Abstract
Endophytes of Panax have the potential to produce their host plant secondary metabolites, ginsenosides. Panax sokpayensis, an endemic traditional medicinal plant of the Sikkim Himalayas was explored for the isolation of endophytic fungi. In the present study, we have isolated 35 endophytic fungal cultures from the rhizome of P. sokpayensis and screened for ginsenosides production by HPLC by comparing the peak retention time with that of standard ginsenosides. The HPLC analysis revealed that out of 35 isolates, the mycelial extracts of four fungal endophytes (PSRF52, PSRF53, PSRF49 and PSRF58) exhibited peaks with a similar retention time of the standard ginsenoside, Compound K (CK). LC–ESI–MS/MS analysis led to the confirmation of ginsenoside CK production by the four fungal endophytes which showed a compound with m/z 639.6278, similar to that of standard ginsenoside CK with yield in potato dextrose broth flask fermentation ranging from 0.0019 to 0.0386 mg/g of mycelial mass in dry weight basis. The four prospective fungal endophyte isolates were identified as Thermothielavioides terrestris PSRF52, Aspergillus sp. PSRF49, Rutstroemiaceae sp. strain PSRF53, and Phaeosphaeriaceae sp. strain PSRF58 based on ITS sequencing. The present finding highlights the need for further study on growth optimization and other culture parameters to exploit the endophytes as an alternative source for ginsenoside CK production.
Similar content being viewed by others
Introduction
The Panax genus of the Araliaceae family is a well-known medicinal plant with its earliest records found in Traditional Chinese Medicine (TCM) literature. TCM reports the use of Panax rhizome extract as a health supplement and the leaves for preparing tea and other concoctions, which are used mainly as a tonic to invigorate weak bodies1,2. With the advent of time, several studies have attributed the ginsenosides present in Panax rhizome to their medicinal properties3. Its acceptance by the US Pharmacopoeia in 1840 drastically increased the use of Panax rhizome as a medicine worldwide, particularly in the Western World. As Panax grows very slowly and requires 5–7 years to produce a reasonable size of rhizome, it requires specific geographical conditions for its growth, like high elevation and temperate climate of the Himalayan region1,4, and primarily collected from the wild, making the Panax rhizome an expensive commodity1,2. Although the Chinese side of the Himalayan region is well-explored for Panax diversity and its medicinal properties, very few studies have investigated the Indian side of the Himalayas.
The ginsenosides are a group of triterpene saponins excessively produced in the Panax rhizome and are known to have multifaceted pharmacological functions such as anti-ageing, analgesic, antidiabetic, antipyretic, anticarcinogenic, antistress, and antifatigue5,6,7,8. The main ginsenosides present in Panax rhizome such as ginsenosides Rg1, Rd, Rc, Rb1 and Rb2, contain multi-sugar moieties, exhibit little pharmacological activities, and are not readily absorbed by the human body9. The deglycosylation of these inactive ginsenosides into active ginsenosides by intestinal bacteria and digestive enzymes allows their absorption by the human digestive system. Ginsenoside Compound K (CK), a rare protopanaxadiol type of ginsenoside, is known to be the major contributor to the activity of ginsenosides and has been shown to possess anti-inflammation, hepatoprotection, anti-diabetes and anti-cancer activity10,11,12,13. CK is currently synthesized by enzymatic or microbial deglycosylation of the main protopanaxadiol category of ginsenosides (Rb1, Rb2, Rd and Rc)9. However, its manufacturing is limited due to the availability of raw materials, as the Panax plant and the quality rhizome require a long time for cultivation14.
Since the discovery of the anti-cancer compound “taxol”, the ability of endophytes to produce bioactive metabolites in medicinal plants has gained attraction15. Very few studies have reported ginsenoside production by the Panax endophytes. The fungal endophytes of Panax ginseng, Fusarium sp. Pg27 and Aspergillus sp. Pg30 could produce ginsenoside Rb2 and Verticillium sp. Pg42-1 could produce ginsenoside Rc16. A similar study, reported in vitro production of rare ginsenosides Rg3 and Rh2 by endophytic bacteria of P. ginseng, Agrobacterium rhizogenes PDA-217. In this context, screening of ginsenoside-producing endophytes of Panax rhizome is gaining momentum as an alternative sustainable method for industrial ginsenoside production.
Among all the Panax species found worldwide, Panax ginseng, Panax quinquefolius and Panax notoginseng are the most studied for their ginsenosides and therapeutic applications18. Panax sokpayensis is endemic to Sikkim (an Indian part of the Himalayan region) and faces a threat due to excessive harvesting from the wild and illegal marketing19,20,21. An earlier study by Gurung et al., showed the major ginsenoside content in P. sokpayensis which is at par with that of P. ginseng and P. notoginseng20. In the present study, we have screened the fungal endophytes of P. sokpayensis for their ability to produce ginsenosides under in vitro conditions.
Materials and methods
Sampling
Five rhizome samples of P. sokpayensis were collected from Jorbotay (Latitude: 27°16′54.84″ N; Longitude: 88°5′8.12″E; Altitude: 2297 m), Uttarey, Gyalshing district of Sikkim, India (Fig. 1). The samples were collected in a sterile plastic bag and transported to the laboratory in a cooler box, and endophyte isolation was performed within 24 h of collection. The sample collection was carried out according to the guideline of the Sikkim State Government after obtaining the research permit (vide letter no.: 78/GOS/FEWMD/BD-R-2015/CCF(T&HQ)35 dated May 15, 2017) from the office of the Chief Conservator of Forest (T&HQ) cum CWLW, Department of Forest, Environment and Wildlife Management, Government of Sikkim, Deorali, Gangtok–737102, Sikkim. The plant specimen was authenticated by the Botanical Survey of India, Sikkim Himalayan Regional Centre, Gangtok, 737103, Sikkim, Ministry of Environment, Forest and Climate Change, Government of India (vide letter no. SHRC/5/40/2021-Tech-273) and the sample number accorded was IBSD-SC-M11.
Location map showing the sampling site at Jorbotay, Sikkim, India. The green colour star indicates the sample collection site (Map was created using the software ArcGIS 9.3 and 10.8.1; https://arcgis.software.informer.com/9.3/).
Isolation of fungal endophytes from rhizome of P. sokpayensis
With a few minor modifications, the methodology outlined by Mazumder et al. was adopted to isolate endophytic fungi from the rhizome of P. sokpayensis22. The collected rhizome sample (Fig. 2) was rinsed meticulously under running tap water to eliminate adhered soil and dust particles which was followed by washing with sterile distilled water and cut into small pieces (3–4 cm) under sterile conditions. The rhizome fragment was then subjected to surface sterilization under laminar airflow. Surface sterilization is a crucial step to remove any epiphytic microorganism which was carried out by following the standard protocol; the rhizome fragment was first dipped in 70% (v/v) ethanol for 5 min, followed by treatment with sodium hypochlorite (6%) for 5 min and then with 70% (v/v) ethanol for 30 s. Finally, the sterilized rhizome fragment was rinsed with sterile distilled water. The outer tissues were removed using a sterile scalpel after drying the sample aseptically. The sterilized rhizome fragment was cut into small pieces (0.5 × 0.5 cm) and placed on potato dextrose agar (PDA) medium (HiMedia Laboratories Pvt. Ltd, Mumbai, India) amended with streptomycin (50 μg/ml) in Perti plates and incubated at 25 ± 1 °C for 30 days. Following incubation, plates were checked and hyphal tips of fungi, emerging out of the plated rhizome pieces, were picked and grown on PDA medium. The endophytic fungal isolates were purified by hyphal tip culturing, and the isolates were preserved as glycerol stock in cryovials at − 80 °C. As an additional test of surface sterilization, aliquots (100 µl) of the final rinse water of rhizome fragment were also plated onto a PDA medium. No growth on the PDA plate inoculated with final rinse water ensured the efficacy of the surface sterilization process and absence of any epiphytic contamination.
Extraction of ginsenosides from endophytic fungal culture
The pure culture of each fungal endophyte isolate was cultured in potato dextrose broth (PDB) (HiMedia Laboratories Pvt. Ltd, Mumbai, India). Fungal hyphae from freshly grown culture were inoculated into 150 ml sterilized PDB contained in 250 ml Erlenmeyer flask and incubated at 25 ± 1 °C for ten days without shaking. After incubation, culture supernatant and fungal mycelia were separated by centrifugation at 10,000 rpm for 10 min (Sorvall Biofuge Primo) for the extraction of ginsenosides. The fungal mycelia were rinsed with sterile distilled water, dried in an oven at 45 °C for two days and then crushed into powder using liquid nitrogen. One gram of the mycelial powder was extracted using 20 ml methanol by subjecting it to ultrasonication at 30 °C for 30 min. The extracted mixture was filtered using Whatman’s filter paper and the resultant filtrate was concentrated in a rotary evaporator (BUCHI). The methanol extract obtained for each isolate was then transferred to a 2 ml vial and was subjected to drying using a vacuum concentrator (SpeedVac Concentrator, ThermoFisher Scientific) and stored at − 20 °C for further analysis23. The culture supernatant of 20 ml was extracted with an equal volume (20 ml) of ethyl acetate, concentrated using a rotary evaporator (BUCHI) and subjected to drying as above for analysis. One ml of Methanol (HPLC grade) was added to the dried extracts and filtered using Whatman’s 0.3 μm syringe filter which was used for HPLC analysis.
HPLC analysis
The production of ginsenosides by the fungal endophytes was analyzed by using reverse-phase HPLC (LC-20AT, Shimadzu) with Luna 5 μm C18 (250 × 4.6 mm) Column (Phenomenex), using Photodiode Array Detector (SPD-M 20A) with an oven temperature of 60 °C. The wavelength of the PAD detector was kept at 205 nm, and the flow rate was maintained at 1 ml/min with an injection volume of 20 μl. Acetonitrile (Pump A) and (Pump B) HPLC grade water in binary gradient mode was used as mobile phase in the ratios of the solvents A: B at 25% B held for 3 min; to 85% B in 12 min; 85% B held for 1 min respectively. The system then returned to the initial condition in 1 min after 100% of B. The reference ginsenosides viz. CK, Rb1, Rb2, Rb3, Rd, Re, Rf, Rg1, Rg2, Rg3 and Rh2 (ChromaDex, Irvine, USA) were used as standards during the analysis. The standard solutions of all reference ginsenosides were prepared using 100% methanol (HPLC grade) at 1 mg/ml concentration while the standard solution of Rg3 at 1 mg/ml concentration was prepared using dimethyl sulfoxide (DMSO) as per the manual provided by Chromadex. 20 μl each of the different standard solutions were injected into the HPLC system and run separately for 20 min. Ginsenosides' presence in the extract samples was analyzed by comparing the retention time with the standards by following the method of detection described by Sarang et al.23. The samples spiked with ginsenoside standard were run for confirmation24,25.
LC–ESI–MS/MS detection of ginsenosides
The mycelial extracts of the fungal endophytes were further subjected to LC–ESI–MS/MS analysis for the presence of ginsenosides detected tentatively by the HPLC. The HPLC (Dionex Ultimate 3000, ThermoFisher Scientific) with C18 (250 × 4.6 mm) column (Syncromis), Acetonitrile and Water with a flow rate of 0.4 ml/min, and Impact HD mass spectrometer having ESI and Q-TOF (Bruker) was used to perform the mass spectral analyses. Nitrogen was used as both nebulizing gas and desolvation gas at flow rates of 3 bar and 12 L/min, respectively. The temperatures of both the electrospray source and desolvation gas were kept at 180 °C, and capillary potential was set at 3.0 kV. After selecting precursor ions by the first quadrupole mass analyzer, Collision RF was carried out in the range of 300–1500 vpp in the hexapole collision cell. The quadrupole ion energy used was 5 eV, and the collision cell energy used was 10 eV24,25. High-resolution mass spectra were acquired in both positive and negative-ion modes by scanning over the m/z range of 40–2000. The analyses of the mass of the ginsenosides were repeated twice for each fungal culture by comparing them with the ginsenoside standards.
Molecular identification of the ginsenoside-producing fungal endophytes
Out of the 35 endophytic fungal isolates screened, four isolates capable of producing ginsenosides in in vitro conditions were identified by molecular method by ITS rDNA sequencing. The four endophytic fungal isolates viz. PSRF52, PSRF53, PSRF49 and PSRF58 were freshly cultured separately in 150 ml sterilized PDB in 250 ml Erlenmeyer flask and incubated for 10 days in a rotary shaker maintained at 25 ± 1 °C with 140 rpm. The fungal mycelia were separated by centrifugation at 10,000 rpm for 10 min (Sorvall Biofuge Primo), subjected to freeze-drying and DNA extraction was performed by adopting the Cetyltrimethylammonium bromide method as described by Vainio et al.26. The primer pair ITS1 (5′-TCCGTAGGTGAACCTGCGG-3′) and ITS4 (5′-TCCTCCGCTTATTGATATGC-3′) as per White et al.27 was used for ITS gene amplification using a thermal cycler (Bio-Rad) by following PCR parameters: denaturation at 94 °C for 3 min which was followed by 30 cycles of 94 °C for 15 s, 50 °C for 1 min, 72 °C for 45 s, and final extension at 72 °C for 7 min. The QIA quick PCR purification kit (Qiagen) was used to purify the PCR product, further sequenced using an ABI 3730xl Genetic Analyzer (Eurofins Genomics India Pvt. Ltd., Bangalore, India). The NCBI-BLAST search was carried out by using the rRNA/ITS database containing internal transcribed spacer region (ITS) from fungal type and reference materials sequences to obtain the relatedness of the endophytic fungal sequences28. The sequence having maximum query coverage, maximum identity score and highest homology was taken as a reference to assign the identity of the new fungal endophyte isolate. Observing the maximum identity score, the closest reference sequences were chosen and aligned using the multiple alignment software program Clustal W29. The Maximum Likelihood method30 based on the Kimura 2-parameter model was adopted for the construction of phylogenetic trees31, applying the Neighbor-Joining approach, and by taking bootstrap testing32 of 1000 replicates using MEGA633. The ITS sequences of these four endophytic fungal isolates (PSRF52, PSRF53, PSFR49 and PSRF58) were deposited to NCBI GenBank and their accession numbers are ON963979, MN888777, MN888810 and MN888811 respectively.
Results
HPLC analysis evidences ginsenoside production by fungal endophytes of P. sokpayensis
We have screened 35 endophytic fungal isolates of Panax sokpayensis rhizome for ginsenosides production by HPLC by comparing the peak retention time with known ginsenosides standards. The HPLC analysis of the mycelial extract of six fungal endophytes exhibited peaks with a similar retention time of the ginsenosides Rg1 (7.8 min), CK (10.8 min), Rf (8.5 min) and Rg2 (8.4 min). The culture supernatant extract of the fungal isolates did not show peaks having an equal retention time as that of the ginsenoside standards analyzed. Further, we spiked the mycelial extracts with the ginsenoside standards Rg1, CK, Rf and Rg2 to validate the similarity in retention times of the detected ginsenoside compounds. Only four fungal endophytes (PSRF52, PSRF53, PSRF49 and PSRF58) showed overlapping single peaks of ginsenoside Rg1 (7.8 min), Rf (8.5 min) and CK (10.8 min) in the spiked HPLC run (Fig. 3), which were subjected to mass spectrometry for confirmation.
Overlay HPLC chromatogram of Ginsenoside Compound K peak with retention time of 10.8 min (denoted in black), Spiked samples: mycelial extract of endophytic isolate PSRF53+ Compound K (denoted in pink), and the mycelial extract of endophytic isolate PSRF53 (denoted in blue). The arrow indicates the peak of Ginsenoside Compound K.
LC–ESI–MS/MS analysis confirms ginsenoside-CK production by fungal endophytes
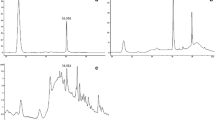
To confirm the ginsenosides detected by HPLC analysis, we subjected the mycelial extracts of the four endophytic fungi (PSRF52, PSRF53, PSRF49 and PSRF58) to LC–ESI–MS/MS analysis. The ginsenoside standards have a neutral mass of m/z 783.4885, 784.4885 and 621.4368 for Rf, Rg2 and CK respectively. The m/z profile of the mycelial extract of four fungal endophyte isolates (PSRF49, PSRF52, PSRF53 and PSRF58) did not coincide with those of ginsenoside standards Rf and Rg2. However, molecular ion peaks of CK[M + NH4]+ (m/z 639.6278) were observed (Fig. 4). The presence of CK was identified by comparing the retention time of 16.4 min and the most abundant ion of 639.6260 in the reference standard and the mycelial extracts of the endophytes. In MS/MS fragmentation, the ion first produced fragment ions at m/z 329.32, m/z 167 and m/z 93. This fragmentation pattern was identical to that of the standard CK. The concentration of ginsenoside CK production by the fungal cultures was calculated based on the LC–MS signal intensity of peak at the retention time of 16.4 min with m/z of 639.6260 in comparison to the known concentration of standard ginsenoside CK (ChromaDex, Irvine, USA). The ginsenoside CK production in PDB flask fermentation by the four endophytic fungal cultures was PSRF49 (0.0386 mg/g), PSRF52 (0.0028 mg/g), PSRF53 (0.0337 mg/g) and PSRF58 (0.0019 mg/g) of mycelial mass on a dry weight basis.
LC–ESI–MS/MS spectra. (A) The first mass spectra of mycelial extract of endophytic isolate PSRF53, the arrow indicates the molecular ion of ginsenoside Compound K (CK) at m/z 639.6260. (B) The MS/MS spectra of m/z 639.6260 of endophyte isolate PSRF53. (C) The MS/MS spectra of m/z 639.6278 of ginsenoside Compound K standard.
Molecular identification of ginsenoside CK-producing fungal endophytes
We used ITS amplicon sequencing to identify the four potential CK-producing fungal endophyte isolates (PSRF49, PSRF52, PSRF53, and PSRF58). The ITS sequence of the four fungal endophyte isolates were compared separately with the related reference sequences of type strains in the GenBank database. The sequence similarity analysis by NCBI-BLAST resulted in 99.64% sequence similarity for isolate PSRF52 with Thermothielavioides terrestris (CBS 117535), 99.63% sequence similarity for isolate PSRF49 with Aspergillus austroafricanus NRRL 233 (NR_135443.1) and 99.29% with Aspergillus versicolor ATCC 9577 (NR_131277.1). The isolate PSRF53 showed less than 96% similarity with Bicornispora seditiosa CBS 135998 of the family Rutstroemiaceae of potentially novel species. Similarly, the isolate PSRF58 showed less than 93% sequence similarity with Edenia gomezpompae CBS 124106 (NR_156217.1) and Setophoma syzygii CBS 146976 (NR_173056.1) of the family Phaeosphaeriaceae. As the isolate PSRF 53 and PSRF 58 showed novelty in their sequence with less than 96% sequence similarity with the available type strains reference sequences, their identity was limited at the family level. Thus, the four fungal endophyte isolates viz. PSRF52, PSRF49, PSRF53 and PSRF58 have been identified as Thermothielavioides terrestris PSRF52, Aspergillus sp. strain PSRF49, Rutstroemiaceae sp. strain PSRF53 and Phaeosphaeriaceae sp. strain PSRF58 respectively. The phylogenetic tree based on the Maximum Likelihood method shows the taxonomic position of these four endophytic fungal isolates in Fig. 5.
Discussion
Studies on the endophytes of Panax spp. mostly focus on the diversity, biotransformation of ginsenosides and changes in the activity of ginsenosides34,35,36,37. These include fungi namely Phoma, Fusarium, Trichoderma, and Setophoma in P. ginseng38; Trichoderma, Chaetomium, Fusarium, Aspergillus, Penicillium, Drechmeria, Emericella, Myrothecium and Preussia in P. notoginseng18,39,40,41; and Cladosporium in P. quinquefolius42 with different bioactivities43,44,45. All these previous reports underscore the difference in the pattern of dominance of the fungal species, indicating the specificity of the distribution of endophytes depends on different geographical locations, environmental conditions as well as plant species46. Among these, only a few studies have been conducted on the ability of the fungal endophytes of Panax to produce ginsenosides. The fungal endophytes of Panax ginseng, particularly Aspergillus spp. cultures reported with triterpenoids saponins production up to 144 mg/L in the flask fermentation16. Agrobacterium, an endophytic bacterium of P. ginseng reported with ginsenosides Rg3, Rb1 and Rh2 production with a concentration ranging from 18 to 62 mg/L in the culture media17. The present study confirmed the ginsenoside CK production by four fungal endophytes of P. sokpayensis in PDB flask fermentation, with a maximum ginsenoside CK yield of 38.6 mg/kg in mycelial mass on a dry weight basis by Aspergillus sp. PSRF49. This is the first report of ginsenoside CK production by the fungal endophytes of Panax and the first investigation on endophytes of Panax sokpayensis.
Our study showed that four fungal endophytes of Panax sokpayensis, namely Thermothielavioides terrestris PSRF52, Aspergillus sp. strain PSRF49, Rutstroemiaceae sp. strain PSRF53 and Phaeosphaeriaceae sp. strain PSRF58 could produce the ginsenoside CK under in vitro conditions. Thielavia terrestris is known for producing industrially important enzymes and bioactive compounds known as thielavialides46. Until now, Thielavia has been reported as an endophyte only in the medicinal plant Physalis alkekengi47. The species of Aspergillus, particularly Aspergillus versicolor, as a fungal endophyte, has been found in several medicinal plants like Anoectochilus roxburghii, Lycoris radiata, Euphorbia royleana, Pulicaria crispa and Avicennia marina, and has been shown to possess the ability to synthesize many anti-microbial compounds and various pharmacologically important compounds48. The member Rutstroemiaceae, particularly Lambertella corni-maris was once reported as an endophyte in the leaves of Abies koreana49. However, the members of Phaeosphaeriaceae, particularly Setophoma spp. have been reported as an endophyte of Panax ginseng and several other plants50,51,52.
Among the ginsenosides discovered, CK is considered the main bioactive compound that brings about the various therapeutic activities of ginsenosides12,53,54. CK is a protopanaxadiol type of ginsenoside that has gained importance due to its high permeability leading to neuroprotective, immune-modulatory, anti-cancer properties and regulatory functions of the hypothalamic–pituitary–adrenal axis9,10,11,55,56. However, CK being a minor ginsenoside is found in low amounts in the Panax rhizome and leaves57. The plant itself is a slow growing, perennial herb that is usually harvested only after 4–6 years and is challenged by sensitivity to various abiotic and biotic factors leading to low yield and variable ginsenoside content58. Panax spp. are grown in greenhouses and controlled environments to meet the demand for ginsenosides but are less preferred than those that are found naturally59,60. The high demands for the plant have led to widespread overexploitation of the plant in its natural habitat making it number negligible in the wild61.
Several methods for ginsenoside production are being sourced to meet this demand. Presently, the methods employed for CK production include microbial engineering, biotransformation, enzymatic and chemical processes57,62,63,64. The latter two processes are deemed expensive and low-yielding, bringing the focus on the former two with a preference for biotransformation. The biotransformation of major ginsenosides Rb1, Rd, Rc etc. to CK by human intestinal bacteria was well studied but the availability of CK was found to be highly influenced by the individual’s dietary and gut microflora65. Deglycosylation by endophytic glucosidases of Arthrinium sp. GE 17–18 from P. ginseng and F. oxysporum and Coniochaeta sp. isolated from P. notoginseng have also enhanced the yield of CK41,63. As new data sets of genomic studies on the biosynthetic gene clusters and metabolic pathways of the host plant and endophytes emerge, information on the possibility of microbial engineering, gene-knockout systems, and heterologous expression in yeasts is being explored65,66,67,68. Some endophytes can directly synthesize rare ginsenosides like Rg3 by Chaetomium69. The discovery of endophytes that could synthesize CK under in vitro was a possibility that researchers believed could enhance CK production. In this regard, our finding on the fungal endophytes of P. sokpayensis having CK production potential confirms that more diversity and screening studies on wild, endemic species of Panax can be employed for the discovery of endophytes that synthesize rare bioactive ginsenosides70. Further study on optimizing and improving the yield of CK by potential fungal endophytes is a prospect for future research work.
Conclusion
In the present study, we report for the first time the fungal endophytes of P. sokpayensis, an endemic medicinal plant of Sikkim, India. Four fungal endophytes of P. sokpayensis exhibited the capability to produce rare ginsenoside CK under in vitro conditions, confirmed by using LC–ESI–MS/MS analysis which had never been discovered earlier. CK is of interest because of its high permeability and several therapeutic uses. The identification of a new source of the ginsenoside from the endophyte offers promise in exploiting the fungus as an alternative source for CK. Further studies are however required for the growth optimization as well as other culture conditions that would not only predispose the endophytes in stable production of the ginsenosides but also without any noticeable attenuation in the production over sub-culture generations. It could pave the way forward towards possible utilization of the endophytes to harness the medicinal property of the host plant without harvesting the rare plant from nature, which can restore the endemic plant in the wild and produce ginsenosides sustainably.
Data availability
The datasets generated and analyzed in the present study are available from the corresponding author upon reasonable written request.
References
Goldstein, B. Ginseng: Its history, dispersion, and folk tradition. Am. J. Chin. Med. 3, 223–234. https://doi.org/10.1142/S0192415X75000244 (1975).
Hu, S. Y. A contribution to our knowledge of ginseng. Am. J. Chin. Med. 5, 1–23. https://doi.org/10.1142/S0192415X77000026 (1977).
Tang, W. & Eisenbrand, G. Panax ginseng CA Mey. In Chinese Drugs of Plant Origin 711–737 (Springer, 1992).
Uchendu, E. E., Paliyath, G., Brown, D. C. W. & Saxena, P. K. In vitro propagation of North American ginseng (Panax quinquefolius L.). In Vitro Cell. Dev. Biol. Plant 47, 710–718. https://doi.org/10.1007/s11627-011-9379-y (2011).
Lyu, X. et al. Ginsenoside Rh1 inhibits colorectal cancer cell migration and invasion in vitro and tumor growth in vivo. Oncol. Lett. 18, 4160–4166. https://doi.org/10.3892/ol.2019.10742 (2019).
Zhang, H. et al. Anticancer effects and potential mechanisms of ginsenoside Rh2 in various cancer types. Oncol. Rep. 45, 1–10. https://doi.org/10.3892/or.2021.7984 (2021).
Zhao, A. et al. A review of neuroprotective effects and mechanisms of ginsenosides from Panax ginseng in treating ischemic stroke. Front. Pharmacol. 13, 946752. https://doi.org/10.3389/fphar.2022.946752 (2022).
You, L., Cha, S., Kim, M. Y. & Cho, J. Y. Ginsenosides are active ingredients in Panax ginseng with immunomodulatory properties from cellular to organismal levels. J. Ginseng Res. 46, 711–721. https://doi.org/10.1016/j.jgr.2021.12.007 (2022).
Chu, L. L., Huy, N. Q. & Tung, N. H. Microorganisms for ginsenosides biosynthesis: Recent progress, challenges, and perspectives. Molecules 28, 1437. https://doi.org/10.3390/molecules28031437 (2023).
Baik, I. H., Kim, K. H. & Lee, K. A. Antioxidant, anti-inflammatory and antithrombotic effects of ginsenoside compound K enriched extract derived from ginseng sprouts. Molecules 26, 4102. https://doi.org/10.3390/molecules26134102 (2021).
Zhou, L., Li, Z. K., Li, C. Y., Liang, Y. Q. & Yang, F. Anticancer properties and pharmaceutical applications of ginsenoside compound K: A review. Chem. Biol. Drug Des. 99, 286–300. https://doi.org/10.1111/cbdd.13983 (2022).
Liu, T., Zhu, L. & Wang, L. A narrative review of the pharmacology of ginsenoside compound K. Ann. Transl. Med. 10, 234–234. https://doi.org/10.21037/atm-22-501 (2022).
Yang, X. D., Yang, Y. Y., Ouyang, D. S. & Yang, G. P. A review of biotransformation and pharmacology of ginsenoside compound K. Fitoterapia 100, 208–220. https://doi.org/10.1016/j.fitote.2014.11.019 (2015).
Kharwanlang, L., Das, M. C., Kumaria, S. & Tandon, P. Histological and SEM studies on somatic embryogenesis in rhizome-derived callus of Panax assamicus. Ban. J. Pharm. Innov. 5(4), 93–99 (2016).
Zhou, X., Zhu, H., Liu, L., Lin, J. & Tang, K. A review: Recent advances and prospects of taxol-producing endophytic fungi. Appl. Microbiol. Biotechnol. 86, 1707–1717. https://doi.org/10.1007/s00253-010-2546-y (2010).
Wu, H., Yang, H. Y., You, X. L. & Li, Y. H. Diversity of endophytic fungi from roots of Panax ginseng and their saponin yield capacities. Springerplus 2, 1–9. https://doi.org/10.1186/2193-1801-2-107 (2013).
Yan, H., Jin, H., Fu, Y., Yin, Z. & Yin, C. Production of rare ginsenosides Rg3 and Rh2 by endophytic bacteria from Panax ginseng. J. Agric. Food Chem. 67, 8493–8499. https://doi.org/10.1021/acs.jafc.9b03159 (2019).
Toghueo, K. R. M., Youmbi, D. Y. & Boyom, F. F. Endophytes from Panax species. Biocatal. Agric. Biotechnol. 31, 101882. https://doi.org/10.1016/j.bcab.2020.101882 (2021).
Sharma, S. K. & Pandit, M. K. A new species of Panax L. (Araliaceae) from Sikkim Himalaya, India. Syst. Bot. 34, 434–438. https://doi.org/10.1600/036364409788606235 (2009).
Gurung, B., Bhardwaj, P. K., Rai, A. K. & Sahoo, D. Major ginsenoside contents in rhizomes of Panax sokpayensis and Panax bipinnatifidus. Nat. Prod. Res. 32, 234–238. https://doi.org/10.1080/14786419.2017.1343322 (2018).
Gurung, B. et al. Molecular cloning and characterization of farnesyl pyrophosphate synthase gene from Panax sokpayensis, a new Panax species from Sikkim Himalaya. J. Appl. Res. Med. Aromat. Plants 14, 100215. https://doi.org/10.1016/j.jarmap.2019.100215 (2019).
Mazumder, S., Singh, L. S. & Bora, T. Endophytic fungi associated with medicinal plants of Gibbon Wild Life Sanctuary, India, and their antagonistic properties. In Advances in Life Sciences (eds Tayung, K. et al.) 213–224 (Studium Press, 2012).
Sarang, H. et al. An endophytic fungus, Gibberella moniliformis from Lawsonia inermis L. produces lawsone, an orange-red pigment. Antonie van Leeuwenhoek 110, 853–862. https://doi.org/10.1007/s10482-017-0858-y (2017).
Miao, X. S., Metcalfe, C. D., Hao, C. & March, R. E. Electrospray ionization mass spectrometry of ginsenosides. J. Mass Spectrom. 37, 495–506. https://doi.org/10.1002/jms.309 (2002).
Wang, C. Z. et al. Ultra-performance liquid chromatography and time-of-flight mass spectrometry analysis of ginsenoside metabolites in human plasma. Am. J. Chin. Med. 39, 1161–1171. https://doi.org/10.1142/S0192415X11009470 (2011).
Vainio, E. J., Korhonen, K. & Hantula, J. Genetic variation in Phlebiopsis gigantea as detected with random amplified microsatellite (RAMS) markers. Mycol. Res. 102, 187–192. https://doi.org/10.1017/S0953756297004577 (1998).
White, T. J., Bruns, T., Lee, S. J. W. T. & Taylor, J. Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. In PCR Protocols: A Guide to Methods and Applications Vol. 18 (eds Innis, M. A. et al.) 315–322 (Academic Press, 1990).
Koetschan, C. et al. The ITS2 Database III sequences and structures for phylogeny. Nucleic Acids Res. 38, D275–D279. https://doi.org/10.1093/nar/gkp966 (2009).
Thompson, J. D., Higgins, D. G. & Gibson, T. J. CLUSTAL W: Improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res. 22, 4673–4680. https://doi.org/10.1093/nar/22.22.4673 (1994).
Felsenstein, J. Evolutionary trees from DNA sequences: A maximum likelihood approach. J. Mol. Evol. 17, 368–376 (1981).
Kimura, M. A simple method for estimating evolutionary rate of base substitutions through comparative studies of nucleotide sequences. J. Mol. Evol. 16, 111–120 (1980).
Felsenstein, J. Confidence limits on phylogenies: An approach using the Bootstrap. Evolution 39, 783–789 (1985).
Tamura, K., Stecher, G., Peterson, D., Filipski, A. & Kumar, S. MEGA6: Molecular evolutionary genetics analysis version 6.0. Mol. Biol. Evol. 30, 2725–2729 (2013).
Fu, Y., Yin, Z. H. & Yin, C. Y. Biotransformation of ginsenoside Rb1 to ginsenoside Rg3 by endophytic bacterium Burkholderia sp. GE 17–7 isolated from Panax ginseng. J. Appl. Microbiol. 122, 1579–1585. https://doi.org/10.1111/jam.13435 (2017).
Zhang, C. et al. Biotransformation of ginsenoside Rc to Rd by endophytic bacterium Bacillus sp G9y isolated from Panax quinquefolius. Int. J. Gen. Mol. Microbiol. 114, 437–444. https://doi.org/10.1007/s10482-021-01529-3 (2021).
Li, R. et al. Diversity and correlation analysis of endophytes and metabolites of Panax quinquefolius L. in various tissues. BMC Plant Biol. 23, 275. https://doi.org/10.1186/s12870-023-04282-z (2023).
An, C. et al. Diversity and ginsenoside biotransformation potential of cultivable endophytic fungi associated with Panax bipinnatifidus var. bipinnatifidus in Qinling mountains, China. Front. Pharmacol. 13, 762862. https://doi.org/10.3389/fphar.2022.762862 (2022).
Goodwin, P. H. The endosphere microbiome of ginseng. Plants 11(3), 415. https://doi.org/10.3390/plants11030415 (2022).
Wei, G. et al. Endophytes isolated from Panax notoginseng converted ginsenosides. Microb. Biotechnol. 14, 1730–1746. https://doi.org/10.1111/1751-7915.13842 (2021).
Liu, S. Y., Yu, Y., Zhang, T. Y., Zhang, M. Y. & Zhang, Y. X. Trichoderma panacis sp. nov., an endophyte isolated from Panax notoginseng. Int. J. Syst. Evol. Microbiol. 70, 3162–3166. https://doi.org/10.1099/ijsem.0.004144 (2020).
Xiong, D. S. et al. Myrothins A–F from endophytic fungus Myrothecium sp. BS-31 harbored in Panax notoginseng. Chem. Biodivers. 18(3), e2000964. https://doi.org/10.1002/cbdv.202000964 (2021).
Li, Y. F. et al. Production of minor ginsenosides from Panax notoginseng flowers by Cladosporium xylophilum. Molecules 27(19), 6615. https://doi.org/10.3390/molecules27196615 (2022).
Park, S. U., Lim, H. S., Park, K. C., Park, Y. H. & Bae, H. Fungal endophytes from three cultivars of Panax ginseng Meyer cultivated in Korea. J. Ginseng Res. 36, 107–113. https://doi.org/10.5142/jgr.2012.36.1.107 (2012).
Dang, L. et al. Chemical constituents from the endophytic fungus Trichoderma ovalisporum isolated from Panax notoginseng. Ann. Microbiol. 60, 317–320. https://doi.org/10.1007/s13213-010-0043-2 (2010).
Xing, X., Guo, S. & Fu, J. Biodiversity and distribution of endophytic fungi associated with Panax quinquefolium L. cultivated in a forest reserve. Symbiosis 51, 161–166. https://doi.org/10.1007/s13199-010-0062-6 (2010).
Ibrahim, S. R. M., Altyar, A. E., Mohamed, S. G. A. & Mohamed, G. A. Genus Thielavia: Phytochemicals, industrial importance and biological relevance. Nat. Prod. Res. 36, 5108–5123. https://doi.org/10.1080/14786419.2021.1919105 (2021).
Wijeratne, E. M. K., Espinosa-Artiles, P., Gruener, R. & Gunatilaka, A. A. L. Thielavialides A–E, nor-spiro-azaphilones, and a bis-spiro-azaphilone from Thielavia sp. PA0001, an endophytic fungus isolated from aeroponically grown Physalis alkekengi. J. Nat. Prod. 77, 1467–1472. https://doi.org/10.1021/np500237h (2014).
Hagag, A., Abdelwahab, M. F., Abd El-kader, A. M. & Fouad, M. A. The endophytic Aspergillus strains: A bountiful source of natural products. J. Appl. Microbiol. 132, 4150–4169. https://doi.org/10.1111/jam.15489 (2022).
Choi, J. W., Park, E. & Eo, J. K. An unrecorded genus Lambertella Höhn. (Rutstroemiaceae) and its unrecorded species in Korea. Korean J. Mycol. 49(1), 127–131. https://doi.org/10.4489/KJM.20210013 (2021).
Plaszko, T. et al. Volatile organic compounds (Vocs) of endophytic fungi growing on extracts of the host, horseradish (Armoracia rusticana). Metabolites 10, 1–15. https://doi.org/10.3390/metabo10110451 (2020).
Poveda, J. et al. Brassica oleracea var. acephala (kale) improvement by biological activity of root endophytic fungi. Sci. Rep. 10, 20224. https://doi.org/10.1038/s41598-020-77215-7 (2020).
Cao, Y. et al. Tobacco root microbial community composition significantly associated with root-knot nematode infections: Dynamic changes in microbiota and growth stage. Front. Microbiol. 13, 807057. https://doi.org/10.3389/fmicb.2022.807057 (2022).
Sharma, A. & Lee, H. J. Ginsenoside compound K: Insights into recent studies on pharmacokinetics and health-promoting activities. Biomolecules 10, 1–40. https://doi.org/10.3390/biom10071028 (2020).
Wang, H. P. et al. Comprehensive identification of ginsenosides in the roots and rhizomes of Panax ginseng based on their molecular features-oriented precursor ions selection and targeted MS/MS analysis. Molecules 28, 941. https://doi.org/10.3390/molecules28030941 (2023).
Choi, E. et al. AKT1-targeted proapoptotic activity of compound K in human breast cancer cells. J. Ginseng Res. 43, 692–698. https://doi.org/10.1016/j.jgr.2019.07.001 (2019).
Im, D. S. Pro-resolving effect of ginsenosides as an anti-inflammatory mechanism of Panax ginseng. Biomolecules 10, 444. https://doi.org/10.3390/biom10030444 (2020).
Noh, K. H., Son, J. W., Kim-, H. J. & Ohy, D. K. Ginsenoside compound K production from ginseng root extract by a thermostable β-glycosidase from Sulfolobus solfataricus. Biosci. Biotechnol. Biochem. 73, 316–321. https://doi.org/10.1271/bbb.80525 (2009).
Fang, X. et al. Effects of growth years on ginsenoside biosynthesis of wild ginseng and cultivated ginseng. BMC Genom. 23, 325. https://doi.org/10.1186/s12864-022-08570-0 (2022).
Proctor, J. T. A. & Palmer, J. W. Optimal light for greenhouse culture of American Ginseng seedlings. J. Ginseng Res. 41, 370–372. https://doi.org/10.1016/j.jgr.2016.04.002 (2017).
Adil, M. & Jeong, B. R. In vitro cultivation of Panax ginseng C.A. Meyer. Ind. Crops Prod. 122, 239–251. https://doi.org/10.1016/j.indcrop.2018.05.076 (2018).
Zhuravlev, Y. N. et al. Panax ginseng natural populations: Their past, current state and perspectives. Acta Pharmacol. Sin. 29, 1127–1136. https://doi.org/10.1111/j.1745-7254.2008.00866.x (2008).
Yao, L., Wang, J., He, J., Huang, L. & Gao, W. Endophytes, biotransforming microorganisms, and engineering microbial factories for triterpenoid saponins production. Crit. Rev. Biotechnol. 41, 249–272. https://doi.org/10.1080/07388551.2020.1869691 (2021).
Sofian, F. F. et al. The 2,3-epoxy naphthoquinol produced by endophyte Arthrinium marii M-211. Nat. Prod. Res. 37(7), 1060–1066. https://doi.org/10.1080/14786419.2021.1998899 (2023).
Chu, L. L. et al. Compound K production: Achievements and perspectives. Life 13(7), 1565. https://doi.org/10.3390/life13071565 (2023).
Sun, W. et al. Novel trends for producing plant triterpenoids in yeast. Crit. Rev. Biotechnol. 39, 618–632. https://doi.org/10.1080/07388551.2019.1608503 (2019).
Sun, Y. et al. Discrepancy study of the chemical constituents of Panax ginseng from different growth environments with UPLC-MS-based metabolomics strategy. Molecules 28(7), 2928. https://doi.org/10.3390/molecules28072928 (2023).
Noushahi, H. A. et al. Biosynthetic pathways of triterpenoids and strategies to improve their biosynthetic efficiency. Plant Growth Regul. 97, 439–454. https://doi.org/10.1007/s10725-022-00818-9 (2022).
Zhao, M. et al. Transcriptome analysis identifies strong candidate genes for ginsenoside biosynthesis and reveals its underlying molecular mechanism in Panax ginseng C.A. Meyer. Sci. Rep. 9, 615. https://doi.org/10.1038/s41598-018-36349-5 (2019).
dan Xu, X. et al. Production of ginsenoside by Chaetomium sp. and its effect on enhancing the contents of ginsenosides in Panax ginseng adventitious roots. Biochem. Eng. J. 174, 108100. https://doi.org/10.1016/j.bej.2021.108100 (2021).
Chu, L. L. & Bae, H. Bacterial endophytes from ginseng and their biotechnological application. J. Ginseng Res. 46(1), 1–10. https://doi.org/10.1016/j.jgr.2021.04.004 (2022).
Acknowledgements
The Department of Biotechnology (DBT), Government of India, New Delhi [DBT/NER/Agri/24/2013] supported the work. LSS thank DBT for the project grant. The authors acknowledge the help and assistance provided by P. Rajani and Dr. M. M. Vasantha, Dept. of Crop Physiology, UAS, Bangalore for HPLC, Mr. Muralidhar Nayak, IISc., Bangalore, for LC-ESI-MS/MS analysis and Dr. Narpati Sharma, Sikkim State Remote Sensing Application Centre, DST, Government of Sikkim for help in map preparation. We thank the Department of Forests, Environment and Wildlife Management, Government of Sikkim for cooperation in sample collection. The Manuscript number provided by IBSD for this communication is IBSD/MS/2020/01/117.
Author information
Authors and Affiliations
Contributions
LSS and RUS conceived the idea and designed the experiments; LSS and SR conducted the experiments and collected the data. SR, LSS and RUS performed the analysis (analyzed the data). SR, LSS and KJ wrote the paper (contributed for manuscript writing). LSS, TP and KJ, edited the manuscript. DS supervised the work. All the authors discussed the results and contributed to the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Rai, S., Singh, L.S., Shaanker, R.U. et al. Endophytic fungi of Panax sokpayensis produce bioactive ginsenoside Compound K in flask fermentation. Sci Rep 14, 9318 (2024). https://doi.org/10.1038/s41598-024-56441-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-024-56441-3
- Springer Nature Limited