Abstract
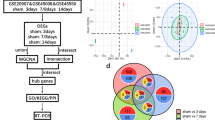
The mechanism of spinal cord injury (SCI) is highly complex, and an increasing number of studies have indicated the involvement of pyroptosis in the physiological and pathological processes of secondary SCI. However, there is limited bioinformatics research on pyroptosis-related genes (PRGs) in SCI. This study aims to identify and validate differentially expressed PRGs in the GEO database, perform bioinformatics analysis, and construct regulatory networks to explore potential regulatory mechanisms and therapeutic targets for SCI. We obtained high-throughput sequencing datasets of SCI in rats and mice from the GEO database. Differential analysis was conducted using the “limma” package in R to identify differentially expressed genes (DEGs). These genes were then intersected with previously reported PRGs, resulting in a set of PRGs in SCI. GO and KEGG enrichment analyses, as well as correlation analysis, were performed on the PRGs in both rat and mouse models of SCI. Additionally, a protein–protein interaction (PPI) network was constructed using the STRING website to examine the relationships between proteins. Hub genes were identified using Cytoscape software, and the intersection of the top 5 hub genes in rats and mice were selected for subsequent experimentally validated. Furthermore, a competing endogenous RNA (ceRNA) network was constructed to explore potential regulatory mechanisms. The gene expression profiles of GSE93249, GSE133093, GSE138637, GSE174549, GSE45376, GSE171441_3d and GSE171441_35d were selected in this study. We identified 10 and 12 PRGs in rats and mice datasets respectively. Six common DEGs were identified in the intersection of rats and mice PRGs. Enrichment analysis of these DEGs indicated that GO analysis was mainly focused on inflammation-related factors, while KEGG analysis showed that the most genes were enriched on the NOD-like receptor signaling pathway. We constructed a ceRNA regulatory network that consisted of five important PRGs, as well as 24 miRNAs and 34 lncRNAs. This network revealed potential regulatory mechanisms. Additionally, the three hub genes obtained from the intersection were validated in the rat model, showing high expression of PRGs in SCI. Pyroptosis is involved in secondary SCI and may play a significant role in its pathogenesis. The regulatory mechanisms associated with pyroptosis deserve further in-depth research.
Similar content being viewed by others
Introduction
Spinal cord injury (SCI) refers to the disruption of the integrity or continuity of the spinal cord caused by various etiologies, resulting in temporary or permanent loss of sensory, motor, and other functions below the level of injury segment1,2,3. Despite remarkable advances in medical technology, treatment options for SCI, including surgery, medication, and rehabilitation, as well as emerging therapies such as stem cell transplantation and biomaterial therapy, are still unable to achieve complete clinical cure4,5,6,7,8,9,10. The complications and sequelae of SCI cause significant physical and psychological impairments to patients and impose a substantial burden on families and society1,11. Recent epidemiological studies indicate that the incidence of traumatic SCI is approximately 10.5 per 100,000 individuals, with nearly 800,000 new cases occurring globally each year6,11,12,13. Therefore, the investigation of the mechanisms underlying SCI and the exploration of novel therapeutic strategies hold important practical significance.
The mechanism underlying SCI is highly complex and broadly can be categorized into two types: primary and secondary injury11. Primary injury results from the immediate impact of the injurious factor on the spinal cord, leading to damage of spinal cord tissue13. In contrast, secondary injury is a cascade of complex physiological and pathological changes that occur following the primary injury. These changes include edema, vascular rupture, electrolyte imbalances, oxidative stress, and various forms of cell death11,13,14,15. Secondary injury is modifiable and thus forms the basis for numerous intervention strategies that aim to improve clinical outcomes. Recent studies have demonstrated that pyroptosis, a newly identified type of programmed cell death, contributes to the secondary injury of the spinal cord16,17,18. Pyroptosis is distinct from both apoptotic and necroptosis and is characterized by the formation of cytoplasmic membrane pores, cell swelling, cell membrane rupture, massive release of inflammatory factors, and cell death18. Pyroptosis is involved in various physiological and pathological processes and plays a regulatory role in the development and progression of a wide range of diseases, including infections, neurodegenerative disorders, brain injury, ischemia–reperfusion injury, atherosclerosis, metabolic disorders, and tumors, among others19,20,21,22. Although some research has explored the role of pyroptosis in SCI, there has been limited exploration of the specific molecular mechanisms and components of this form of programmed cell death in the context of SCI23,24. Further investigation in this area has the potential to improve our understanding of the pathogenesis of SCI and facilitate the development of novel treatment strategies.
The aim of this study is to collate and analyze the spinal cord tissue sequencing data from rat and mouse models of SCI in the GEO (gene expression omnubus) database (https://www.ncbi.nlm.nih.gov). The differentially expressed genes (DEGs) will be integrated with the pyroptosis-related genes (PRGs) to identify the PRGs network involved in the differential expression of SCI. This approach will enhance our understanding of the role of PRGs in SCI and help to elucidate the specific mechanisms of injury. Additionally, the identification of these genes may provide new targets for intervention in the treatment of SCI.
Materials and methods
Data source and data processing
We downloaded four high-throughput sequencing datasets of SCI in Rattus norvegicus (GSE93249, GSE133093, GSE138637 and GSE174549) and two high-throughput sequencing datasets in Mus musculus (GSE45376 and GSE171441) from the GEO database. In these datasets, we exclusively analyzed the sequencing results of the pure SCI group and the sham surgery group, excluding data from other treatment groups such as drug administration or electrical stimulation. Additionally, in the GSE171441 dataset, samples were collected for sequencing at 3 days and 35 days after SCI. We separated and compared these samples separately with the sham surgery group during data processing, and named the dataset as GSE171441_3d and GSE171441_35d respectively. PRGs data set was identified and synthesized from previous literature18,25,26,27,28,29 through a thorough review of full texts and corresponding references (Supplementary Table 1).
Gene differential expression analysis
The “limma” R package was utilized to perform the differential expression analysis between SCI group and sham surgery group from GEO databases. |Log2(fold change)| ≥ 0.5 and P value < 0.05 were considered as the cutoff criteria. The volcano plot was drawn to show the DEGs.
Protein–protein interaction (PPI) analysis
To explore the PPI network of the selected genes, they were imported into the STRING database (https://string-db.org), which is a web tool used to explore the interactions between multiple proteins.
Functional enrichment analysis
Gene Ontology (GO) and Kyoto encyclopedia of genes and genomes30,31(KEGG, https://www.kegg.jp/kegg/kegg1.html) analyses were performed with the Database for Annotation, Visualization and Integrated Discovery (DAVID) to explore the mechanisms and pathways of DEGs.
Animal groups and SCI model
All animal study protocols were approved by the Research Ethics Committee of The Second Xiangya Hospital, Central South University, Hunan, P.R. China, and all procedures were performed in accordance with ethical standards and ARRIVE guidelines. Female Sprague–Dawley (SD) rats, aged 10 weeks old, weighting between 200 and 230 g, were chosen as the experimental animal for this study, provided by the Department of laboratory Animals of Central South University. All rats were housed under a 12 h light/dark cycle pathogen-free condition with a controlled temperature of 23 ± 2 °C and 60 ± 5% humidity and free access to food and water. The rats were randomly divided into two groups: the sham surgery group and the SCI group, and 4 rats in each group.
The SCI model was established based on a modified Allen’s method32,33, as described in previous literature. Briefly, rats were anesthetized by 1% pentobarbital sodium (40 mg/kg, intraperitoneal injection). After the disappearance of corneal reflex, a 2 cm incision was made in the posterior skin at the T8-10 vertebral level. The T9 vertebral plate was removed to expose the spinal cord. Subsequently, a 3 mm diameter, 10 g weight rod was freely dropped from a height of 2.5 cm, aiming at the T9 spinal cord. Successful modeling was immediately characterized by congested and edematous spinal cord at the impact site, accompanied by hind limb extension and tail flick reflex in rats. Furthermore, upon awakening from anesthesia, the rats exhibited dragging of the hind limbs. In contrast, the rats in sham group were accepted the T9 laminectomy only without any spinal cord impact procedure. After surgery, rats continued to be bred for a week. Bladder emptying was performed twice daily by applying abdominal pressure until voluntary urination was achieved or euthanasia was carried out.
Total RNA extraction and quantitative real-time PCR
After perfusing the rat with pre-chilled physiological saline solution through the heart, the spinal cord was harvested in an icebox. According to the protocol provided by the manufacturer, total RNA from spinal cord tissues was extracted using TRIzol (Life Technologies, USA). For every 20 mg spinal cord tissue, add 1 ml of TRIzol for RNA extraction. Complementary DNA was synthesized from total RNA with a reverse transcription kit (Thermo Fisher Scientific, USA). Real-time qPCR was executed with SYBR qPCR Master Mix (Vazyme, China). GAPDH was used as an internal reference gene, and 2–ΔΔCt was used to quantify the relative expression level of genes. The primer sequences were listed in the supplementary materials (Supplementary Table 2).
Western blot
In order to assess the expression levels of key genes, we used Western blot (WB) to validate the protein expression levels encoded by those genes. After homogenizing spinal cord tissue obtained from rats, the tissue was lysed using RIPA lysis buffer (Beyotime, China) (20 mg of spinal cord tissue mixed with 200 μl of lysis buffer and 2 μl of protease inhibitor). Total proteins were collected after centrifugation, and the protein concentration was determined using the BCA method (Bioss, China). Subsequently, 40 μg of total protein was loaded onto an SDS-PAGE gel for protein separation. Following that, the proteins were transferred to a PVDF membrane at low temperature. After blocking with 5% skim milk for 2 h, the membrane was incubated with the primary antibody overnight on a shaker at 4 °C. Following three washes with TBST, the membrane was incubated with the secondary antibody for 1 h at room temperature. Subsequently, the membrane was washed three times with TBST. Finally, protein expression levels were visualized and analyzed using enhanced chemiluminescence technology and Image J software. The GAPDH was used as an internal reference gene. The detailed information on the manufacturer and dilution ratio of the primary antibody was listed in the supplementary materials (Supplementary Table 3).
Statistical analysis
All results are presented as the mean ± standard error of the mean (M ± SEM). Statistical analysis was performed in GraphPad Prism 7.0 (GraphPad Software Inc., San Diego, CA). Comparisons between the two groups were performed using Student’s t test or the Mann–Whitney test, as appropriate. Graphs were made using R software (Version 3.6.0). A P value < 0.05 was considered to be statistically significant. *p < 0.05, **p < 0.01, ***p < 0.001, and ****p < 0.0001.
Ethics approval and consent to participate
The study protocol was approved by the Research Ethics Committee of The Second Xiangya Hospital, Central South University, Hunan, P.R. China.
Results
Identification of PRGs in SCI
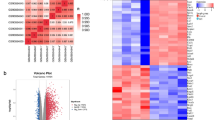
The gene expression profiles of GSE93249, GSE133093, GSE138637, GSE174549, GSE45376, GSE171441_3d and GSE171441_35d were selected in this study. These datasets contained respectively 11, 7, 8, 6, 8, 6, 7 samples. There were 7002, 2464, 3547, 9349, 5356, 5342 and 3762 DEGs after SCI respectively (P < 0.05 and |log FC|≥ 0.5). In the volcano plot, every plot indicated a gene and the blue plots were downregulated, the red plots were upregulated (Fig. 1A–G). We have identified a total of 65 PRGs from the existing literature, which will be used for subsequent comprehensive analysis (Supplementary Table 1). Veen analysis was performed to get the intersection of these DEGs with PRGs (Fig. 1H and I). Results showed that there were 10 intersection genes in rat datasets and 12 in mouse datasets. Then, heatmaps demonstrated that the expression difference of these genes between Sham group and SCI group was very pronounced (Fig. 2A–G). Finally, Venn diagram showed 6 common pyroptosis-related DEGs between rat and mouse (Fig. 2H).
The seven volcano plots show differentially expressed genes (DEGs) between the sham group and SCI group. The criterion of |log2 (fold change) ≥ 0.5| and P value < 0.05 was used. The red, green, and blue points represent up-regulated genes, genes with no significant difference, and down-regulated genes, respectively. (A–D) are derived from rat database in the GEO database, (E–G) are from the mouse dataset. Veen diagrams separately display the intersections between DEGs and pyroptosis-related genes (PRGs) in rats and mice. There are 10 intersections genes (H) in rats and 12 intersections (I) in mice.
GO (gene ontology) and KEGG enrichment analysis of pyroptosis-related DEGs
The differentially expressed PRGs were submitted to Metascape for KEGG enrichment and GO enrichment analysis involving Biologocal Process (BP), cellular composition (CC) and molecular function (MF) and visualized by using R software.
In the rat datasets, the top 15 eligible terms for GO analysis of pyroptosis-related DEGs were shown in Fig. 3A, and the number of enriched genes and adjusted P-value were shown in Fig. 3B. From the figures, it is evident that the pyroptosis-related DEGs were primarily enriched in interleukin-1 production and regulation, inflammatory responses, pyroptosis, inflammasome complex and cysteine-type endopeptidase actively involved in apoptotic process. The relationship of genes and GO terms was shown by Chord plot (Fig. 3C), which allows for a more intuitive visualization of the relationship between genes and pathways. In the KEGG enrichment analysis the pyroptosis-related DEGs were mainly enriched in the following top 10 pathways: NOD-like receptor signaling pathway, Yersinia infection, Salmonella infection, Neutrophil extracellular trap formation, Necroptosis, Legionellosis, Influenza A, Lipid and atherosclerosis and C-type lectin receptor signaling pathway (Fig. 5A). Figure 5C and E, respectively, demonstrate the relationship between genes and pathways, as well as the relationship between pathways themselves. While, in the mouse datasets, similar results were obtained in terms of GO (Fig. 4A–C) and KEGG (Fig. 5B,D and F) enrichment analyses, with only slight variations in the ordering of presentation. From these results, it can be seen that in both rat and mouse SCI, PRGs primarily participate in inflammatory processes, programmed cell death, and defense-related physiological and pathological processes.
GO annotation of 10 pyroptosis-related DEGs of rat datasets following SCI. The bar charts (A) of the top 15 GO terms were drawn on the basis of P-value and the percentage of genes; terms with P-value < 0.01 are statistically significant. The bubble plot visually presents the GO terms in the enrichment analysis, with the size and color coding indicating the degree of enrichment and significance level (B). The chord plot shows the PRGs enriched in different GO terms (C).
GO annotation of 12 pyroptosis-related DEGs of mouse datasets following SCI. The bar charts (A) of the top 15 GO terms were drawn on the basis of P-value and the percentage of genes; terms with P-value < 0.01 are statistically significant. The bubble plot visually presents the GO terms in the enrichment analysis, with the size and color coding indicating the degree of enrichment and significance level (B). The chord plot shows the PRGs enriched in different GO terms (C).
The top 10 KEGG pathways for DEGs in rats and mice datasets. The chord diagram displays the association between the PRGs and KEGG pathway, while also indicating the number of enriched genes ((A) from rat, (B) from mouse). (C) and (D) visualizes the association network between PRGs and their corresponding pathways in rat and mouse respectively. (E) and (F) respectively demonstrate the interconnections between pathways in rats and mice.
Correlation analysis based on pyroptosis-related DEGs and PPI network
The correlation analysis among these pyroptosis-related DEGs expression was shown in Fig. 6. The results demonstrate significant correlations between genes, particularly Casp1 and NRPL3, which exhibit significant and meaningful correlations with other genes. This suggests potential regulatory or synergistic interactions among these PRGs. To explore the interactions between the proteins expressed by the pyroptosis-related DEGs, we constructed a PPI network using the STRING database. Cytoscape (version 3.7.2) was used for visualization of PPI network. The PPI network of rat pyroptosis-related DEGs contained 7 nodes and 27 linkages (Fig. 7A), while 11 nodes and 46 linkages contained in mouse (Fig. 7B). The PPI data from both rats and mouses were separately imported into Cytoscape software to identify hub genes (Fig. 7C and Fig. 7D). The intersection of the top 5 hub genes was selected for subsequent validation experiments. We identified a total of three common hub genes, namely Casp1, Casp4 and Nlrp3 for subsequent study.
PPI network of pyroptosis-related DEGs identified were obtained from the STRING database (A,B). Hub genes identified by Cytoscape software, the higher the ranking, the redder the color (C and D) ((A) and (C) is derived from rat database in the GEO database, (B) and (D) is from the mouse dataset). Pyroptosis-related mRNA-miRNA-lncRNA regulatory networks in SCI (E). The pink circles represent PRGs, the green circles represent miRNAs, and the pale green circles represent lncRNAs. lncRNA long non-coding RNA, miRNA microRNA, SCI spinal cord injury.
Construction of the lncRNA-miRNA-mRNA regulatory network related to pyroptosis
We performed miRNA and lncRNA predictions for the 6 intersecting genes (Casp1, Gbp2, Nlrp3, Pycard, Tlr4 and Casp4) obtained from Venn diagram analysis using the starBase (Version 2.0) website. However, no miRNA or lncRNA predictions were obtained for Casp4 gene. We predicted a total of 28 miRNAs and 34 lncRNAs. These data were imported into Cytoscape software to generate the ceRNA regulatory network (Fig. 7E). This regulatory network elucidated the potential regulatory mechanisms of pyroptosis in SCI.
Validation of hub genes for pyroptosis in SCI by qPCR and WB
The three hub genes (Casp1, Casp4 and Nlrp3) were validated in tissue samples. As shown in Fig. 8, compared to the sham surgery group, the three hub genes in the SCI group exhibited high expression at both the RNA and protein levels, and the differences were statistically significant. These findings are consistent with the results of previous data analysis, indicating an upregulation of PRGs in spinal cord injury and suggesting that inhibiting the expression of PRGs may contribute to the recovery of spinal cord function.
WB (A, C and E) and qPCR (B, D and F) were used to validate the expression of PRGs in rat spinal cord tissue. SCI (spinal cord injury). (The three target gene bands originated from different gels. The gels/blots were cropped, and the original blots are presented in Supplementary Figures. S2–S4) *P < 0.05, **P < 0.01, ***P < 0.001.
Discussion
In this study, we performed bioinformatics analysis of mRNA sequencing data from rat and mouse models of SCI, which were obtained from the GEO database. We integrated these data with PRGs, which have been the focus of recent research. Our analysis revealed differential expression of PRGs in both rat and mouse SCI models, with 10 and 12 DEGs, respectively. The majority of these genes exhibited high expression levels both rat and mouse models of SCI, except for one gene (Dfna5) showing low expression in the mouse injury model. Six common differentially expressed genes were identified in the intersection of rat and mouse PRGs. Enrichment analysis of these DEGs indicated that GO analysis was mainly focused on inflammation-related factors, while KEGG analysis showed that the most genes were enriched on the NOD-like receptor signaling pathway. Our study represents the integration of analysis of SCI and PRGs, and our results suggest that pyroptosis may be one of the specific mechanisms underlying secondary spinal cord injury. Therefore, inhibiting the high expression of PRGs may be a novel therapeutic target for SCI treatment, and our findings have significant implications in this area.
The mechanism of secondary spinal cord injury is highly complex. Currently, there is a considerable amount of evidence indicating the involvement of programmed cell death in the occurrence and progression of secondary spinal cord injury34. These programmed cell death processes include autophagy, apoptosis, ferroptosis, necroptosis, parthanatos, and pyroptosis34. Among these processes, autophagy and apoptosis have been extensively studied and researched in SCI, while pyroptosis, as a relatively recent research focus, has also gained increasing attention. With the elucidation of the mechanisms of pyroptosis in SCI becoming increasingly clear, it is now known that changes in PRGs may have various effects on neuronal cells. Moreover, pyroptosis does not occur as an independent event but often interacts with other forms of programmed cell death, forming a complex network of interactions. Following SCI, a large amount of reactive oxygen species accumulates in cells, which can induce the activation of the NLRP3 inflammasome, subsequently activating Caspase-1. Mature Caspase-1, on one hand, triggers cell pyroptosis by cleaving Gasdermin proteins, and on the other hand, it can weaken the TRIF/Toll-like receptor 4 signaling pathway, inducing cell autophagy35,36. Cell autophagy, in turn, can eliminate damaged organelles (such as mitochondria) and inflammatory factors, thereby negatively regulating NLRP3 and inhibiting cell pyroptosis36. Xu et al.37 found that Toll-like receptor 4 (TLR4) plays an important role in SCI, and inhibiting the expression of TLR4 can alleviate neuron pyroptosis. Additionally, they discovered that the expression levels of NLRP3/GSDMD proteins in the peripheral blood of SCI patients were higher compared to normal patients, and the expression levels were positively correlated with the severity of the injury, suggesting that pyroptosis may aggravate SCI. In this study, we integrated and analyzed multiple datasets from rats and mice, and found differential expression of PRGs in SCI of both species. Furthermore, high expression was observed at days 3 and 35 post-injury in the datasets (GSE171441), indicating that pyroptosis may play an important role in the acute and subacute stages of the disease course. In the correlation analysis (Fig. 6), it can be observed that Caspase-1 exhibits significant correlations with Nlrp3, Pycard, and others, suggesting a possible synergistic or regulatory relationship between them. In conclusion, the role of pyroptosis and its molecular regulation in SCI are highly complex, requiring further in-depth and extensive research.
Pyroptosis is a current research hotspot. It is worth noting that there have been few reports on the study of PRGs in SCI using bioinformatics analysis. In this study, we identified differentially expressed PRGs in SCI through transcriptomic data analysis and joint analysis of multiple datasets. Among them, 10 genes were identified in rats and 12 genes in mice, with an intersection of 6 genes (Fig. S1). Subsequently, GO and KEGG enrich analysis were performed on the PRGs in rats and mice separately to understand their functions and regulatory mechanisms in biological processes. GO analysis indicated that these genes are mainly enriched in pathways related to inflammation, such as the inflammasome complex, interleukin-1 beta production, interleukin-1 production, and positive regulation of interleukin-1 beta production. KEGG analysis suggested that these genes are mainly enriched in the NOD-like receptor signaling pathway, necroptosis, and infection, both in rats and mice. These results indicate that pyroptosis does occur in SCI and may play an important role.
The role of non-coding RNAs in SCI is increasingly being studied, especially miRNAs and lncRNAs. A substantial body of research has demonstrated their widespread involvement in the pathological and physiological processes of SCI, making them potential new targets for therapeutic interventions. miRNAs play an important regulatory role in neuronal apoptosis, the release of inflammatory factors, and oxidative stress following SCI. Liu et al.38 found that the expression level of miRNA-233 significantly increased in a rat SCI model. Intrathecal injection of the miRNA-233 inhibitor, antagomir-223, significantly reduced the expression of apoptosis-related proteins (Bax and cleaved caspase-3), leading to a significant improvement in hindlimb motor function recovery after SCI. Furthermore, studies have shown that miRNAs such as miR-34a, miR-92b-3p, miR-29a, miR-181d-5p, miRNA-99b-5p, miRNA-125b, miRNA-21, miRNA-212-3p, and miRNA-26a are involved in the regulation of neuronal apoptosis following SCI. Fei et al.39 demonstrated that overexpression of miRNA-82 can attenuate the release of inflammatory cytokines (TNF-a, IL-6, IL-1β, etc.) by blocking the IKKβ/NF-kB pathway, thereby improving spinal cord edema and functional recovery in mice after SCI. Wang et al.40 found that miRNA-99a can alleviate oxidative stress and promote functional recovery in rats after SCI by inhibiting the expression of NOX4. The miRNAs exert both positive and negative regulation in SCI, and their mechanisms are highly complex, requiring further in-depth research. It has been discovered that lncRNAs, similar to miRNAs, play an important role in the regulation following SCI and have close interactions with miRNAs41. LncRNAs XIST, BDNF-AS, and MALAT1 are involved in the regulation of neuronal apoptosis after spinal cord injury42,43. Among them, lncRNA XIST can induce neuronal apoptosis by inhibiting the expression of miRNA-7a42. Additionally, similar to miRNAs, lncRNAs can also regulate inflammatory factors and oxidative stress after SCI. In this study, we predicted and identified 28 miRNA and 34 lncRNA through 5 key PRGs. We constructed an mRNA-miRNA-lncRNA regulatory network, which revealed close relationships among them. This regulatory network may represent potential regulatory mechanisms underlying SCI and deserves further investigation.
The involvement of pyroptosis-related signaling pathways in SCI is currently extensive, and the effects of various signals may differ. The PI3K/AKT signaling pathway37,44 and PI3K-AKT-Foxo1 have been found by Hu44 and Xu et al.45, to demonstrate that Taxifolin and CD73 (an immunosuppressive molecule) can alleviate inflammation and microglia-mediated pyroptosis after SCI through the PI3K/AKT signaling pathway, thereby improving spinal cord function. The STING signaling pathway46, discovered by scholars has been found to amplify inflammatory responses and induce pyroptosis46,47. Additionally, Liu and others have found that inhibiting the MAPKs-NF-κB signaling pathway can alleviate neuroinflammation and pyroptosis mediated by microglia after SCI48,49. Of course, many other signaling pathways have been discovered in SCI, such as the JAK2/STAT1 signaling pathway50, AMPK/NLRP3 signaling pathway51, AMPK-mTOR-TFEB signaling pathway52, and AIM2/ASC/Caspase-1 signaling pathway53. In our research, the PRGs enriched the most in the KEGG enrichment analysis were associated with the NOD-like receptor signaling pathway, both in rats and mice data. The NOD-like receptor signaling pathway, within the NOD-like receptor protein family, plays a crucial role in pyroptosis by mediating the assembly of inflammasomes, the activation of Caspase-1, the release of pro-inflammatory cytokines, and inducing cell membrane rupture through GSDMD cleavage. Among the NOD-like receptor protein family, the most relevant inflammasomes to the central nervous system are NLRP1 and NLRP3. Our analyzed data showed significant differences in NLRP3 expression, which closed correlated with SCI.
In conclusion, the specific molecular regulatory mechanisms of pyroptosis in SCI are complex and not yet fully understood. Further research involving extensive molecular exploration and experimental validation is still needed to investigate its role in improving the function of the injured spinal cord.
Conclusions
Pyroptosis is involved in secondary SCI and likely plays a significant role in this process. The majority of PRGs exhibit elevated expression levels in SCI. Inhibiting the expression of PRGs may contribute to the recovery of SCI. The specific regulatory mechanisms of pyroptosis in SCI are complex and require further in-depth research.
Data availability
All data relevant to the study are included in the article or uploaded as supplementary information.
References
McDonald, J. & Sadowsky, C. Spinal-cord injury. Lancet (Lond., Engl.) 359(9304), 417–425 (2002).
Kirshblum, S., Snider, B., Eren, F. & Guest, J. Characterizing natural recovery after traumatic spinal cord injury. J. Neurotrauma 38(9), 1267–1284 (2021).
Dietz, V. & Fouad, K. Restoration of sensorimotor functions after spinal cord injury. Brain J. Neurol. 137, 654–667 (2014).
Stenudd, M., Sabelström, H. & Frisén, J. Role of endogenous neural stem cells in spinal cord injury and repair. JAMA Neurol. 72(2), 235–237 (2015).
Gensel, J. & Zhang, B. Macrophage activation and its role in repair and pathology after spinal cord injury. Brain Res. 1619, 1–11 (2015).
Karsy, M. & Hawryluk, G. Modern medical management of spinal cord injury. Curr. Neurol. Neurosci. Rep. 19(9), 65 (2019).
Cofano, F. et al. Mesenchymal stem cells for spinal cord injury: Current options, limitations, and future of cell therapy. Int. J. Mol. Sci. 20(11), 2698 (2019).
Hutson, T. & Di Giovanni, S. The translational landscape in spinal cord injury: focus on neuroplasticity and regeneration. Nat. Rev. Neurol. 15(12), 732–745 (2019).
Ahuja, C. et al. The leading edge: Emerging neuroprotective and neuroregenerative cell-based therapies for spinal cord injury. Stem Cells Transl. Med. 9(12), 1509–1530 (2020).
Xu, H., Zhao, W., Zhou, Y. & Zhang, L. Multi-omics in spinal cord injury: Diagnosis, prognosis, and treatment. Cell. Mol. Biol. (Noisy-le-Grand, France) 68(11), 58–70 (2022).
Quadri, S. et al. Recent update on basic mechanisms of spinal cord injury. Neurosurg. Rev. 43(2), 425–441 (2020).
Golestani, A. et al. Epidemiology of traumatic spinal cord injury in developing countries from 2009 to 2020: A systematic review and meta-analysis. Neuroepidemiology 56(4), 219–239 (2022).
Sekhon, L. & Fehlings, M. Epidemiology, demographics, and pathophysiology of acute spinal cord injury. Spine 26, S2-12 (2001).
Anjum, A. et al. Spinal cord injury: Pathophysiology, multimolecular interactions, and underlying recovery mechanisms. Int. J. Mol. Sci. 21(20), 7533 (2020).
Sterner, R. & Sterner, R. Immune response following traumatic spinal cord injury: Pathophysiology and therapies. Front. Immunol. 13, 1084101 (2022).
Al Mamun, A. et al. Role of pyroptosis in spinal cord injury and its therapeutic implications. J. Adv. Res. 28, 97–109 (2021).
Hu, X. et al. Role of pyroptosis in traumatic brain and spinal cord injuries. Int. J. Biol. Sci. 16(12), 2042–2050 (2020).
Yin, J., Gong, G., Wan, W. & Liu, X. Pyroptosis in spinal cord injury. Front. Cell. Neurosci. 16, 949939 (2022).
Yu, P. et al. Pyroptosis: Mechanisms and diseases. Signal Transduct. Target. Ther. 6(1), 128 (2021).
Tang, R. et al. Ferroptosis, necroptosis, and pyroptosis in anticancer immunity. J. Hematol. Oncol. 13(1), 110 (2020).
Coll, R., Schroder, K. & Pelegrín, P. NLRP3 and pyroptosis blockers for treating inflammatory diseases. Trends Pharmacol. Sci. 43(8), 653–668 (2022).
Rao, Z. et al. Pyroptosis in inflammatory diseases and cancer. Theranostics 12(9), 4310–4329 (2022).
He, X. et al. Bioinformatics analysis of programmed cell death in spinal cord injury. World Neurosurg. 75, 387 (2023).
Shan, W., Li, S. & Yin, Z. Identification of canonical pyroptosis-related genes, associated regulation axis, and related traditional Chinese medicine in spinal cord injury. Front. Aging Neurosci. 15, 1152297 (2023).
Wang, N. et al. Molecular investigation of candidate genes for pyroptosis-induced inflammation in diabetic retinopathy. Front. Endocrinol. 13, 918605 (2022).
Liu, J. et al. Screening of crosstalk and pyroptosis-related genes linking periodontitis and osteoporosis based on bioinformatics and machine learning. Front. Immunol. 13, 955441 (2022).
Wu, L. et al. The role of pyroptosis and its crosstalk with immune therapy in breast cancer. Front. Immunol. 13, 973935 (2022).
Wu, J., Zhu, Y., Luo, M. & Li, L. Comprehensive analysis of pyroptosis-related genes and tumor microenvironment infiltration characterization in breast cancer. Front. Immunol. 12, 748221 (2021).
Lin, W., Chen, Y., Wu, B., Chen, Y. & Li, Z. Identification of the pyroptosis-related prognostic gene signature and the associated regulation axis in lung adenocarcinoma. Cell Death Discov. 7(1), 161 (2021).
Kanehisa, M. Toward understanding the origin and evolution of cellular organisms. Protein Sci. Publ. Protein Soc. 28(11), 1947–1951 (2019).
Kanehisa, M. & Goto, S. KEGG: Kyoto encyclopedia of genes and genomes. Nucleic Acids Res. 28(1), 27–30 (2000).
Song, Y. X. et al. Antioxidant effect of quercetin against acute spinal cord injury in rats and its correlation with the p38MAPK/iNOS signaling pathway. Life Sci. 92(24–26), 1215–1221 (2013).
Liu, H. T. et al. SARM1 promotes neuroinflammation and inhibits neural regeneration after spinal cord injury through NF-κB signaling. Theranostics 11(9), 4187–4206 (2021).
Shi, Z. et al. Programmed cell death in spinal cord injury pathogenesis and therapy. Cell Prolif. 54(3), e12992 (2021).
Wallisch, J. S. et al. Cerebrospinal fluid NLRP3 is increased after severe traumatic brain injury in infants and children. Neurocrit. Care 27(1), 44–50 (2017).
Lai, M. et al. The NLRP3-Caspase 1 inflammasome negatively regulates autophagy via TLR4-TRIF in prion peptide-infected microglia. Front. Aging Neurosci. 10, 116 (2018).
Xu, S. et al. TLR4 promotes microglial pyroptosis via lncRNA-F630028O10Rik by activating PI3K/AKT pathway after spinal cord injury. Cell Death Dis. 11(8), 693 (2020).
Liu, D. et al. Administration of antagomir-223 inhibits apoptosis, promotes angiogenesis and functional recovery in rats with spinal cord injury. Cell. Mol. Neurobiol. 35(4), 483–491 (2015).
Fei, M. et al. MicroRNA-182 improves spinal cord injury in mice by modulating apoptosis and the inflammatory response via IKKβ/NF-κB. Lab. Investig. J. Tech. Methods Pathol. 101(9), 1238–53 (2021).
Wang, R., Liu, Y. & Jing, L. MiRNA-99a alleviates inflammation and oxidative stress in lipopolysaccharide-stimulated PC-12 cells and rats post spinal cord injury. Bioengineered 13(2), 4248–59 (2022).
Bossi, L. & Figueroa-Bossi, N. Competing endogenous RNAs: A target-centric view of small RNA regulation in bacteria. Nat. Rev. Microbiol. 14(12), 775–784 (2016).
Zhao, Q. et al. Knockdown of long noncoding RNA XIST mitigates the apoptosis and inflammatory injury of microglia cells after spinal cord injury through miR-27a/Smurf1 axis. Neurosci. Lett. 715, 134649 (2020).
Zhou, H. et al. Long noncoding RNA MALAT1 contributes to inflammatory response of microglia following spinal cord injury via the modulation of a miR-199b/IKKβ/NF-κB signaling pathway. Am. J. Physiol. Cell Physiol. 315(1), C52–C61 (2018).
Hu, Z. et al. Taxifolin attenuates neuroinflammation and microglial pyroptosis via the PI3K/Akt signaling pathway after spinal cord injury. Int. Immunopharmacol. 114, 109616 (2023).
Xu, S. et al. CD73 alleviates GSDMD-mediated microglia pyroptosis in spinal cord injury through PI3K/AKT/Foxo1 signaling. Clin. Transl. Med. 11(1), e269 (2021).
Hu, X. et al. Emerging role of STING signalling in CNS injury: Inflammation, autophagy, necroptosis, ferroptosis and pyroptosis. J. Neuroinflamm. 19(1), 242 (2022).
Wang, Y., Shen, D., Zhao, L., Zeng, N. & Hu, T. Sting is a critical regulator of spinal cord injury by regulating microglial inflammation via interacting with TBK1 in mice. Biochem. Biophys. Res. Commun. 517(4), 741–748 (2019).
Liu, Z. et al. Advanced oxidation protein products induce microglia-mediated neuroinflammation via MAPKs-NF-κB signaling pathway and pyroptosis after secondary spinal cord injury. J. Neuroinflamm. 17(1), 90 (2020).
Liu, Z. et al. Pretreatment with kaempferol attenuates microglia-mediate neuroinflammation by inhibiting MAPKs-NF-κB signaling pathway and pyroptosis after secondary spinal cord injury. Free Radic. Biol. Med. 168, 142–154 (2021).
Wang, J. et al. TLR4 aggravates microglial pyroptosis by promoting DDX3X-mediated NLRP3 inflammasome activation via JAK2/STAT1 pathway after spinal cord injury. Clin. Transl. Med. 12(6), e894 (2022).
Yuan, Y., Fan, X., Guo, Z., Zhou, Z. & Gao, W. Metformin protects against spinal cord injury and cell pyroptosis via AMPK/NLRP3 inflammasome pathway. Anal. Cell. Pathol. (Amst.) 2022, 3634908 (2022).
Wu, C. et al. Betulinic acid inhibits pyroptosis in spinal cord injury by augmenting autophagy via the AMPK-mTOR-TFEB signaling pathway. Int. J. Biol. Sci. 17(4), 1138–1152 (2021).
Zhou, Z. et al. Exosome-shuttled miR-672-5p from anti-inflammatory microglia repair traumatic spinal cord injury by inhibiting AIM2/ASC/Caspase-1 signaling pathway mediated neuronal pyroptosis. J. Neurotrauma 39, 1057–1074 (2022).
Funding
This work was supported by the National Natural Science Foundation of China Youth Science Fund Project (81701228 to FBL), the Hunan Provincial Natural Science Foundation (2022JJ70060 to QSZ, 2018JJ3754 to FBL), and the Changsha Natural Science Foundation (kq2208321 to FBL).
Author information
Authors and Affiliations
Contributions
All authors participated in data acquisition. JL and FBL contributed to the conception and design of the study. FSL, HLH, LXD, QSZ and XBW did the data collection and analysis. FSL, HLH and FBL performed experiments. FSL and HLH contributed to the drafting and revision of the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Liu, FS., Huang, HL., Deng, LX. et al. Identification and bioinformatics analysis of genes associated with pyroptosis in spinal cord injury of rat and mouse. Sci Rep 14, 14023 (2024). https://doi.org/10.1038/s41598-024-64843-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-024-64843-6
- Springer Nature Limited