Abstract
The action of the spliceosome depends on the stepwise cooperative assembly and disassembly of its components. Very strong cooperativity was observed for the RES (Retention and Splicing) hetero-trimeric complex where the affinity from binary to tertiary interactions changes more than 100-fold and affects RNA binding. The RES complex is involved in splicing regulation and retention of not properly spliced pre-mRNA with its three components—Snu17p, Pml1p and Bud13p—giving rise to the two possible intermediate dimeric complexes Pml1p-Snu17p and Bud13p-Snu17p. Here we determined the three-dimensional structure and dynamics of the Pml1p-Snu17p and Bud13p-Snu17p dimers using liquid state NMR. We demonstrate that localized as well as global changes occur along the RES trimer assembly pathway. The stepwise rigidification of the Snu17p structure following the binding of Pml1p and Bud13p provides a basis for the strong cooperative nature of RES complex assembly.
Similar content being viewed by others
Introduction
The key step of producing mature and nuclear export ready mRNA particles involves excision of introns in a process termed splicing1. Responsible for the catalysis and orchestration of this process is the spliceosome, a multimegadalton assembly of proteins and snRNAs1. Unlike ribosomes at the onset of translation, each complete and active spliceosome needs to assemble on its substrate de novo during the splicing cycle. Assembly, disassembly and remodelling of the spliceosome is therefore important1,2. As part of this dynamic process various subcomplexes of changing composition are formed. In line with an efficient remodelling of the spliceosome, the spliceosomal proteins are believed not to act independently. Instead cooperative binding3,4,5, resulting in a cooperative cascade, might drive spliceosome formation and thus its function2.
One of the few protein complexes controlling both splicing and export of pre-mRNA is the retention and splicing (RES) complex6. A set of introns, specifically associated with pre-mRNA of proteins engaged in nucleotide metabolism, shows splicing controlled by RES7,8,9. The RES complex is composed of the 17.1 kDa small nuclear ribonucleoprotein-associated protein 17 (Snu17p), the pre-mRNA leakage protein 1 (Pml1p) and the 30.5 kDa bud site selection protein 13 (Bud13p)10,11. RES was shown to interact with U2 snRNA splicing factor 3B (U2 SF3B) proteins and Bud13p showed chemical crosslinks to human sap homolog 155 (Hsh155)12,13. In addition, we showed that Snu17p can be crosslinked to pre-mRNA between the branch point and 3′ splice site12.
Recently, we determined the three-dimensional structure of the core of the RES trimer composed of Snu17p, Pml1p and Bud13p12. We further demonstrated that the RES trimer assembles and forms a ternary complex with RNA in a highly cooperative manner12. In order to obtain further insight into the molecular basis of the cooperative nature of RES assembly, we here present the three-dimensional structures of two dimeric intermediates along the RES assembly pathway. The structures of the two intermediates provide insight into the atomistic details of the rearrangements that are required in order to accommodate the two intrinsically disordered protein fragments of Bud13p and Pml1p and reveal the nature of the conformational plasticity of RES intermediates.
Material and methods
Natural abundance and isotopically labelled cSnu17p (residues 25–138 of Snu17p), cPml1p (residues 22–42 of Pml1p) and hcBud13p (residues 215–255 of Bud13p) were prepared as described previously12. Samples contained either 13C,15N-labelled peptide with natural abundance cSnu17p or 13C,15N-labelled cSnu17p with natural abundance peptide. In case of the hcBud13p–cSnu17p dimer, we also prepared a sample where both components of the dimer were 13C,15N labelled. For measurement of HN residual dipolar couplings (RDCs), samples were aligned by the addition of Pf1 phage (ASLA Biotech).
NMR experiments were carried out at 35 °C on 600, 700, 800 and 900 MHz Bruker spectrometers. Spectrometers were equipped with cryogenically cooled HCN or room temperature HCN probes. In addition to the protein backbones, which were sequence-specifically assigned previously12, the sequence-specific backbone resonance assignment of hcBud13p and cPml1p in their dimeric complexes with cSnu17p was performed using 3D HNCA, HNCOCA, HNCACB, HNCO14 and 15N-edited NOESY-HSQC experiments14. We also assigned the sidechains in the dimeric complexes with the help of 3D HcCH-TOCSY15, 3D HBCBCGCDHD16 and 3D 13C-edited NOESY-HSQC (both aromatic and aliphatic) experiments14. Inter-molecular NOEs were extracted from 13C F1-edited/13C-15N F3-filtered HSQC-NOESY spectra, from the standard Bruker pulse sequence library, which were recorded with a mixing time of 120 ms, recycle delay of 1 s and 32 scans per increment17. A total of 256 points in the indirect 1H dimension and 32 in the indirect 13C dimensions were acquired. FIDs were processed with NMRPipe18 or Topspin (Bruker) and the resulting spectra were analysed using ccpnmr Analysis 2.2.119. The BSD-IPAP-HSQC experiment20 was used to measure HN RDCs. A total of 91 RDCs were obtained for cSnu17p in the cPml1p–cSnu17p dimer and 75 were used for the purpose of structure validation. In addition, 53 RDCs of residues located in well-defined regions were used for validation of the hcBud13p–cSnu17p dimer structure out of the total number of 87 obtained. For pairwise analysis of RDC sets, all 91 RDCs observed in the cPml1p–cSnu17p dimer were compared against values observed in the RES trimer. Moreover, 79 (65 for the core RRM and 14 in the unfolded C-terminal α-helix) RDCs observed in hcBud13p–cSnu17p dimer were compared against cRES. Finally, 87 RDCs were compared between the hcBud13p–cSnu17p and cPml1p–cSnu17p dimers. The software PALES was used for RDC analysis21.
Structures were calculated in CYANA 3.022 and refined using Xplor-NIH 2.3.423. The maximum upper distance limit was set to 6.5 Å and the reference distance to 4.25 Å. Structure calculations were supplemented by dihedral angle restraints that were derived from backbone chemical shifts using TALOS-N24. Only high-confidence values (labelled by TALOS-N as „strong“) were used. Eight cycles of structure calculation using CYANA’s noeassig.py protocol were carried out. Intermolecular NOE contacts, which were extracted from filtered/edited NOESY experiments, were treated separately to the automatic CYANA protocols and were manually refined in an iterative manner. H-bonds in α-helices and β-sheets were identified from the initial structural ensemble and confirmed by H-D exchange in combination with NOEs patterns. The structure, which was closest to the mean, was used as representative of an ensemble.
Unless stated otherwise, statistics and structural comparisons were determined using the well-structured parts of the three RES components, that is residues 32–62/74–108 of cSnu17p and residues 223–238 of hcBud13p in the hcBud13p–cSnu17p dimer and residues 32–126 of cSnu17p and residues 26–39 of cPml1p in the cPml1p–cSnu17p dimer. Structural statistics were calculated using Xplor-NIH23 and the ICING server25. APBS cut-offs were set to ±2 kT/eV in order to visualize the electrostatic potential on a solvent accessible surface26. Figures were prepared in PyMOL (http://www.pymol.org/) and VMD-xplor27.
Results and Discussion
Three-dimensional structures of cPml1p-cSnu17p and hcBud13p-cSnu17p complexes
The minimal regions required for binding to residues 25–138 (core Snu17p, cSnu17p) of the 148-residue protein Snu17p are residues 22–42 of Pml1p (cPml1p) and residues 215–245 of Bud13p12. The selected regions are sufficient to reproduce the binding affinities of the full-length proteins and thus represent the core of the RES trimer11,12. In the current study, we further included residues 246–255 of Bud13p (resulting in the fragment hcBud13p which comprises residues 215–255 of Bud13p) as Collinet et al. reported that residues 246–255 form an α-helix28. Using a variety of multidimensional NMR experiments the sequence-specific assignment of the cPml1p–cSnu17p dimer as well as the hcBud13p–cSnu17p dimer was achieved. Based on the assignment of 88.0% (hcBud13p–cSnu17p dimer) and 91.2% (cPml1p–cSnu17p dimer) of all 1H proton resonances, we collected a large number of unambiguous intramolecular and intermolecular cSnu17p-cBud13p and cSnu17p-cPml1p NOE distance restraints (Table 1). They defined the structure of the cPml1p–cSnu17p dimer and the hcBud13p–cSnu17p dimer at high resolution (Figs 1 and 2A,C). The final ensembles displayed RMSDs for all heavy atoms of 1.21 Å and 1.31 Å, respectively, with Ramachandran plot statistics of 90.3, 8.5, 1.0, 0.1% (hcBud13p–cSnu17p dimer) and 88.2, 11.7, 0.1, 0.0% (cPml1p–cSnu17p dimer) for core, allowed, generous and disallowed regions, respectively (Table 1). The quality of the structures was further validated by RDCs (Fig. 3A). Notably, the molecular alignment of Snu17p in Pf1 phage differed strongly between the hcBud13p-cSnu17p dimer and the cRES trimer, as well as between hcBud13p-cSnu17p and cPml1p-cSnu17p dimer, consistent with a release of the C-terminal helix in the hcBud13p-cSnu17p dimer (Supplementary Fig. 1C–E). At the same time, the alignment was nearly identical between cPml1p-cSnu17p dimer and the cRES trimer, in agreement with the stabilization of the C-terminal helix of cSnu17p in the two complexes.
Three-dimensional structures of intermediates during cRES complex assembly.
(A) hcBud13p–cSnu17p dimer. 20 lowest-energy structures (backbone atoms only) are shown. Residues 116–131 of Snu17p, which fold into an α-helix in the cRES trimer12, remain flexible in the hcBud13p–cSnu17p dimer. In addition, the relative orientation of the C-terminal α-helix of hcBud13p (marked by a dashed ellipsoid) is flexible. Red, cSnu17p; pink hcBud13p. (B) cPml1p–cSnu17p dimer. Shown are the backbones of the 20 lowest-energy structures of the NMR ensemble. Blue, cSnu17p; cyan cPml1p.
Comparison of the 3D structures of the RES core complex, the cPml1p–cSnu17p dimer and the hcBud13p–cSnu17p dimer.
(A) hcBud13p (pink) in complex with –cSnu17p (red). (B) NMR ensemble of the RES core complex (PDB code: 2MKC12). Gray, cSnu17p; black cBud13p and cPml1p. (C) Structure of cSnu17p (blue) in complex with cPml1p (cyan). 20 lowest-energy structures (backbone atoms only) are shown. The L63 – F73 loop is encircled in gray and the C-terminal region of cSnu17p, when folded into an α-helix, in black and the disordered C-terminal part of cPml1p, in red. (D) Detailed view of the L63–F73 loop of cSnu17p (blue) and residues 205–210 of cPml1p (cyan) in the cPml1p–cSnu17p dimer (upper panel). The same loop is shown below for the hcBud13p–cSnu17p dimer. Experimentally observed NOE contacts are represented with black lines. (E) Residues 106–115 of Snu17p in the hcBud13p–cSnu17p dimer. Residues 106–115 are shown as an ensemble in light-blue and the rest of the hcBud13p–cSnu17p dimer as a single structure in red. Experimental NOE contacts between Y109 (orange), P111 (green) and I61 are schematically indicated with dashed lines. (F) Regions of Snu17p, which are dynamic in the hcBud13p–cSnu17p dimer, were mapped onto the 3D structure of the cPml1p–cSnu17p dimer. Residues 35, 37, 40, 43 – 46, 75, 98, 99, 101, 102, 107, 110, 112 of cSnu17p were marked in red as they showed Rex values exceeding 10 Hz in NMR relaxation measurements12 and/or experienced line broadening of 10 Hz above the average value among the folded part of cSnu17p in 1H-15N HSQC experiments12. L63–F73 of cSnu17p was also marked in black to highlight the sparse NOE network as shown in (D) and therefore higher RMSD values as presented in (A) and consistent with lower than average heteronuclear NOE values reported in10. Flexible N- and C-terminal residues (20–32 and 113–138) were excluded from this analysis. α-helix 3 of Snu17p, which is not formed in the hcBud13p–cSnu17p dimer, is labelled and constitutes the most dynamic element in the hcBud13p–cSnu17p dimer structure (as seen in (A)).
Analysis of HN RDCs observed in cSnu17p when part of the cPml1p–cSnu17p dimer, the hcBud13p–cSnu17p dimer and cRES.
(A,B) Comparison of experimental RDCs with values back-calculated from the 3D structure of the cPml1p–cSnu17p dimer (A) and the hcBud13p–cSnu17p dimer (B). (C,D) Comparison of RDCs observed in cSnu17p in cRES with RDCs observed in cSnu17p when part of the hcBud13p–cSnu17p dimer (C) or the cPml1p–cSnu17p dimer (D). Black dots indicate disordered C-terminal residues in the hcBud13p–cSnu17p dimer. (E) Comparison of RDCs observed in cSnu17p in complex with cSnu17p and in complex with hcBud13p. Black dots indicated disordered C-terminal residues in hcBud13p–cSnu17p dimer. (F) Residue-specific comparison of RDCs observed in dimeric complexes of cSnu17p with cPml1p and with hcBud13p. In (F), RDC values of the hcBud13p–cSnu17p dimer were normalized to the magnitude of the alignment tensor of the cPml1p–cSnu17p dimer.
The structure of the cPml1p-cSnu17p dimer and the hcBud13p-cSnu17p dimer retain the β1α1β2β3α2β4 topology of RRMs (Figs 1 and 2) and the domain characteristics of Snu17p seen in the structure of the cRES trimer12. Despite the apparent similarity of the Snu17p and Bud13p complex structure to prototypical U2AF homology motif (UHM) and UHM ligand motif ULM interactions, the mode of interaction appears to be different10,12. Whereas in classical ULM-UHM complexes a central tryptophan is positioned in a deep hydrophobic pocket provided by the RRM domain, tryptophan 232 of Bud13p is found in a shallow space approximately 11 Å away from the canonical site in Snu17p. This is the case for both the cRES trimer as well as hcBud13p–cSnu17p dimer, despite the lack of steric obstruction provided by cPml1p in the latter case. The charge distribution over all three structures appears to be similar (Supplementary Fig. 1) although, the C-terminal region of Snu17p, which only forms an α-helix in the cPml1p-cSnu17p dimer and the cRES trimer but not in the hcBud13p-cSnu17p dimer (Fig. 1A), is partially positively charged and might contribute to RNA binding. The overall similar charge distribution suggests that optimization of the electrostatic interaction might not be the major contributor to the cooperativity, which was observed for binding of cRES to RNA when compared to monomeric Snu17p and the two dimers12.
Molecular motions in intermediate structures of the RES complex assembly pathway
In a recently solved structure of residues 25–113 of Snu17p in complex with residues 222–256 of Bud13p10 the C-terminal region of Snu17p, which forms an α-helix in the cRES trimer and contributes to RNA binding12, was not present and therefore did not allow analysis of this functionally important region in the dimeric complex with Bud13p. Based on chemical shift and 15N spin relaxation data, we predicted that cSnu17p residues beyond 115 would be unstable in the cBud13p–cSnu17p dimer12. Consistent with this prediction, the three-dimensional structure of the hcBud13p–cSnu17p dimer, revealed the C-terminal part of cSnu17p to be disordered and to sample a large conformational space (Figs 1A, 2A). Due to this pronounced mobility, RDC values in this region were efficiently averaged to near zero values (Fig. 3F). However, the long loop, which is formed by residues 106–115 of Snu17p, connects the C-terminal part to the core of Snu17p and traverses its β-sheet in the cRES trimer, remains partially in place in the absence of cPml1p (Fig. 2E and Supplementary Fig. 2). The partial attachment of this region to the Snu17p β-sheet provides a structural basis for the finding that the ability of the cBud13p–cSnu17p dimer to bind to RNA was diminished but not abolished12.
Additional mobility in the hcBud13p–cSnu17p dimer when compared to cRES was observed for the loop between L63 and F73 of cSnu17p, which samples a larger conformational space when cPml1p is absent (Figs 1A and 2A,C) This can be tracked back to a lack of interactions between residues R64–E66 of cSnu17p and I26, I28 and D31 of cPml1p as well as sparse intra-loop contacts (Fig. 2D). Altogether, it gives rise to an at least three times lower amount of NOE contacts when cPml1p is absent (Fig. 2D). Moreover, the L63-F73 loop of cSnu17p was reported to have lower than average heteronuclear NOE values pointing to increased pico-to-nanosecond motions10. Although the chemical exchange contribution to the R2 15N relaxation rate (Rex) in this region was not elevated, the adjacent loop (V40–E46) showed increased Rex values when compared to the cPml1p–cSnu17p dimer and the cRES trimer12. In addition, the β-turn adjacent to V40–E46 showed an elevated Rex contribution and was affected by NMR line broadening12. The two loops and the adjacent β-turn are the site of cPml1p binding, together with the C-terminal region of Snu17p, which folds into an α-helix upon binding of Pml1p (Fig. 2F). The ensemble of Snu17p conformations in this region is therefore ready to accept the incoming Pml1p. On the other hand, we did not detect a structural perturbation of hcBud13p in the dimeric complex with Snu17p when compared to the cRES trimer. Small 1H-15N HSQC chemical shift changes (Fig. 4A) were probably caused by a change in the environment associated with the lack of cPml1p and the unfolding of the C-terminal region of Snu17p (Fig. 4B,C).
Localized changes in the structure and dynamics during RES core complex assembly.
(A) Comparison between 1H-15N HSQC spectra of cBud13p in cRES (dark gray) and the hcBud13p–cSnu17p dimer (pink). Residues experiencing significant chemical shift perturbation are indicated and some of the corresponding regions are enlarged below. (B) Superposition of the 3D structures of the cRES trimer (gray and graphite) and the hcBud13p–cSnu17p dimer (pink and red). Residues experiencing chemical shift perturbation in a) are indicated with spheres. (C) Superposition of the 3D structures of the cRES trimer (PDB code: 2MKC12; gray and black) and cPml1p–cSnu17p dimer (blue and cyan). The three leucine residues, which strongly shift in (E), are highlighted with spheres. The red circle indicates the part of the cPml1p structure that is partially disordered in the cPml1p–cSnu17p dimer. Structures are colored as follows: graphite, cBud13p and cPml1p in cRES; gray, cRES; pink cBud13p in hcBud13p–cSnu17p dimer; red cSnu17p in hcBud13p–cSnu17p dimer; cyan cPml1p in cPml1p–cSnu17p dimer; blue cSnu17p in cPml1p–cSnu17p dimer. (D) Comparison between the α-helix content of cPml1p in the cRES trimer (graphite) and the cPml1p–cSnu17p dimer (cyan). The α-helix content was estimated on the basis of backbone chemical shifts using TALOS-N24. (E) Superposition of 1H-15N HSQC spectra of cPml1p in cRES (dark gray) and the cPml1p-cSnu17p dimer (cyan). Three leucine residues experiencing a strong perturbation are marked.
Next, we analyzed the differences between the structure of the cPml1p–cSnu17p dimer and the cRES trimer (Fig. 4C). Comparison of the two structures revealed increased disorder of the C-terminal part of cPml1p associated with a loss of α-helical character (Figs 2B,C and 4C,D). Moreover, the cPml1p position was slightly modified in response to cBud13p binding (Fig. 4C). The changes observed in the structure of the cPml1p–cSnu17p dimer were supported by 1H-15N HSQC spectra: chemical shifts of backbone amides at the C-terminal part of cPml1p differ depending on the presence of cBud13p (Fig. 4E) and reflect both changes in the level of disorder and structural changes.
Both cPml1p and hcBud13p represent largely the fragments that are necessary for binding to cSnu17p. However, the rest of each sequence could, in the context of the spliceosome, play a role in modulating the assembly of the RES complex. For example, the FHA domain of Pml1p is separated by only a short, six-residue linker from the Pml1p region, which binds to Snu17p. Notably, the phosphopeptide binding site of the FHA domain occurs in proximity to this linker. Alternatively, additional parts of Bud13p, which is intrinsically disordered along its complete sequence, might fold upon binding to other spliceosomal proteins in the context of the spliceosome. Currently, the order of RES complex assembly is not known, but given the two orders of magnitude higher affinity of Bud13p to Snu17p one can speculate that Bud13p might bind first, followed by Pml1p.
Continuum of UHM-ULM-like interactions
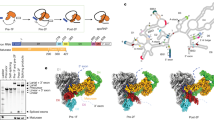
We then compared the non-canonical position of tryptophan 232 of cBud13p as observed in both the dimeric complex with cSnu17p and the cRES trimer (Fig. 5) and10,12, with other known RRM-peptide interactions. A canonical UHM–ULM interaction, in which a tryptophan residue is buried in a hydrophobic pocket of the RRM domain was observed for example for the complexes of splicing factor 3b (SF3b155(ULM5)) with alternative splicing factor 45 (SPF45) and of splicing factor 1 (SF1) with the large subunit of U2 snRNP auxiliary factor (U2AF6529,30). On the other hand, the complex structures of Acinus with RNA binding protein with serine rich domain 1 (RNPS131), eukaryotic translation initiation factors 3 J and 3b (EIF3J–EIF3b32), infected cell protein 27 (ICP27) with RNA export factor2 (REF233) and immediate-early phosphoprotein from Saimiriine Herpes Virus ORF57 with REF234 do not have this canonical interaction. In these complexes—as well as in the Snu17p-Bud13p interaction — a conserved tryptophan residue is important for RRM binding, but its position is variable (Fig. 5A). In addition, the part of the protein, which is in contact with the RRM domain, samples a range of conformations (Fig. 5B). Most similar to the hcBud13p–cSnu17p recognition mode is the REF2–ICP27 complex, where the tryptophan side chain occupies a region near the C-terminus of α-helix 2 of the RRM domain (Fig. 5A). Intriguingly, ORF57, another REF2 binder, bares a degree of structural similarity to cPml1p as observed in the cPml1p–cSnu17p dimer structure. Notably, both Pml1p and REF2 are proteins involved in mRNA export35,36,37. The analysis suggests that there is a structural continuum of how tryptophan containing motifs bind to RRM domains.
Architecture of central tryptophan containing motifs (CTCMs) in RRMs.
(A) Comparison between the side chain position of a key tryptophan in different CTCMs bound to RRMs. (B) Comparison between backbone traces of CTCMs bound to different RRM domains. CTCMs are labelled 1, dark green, D-Acinus–RNPS1 (PDB code: 4A8X)31; 2, red, SF3b155(ULM5)–SPF45 (PDB code: 2PEH)29; 3, orange, SF1–U2AF65 (PDB code: 1OPI)30; 4, light green, EIF3J–EIF3b (PDB code: 2KRB)32; 5, yellow, ICP27–REF2 (PDB code: 2KT5)33; 6, light blue, ORF57–REF2 (PDB code: 2YKA)34; 7, dark blue, hcBud13p–cSnu17p dimer. Only α-helices 1 and 2 of the RRM are shown for clarity. (C) Schematic representation of the folding-upon-binding mechanism as seen in the assembly of the RES complex.
In summary, we provided high-resolution structural evidence that the dimeric intermediates along the RES assembly pathway are not a simple structural equivalent of subtracting a given binding partner (Bud13p or Pml1p) of Snu17p from the RES trimeric complex. Instead, a number of localized structural changes are required for successive binding. The local structural changes are further accompanied by the large-scale rearrangement of the C-terminal part of Snu17p, which only folds into a stable α-helix upon interaction with Pml1p. The stepwise rigidification of the Snu17p structure upon binding of Bud13p and Pml1p provides a basis for the strong cooperative nature of RES assembly and RNA binding (Fig. 5C).
Additional Information
How to cite this article: Wysoczanski, P. et al. Structures of intermediates during RES complex assembly. Sci. Rep. 5, 12545; doi: 10.1038/srep12545 (2015).
References
Will, C. L. & Luhrmann, R. Spliceosome structure and function. Cold Spring Harb. Perspect. Biol. 3, a003707 (2011).
Brow, D. A. Allosteric cascade of spliceosome activation. Annu. Rev. Genet. 36, 333–60 (2002).
Williams, S. G. & Hall, K. B. Linkage and Allostery in snRNP Protein/RNA Complexes. Biochemistry 53, 3529–39 (2014).
Williams, S. G. & Hall, K. B. Binding Affinity and Cooperativity Control U2B”/snRNA/U2A’ RNP Formation. Biochemistry 53, 3727–37 (2014).
Hennig, J. et al. Structural basis for the assembly of the Sxl-Unr translation regulatory complex. Nature 515, 287–290 (2014).
Dziembowski, A. et al. Proteomic analysis identifies a new complex required for nuclear pre-mRNA retention and splicing. EMBO J. 23, 4847–56 (2004).
Zhou, Y., Chen, C. & Johansson, M. J. The pre-mRNA retention and splicing complex controls tRNA maturation by promoting TAN1 expression. Nucleic Acids Res. 41, 5669–78 (2013).
Tuo, S., Nakashima, K. & Pringle, J. R. Apparent defect in yeast bud-site selection due to a specific failure to splice the pre-mRNA of a regulator of cell-type-specific transcription. PloS one 7, e47621 (2012).
Scherrer, F. W., Jr. & Spingola, M. A subset of Mer1p-dependent introns requires Bud13p for splicing activation and nuclear retention. RNA 12, 1361–72 (2006).
Tripsianes, K. et al. A Novel Protein-Protein Interaction in the RES (REtention and Splicing) Complex. The Journal of biological chemistry 289, 28640–50 (2014).
Trowitzsch, S., Weber, G., Luhrmann, R. & Wahl, M. C. An unusual RNA recognition motif acts as a scaffold for multiple proteins in the pre-mRNA retention and splicing complex. J. Biol. Chem. 283, 32317–27 (2008).
Wysoczanski, P. et al. Cooperative structure of the heterotrimeric pre-mRNA retention and splicing complex. Nat. Struct. Mol. Biol. 21, 911–8 (2014).
Wang, Q., He, J., Lynn, B. & Rymond, B. C. Interactions of the yeast SF3b splicing factor. Mol. Cell. Biol. 25, 10745–54 (2005).
Sattler, M., Schleucher, J. & Griesinger, C. Heteronuclear multidimensional NMR experiments for the structure determination of proteins in solution employing pulsed field gradients. Prog. Nucl. Magn. Reson. Spectrosc. 34, 93–158 (1999).
Bax, A., Clore, G. M. & Gronenborn, A. M. 1H-1H correlation via isotropic mixing of 13C magnetization, a new three-dimensional approach for assigning 1H and 13C spectra of 13C-enriched proteins. J. Magn. Reson. 88, 425–431 (1990).
Yamazaki, T., Forman-Kay, J. D. & Kay, L. E. Two-dimensional NMR experiments for correlating carbon-13.beta. and proton.delta./.epsilon. chemical shifts of aromatic residues in 13C-labeled proteins via scalar couplings. J. Am. Chem. Soc. 115, 11054–11055 (1993).
Zwahlen, C. et al. Methods for Measurement of Intermolecular NOEs by Multinuclear NMR Spectroscopy: Application to a Bacteriophage λ N-Peptide/boxB RNA Complex. J. Am. Chem. Soc. 119, 6711–6721 (1997).
Delaglio, F. et al. NMRPipe: a multidimensional spectral processing system based on UNIX pipes. J. Biomol. NMR 6, 277–93 (1995).
Vranken, W. F. et al. The CCPN data model for NMR spectroscopy: Development of a software pipeline. Proteins 59, 687–696 (2005).
Yao, L., Ying, J. & Bax, A. Improved accuracy of 15N-1H scalar and residual dipolar couplings from gradient-enhanced IPAP-HSQC experiments on protonated proteins. J. Biomol. NMR 43, 161–70 (2009).
Zweckstetter, M. NMR: prediction of molecular alignment from structure using the PALES software. Nature protocols 3, 679–90 (2008).
Guntert, P., Mumenthaler, C. & Wuthrich, K. Torsion angle dynamics for NMR structure calculation with the new program DYANA. J. Mol. Biol. 273, 283–298 (1997).
Schwieters, C. D., Kuszewski, J. J., Tjandra, N. & Clore, G. M. The Xplor-NIH NMR molecular structure determination package. J. Magn. Reson. 160, 65–73 (2003).
Shen, Y. & Bax, A. Protein backbone and sidechain torsion angles predicted from NMR chemical shifts using artificial neural networks. J. Biomol. NMR 56, 227–41 (2013).
Doreleijers, J. F. et al. CING: an integrated residue-based structure validation program suite. J. Biomol. NMR 54, 267–283 (2012).
Baker, N. A., Sept, D., Joseph, S., Holst, M. J. & McCammon, J. A. Electrostatics of nanosystems: application to microtubules and the ribosome. Proc. Natl. Acad. Sci. USA 98, 10037–41 (2001).
Schwieters, C. D. & Clore, G. M. The VMD-XPLOR visualization package for NMR structure refinement. J. Magn. Reson. 149, 239–44 (2001).
Collinet, B. et al. Strategies for the structural analysis of multi-protein complexes: lessons from the 3D-Repertoire project. J. Struct. Biol. 175, 147–58 (2011).
Corsini, L. et al. U2AF-homology motif interactions are required for alternative splicing regulation by SPF45. Nat. Struct. Mol. Biol. 14, 620–9 (2007).
Selenko, P. et al. Structural basis for the molecular recognition between human splicing factors U2AF65 and SF1/mBBP. Mol. Cell 11, 965–76 (2003).
Murachelli, A. G., Ebert, J., Basquin, C., Le Hir, H. & Conti, E. The structure of the ASAP core complex reveals the existence of a Pinin-containing PSAP complex. Nat. Struct. Mol. Biol. 19, 378–86 (2012).
Elantak, L. et al. The indispensable N-terminal half of eIF3j/HCR1 cooperates with its structurally conserved binding partner eIF3b/PRT1-RRM and with eIF1A in stringent AUG selection. J. Mol. Biol. 396, 1097–116 (2010).
Tunnicliffe, R. B. et al. Structural basis for the recognition of cellular mRNA export factor REF by herpes viral proteins HSV-1 ICP27 and HVS ORF57. PLoS Path. 7, e1001244 (2011).
Tunnicliffe, R. B., Hautbergue, G. M., Wilson, S. A., Kalra, P. & Golovanov, A. P. Competitive and cooperative interactions mediate RNA transfer from herpesvirus saimiri ORF57 to the mammalian export adaptor ALYREF. PLoS Path. 10, e1003907 (2014).
Chekanova, J. A., Abruzzi, K. C., Rosbash, M. & Belostotsky, D. A. Sus1, Sac3 and Thp1 mediate post-transcriptional tethering of active genes to the nuclear rim as well as to non-nascent mRNP. RNA 14, 66–77 (2008).
Palancade, B., Zuccolo, M., Loeillet, S., Nicolas, A. & Doye, V. Pml39, a novel protein of the nuclear periphery required for nuclear retention of improper messenger ribonucleoparticles. Mol. Biol. Cell 16, 5258–68 (2005).
Palancade, B. & Doye, V. Sumoylating and desumoylating enzymes at nuclear pores: underpinning their unexpected duties? Trends Cell Biol. 18, 174–83 (2008).
Acknowledgements
We thank K. Giller for the preparation of expression constructs. This work was supported by the German Science Foundation [Collaborative Research Center 860 project B2 to M.Z.].
Author information
Authors and Affiliations
Contributions
P.W. performed NMR measurements, data analysis and structure calculation. P.W. and M.Z. wrote the manuscript. S.B. and M.Z. supervised the study.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Wysoczanski, P., Becker, S. & Zweckstetter, M. Structures of intermediates during RES complex assembly. Sci Rep 5, 12545 (2015). https://doi.org/10.1038/srep12545
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep12545
- Springer Nature Limited