gallate, one of the most important organic substances of green tea, reduces stemness and oncogenicity of the human lung cancer cells by inhibiting AXL receptors [151].
A significant obstacle to the development of rational cancer therapies is mismatch between the model systems and the patient’s body. To solve this problem, Fendler et al. used CSCs to create tumor spheroids, three-dimensional tumor organoids, and xenografts of the light cell renal cell cancer. Treatment with the Wnt and Notch signaling pathway inhibitors blocked proliferation and self-renewal of CSCs in the spheroids and organoids and inhibited tumor growth in the patient-derived xenografts in mice [152].
Thus, use of these model systems expands the possibilities for creating a screening platform for drug testing. In addition, use of 3D models opens up prospects for the development of personalized medicine. And the study of CSCs reveals their molecular mechanisms and contributes to the search for a therapeutic target for innovative therapies aimed at eliminating CSCs.
ENRICHMENT OF THE POPULATION OF CANCER STEM CELLS IN SPHEROIDS
One of the in vitro methods commonly used to identify CSCs and study their properties is the spherulation assay. The sphere-forming cells are stem cells. Spheroids are associated with the in vitro cancer stemness studies and are considered as one of the criteria to confirm the presence of CSCs [153]. It should be noted that in the literature both sphere and spheroid names are used for this CSCs analysis. For example, Qureshi-Baig et al. cultured spheroids obtained from the clinical specimens of colorectal carcinoma. These spheroids were shown to phenotypically resemble a native tumor, retain several similar mutations related to colorectal cancer, and express SOX2, OCT4, and NANOG. After comparing the tissue and cell line spheroids, it was found that both types of spheroids showed similar self-renewal ability and similarly formed tumors in immunodeficient mice [154]. Human ovarian CSCs were also characterized, which demonstrated increased chemoresistance to cisplatin and paclitaxel and expression of the stem cell genes (Notch-1, NANOG, NES, ABCG2, and OCT4) [155].
There is a fairly large number of CSCs studies related to the use of spheroids. For example, combining mathematical modeling and RNA sequencing of the ovarian tumor spheroids formed by the “hanging drop” method allowed revealing genes associated with the CSCs appearance and support of chemoresistance and malignant phenotype. Emergence of the CSC populations in the spheroids has been shown [156]. Bahmad et al. described protocols and a list of necessary reagents for creation of prostatic tumor spheroids and various manipulations with them [157]. The prostate tumor spheroids have been shown to survive more than 30 passages without noticeable loss of their stemness. Spheroids proliferated more slowly with the significant decrease in activation of the PI3K/AKT signaling pathway compared to the DU145 monolayer cells [158].
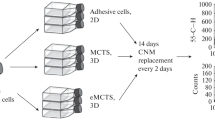
Various factors can influence enrichment of the CSCs population in spheroids, which can be used in studies. For example, serum-free culture conditions have been used to enrich SCs spheroids. Herheliuk et al. showed the possibility of enriching a population of tumor cells [in this case, multicellular tumor spheroids (MTS)] with CSCs. Standardization of culturing protocols for the enriched MTS (eMTS) may provide an opportunity to use these cultures for identification of the drugs that can inhibit proliferation of CSCs. Serum-free medium with addition of the fibroblast growth factor (FGF), EGF, insulin, and hydrocortisone was used to enrich CSCs in spheroids. It was shown that the highest expression of CD133, CD44, CD24, and BMI-1 receptors was detected in the eMTS. eMTS were less sensitive to anticancer drugs (cisplatin, methotrexate, and doxorubicin) than the adherent cell cultures and MTS cultured under standard conditions in complete nutrient medium [159].
Similarly, human hepatoma cell lines PLC/PRF/5 and HepG2 were cultured under the conditions used for CSCs. The culture medium included serum-free DMEM/F12 medium supplemented with 100 IU/ml penicillin, 100 μg/ml streptomycin, 20 ng/ml recombinant human EGF, 10 ng/ml recombinant basic human FGF and B27, and N-2 supplements. The cells were then cultured in 6-well non-adhesive plates at a density of no more than 5000 cells/well. As a result, the PLC/PRF/5 spherule-forming cells exhibited the properties of CSCs: self-renewal, extensive proliferation, drug resistance, and overexpression of OCT3/4, OV6, EpCAM (epithelial cell adhesion molecule), CD133, and CD44. Even a small number of cells were capable of forming tumors in the immunodeficient mice, and did not lose this property after repeated passaging [160]. It was also found that the changes in the components of the environment can play a role in the stem cell phenotype. Maliszewska-Olejniczak et al. showed that adding xeno-free or low-serum media to the 3D renal cell carcinoma culture increased expression of the genes encoding stemness factors, including ECAD (E-cadherin), N-cadherin (NCAD), VEGF, SOX2, PAX2, and NESTIN [161].
Different matrices can influence increase in the number of CSCs in spheroids. For example, culturing MCF-7 cells on the collagen matrices increased cell proliferation and generated a population of cells with CSCs properties [162]. Moreover, small leucine-rich proteoglycans of the extracellular matrix can play a role in regulation of the signaling pathways, drug resistance, and plasticity of the CSC-like cells [163]. It has been also shown that agar, which can be used to create non-adhesive conditions when generating spheroids, can play a role in the enriching spheroids with CSCs. This has been confirmed by the increased expression of CSCs markers, presence of the CD133+/CD44+, or SORE6+ cells, and increased capacity for self-renewal and carcinogenesis in vivo [164]. The same coating (agar) was used to cultivate spheroids from the oral squamous cell carcinoma cells, which showed expression of the stem cell markers and manifested chemoradiotherapy resistance in addition to the ability to initiate tumor formation and self-renewal [165]. Furthermore, additives such as methylcellulose and gellan gum were used to create a new culturing system based on the traditional system. The developed culturing system offers an attractive alternative method for in vitro CSC amplification with the potential for extensive use in CSC research and drug screening [166].
To create spheroids with the increased CSCs potential, a single cell cloning method can be applied. In particular, Muenzner et al. created single-cell HepG2 clones that formed 3D clusters of cells after 8 days of incubation that were individually transferred to the wells of a 12-well plate for cell culture. The clone no. 5 has been shown to exhibit more highly aggressive and invasive characteristics, most likely related to the significantly higher expression of the clinically relevant stem cell markers and more pronounced mesenchymal phenotype. In addition, higher metastatic potential was detected for the clone no. 5 cells compared to the parental HepG2 cell line in vivo using fluorescence imaging. Hence, the CSCs-enriched HepG2 clones certainly represent suitable model systems to study the role of CSCs during initiation, progression, and drug resistance of hepatocellular carcinoma [167].
Interestingly, it is possible to study the effect of MSCs on the tumor by creating hybrids with gastric cancer cells using polyethylene glycol (PEG) in vitro. Such hybrids were characterized by the changes in EMT with downregulation of ECAD and upregulation of vimentin, NCAD, α-smooth muscle actin (α-SMA), and fibroblast activation protein (FAP). The hybrids had increased expression of the stemness transcription factors OCT4, NANOG, SOX2, and LIN28. Expression of CD44 and CD133 was higher in the hybrid cells than in the parental gastric cancer cells. Enhanced migratory properties and division of the hybrid cells were observed. In addition, heterotypic hybrids promoted gastric xenograft tumor growth in vivo. Probably, fusion between the MSCs and gastric cancer cells promotes formation of oncogenic hybrids with EMT and stem cell properties that can provide an opportunity to study the role of MSCs in gastric cancer [168].
Thus, main feature of the tumor-derived spheroids is their enrichment with CSCs or cells with stem cell-related characteristics. Since eradication of CSCs is likely to be of clinical importance because of their association with the malignant nature of tumor cells such as carcinogenicity or chemoresistance, the study of tumor-derived spheroids could provide invaluable clues for the fight against cancer. The tumor-derived spheroids could be useful for the detection of new CSCs markers, become a convenient model for better assessment of tumor cell response to drugs, cell proliferation, and morphology [161, 169]. In addition, increase of the spheroid stemness can contribute to the improvement of the screening platform of antitumor therapeutics, identification of new therapy targets, and study of CSCs biology.
CONCLUSION
Survival rates of the patients with malignant neoplasms have increased significantly over the past decades due to development of new antitumor therapeutic strategies, which include improved chemotherapeutic agents and targeted therapies. Nevertheless, therapy of the advanced stages of cancer is complicated with high cellular heterogeneity due to genetic mutations, tumor stroma, and presence of CSCs, angiogenicity, and tumor metastasis. In the tumor microenvironment, CSCs support tumor growth through activation of various signaling pathways thus mediating supply of nutrients and metabolites. Due to plasticity and self-renewal, CSCs form a group of resistant clones, thus contributing to the development of drug resistance and tumor metastases. Hence, CSCs are an important therapeutic target in the creation of new antitumor treatment strategies. Better understanding of the biology of CSCs and their interaction with microenvironment will allow significant advances in the fight against tumor progression, recurrence, and resistance to therapy. A limiting factor in the development of effective therapies is structural complexity of the tumor, which is difficult to reproduce in the in vitro model systems. Modeling TME is particularly difficult, and therefore the use of 3D models can improve selection of drug candidates at the stage of their development. One such model is spheroids, which are of great interest because of their ability to recreate tumor pathogenesis through flexible modification. The use of spheroids to study CSCs will enhance prognostic potential of the cancer research and provide new opportunities for cancer therapy.
Abbreviations
- ALDH:
-
aldehyde dehydrogenase
- CSCs:
-
cancer stem cells
- ECM:
-
extracellular matrix
- EGF:
-
epidermal growth factor
- EMT:
-
epithelial-mesenchymal transition
- EpCAM:
-
epithelial cell adhesion molecule
- EV:
-
extracellular vesicles
- JAK/STAT:
-
janus kinase and transcription pathway activator
- IL:
-
interleukin
- MSC:
-
mesenchymal stem cells
- PI3K/AKT/mTOR:
-
phosphatidylinositol-3-kinase and serine-threonine protein kinase
- TME:
-
tumor microenvironment
References
Wei, R., Liu, S., Zhang, S., Min, L., and Zhu, S. (2020) Cellular and extracellular components in tumor microenvironment and their application in early diagnosis of cancers, Anal. Cell Pathol. (Amst), 2020, 6283796, https://doi.org/10.1155/2020/6283796.
Bozyk, A., Wojas-Krawczyk, K., Krawczyk, P., and Milanowski, J. (2022) Tumor microenvironment – a short review of cellular and interaction diversity, Biology (Basel), 11, 929, https://doi.org/10.3390/biology11060929.
Ribeiro Franco, P. I., Rodrigues, A. P., de Menezes, L. B., and Pacheco Miguel, M. (2020) Tumor microenvironment components: allies of cancer progression, Pathol. Res. Pract., 216, 152729, https://doi.org/10.1016/j.prp.2019.152729.
Batlle, E., and Clevers, H. (2017) Cancer stem cells revisited, Nat. Med., 23, 1124-1134, https://doi.org/10.1038/nm.4409.
Capp, J. P. (2019) Cancer stem cells: from historical roots to a new perspective, J. Oncol., 2019, 5189232, https://doi.org/10.1155/2019/5189232.
Reya, T., Morrison, S. J., Clarke, M. F., and Weissman, I. L. (2001) Stem cells, cancer, and cancer stem cells, Nature, 414, 105-111, https://doi.org/10.1038/35102167.
Brooks, M. D., Burness, M. L., and Wicha, M. S. (2015) Therapeutic implications of cellular heterogeneity and plasticity in breast cancer, Cell Stem Cell, 17, 260-271, https://doi.org/10.1016/j.stem.2015.08.014.
Sell, S. (2010) On the stem cell origin of cancer, Am. J. Pathol., 176, 2584-2594, https://doi.org/10.2353/ajpath.2010.091064.
Sancho, P., Barneda, D., and Heeschen, C. (2016) Hallmarks of cancer stem cell metabolism, Br. J. Cancer, 114, 1305-1312, https://doi.org/10.1038/bjc.2016.152.
Clarke, M. F., Dick, J. E., Dirks, P. B., Eaves, C. J., Jamieson, C. H. M., Jones, D. L., Visvader, J., Weissman, I. L., and Wahl, G. M. (2006) Cancer stem cells – perspectives on current status and future directions: AACR Workshop on cancer stem cells, Cancer Res., 66, 9339-9344, https://doi.org/10.1158/0008-5472.can-06-3126.
Cabrera, M. C., Hollingsworth, R. E., and Hurt, E. M. (2015) Cancer stem cell plasticity and tumor hierarchy, World J. Stem Cells, 7, 27-36, https://doi.org/10.4252/wjsc.v7.i1.27.
Chaffer, C. L., Brueckmann, I., Scheel, C., Kaestli, A. J., Wiggins, P. A., Rodrigues, L. O., Brooks, M., Reinhardt, F., Su, Y., Polyak, K., Arendt, L. M., Kuperwasser, C., Bierie, B., and Weinberg, R. A. (2011) Normal and neoplastic nonstem cells can spontaneously convert to a stem-like state, Proc. Natl. Acad. Sci. USA, 108, 7950-7955, https://doi.org/10.1073/pnas.1102454108.
Borovski, T., Melo, F. D. E., Vermeulen, L., and Medema, J. P. (2011) Cancer stem cell niche: the place to be, Cancer Res., 71, 634-639, https://doi.org/10.1158/0008-5472.can-10-3220.
Plaks, V., Kong, N. W., and Werb, Z. (2015) The cancer stem cell niche: how essential is the niche in regulating stemness of tumor cells? Cell Stem Cell, 16, 225-238, https://doi.org/10.1016/j.stem.2015.02.015.
Kim, W. T., and Ryu, C. J. (2017) Cancer stem cell surface markers on normal stem cells, Bmb Rep., 50, 285-298, https://doi.org/10.5483/BMBRep.2017.50.6.039.
Bonnet, D., and Dick, J. E. (1997) Human acute myeloid leukemia is organized as a hierarchy that originates from a primitive hematopoietic cell, Nat. Med., 3, 730-737, https://doi.org/10.1038/nm0797-730.
Singh, S. K., Hawkins, C., Clarke, I. D., Squire, J. A., Bayani, J., Hide, T., Henkelman, R. M., Cusimano, M. D., and Dirks, P. B. (2004) Identification of human brain tumour initiating cells, Nature, 432, 396-401, https://doi.org/10.1038/nature03128.
Liu, G. T., Yuan, X. P., Zeng, Z. H., Tunici, P., Ng, H. S., Abdulkadir, I. R., Lu, L. Z., Irvin, D., Black, K. L., and Yu, J. S. (2006) Analysis of gene expression and chemoresistance of CDI33+ cancer stem cells in glioblastoma, Mol. Cancer, 5, 67, https://doi.org/10.1186/1476-4598-5-67.
Medema, J. P. (2013) Cancer stem cells: the challenges ahead, Nat. Cell Biol., 15, 338-344, https://doi.org/10.1038/ncb2717.
Gimple, R. C., Yang, K., Halbert, M. E., Agnihotri, S., and Rich, J. N. (2022) Brain cancer stem cells: resilience through adaptive plasticity and hierarchical heterogeneity, Nat. Rev. Cancer, 22, 497-514, https://doi.org/10.1038/s41568-022-00486-x.
De Blank, P., Fouladi, M., and Huse, J. T. (2020) Molecular markers and targeted therapy in pediatric low-grade glioma, J. Neurooncol., 150, 5-15, https://doi.org/10.1007/s11060-020-03529-1.
Di, W., Fan, W., Wu, F., Shi, Z., Wang, Z., Yu, M., Zhai, Y., Chang, Y., Pan, C., Li, G., Kahlert, U. D., and Zhang, W. (2022) Clinical characterization and immunosuppressive regulation of CD161 (KLRB1) in glioma through 916 samples, Cancer Sci., 113, 756-769, https://doi.org/10.1111/cas.15236.
Birko, Z., Nagy, B., Klekner, A., and Virga, J. (2020) Novel molecular markers in glioblastoma-benefits of liquid biopsy, Int. J. Mol. Sci., 21, 7522, https://doi.org/10.3390/ijms21207522.
Srivastava, M., Ahlawat, N., and Srivastava, A. (2021) Ovarian cancer stem cells: newer horizons, J. Obstet. Gynaecol. India, 71, 115-117, https://doi.org/10.1007/s13224-020-01412-7.
Barani, M., Bilal, M., Sabir, F., Rahdar, A., and Kyzas, G. Z. (2021) Nanotechnology in ovarian cancer: diagnosis and treatment, Life Sci., 266, 118914, https://doi.org/10.1016/j.lfs.2020.118914.
Zhang, R., Siu, M. K. Y., Ngan, H. Y. S., and Chan, K. K. L. (2022) Molecular biomarkers for the early detection of ovarian cancer, Int. J. Mol. Sci., 23, 12041, https://doi.org/10.3390/ijms231912041.
Escudero-Lourdes, C., Alvarado-Morales, I., and Tokar, E. J. (2022) Stem cells as target for prostate cancer therapy: opportunities and challenges, Stem Cell Rev. Rep., 18, 2833-2851, https://doi.org/10.1007/s12015-022-10437-6.
Kerr, B. A., Miocinovic, R., Smith, A. K., West, X. Z., Watts, K. E., Alzayed, A. W., Klink, J. C., Mir, M. C., Sturey, T., Hansel, D. E., Heston, W. D., Stephenson, A. J., Klein, E. A., and Byzova, T. V. (2015) CD117+ cells in the circulation are predictive of advanced prostate cancer, Oncotarget, 6, 1889-1897, https://doi.org/10.18632/oncotarget.2796.
Zhang, K., Zhou, S., Wang, L., Wang, J., Zou, Q., Zhao, W., Fu, Q., and Fang, X. (2016) Current stem cell biomarkers and their functional mechanisms in prostate cancer, Int. J. Mol. Sci., 17, 1163, https://doi.org/10.3390/ijms17071163.
Gupta, R., Bhatt, L. K., Johnston, T. P., and Prabhavalkar, K. S. (2019) Colon cancer stem cells: potential target for the treatment of colorectal cancer, Cancer Biol. Ther., 20, 1068-1082, https://doi.org/10.1080/15384047.2019.1599660.
Chen, K., Huang, Y. H., and Chen, J. L. (2013) Understanding and targeting cancer stem cells: therapeutic implications and challenges, Acta Pharmacol. Sin., 34, 732-740, https://doi.org/10.1038/aps.2013.27.
Ishiwata, T., Matsuda, Y., Yoshimura, H., Sasaki, N., Ishiwata, S., Ishikawa, N., Takubo, K., Arai, T., and Aida, J. (2018) Pancreatic cancer stem cells: features and detection methods, Pathol. Oncol. Res., 24, 797-805, https://doi.org/10.1007/s12253-018-0420-x.
Immervoll, H., Hoem, D., Steffensen, O. J., Miletic, H., and Molven, A. (2011) Visualization of CD44 and CD133 in normal pancreas and pancreatic ductal adenocarcinomas: non-overlapping membrane expression in cell populations positive for both markers, J. Histochem. Cytochem., 59, 441-455, https://doi.org/10.1369/0022155411398275.
Patil, K., Khan, F. B., Akhtar, S., Ahmad, A., and Uddin, S. (2021) The plasticity of pancreatic cancer stem cells: implications in therapeutic resistance, Cancer Metastasis Rev., 40, 691-720, https://doi.org/10.1007/s10555-021-09979-x.
Gu, Y., Zheng, X., and Ji, J. (2020) Liver cancer stem cells as a hierarchical society: yes or no? Acta Biochim. Biophys. Sin. (Shanghai), 52, 723-735, https://doi.org/10.1093/abbs/gmaa050.
Fang, X., Yan, Q., Liu, S., and Guan, X. Y. (2022) Cancer stem cells in hepatocellular carcinoma: intrinsic and extrinsic molecular mechanisms in stemness regulation, Int. J. Mol. Sci., 23, 12327, https://doi.org/10.3390/ijms232012327.
Nio, K., Yamashita, T., and Kaneko, S. (2017) The evolving concept of liver cancer stem cells, Mol. Cancer, 16, 4, https://doi.org/10.1186/s12943-016-0572-9.
Joshi, P., Ghadi, D. S., and Waghmare, S. K. (2022) Isolation of cancer stem cells from skin squamous cell carcinoma, Methods Cell Biol., 171, 63-80, https://doi.org/10.1016/bs.mcb.2022.06.002.
Yin, Q., Shi, X., Lan, S., Jin, H., and Wu, D. (2021) Effect of melanoma stem cells on melanoma metastasis, Oncol. Lett., 22, 566, https://doi.org/10.3892/ol.2021.12827.
Zheng, Y., Wang, L., Yin, L., Yao, Z., Tong, R., Xue, J., and Lu, Y. (2022) Lung cancer stem cell markers as therapeutic targets: an update on signaling pathways and therapies, Front. Oncol., 12, 873994, https://doi.org/10.3389/fonc.2022.873994.
Rowbotham, S. P., Goruganthu, M. U. L., Arasada, R. R., Wang, W. Z., Carbone, D. P., and Kim, C. F. (2022) Lung cancer stem cells and their clinical implications, Cold Spring Harb. Perspect. Med., 12, a041270, https://doi.org/10.1101/cshperspect.a041270.
Pustovalova, M., Blokhina, T., Alhaddad, L., Chigasova, A., Chuprov-Netochin, R., Veviorskiy, A., Filkov, G., Osipov, A. N., and Leonov, S. (2022) CD44+ and CD133+ non-small cell lung cancer cells exhibit DNA damage response pathways and dormant polyploid giant cancer cell enrichment relating to their p53 status, Int. J. Mol. Sci., 23, 4922, https://doi.org/10.3390/ijms23094922.
Butti, R., Gunasekaran, V. P., Kumar, T. V. S., Banerjee, P., and Kundu, G. C. (2019) Breast cancer stem cells: Biology and therapeutic implications, Int. J. Biochem. Cell Biol., 107, 38-52, https://doi.org/10.1016/j.biocel.2018.12.001.
Wright, M. H., Calcagno, A. M., Salcido, C. D., Carlson, M. D., Ambudkar, S. V., and Varticovski, L. (2008) Brca1 breast tumors contain distinct CD44+/CD24– and CD133+ cells with cancer stem cell characteristics, Breast Cancer Res., 10, R10, https://doi.org/10.1186/bcr1855.
Farace, C., Pisano, A., Grinan-Lison, C., Solinas, G., Jimenez, G., Serra, M., Carrillo, E., Scognamillo, F., Attene, F., Montella, A., Marchal, J. A., and Madeddu, R. (2020) Deregulation of cancer-stem-cell-associated miRNAs in tissues and sera of colorectal cancer patients, Oncotarget, 11, 116-130, https://doi.org/10.18632/oncotarget.27411.
Aponte, P. M., and Caicedo, A. (2017) Stemness in cancer: stem cells, cancer stem cells, and their microenvironment, Stem Cells Int., 2017, 5619472, https://doi.org/10.1155/2017/5619472.
Visvader, J. E., and Clevers, H. (2016) Tissue-specific designs of stem cell hierarchies, Nat. Cell Biol., 18, 349-355, https://doi.org/10.1038/ncb3332.
Kaseb, H. O., Fohrer-Ting, H., Lewis, D. W., Lagasse, E., and Gollin, S. M. (2016) Identification, expansion and characterization of cancer cells with stem cell properties from head and neck squamous cell carcinomas, Exp. Cell Res., 348, 75-86, https://doi.org/10.1016/j.yexcr.2016.09.003.
Todaro, M., Alea, M. P., Di Stefano, A. B., Cammareri, P., Vermeulen, L., Lovino, F., Tripodo, C., Russo, A., Gulotta, G., Medema, J. P., and Stassi, G. (2007) Colon cancer stem cells dictate tumor growth and resist cell death by production of interleukin-4, Cell Stem Cell, 1, 389-402, https://doi.org/10.1016/j.stem.2007.08.001.
Li, C. W., Wu, J. J., Hynes, M., Dosch, J., Sarkar, B., Welling, T. H., di Magliano, M. P., and Simeone, D. M. (2011) c-Met is a marker of pancreatic cancer stem cells and therapeutic target, Gastroenterology, 141, 2218-2227.e5, https://doi.org/10.1053/j.gastro.2011.08.009.
Nassar, D., and Blanpain, C. (2016) Cancer stem cells: basic concepts and therapeutic implications, Annu. Rev. Pathol. Mech. Dis., 11, 47-76, https://doi.org/10.1146/annurev-pathol-012615-044438.
Laplane, L., and Solary, E. (2019) Towards a classification of stem cells, Elife, 8, e46563, https://doi.org/10.7554/eLife.46563.
Yang, L., Shi, P., Zhao, G., Xu, J., Peng, W., Zhang, J., Zhang, G., Wang, X., Dong, Z., Chen, F., and Cui, H. (2020) Targeting cancer stem cell pathways for cancer therapy, Signal Transduct. Target. Ther., 5, 8, https://doi.org/10.1038/s41392-020-0110-5.
Jerabek, S., Merino, F., Scholer, H. R., and Cojocaru, V. (2014) OCT4: dynamic DNA binding pioneers stem cell pluripotency, Biochim. Biophys. Acta, 1839, 138-154, https://doi.org/10.1016/j.bbagrm.2013.10.001.
Murakami, S., Ninomiya, W., Sakamoto, E., Shibata, T., Akiyama, H., and Tashiro, F. (2015) SRY and OCT4 are required for the acquisition of cancer stem cell-like properties and are potential differentiation therapy targets, Stem Cells, 33, 2652-2663, https://doi.org/10.1002/stem.2059.
Du, Z., Jia, D., Liu, S., Wang, F., Li, G., Zhang, Y., Cao, X., Ling, E. A., and Hao, A. (2009) Oct4 is expressed in human gliomas and promotes colony formation in glioma cells, Glia, 57, 724-733, https://doi.org/10.1002/glia.20800.
Song, B., Kim, D. K., Shin, J., Bae, S. H., Kim, H. Y., Won, B., Kim, J. K., Youn, H. D., Kim, S. T., Kang, S. W., and Jang, H. (2018) OCT4 directly regulates stemness and extracellular matrix-related genes in human germ cell tumours, Biochem. Biophys. Res. Commun., 503, 1980-1986, https://doi.org/10.1016/j.bbrc.2018.07.145.
Fujino, S., and Miyoshi, N. (2019) Oct4 gene expression in primary colorectal cancer promotes liver metastasis, Stem Cells Int., 2019, 7896524, https://doi.org/10.1155/2019/7896524.
Lu, C. S., Shiau, A. L., Su, B. H., Hsu, T. S., Wang, C. T., Su, Y. C., Tsai, M. S., Feng, Y. H., Tseng, Y. L., Yen, Y. T., Wu, C. L., and Shieh, G. S. (2020) Oct4 promotes M2 macrophage polarization through upregulation of macrophage colony-stimulating factor in lung cancer, J. Hematol. Oncol., 13, 62, https://doi.org/10.1186/s13045-020-00887-1.
Hagey, D. W., Klum, S., Kurtsdotter, I., Zaouter, C., Topcic, D., Andersson, O., Bergsland, M., and Muhr, J. (2018) SOX2 regulates common and specific stem cell features in the CNS and endoderm derived organs, PLoS Genet., 14, e1007224, https://doi.org/10.1371/journal.pgen.1007224.
Schaefer, T., and Lengerke, C. (2020) SOX2 protein biochemistry in stemness, reprogramming, and cancer: the PI3K/AKT/SOX2 axis and beyond, Oncogene, 39, 278-292, https://doi.org/10.1038/s41388-019-0997-x.
Liu, P., Tang, H., Song, C., Wang, J., Chen, B., Huang, X., Pei, X., and Liu, L. (2018) SOX2 promotes cell proliferation and metastasis in triple negative breast cancer, Front. Pharmacol., 9, 942, https://doi.org/10.3389/fphar.2018.00942.
Maurizi, G., Verma, N., Gadi, A., Mansukhani, A., and Basilico, C. (2018) Sox2 is required for tumor development and cancer cell proliferation in osteosarcoma, Oncogene, 37, 4626-4632, https://doi.org/10.1038/s41388-018-0292-2.
Han, S., Huang, T., Wu, X., Wang, X., Liu, S., Yang, W., Shi, Q., Li, H., and Hou, F. (2019) Prognostic value of CD133 and SOX2 in advanced cancer, J. Oncol., 2019, 3905817, https://doi.org/10.1155/2019/3905817.
Heurtier, V., Owens, N., Gonzalez, I., Mueller, F., Proux, C., Mornico, D., Clerc, P., Dubois, A., and Navarro, P. (2019) The molecular logic of Nanog-induced self-renewal in mouse embryonic stem cells, Nat. Commun., 10, 1109, https://doi.org/10.1038/s41467-019-09041-z.
Chiou, S. H., Wang, M. L., Chou, Y. T., Chen, C. J., Hong, C. F., Hsieh, W. J., Chang, H. T., Chen, Y. S., Lin, T. W., Hsu, H. S., and Wu, C. W. (2010) Coexpression of Oct4 and Nanog enhances malignancy in lung adenocarcinoma by inducing cancer stem cell-like properties and epithelial-mesenchymal transdifferentiation, Cancer Res., 70, 10433-10444, https://doi.org/10.1158/0008-5472.CAN-10-2638.
Lin, T., Ding, Y. Q., and Li, J. M. (2012) Overexpression of Nanog protein is associated with poor prognosis in gastric adenocarcinoma, Med. Oncol., 29, 878-885, https://doi.org/10.1007/s12032-011-9860-9.
De Vicente, J. C., Rodriguez-Santamarta, T., Rodrigo, J. P., Allonca, E., Vallina, A., Singhania, A., Donate-Perez Del Molino, P., and Garcia-Pedrero, J. M. (2019) The emerging role of NANOG as an early cancer risk biomarker in patients with oral potentially malignant disorders, J. Clin. Med., 8, 1376, https://doi.org/10.3390/jcm8091376.
Dehghan Harati, M., Rodemann, H. P., and Toulany, M. (2019) Nanog signaling mediates radioresistance in ALDH-positive breast cancer cells, Int. J. Mol. Sci., 20, 1151, https://doi.org/10.3390/ijms20051151.
Wang, X., Jin, J., Wan, F., Zhao, L., Chu, H., Chen, C., Liao, G., Liu, J., Yu, Y., Teng, H., Fang, L., Jiang, C., Pan, W., Xie, X., Li, J., Lu, X., Jiang, X., Ge, X., Ye, D., and Wang, P. (2019) AMPK promotes SPOP-mediated NANOG degradation to regulate prostate cancer cell stemness, Dev. Cell, 48, 345-360.e347, https://doi.org/10.1016/j.devcel.2018.11.033.
Ghaleb, A. M., and Yang, V. W. (2017) Kruppel-like factor 4 (KLF4): what we currently know, Gene, 611, 27-37, https://doi.org/10.1016/j.gene.2017.02.025.
Hsieh, M. H., Chen, Y. T., Chen, Y. T., Lee, Y. H., Lu, J., Chien, C. L., Chen, H. F., Ho, H. N., Yu, C. J., Wang, Z. Q., and Teng, S. C. (2017) PARP1 controls KLF4-mediated telomerase expression in stem cells and cancer cells, Nucleic Acids Res., 45, 10492-10503, https://doi.org/10.1093/nar/gkx683.
Riverso, M., Montagnani, V., and Stecca, B. (2017) KLF4 is regulated by RAS/RAF/MEK/ERK signaling through E2F1 and promotes melanoma cell growth, Oncogene, 36, 3322-3333, https://doi.org/10.1038/onc.2016.481.
Qi, X. T., Li, Y. L., Zhang, Y. Q., Xu, T., Lu, B., Fang, L., Gao, J. Q., Yu, L. S., Zhu, D. F., Yang, B., He, Q. J., and Ying, M. D. (2019) KLF4 functions as an oncogene in promoting cancer stem cell-like characteristics in osteosarcoma cells, Acta Pharmacol. Sin., 40, 546-555, https://doi.org/10.1038/s41401-018-0050-6.
Ding, X., Zhong, T., Jiang, L., Huang, J., Xia, Y., and Hu, R. (2018) miR-25 enhances cell migration and invasion in non-small-cell lung cancer cells via ERK signaling pathway by inhibiting KLF4, Mol. Med. Rep., 17, 7005-7016, https://doi.org/10.3892/mmr.2018.8772.
Wang, B., Shen, A., Ouyang, X., Zhao, G., Du, Z., Huo, W., Zhang, T., Wang, Y., Yang, C., Dong, P., Watari, H., Pfeffer, L. M., and Yue, J. (2017) KLF4 expression enhances the efficacy of chemotherapy drugs in ovarian cancer cells, Biochem. Biophys. Res. Commun., 484, 486-492, https://doi.org/10.1016/j.bbrc.2017.01.062.
Dang, C. V. (2012) MYC on the path to cancer, Cell, 149, 22-35, https://doi.org/10.1016/j.cell.2012.03.003.
Galardi, S., Savino, M., Scagnoli, F., Pellegatta, S., Pisati, F., Zambelli, F., Illi, B., Annibali, D., Beji, S., Orecchini, E., Alberelli, M. A., Apicella, C., Fontanella, R. A., Michienzi, A., Finocchiaro, G., Farace, M. G., Pavesi, G., Ciafre, S. A., and Nasi, S. (2016) Resetting cancer stem cell regulatory nodes upon MYC inhibition, EMBO Rep., 17, 1872-1889, https://doi.org/10.15252/embr.201541489.
Lourenco, C., Kalkat, M., Houlahan, K. E., De Melo, J., Longo, J., Done, S. J., Boutros, P. C., and Penn, L. Z. (2019) Modelling the MYC-driven normal-to-tumour switch in breast cancer, Dis. Model. Mech., 12, dmm038083, https://doi.org/10.1242/dmm.038083.
Dong, H., Hu, J., Wang, L., Qi, M., Lu, N., Tan, X., Yang, M., Bai, X., Zhan, X., and Han, B. (2019) SOX4 is activated by C-MYC in prostate cancer, Med. Oncol., 36, 92, https://doi.org/10.1007/s12032-019-1317-6.
Fabregat, I., Malfettone, A., and Soukupova, J. (2016) New insights into the crossroads between EMT and stemness in the context of cancer, J. Clin. Med., 5, 37, https://doi.org/10.3390/jcm5030037.
Katoh, M. (2007) Networking of WNT, FGF, Notch, BMP, and Hedgehog signaling pathways during carcinogenesis, Stem Cell Rev., 3, 30-38, https://doi.org/10.1007/s12015-007-0006-6.
Visvader, J. E., and Lindeman, G. J. (2012) Cancer stem cells: current status and evolving complexities, Cell Stem Cell, 10, 717-728, https://doi.org/10.1016/j.stem.2012.05.007.
Clements, W. M., Wang, J., Sarnaik, A., Kim, O. J., MacDonald, J., Fenoglio-Preiser, C., Groden, J., and Lowy, A. M. (2002) beta-Catenin mutation is a frequent cause of Wnt pathway activation in gastric cancer, Cancer Res., 62, 3503-3506.
Abd El-Rehim, D., and Ali, M. M. (2009) Aberrant expression of beta-catenin in invasive ductal breast carcinomas, J. Egypt. Natl. Canc. Inst., 21, 185-195.
Kudo, J., Nishiwaki, T., Haruki, N., Ishiguro, H., Shibata, Y., Terashita, Y., Sugiura, H., Shinoda, N., Kimura, M., Kuwabara, Y., and Fujii, Y. (2007) Aberrant nuclear localization of β-catenin without genetic alterations in beta-catenin or Axin genes in esophageal cancer, World J. Surg. Oncol., 5, 21, https://doi.org/10.1186/1477-7819-5-21.
Zhao, X., Jiang, C., Xu, R., Liu, Q., Liu, G., and Zhang, Y. (2020) TRIP6 enhances stemness property of breast cancer cells through activation of Wnt/beta-catenin, Cancer Cell Int., 20, 51, https://doi.org/10.1186/s12935-020-1136-z.
Zhu, L., Pan, R., Zhou, D., Ye, G., and Tan, W. (2019) BCL11A enhances stemness and promotes progression by activating Wnt/beta-catenin signaling in breast cancer, Cancer Manag. Res., 11, 2997-3007, https://doi.org/10.2147/CMAR.S199368.
Zhang, L., Dong, X., Yan, B., Yu, W., and Shan, L. (2020) CircAGFG1 drives metastasis and stemness in colorectal cancer by modulating YY1/CTNNB1, Cell Death Dis., 11, 542, https://doi.org/10.1038/s41419-020-2707-6.
Xiang, X., Xiong, R., Yu, C., Deng, L., Bie, J., Xiao, D., Chen, Z., Zhou, Y., Li, X., Liu, K., and Feng, G. (2019) Tex10 promotes stemness and EMT phenotypes in esophageal squamous cell carcinoma via the Wnt/betacatenin pathway, Oncol. Rep., 42, 2600-2610, https://doi.org/10.3892/or.2019.7376.
Stylianou, S., Clarke, R. B., and Brennan, K. (2006) Aberrant activation of notch signaling in human breast cancer, Cancer Res., 66, 1517-1525, https://doi.org/10.1158/0008-5472.CAN-05-3054.
Li, L., Tang, P., Li, S., Qin, X., Yang, H., Wu, C., and Liu, Y. (2017) Notch signaling pathway networks in cancer metastasis: a new target for cancer therapy, Med. Oncol., 34, 180, https://doi.org/10.1007/s12032-017-1039-6.
Katoh, M., and Katoh, M. (2020) Precision medicine for human cancers with Notch signaling dysregulation (Review), Int. J. Mol. Med., 45, 279-297, https://doi.org/10.3892/ijmm.2019.4418.
Amantini, C., Morelli, M. B., Nabissi, M., Cardinali, C., Santoni, M., Gismondi, A., and Santoni, G. (2016) Capsaicin triggers autophagic cell survival which drives epithelial mesenchymal transition and chemoresistance in bladder cancer cells in an Hedgehog-dependent manner, Oncotarget, 7, 50180-50194, https://doi.org/10.18632/oncotarget.10326.
Villegas, V. E., Rondon-Lagos, M., Annaratone, L., Castellano, I., Grismaldo, A., Sapino, A., and Zaphiropoulos, P. G. (2016) Tamoxifen treatment of breast cancer cells: impact on Hedgehog/GLI1 signaling, Int. J. Mol. Sci., 17, 308, https://doi.org/10.3390/ijms17030308.
Jeng, K. S., Chang, C. F., and Lin, S. S. (2020) Sonic hedgehog signaling in organogenesis, tumors, and tumor microenvironments, Int. J. Mol. Sci., 21, 758, https://doi.org/10.3390/ijms21030758.
Petrova, R., and Joyner, A. L. (2014) Roles for Hedgehog signaling in adult organ homeostasis and repair, Development, 141, 3445-3457, https://doi.org/10.1242/dev.083691.
Po, A., Silvano, M., Miele, E., Capalbo, C., Eramo, A., Salvati, V., Todaro, M., Besharat, Z. M., Catanzaro, G., Cucchi, D., Coni, S., Di Marcotullio, L., Canettieri, G., Vacca, A., Stassi, G., De Smaele, E., Tartaglia, M., Screpanti, I., De Maria, R., and Ferretti, E. (2017) Noncanonical GLI1 signaling promotes stemness features and in vivo growth in lung adenocarcinoma, Oncogene, 36, 4641-4652, https://doi.org/10.1038/onc.2017.91.
Zhu, R., Gires, O., Zhu, L., Liu, J., Li, J., Yang, H., Ju, G., Huang, J., Ge, W., Chen, Y., Lu, Z., and Wang, H. (2019) TSPAN8 promotes cancer cell stemness via activation of sonic Hedgehog signaling, Nat. Commun., 10, 2863, https://doi.org/10.1038/s41467-019-10739-3.
Hayden, M. S., and Ghosh, S. (2008) Shared principles in NF-kappaB signaling, Cell, 132, 344-362, https://doi.org/10.1016/j.cell.2008.01.020.
Prasad, S., Rarnachandran, S., Gupta, N., Kaushik, I., and Srivastava, S. K. (2020) Cancer cells stemness: A doorstep to targeted therapy, Biochim. Biophys. Acta Mol. Bas. Dis., 1866, 165424, https://doi.org/10.1016/j.bbadis.2019.02.019.
Prasad, S., Ravindran, J., and Aggarwal, B. B. (2010) NF-kappaB and cancer: how intimate is this relationship, Mol. Cell. Biochem., 336, 25-37, https://doi.org/10.1007/s11010-009-0267-2.
Van der Zee, M., Sacchetti, A., Cansoy, M., Joosten, R., Teeuwssen, M., Heijmans-Antonissen, C., Ewing-Graham, P. C., Burger, C. W., Blok, L. J., and Fodde, R. (2015) IL6/JAK1/STAT3 Signaling blockade in endometrial cancer affects the ALDHhi/CD126+ stem-like component and reduces tumor burden, Cancer Res., 75, 3608-3622, https://doi.org/10.1158/0008-5472.CAN-14-2498.
Yang, L., Dong, Y., Li, Y., Wang, D., Liu, S., Wang, D., Gao, Q., Ji, S., Chen, X., Lei, Q., Jiang, W., Wang, L., Zhang, B., Yu, J. J., and Zhang, Y. (2019) IL-10 derived from M2 macrophage promotes cancer stemness via JAK1/STAT1/NF-kappaB/Notch1 pathway in non-small cell lung cancer, Int. J. Cancer, 145, 1099-1110, https://doi.org/10.1002/ijc.32151.
Park, S. Y., Lee, C. J., Choi, J. H., Kim, J. H., Kim, J. W., Kim, J. Y., and Nam, J. S. (2019) The JAK2/STAT3/CCND2 axis promotes colorectal cancer stem cell persistence and radioresistance, J. Exp. Clin. Cancer Res., 38, 399, https://doi.org/10.1186/s13046-019-1405-7.
Toh, T. B., Lim, J. J., Hooi, L., Rashid, M., and Chow, E. K. (2020) Targeting Jak/Stat pathway as a therapeutic strategy against SP/CD44+ tumorigenic cells in Akt/beta-catenin-driven hepatocellular carcinoma, J. Hepatol., 72, 104-118, https://doi.org/10.1016/j.jhep.2019.08.035.
Chambers, I. (2004) The molecular basis of pluripotency in mouse embryonic stem cells, Cloning Stem Cells, 6, 386-391, https://doi.org/10.1089/clo.2004.6.386.
Zhou, J., Wulfkuhle, J., Zhang, H., Gu, P., Yang, Y., Deng, J., Margolick, J. B., Liotta, L. A., Petricoin, E., 3rd, and Zhang, Y. (2007) Activation of the PTEN/mTOR/STAT3 pathway in breast cancer stem-like cells is required for viability and maintenance, Proc. Natl. Acad. Sci. USA, 104, 16158-16163, https://doi.org/10.1073/pnas.0702596104.
Kaowinn, S., Kaewpiboon, C., Koh, S. S., Kramer, O. H., and Chung, Y. H. (2018) STAT1HDAC4 signaling induces epithelialmesenchymal transition and sphere formation of cancer cells overexpressing the oncogene, CUG2, Oncol. Rep., 40, 2619-2627, https://doi.org/10.3892/or.2018.6701.
Dey, N., De, P., and Leyland-Jones, B. (2017) PI3K-AKT-mTOR inhibitors in breast cancers: from tumor cell signaling to clinical trials, Pharmacol. Ther., 175, 91-106, https://doi.org/10.1016/j.pharmthera.2017.02.037.
Tan, A. C. (2020) Targeting the PI3K/Akt/mTOR pathway in non-small cell lung cancer (NSCLC), Thorac. Cancer, 11, 511-518, https://doi.org/10.1111/1759-7714.13328.
Karami Fath, M., Ebrahimi, M., Nourbakhsh, E., Zia Hazara, A., Mirzaei, A., Shafieyari, S., Salehi, A., Hoseinzadeh, M., Payandeh, Z., and Barati, G. (2022) PI3K/Akt/mTOR signaling pathway in cancer stem cells, Pathol. Res. Pract., 237, 154010, https://doi.org/10.1016/j.prp.2022.154010.
Nepstad, I., Hatfield, K. J., Gronningsaeter, I. S., and Reikvam, H. (2020) The PI3K-Akt-mTOR signaling pathway in human acute myeloid leukemia (AML) cells, Int. J. Mol. Sci., 21, 2907, https://doi.org/10.3390/ijms21082907.
Madsen, R. R. (2020) PI3K in stemness regulation: from development to cancer, Biochem. Soc. Trans., 48, 301-315, https://doi.org/10.1042/BST20190778.
Fitzgerald, T. L., Lertpiriyapong, K., Cocco, L., Martelli, A. M., Libra, M., Candido, S., Montalto, G., Cervello, M., Steelman, L., Abrams, S. L., and McCubrey, J. A. (2015) Roles of EGFR and KRAS and their downstream signaling pathways in pancreatic cancer and pancreatic cancer stem cells, Adv. Biol. Regul., 59, 65-81, https://doi.org/10.1016/j.jbior.2015.06.003.
Nangia-Makker, P., Hogan, V., and Raz, A. (2018) Galectin-3 and cancer stemness, Glycobiology, 28, 172-181, https://doi.org/10.1093/glycob/cwy001.
Li, Y., Hu, H., Wang, Y., Fan, Y., Yang, Y., Guo, B., Xie, X., Lian, J., Jiang, B., Han, B., Wang, Y., Shao, C., and Gong, Y. (2020) CUL4B contributes to cancer stemness by repressing tumor suppressor miR34a in colorectal cancer, Oncogenesis, 9, 20, https://doi.org/10.1038/s41389-020-0206-3.
Ramadoss, S., Sen, S., Ramachandran, I., Roy, S., Chaudhuri, G., and Farias-Eisner, R. (2017) Lysine-specific demethylase KDM3A regulates ovarian cancer stemness and chemoresistance, Oncogene, 36, 1537-1545, https://doi.org/10.1038/onc.2016.320.
Kim, H. Y., Kim, D. K., Bae, S. H., Gwak, H., Jeon, J. H., Kim, J. K., Lee, B. I., You, H. J., Shin, D. H., Kim, Y. H., Kim, S. Y., Han, S. S., Shim, J. K., Lee, J. H., Kang, S. G., and Jang, H. (2018) Farnesyl diphosphate synthase is important for the maintenance of glioblastoma stemness, Exp. Mol. Med., 50, 1-12, https://doi.org/10.1038/s12276-018-0166-2.
Mei, Y., Cai, D., and Dai, X. (2020) Modulating cancer stemness provides luminal a breast cancer cells with HER2 positive-like features, J. Cancer, 11, 1162-1169, https://doi.org/10.7150/jca.37117.
Ouspenskaia, T., Matos, I., Mertz, A. F., Fiore, V. F., and Fuchs, E. (2016) WNT-SHH antagonism specifies and expands stem cells prior to niche formation, Cell, 164, 156-169, https://doi.org/10.1016/j.cell.2015.11.058.
Wahlster, L., and Daley, G. Q. (2016) Progress towards generation of human haematopoietic stem cells, Nat. Cell. Biol., 18, 1111-1117, https://doi.org/10.1038/ncb3419.
Beck, B., Driessens, G., Goossens, S., Youssef, K. K., Kuchnio, A., Caauwe, A., Sotiropoulou, P. A., Loges, S., Lapouge, G., Candi, A., Mascre, G., Drogat, B., Dekoninck, S., Haigh, J. J., Carmeliet, P., and Blanpain, C. (2011) A vascular niche and a VEGF-Nrp1 loop regulate the initiation and stemness of skin tumours, Nature, 478, 399-403, https://doi.org/10.1038/nature10525.
Zhang, Z., Dong, Z., Lauxen, I. S., Filho, M. S., and Nor, J. E. (2014) Endothelial cell-secreted EGF induces epithelial to mesenchymal transition and endows head and neck cancer cells with stem-like phenotype, Cancer Res., 74, 2869-2881, https://doi.org/10.1158/0008-5472.CAN-13-2032.
Yu, Y., Xiao, C. H., Tan, L. D., Wang, Q. S., Li, X. Q., and Feng, Y. M. (2014) Cancer-associated fibroblasts induce epithelial-mesenchymal transition of breast cancer cells through paracrine TGF-beta signalling, Br. J. Cancer, 110, 724-732, https://doi.org/10.1038/bjc.2013.768.
Bao, B., Azmi, A. S., Ali, S., Ahmad, A., Li, Y., Banerjee, S., Kong, D., and Sarkar, F. H. (2012) The biological kinship of hypoxia with CSC and EMT and their relationship with deregulated expression of miRNAs and tumor aggressiveness, Biochim. Biophys. Acta, 1826, 272-296, https://doi.org/10.1016/j.bbcan.2012.04.008.
Joseph, J. V., Conroy, S., Pavlov, K., Sontakke, P., Tomar, T., Eggens-Meijer, E., Balasubramaniyan, V., Wagemakers, M., den Dunnen, W. F., and Kruyt, F. A. (2015) Hypoxia enhances migration and invasion in glioblastoma by promoting a mesenchymal shift mediated by the HIF1alpha-ZEB1 axis, Cancer Lett., 359, 107-116, https://doi.org/10.1016/j.canlet.2015.01.010.
Zhang, M., Xu, C., Wang, H. Z., Peng, Y. N., Li, H. O., Zhou, Y. J., Liu, S., Wang, F., Liu, L., Chang, Y., Zhao, Q., and Liu, J. (2019) Soft fibrin matrix downregulates DAB2IP to promote Nanog-dependent growth of colon tumor-repopulating cells, Cell Death Dis., 10, 151, https://doi.org/10.1038/s41419-019-1309-7.
Valadão, I. C., Ralph, A. C. L., Bordeleau, F., Dzik, L. M., Borbely, K. S. C., Geraldo, M. V., Reinhart-King, C. A., and Freitas, V. M. (2020) High type I collagen density fails to increase breast cancer stem cell phenotype, PeerJ, 8, e9153, https://doi.org/10.7717/peerj.9153.
Zhang, C., Ma, K., and Li, W. Y. (2019) IL-6 promotes cancer stemness and oncogenicity in U2OS and MG-63 osteosarcoma cells by upregulating the OPN-STAT3 pathway, J. Cancer, 10, 6511-6525, https://doi.org/10.7150/jca.29931.
Li, Y., Wang, L., Pappan, L., Galliher-Beckley, A., and Shi, J. (2012) IL-1beta promotes stemness and invasiveness of colon cancer cells through Zeb1 activation, Mol. Cancer, 11, 87, https://doi.org/10.1186/1476-4598-11-87.
Hong, H. S., Akhavan, J., Lee, S. H., Kim, R. H., Kang, M. K., Park, N. H., and Shin, K. H. (2020) Proinflammatory cytokine TNFalpha promotes HPV-associated oral carcinogenesis by increasing cancer stemness, Int. J. Oral Sci., 12, 3, https://doi.org/10.1038/s41368-019-0069-7.
Bronisz, A., Wang, Y., Nowicki, M. O., Peruzzi, P., Ansari, K. I., Ogawa, D., Balaj, L., De Rienzo, G., Mineo, M., Nakano, I., Ostrowski, M. C., Hochberg, F., Weissleder, R., Lawler, S. E., Chiocca, E. A., and Godlewski, J. (2014) Extracellular vesicles modulate the glioblastoma microenvironment via a tumor suppression signaling network directed by miR-1, Cancer Res., 74, 738-750, https://doi.org/10.1158/0008-5472.Can-13-2650.
Hu, Y. B., Yan, C., Mu, L., Huang, K. Y., Li, X. L., Tao, D. D., Wu, Y. Q., and Qin, J. C. (2015) Fibroblast-derived exosomes contribute to chemoresistance through priming cancer stem cells in colorectal cancer, PLoS One, 10, e0125625, https://doi.org/10.1371/journal.pone.0125625.
Bruno, S., Collino, F., Deregibus, M. C., Grange, C., Tetta, C., and Camussi, G. (2013) Microvesicles derived from human bone marrow mesenchymal stem cells inhibit tumor growth, Stem Cells Dev., 22, 758-771, https://doi.org/10.1089/scd.2012.0304.
Zhang, X., Tu, H., Yang, Y., Fang, L., Wu, Q., and Li, J. (2017) Mesenchymal stem cell-derived extracellular vesicles: roles in tumor growth, progression, and drug resistance, Stem Cells Int., 2017, 1758139, https://doi.org/10.1155/2017/1758139.
Weiswald, L. B., Bellet, D., and Dangles-Marie, V. (2015) Spherical cancer models in tumor biology, Neoplasia, 17, 1-15, https://doi.org/10.1016/j.neo.2014.12.004.
Gupta, P. B., Onder, T. T., Jiang, G., Tao, K., Kuperwasser, C., Weinberg, R. A., and Lander, E. S. (2009) Identification of selective inhibitors of cancer stem cells by high-throughput screening, Cell, 138, 645-659, https://doi.org/10.1016/j.cell.2009.06.034.
Oskarsson, T., Acharyya, S., Zhang, X. H., Vanharanta, S., Tavazoie, S. F., Morris, P. G., Downey, R. J., Manova-Todorova, K., Brogi, E., and Massague, J. (2011) Breast cancer cells produce tenascin C as a metastatic niche component to colonize the lungs, Nat. Med., 17, 867-874, https://doi.org/10.1038/nm.2379.
Ishiguro, T., Ohata, H., Sato, A., Yamawaki, K., Enomoto, T., and Okamoto, K. (2017) Tumor-derived spheroids: relevance to cancer stem cells and clinical applications, Cancer Sci., 108, 283-289, https://doi.org/10.1111/cas.13155.
Singh, S. K., Clarke, I. D., Terasaki, M., Bonn, V. E., Hawkins, C., Squire, J., and Dirks, P. B. (2003) Identification of a cancer stem cell in human brain tumors, Cancer Res., 63, 5821-5828.
Ponti, D., Costa, A., Zaffaroni, N., Pratesi, G., Petrangolini, G., Coradini, D., Pilotti, S., Pierotti, M. A., and Daidone, M. G. (2005) Isolation and in vitro propagation of tumorigenic breast cancer cells with stem/progenitor cell properties, Cancer Res., 65, 5506-5511, https://doi.org/10.1158/0008-5472.CAN-05-0626.
Gilazieva, Z., Ponomarev, A., Rutland, C., Rizvanov, A., and Solovyeva, V. (2020) Promising applications of tumor spheroids and organoids for personalized medicine, Cancers (Basel), 12, 2727, https://doi.org/10.3390/cancers12102727.
Han, S. J., Kwon, S., and Kim, K. S. (2021) Challenges of applying multicellular tumor spheroids in preclinical phase, Cancer Cell Int., 21, 152, https://doi.org/10.1186/s12935-021-01853-8.
Hirschhaeuser, F., Menne, H., Dittfeld, C., West, J., Mueller-Klieser, W., and Kunz-Schughart, L. A. (2010) Multicellular tumor spheroids: an underestimated tool is catching up again, J. Biotechnol., 148, 3-15, https://doi.org/10.1016/j.jbiotec.2010.01.012.
Costa, E. C., Moreira, A. F., de Melo-Diogo, D., Gaspar, V. M., Carvalho, M. P., and Correia, I. J. (2016) 3D tumor spheroids: an overview on the tools and techniques used for their analysis, Biotechnol. Adv., 34, 1427-1441, https://doi.org/10.1016/j.biotechadv.2016.11.002.
Jensen, C., and Teng, Y. (2020) Is it time to start transitioning from 2D to 3D cell culture? Front. Mol. Biosci., 7, 33, https://doi.org/10.3389/fmolb.2020.00033.
Forte, E., Chimenti, I., Rosa, P., Angelini, F., Pagano, F., Calogero, A., Giacomello, A., and Messina, E. (2017) EMT/MET at the crossroad of stemness, regeneration and oncogenesis: the ying-yang equilibrium recapitulated in cell spheroids, Cancers, 9, 98, https://doi.org/10.3390/cancers9080098.
Eyler, C. E., Foo, W. C., Lafiura, K. M., McLendon, R. E., Hjelmeland, A. B., and Rich, J. N. (2008) Brain cancer stem cells display preferential sensitivity to Akt inhibition, Stem Cells, 26, 3027-3036, https://doi.org/10.1634/stemcells.2007-1073.
Lindemann, R. K. (2008) Stroma-initiated hedgehog signaling takes center stage in B-cell lymphoma, Cancer Res., 68, 961-964, https://doi.org/10.1158/0008-5472.can-07-5500.
Namiki, K., Wongsirisin, P., Yokoyama, S., Sato, M., Rawangkan, A., Sakai, R., Iida, K., and Suganuma, M. (2020) (–)-Epigallocatechin gallate inhibits stemness and tumourigenicity stimulated by AXL receptor tyrosine kinase in human lung cancer cells, Sci. Rep., 10, 2444, https://doi.org/10.1038/s41598-020-59281-z.
Fendler, A., Bauer, D., Busch, J., Jung, K., Wulf-Goldenberg, A., Kunz, S., Song, K., Myszczyszyn, A., Elezkurtaj, S., Erguen, B., Jung, S., Chen, W., and Birchmeier, W. (2020) Inhibiting WNT and NOTCH in renal cancer stem cells and the implications for human patients, Nat. Commun., 11, 929, https://doi.org/10.1038/s41467-020-14700-7.
Stewart, M., and Fox, S. E. (1989) Firing relations of medial septal neurons to the hippocampal theta-rhythm in urethane anesthetized rats, Exp. Brain Res., 77, 507-516, https://doi.org/10.1007/BF00249604.
Qureshi-Baig, K., Ullmann, P., Rodriguez, F., Frasquilho, S., Nazarov, P. V., Haan, S., and Letellier, E. (2016) What do we learn from spheroid culture systems? Insights from tumorspheres derived from primary colon cancer tissue, PLoS One, 11, e0146052, https://doi.org/10.1371/journal.pone.0146052.
Zhang, S., Balch, C., Chan, M. W., Lai, H. C., Matei, D., Schilder, J. M., Yan, P. S., Huang, T. H. M., and Nephew, K. P. (2008) Identification and characterization of ovarian cancer-initiating cells from primary human tumors, Cancer Res., 68, 4311-4320, https://doi.org/10.1158/0008-5472.can-08-0364.
Rashidi, M. R. W., Mehta, P., Bregenzer, M., Raghavan, S., Fleck, E. M., Horst, E. N., Harissa, Z., Ravikumar, V., Brady, S., Bild, A., Rao, A., Buckanovich, R. J., and Mehta, G. (2019) Engineered 3D Model of Cancer Stem Cell Enrichment and Chemoresistance, Neoplasia, 21, 822-836, https://doi.org/10.1016/j.neo.2019.06.005.
Bahmad, H. F., Cheaito, K., Chalhoub, R. M., Hadadeh, O., Monzer, A., Ballout, F., El-Hajj, A., Mukherji, D., Liu, Y. N., Daoud, G., and Abou-Kheir, W. (2018) Sphere-formation assay: three-dimensional in vitro culturing of prostate cancer stem/progenitor sphere-forming cells, Front. Oncol., 8, 347, https://doi.org/10.3389/fonc.2018.00347.
Rybak, A. P., He, L. Z., Kapoor, A., Cutz, J. C., and Tang, D. (2011) Characterization of sphere-propagating cells with stem-like properties from DU145 prostate cancer cells, Biochim. Biophys. Acta Mol. Cell Res., 1813, 683-694, https://doi.org/10.1016/j.bbamcr.2011.01.018.
Herheliuk, T., Perepelytsina, O., Ugnivenko, A., Ostapchenko, L., and Sydorenko, M. (2019) Investigation of multicellular tumor spheroids enriched for a cancer stem cell phenotype, Stem Cell Invest., 6, 21-21, https://doi.org/10.21037/sci.2019.06.07.
Cao, L., Zhou, Y. M., Zhai, B. B., Liao, J., Xu, W., Zhang, R. X., Li, J., Zhang, Y., Chen, L., Qian, H. H., Wu, M. C., and Yin, Z. F. (2011) Sphere-forming cell subpopulations with cancer stem cell properties in human hepatoma cell lines, BMC Gastroenterol., 11, 71, https://doi.org/10.1186/1471-230x-11-71.
Maliszewska-Olejniczak, K., Brodaczewska, K. K., Bielecka, Z. F., Solarek, W., Kornakiewicz, A., Szczylik, C., Porta, C., and Czarnecka, A. M. (2019) Development of extracellular matrix supported 3D culture of renal cancer cells and renal cancer stem cells, Cytotechnology, 71, 149-163, https://doi.org/10.1007/s10616-018-0273-x.
Chen, L., Xiao, Z. F., Meng, Y., Zhao, Y. N., Han, J., Su, G. N., Chen, B., and Dai, J. W. (2012) The enhancement of cancer stem cell properties of MCF-7 cells in 3D collagen scaffolds for modeling of cancer and anti-cancer drugs, Biomaterials, 33, 1437-1444, https://doi.org/10.1016/j.biomaterials.2011.10.056.
Farace, C., Oliver, J. A., Melguizo, C., Alvarez, P., Bandiera, P., Rama, A. R., Malaguarnera, G., Ortiz, R., Madeddu, R., and Prados, J. (2015) Microenvironmental modulation of decorin and lumican in temozolomide-resistant glioblastoma and neuroblastoma cancer stem-like cells, PLoS One, 10, e0134111, https://doi.org/10.1371/journal.pone.0134111.
Gao, W. J., Wu, D. L., Wang, Y. L., Wang, Z., Zou, C., Dai, Y., Ng, C. F., Teoh, J. Y. C., and Chan, F. L. (2018) Development of a novel and economical agar-based non-adherent three-dimensional culture method for enrichment of cancer stem-like cells, Stem Cell Res. Ther., 9, 243, https://doi.org/10.1186/s13287-018-0987-x.
Chen, S. F., Chang, Y. C., Nieh, S., Liu, C. L., Yang, C. Y., and Lin, Y. S. (2012) Nonadhesive culture system as a model of rapid sphere formation with cancer stem cell properties, PLoS One, 7, e31864, https://doi.org/10.1371/journal.pone.0031864.
Zhu, Z. W., Chen, L., Liu, J. X., Huang, J. W., Wu, G., Zheng, Y. F., and Yao, K. T. (2018) A novel three-dimensional tumorsphere culture system for the efficient and low-cost enrichment of cancer stem cells with natural polymers, Exper. Ther. Med., 15, 85-92, https://doi.org/10.3892/etm.2017.5419.
Muenzner, J. K., Kunze, P., Lindner, P., Polaschek, S., Menke, K., Eckstein, M., Geppert, C. I., Chanvorachote, P., Baeuerle, T., Hartmann, A., and Schneider-Stock, R. (2018) Generation and characterization of hepatocellular carcinoma cell lines with enhanced cancer stem cell potential, J. Cell. Mol. Med., 22, 6238-6248, https://doi.org/10.1111/jcmm.13911.
Xue, J. G., Zhu, Y., Sun, Z. X., Ji, R. B., Zhang, X., Xu, W. R., Yuan, X., Zhang, B., Yan, Y. M., Yin, L., Xu, H. J., Zhang, L. L., Zhu, W., and Qian, H. (2015) Tumorigenic hybrids between mesenchymal stem cells and gastric cancer cells enhanced cancer proliferation, migration and stemness, BMC Cancer, 15, 793, https://doi.org/10.1186/s12885-015-1780-1.
Nath, S., and Devi, G. R. (2016) Three-dimensional culture systems in cancer research: focus on tumor spheroid model, Pharmacol. Ther., 163, 94-108, https://doi.org/10.1016/j.pharmthera.2016.03.013.
Acknowledgments
This work was supported by the Strategic Academic Leadership Program of Kazan (Volga Region) Federal University (PRIORITY-2030).
Funding
This work was supported by the Russian Science Foundation (grant no. 21-74-10021).
Author information
Authors and Affiliations
Contributions
V.V.S., A.A.R. – concept and management of the study; A.S.P., Z.E.G. – text writing and text editing.
Corresponding author
Ethics declarations
The authors declare no conflict of interest in financial or any other sphere. This article does not describe any research involving humans or animals performed by any of the authors.
Rights and permissions
Open access. This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution, and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit https://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Ponomarev, A.S., Gilazieva, Z.E., Solovyova, V.V. et al. Molecular Mechanisms of Tumor Cell Stemness Modulation during Formation of Spheroids. Biochemistry Moscow 88, 979–994 (2023). https://doi.org/10.1134/S0006297923070106
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1134/S0006297923070106