Abstract
Maize is gaining impetus in non-traditional and non-conventional seasons such as off-season, primarily due to higher demand and economic returns. Maize varieties directed for growing in the winter season of South Asia must have cold resilience as an important trait due to the low prevailing temperatures and frequent cold snaps observed during this season in most parts of the lowland tropics of Asia. The current study involved screening of a panel of advanced tropically adapted maize lines to cold stress during vegetative and flowering stage under field conditions. A suite of significant genomic loci (28) associated with grain yield along and agronomic traits such as flowering (15) and plant height (6) under cold stress environments. The haplotype regression revealed 6 significant haplotype blocks for grain yield under cold stress across the test environments. Haplotype blocks particularly on chromosomes 5 (bin5.07), 6 (bin6.02), and 9 (9.03) co-located to regions/bins that have been identified to contain candidate genes involved in membrane transport system that would provide essential tolerance to the plant. The regions on chromosome 1 (bin1.04), 2 (bin 2.07), 3 (bin 3.05–3.06), 5 (bin5.03), 8 (bin8.05–8.06) also harboured significant SNPs for the other agronomic traits. In addition, the study also looked at the plausibility of identifying tropically adapted maize lines from the working germplasm with cold resilience across growth stages and identified four lines that could be used as breeding starts in the tropical maize breeding pipelines.
Similar content being viewed by others
Introduction
Maize acreage has been increasing due to its higher demand and profitability owing to its diverse uses such as feed, food, fuel, and industrial raw material1.In Asian tropics, maize is largely grown as a rainfed crop in the rainy season2. Over the recent years, winter season maize is gaining importance in areas with assured irrigation facilities, due to higher yield potential and low incidence of pest and diseases. However, maize cultivation during winter season particularly in Asian tropics is constrained when the seasons temperatures drops below 10 °C. Temperatures below 20 °C is known to restrict growth and development of maize3. Sub-optimal temperature in crops affects the growth from delayed emergence to poor seed-setting4,5 and could lead to a complete loss of crop yield. Low temperatures during early seeding stages diminishes leaf development6,7,8, while the stress at vegetative stage often decreases plant height and causes chlorosis and reduces photosynthetic efficiency9,10,11,12 and the losses. At reproductive stage the stress delays anthesis, increases anthesis silking interval (ASI), decreases pollen shed and eventually leads low seed set13. Use of cold tolerant hybrids along with suitable agronomic practices14,15 might help mitigate the adversities of low temperatures16,17.
One of the major challenges in breeding for cold stress is availability of a reliable high throughput screening facility for imposing the stress on large-scale screening materials. Screening of large number of germplasm particularly in field would entail adjusting the time of planting, such that the targeted crop stage is exposed to the stress18,19. However, it is often difficult to predict with accuracy and duration of the low temperature regime that matches the targeted crop stage(s) 20. Using growth chambers for smaller studies might be a feasible option, but such conditions may not be suitable for large-scale screening of germplasm essential to any breeding program particularly due to the intensive cost and labour. Further, the complexity of the stress in addition to the uncertain relationship between these artificial and natural screens and varied target population of environments (TPEs) could often hamper in obtaining tangible outputs of the pipeline18,21,22.
In addition, most of these studies were limited to improving cold tolerance at germination and seedling stage23,24 and/or were largely based on maize germplasm adapted to temperate climates. It is comparatively easier to mitigate the germination/ seedling stage cold stress through alterations to planting dates25,26, however the uncertainty of the onset of cold waves/ cold snaps during winter season at later stages of crop growth (Vegetative or flowering) necessitates the use of germplasm that show cold tolerance at these key growth stages.
Complexity of the cold tolerant traits along with high genotype × environment interaction suggests the use of molecular markers in breeding for cold stress tolerance27. Molecular markers would largely assist breeders in developing materials with reasonable level of cold tolerance that are suitable for winter season cultivations by circumventing the need for large scale evaluations of germplasm in breeding pipeline. Earliest report of QTL mapping study on maize seedling tolerance to cold stress was by Frashboud et al.28 on a set of recombinant inbred lines. QTLs for cold stress at seedling stage have since been identified in several studies 27,28,29,30,31,32,33,34,35,36. A meta QTL analysis34 involving seven studies identified major regions on chromosomes 2,4 and 8 regulating early development of maize seedlings under cold conditions. The differences in numbers, effects, and positions of QTL in various studies indicate that cold stress tolerance is a very complex trait and is dependent on the population, environment and the stage of the crop that is exposed to the stress. Leipner et al.27 identified QTLs for flowering time, plant height and biomass at harvest on the mapping population used in a previous study screened under suboptimal conditions in field. The study, however, did not co-map any major QTL that was associated with early seedling stage chilling tolerance to the later stage traits, suggesting that maize has higher compensation capacity and the conditions during later stage of crop may be more important in determining the final biomass accumulation. Further, studies involving transcriptome analysis have also been done to understand the molecular basis of the cold stress in maize26,37,38,39.
Considering the limitations and low resolution of QTLs detected through linkage mapping, Genome wide association study (GWAS) that exploits historical recombination was conducted on a set of North American and European dent and European Flint lines exposed to cold stress during seedling stage21. The study revealed three important genomic regions for photosynthetic capacity traits for seedling stage cold tolerance on chromosomes 1, 5 and 7. A similar study on two major maize panels40 identified few major genomic regions (Chromosomes 1,3,4,5 and 7) for photosynthetic capacity and early vigor under cold stress. A study on early-stage cold tolerance revealed SNPs on chromosomes 1, 2, 4 and 7 for germination related traits41. Similar study on a panel of 283 maize lines was conducted by Hu42 and identified 17 SNPs associated with germination traits under cold stress environments. Zhang et al.43 also reported several genomic regions for fourteen germination traits associated with cold tolerance on a panel of 222 maize inbred lines. A recent GWAS on 406 inbred lines44 derived from multiparent advanced generations inter-cross (MAGIC) population revealed the significance of chromosome 10 (bin10.04) as hot spot for photosynthetic traits associated with early-stage cold stress in maize. The most recent study by Yi et al.17 on a panel of 836 diverse maize lines revealed close to 159 loci associated with cold stress related germination and seedling traits, few of the QTLs identified in this study co-mapped to previous QTL mapping studies. Although several genomic regions have been identified for early crop stage cold tolerance in temperate adapted maize, little to no information is available on vegetative/ flowering stage cold stress tolerance particularly for cold snaps45 that are frequented in winter season maize in tropical maize. In the current study, the test crosses from a panel of 297 Asia adapted tropical maize lines were evaluated for cold stress tolerance at early various growth stages. The study envisaged i) identifying putative regions for cold tolerance associated traits such as grain yield, flowering and plant height across different stages of crop growth in this working germplasm and ii) comparing these results to regions previously identified for the traits associated with early-stage cold stress tolerance.
Results
Trait and genotypic variability
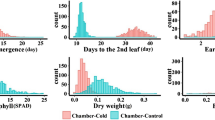
Significant effects of cold stress were observed among the entries exposed to cold stress at different growth stages across the test locations (Table1). Substantial genotypic variation (> 70%) was recorded amongst the entries for flowering date followed by the trait plant height and grain weight (Fig. 1 and Table 2). Cold stress prolonged the vegetative state of the planted crop and invariably extended flowering time of the crop (> 90 days). The most substantial effect of cold stress environments could be observed in the trial planted at L4 (Ludhiana_2019), where the average days to 50 percent flowering in the entries was 160 days followed by L5 (Varanasi_2019). The stress affected both male and female flowering in the crop and as expected the variation was highest in L4, where the trials were exposed to cold stress and remained suspended at vegetative stage for a very long duration. The mean grain yield in the trials at L2 and L3 (> 6.0 t/ha) was higher than that observed for in L4 and L5 (~ 5 t/ha). Similarly, the average plant height at this location (L4) was high, with large number of entries planted reaching above 150 cm. However, the heritability for the trait in L4 was low (0.24). Most decipherable effect of cold stress in field condition was recorded on the plant stand in L1 (Ludhiana_2018). The trial at this location was planted in sub optimal temperatures to study the effect of low temperatures on the germination ability of the entries (58 percent). Agronomic traits such as days to 50 percent flowering, plant height or yield were not recorded in this trial due to the poor plant stand. Among the entries four most stable and adaptable entries were identified based on stability parameters and their performance across the cold stress tolerance at different crop stages are presented in Table 3. The coefficient of regression recorded for the entries along with standard deviation are presented in supplementary Table 1. Most of the traits observed under cold stress showed near normal distribution across locations with a slight skew for plant height which showed strong association with grain yield (R2 = 0.50). The association of the flowering traits among themselves was positive and significant (R2 > 0.98) while this relationship was negative and weak with grain yield (R2 < − 0.20) (Fig. 2).
Population structure and Linkage disequilibrium
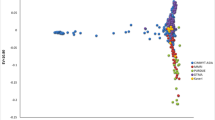
The population showed a high LD decay of 1.5 Kb at r2 of 0.2 and 4.27 Kb at r2 of 0.1 (Fig. 3). SNP associations identified were corrected for structure using three principal components cumulatively accounting for close to 15% of the variation (Supplementary Material Fig. 1). GWAS analysis was done on the primary trait grain yield under cold stress at the four test locations using the multi-locus models (MLM, mrMLM, MLMM, BLINK and Farm CPU). Significantly associated SNPs were identified and compiled through multi locus model “BLINK” as it had the best fit Q–Q plots (Supplementary Material Fig. 2a and 2b with least amount of genomic inflation for the primary trait- Grain yield across the environments. Considering that the test environments in this study were varying with significantly high environmental variance, the GWAS analysis was done on three combined datasets involving vegetative stage cold stress environments (L4 and L5), flowering stage cold stress environments (L2 and L3) and across all environments. The results were then compared to identify common significant SNPs across these datasets. For the study we used a threshold of P < 1 × 10−5, against a holm and bonferroni P value (< 0.01) which was too stringent in identifying top SNP association across the different environment data sets.
Association mapping
Large number of significant SNP associations at P value P < 1 × 10−5 were identified for the traits studied across the different environments (Supplementary Material Table 2, Suppl. Figure 3), however, several of the most significant associations (P value < 0.01) across the traits and locations congregated on chromosomes 3 (bin 3.05–3.06) (Table 4). Among the traits studied, most significant SNPs for percent germination were found on chromosomes 4 (bin 4.08) and 8 (bin8.03). The SNP on chromosome 8 (S8_21830966) recorded the highest allelic effect (> 3% germination). Significant SNPs for flowering traits (AD, ASI and SD) were also found on most of the chromosomes with most significant SNPs accounting for more than 10% percent of phenotypic variation in vegetative stage cold stress environments on chromosomes 1 (bin 1.04), 3 (bin 3.06), 7 (7.02) and 8 (8.06) and on chromosomes 1 (bin1.02, 1.09), 5 (5.06) and 9 (bin9.07) for flowering stage cold stress locations. The SNPs identified for male flowering on chromosomes 5 (bin 5.06) and 8 (bin8.06) co-localised with the identified significant SNPs for female flowering accounting for more than 10 percent of the phenotypic variation across the two datasets.
Significant SNPs for plant height were found distributed across the chromosomes for flowering and across environment datasets. The SNP on chromosome 3 (S3_153769448) and 5 (S5_206305411) accounted for more than 15% of the phenotypic variation. Significant SNPs for the trait grain yield (adjusted for plant height) were also found distributed across most of the chromosomes, however the SNPs on bins 2.02 (S2_14539348) and 2.04 (S2_45988012) for flowering stage cold stress datasets and bin 2.07 (S2_213163019) for vegetative stage cold stress are of particular interest as they accounted for more than 20% of the phenotypic variation (Table 4, Fig. 4). The complexity of the trait grain yield is evident as the study did not identify any common SNPs across the datasets. Hot spots with significant QTNs (P value < 0.5 × 10–5) for multiple traits were also observed in the study (Table5). The region on bin 3.05–3.06 and 8.05–8.06 are of particular interest, as QTNs of multiple traits congregated to these region.
Haplotype regression
A subset of SNPs was selected to perform a haplotype trend regression on the grain yield dataset of the current study. A set of 292 SNPs with P value ≤ 1.0 × 10−3 in the association tests for grain yield across location dataset formed 50 haplotype blocks. The regression analysis identified 6 of these haplotype blocks with Bonferroni p value of less than ≤ 0.05 (Table 6). These blocks were found congregated on chromosomes 2, 5, 6 and 9. The highest phenotypic variation was accounted for by the blocks on chromosome 5 (bin 5.07 and bin 5.05) followed by on chromosome2 (bin2.02).
Discussion
Cold stress is one of the major abiotic stresses encountered by tropical maize cultivated during off-season (Winter season) particularly in Asian tropics, causing irreversible physiological damages and severe yield losses26,46,47,48,49,50,51,52. The unpredictability of cold snaps during the winter season is a major impediment to adoption of maize for cultivation. Consequently, it is pertinent to have germplasm resilient to cold stress particularly for products that are targeted to this season.
Variability for cold stress tolerance within tropical germplasm has been reported in previous studies19. The current study also showed substantial variation for this stress except for the trait germination percentage with most entries in L1 showing poor plant stand. The poor germination percentage (< 50 percent) in the trials sown in December at L1 is attributed to the low temperatures prevailing during the planting of the trial. The effect of low temperatures on germination in maize has been extensively reviewed in several studies14,53,54. The poor germination rates observed in the current study however hindered reliable estimation of other morpho-physiological traits such as grain yield plant height or days to 50 percent flowering, and this trial was limited to studying the effect of cold stress on seedling germination. In addition, cold tolerance to germination is unlikely a breeding target for off season planting in tropical environments, as this stress can also be mitigated largely through changes in planting time14. Apart from germination, the most evident effect of cold stress was observed on the trait days to 50% flowering. In both the testing years, the flowering stage of the trials were delayed. However, the earliest flowering (< 100 days) was recorded in trials planted during 2018 (L2 and L3), wherein the vegetative stage (V1 to V5) of the crop was not exposed to low temperatures and sub-optimal temperatures were primarily reported during V9 to VT stage of the crop. Cold stress delays flowering of most of the crops55, however, sub-optimal temperatures just before or at the initiation of floral transition is reported to inhibit branch meristem development and number of spikelet pairs56 in maize. These locations, however reported comparatively higher grain yields than L4 and L5 where the crop was exposed to cold stress during vegetative stage. The entries in these trials planted at L4 and L5 remained suspended in vegetative state (V3 to V6) for longer duration further delaying the flowering till the temperatures improved. Interestingly, the flowering initiation in the entries were slow initially due to cold stress and hastened particularly in L4 as the temperature improved during later part of the crop cycle. All the entries in this location (L4) flowered within an interval of 10 days as against more than 20 days observed in other trials. Zaidi et al.19 suggested that the genotypes which required more days to complete their vegetative growth had fewer days for reproductive growth, resulting in lower per day yield, and eventually yielded very low.
Cold stress also impedes the cell cycle and rate of cell division57 there by slowing the growth rate of crops. The rate and duration of seed filling are also reported to decrease with prolonged exposure to cold stress at early grain filling58. Even short spells of cold nearing 5 °C at the V3–V5 growth stages are reported to cause photoinhibition51, chlorosis, membrane damage, and eventually necrosis59 or even plant death52. As expected, reduced plant height was observed in most of the test locations where the crop was exposed to cold stress, however this effect was not prominent in L4 as the entries were taller in this location. This difference might possibly be a result of elongated internodes due to poor sun-light intensity at this location during winter season. Severe cold stress in general is reported to reduce the plant height along with total crop biomass in maize19,60. However, it is reported that moderate temperatures for few days (temperatures 10–12 °C) preceding severe cold stress slows the growth and development of the crop but does not cause permanent injury26,61. In L4, the initial daily temperatures (Tmin) were observed to 10–12 °C for nearly a month prior to prolonged severe cold stress possibly contributing to cold acclimation of the crop. Plant height has been suggested in few previous studies as one of the primary candidates for breeding for stability due to its non-relevant interactions with environments and positive effects on yield62. Considering the high variation observed among the different environments, the performance of entries particularly for grain yield varied. The identification of only four stable and adaptable entries out of 300 entries screened across environments, suggests that resilience do exist, though at very low frequency in the tropical germplasm for cold stress. Expanding the trial size and accommodating more germplasm in screens may result in identifying more of stable germplasm for tolerance to cold stress. The test crosses from these lines performed reasonably well in all the cold stressed environments and had showed reasonable levels of gemination percentages. However, considering these lines are not elite, they would be most suitable as donor lines or sources of enriching favourable alleles in populations that exist for developing elite cold resilient inbred lines in the program.
Mapping of QTLs in biparental families and genome wide association studies on diversity panels are the two most used approaches in identifying genomic regions associated with the trait of interest. GWAS approach uses historical recombination events and larger allelic diversity compared to family-based mapping approach to identify potential trait -marker associations. In the current study, we used GWAS approach to identify SNP markers associated with cold stress tolerance in a panel of tropical maize lines screened under field conditions. The current panel of lines had a high LD decay which is typical of tropical germplasm63,64,65, which indicates the diversity among the lines involved and the amenability to high resolution mapping studies.
The current study identified a suite of significant SNPs for various traits recorded under cold stress at different crop stages across various locations. Poor germination in seedling stage exposed to cold stress is one of the major traits that could adversely affect the final plant stand in the field there by impacting the realised yield. In the current study, SNPs have been identified for this trait in most of the chromosomes The SNP with the lowest P value (4.32 × 10–6) was observed on chromosome 4 (S4_198565168) bin 4.08. A large number of SNPs associated with the trait also colocalized (< 20 Mb) to this region. This SNP was in close proximity to QTLs identified in previous studies for specific leaf area21 and chlorophyll content40 in maize seedlings exposed to chilling temperatures. Further the gene model associated within this region also mapped to a gene dek44 (defective kernel44), encoding a mitochondrial ribosomal protein associated with seed development66,67. The SNP identified on bin2.04 (S2_47606952) and bin 7.00 (S7_2582951) were in close proximity to the SNPs identified for relative germination index, relative days to 50% root germination 42 and relative simple vitality index43 on a large panel of maize lines screened under low temperatures.
Cold stress is known to affect flowering machinery of maize leading to poor seed set14. As expected, in the current study cold stress delayed the flowering of the crop. However, this delay was not consistent across locations, and the flowering was delayed the most in L5 (Ludhiana_2019) followed by L4 (Varanasi_2019). These variations might be a result of possible differences in genetic mechanisms associated with delayed flowering in response to cold stress at different stages. The few co-localized SNPs identified for flowering across the environments is possibly a consequence of the profound variation observed in each of the environments. QTLs for flowering in maize have been reviewed extensively68,69,70,71with few of the loci also cloned or fine mapped72,73. In the current study, SNPs identified for male flowering across datasets were found on chromosome 3 (S3_186214261; bin3.06) and chromosome 9 (S9_90843159, bin9.03). In addition, the region on bin3.05–3.06 seems to be very important as a large number of SNPs co-localized to this bin. The gene models associated with the significant SNPs found with in this region were found to code for S-formylglutathion hydrolase and GDSL esterase/lipase (LIP4). GDSL esterase are largely involved in various plant developmental processes74, and play a major role in reproductive structures such as anther development and pollen dehiscensce in maize and Arabidopsis75,76,77,78. In addition, a cluster of SNPs (more than 7) were also identified for the across dataset on chromosome 2 (bin 2.03) and 8 (bin8.05) which were in close proximity (< 10 Mb) to identified genes for flowering zfl2 and vgt173,79. The gene model associated with the S9_90843159 (GRMZM2G056093) was associated with Tetratricopeptide repeat (TPR)-like superfamily proteins. Tetratricopeptide repeats are known to mediate protein interactions with partner proteins and are involved in plant stress and hormone signalling. Role of few TPRs during temperature stress for mediating removal of damaged proteins in Arabidopsis and maintaining homeostasis is well studied80,81. Two of the SNPs identified for female flowering on chromosomes 4 (S4_226942041) and 10 (S10_145198108) were near the QTNs identified in a previous study33 for female flowering in maize under cold stress. The SNP on chromosome 10 mapped to a region close to a flowering gene zfl182.
Plant height (PH) is also one of the major selection factors in any breeding program. In the current study, reduction in plant height have been reported under cold stress across most of the locations. Several QTLs for plant height have been identified and genes for this trait cloned in maize83. Numerous SNPs for plant height across vegetative and flowering locations co-located in a region on chromosome 3 (bin 3.05-bin3.07), followed by SNPs on chromosome 1 (bin 1.01; 1.03–1.04) chromosome 2 (bin2.08) and chromosome5 (bin5.05). The region on chromosome 3 (S3_153769448) is known to harbour recessive semi-dwarf gene (sdw2)84. Incidentally this SNP also accounted for a large proportion (> 60%) of the phenotypic variation for the trait under vegetative stage cold stress tolerance. Most of the identified SNPs for plant height collocated to previously identified significant QTNs11,85.
Grain yield under cold stress being the primary trait was also subjected to GWAS in the current study and as expected, several significant SNPs were identified for this trait across locations. However, the study did not find any congruent SNPs across the four test locations revealing the complexity of the trait. The most significant SNPs for flowering stage cold stress tolerance dataset were identified on chromosomes1 (bin 1.11), 2 (bin2.08), 3 (bin3.05) and 7 (bin7.04) and on chromosomes 2 (bin2.02 &2.04) and 3 (bin3.06) for vegetative stage cold stress tolerance. The SNPs on chromosome 2 (bin 2.04 S2_45988012 and 2.08 S2_213163019) contributed substantially to the phenotypic variation under vegetative and flowering stage cold stress. As several of the significant SNPs were in LD, a haplotype trend regression was performed on the across yield dataset. The haplotype regression on grain yield revealed 6 haplotypes with a Bonferroni threshold (P value ≤ 0.05) on chromosome 2, 5, 6 and 9. The haplotypes identified on chromosome5 (bin5.05 and 5.07) had the lowest FDR. In addition, several SNPs associated with flowering and grain yield traits at most of the test location also collocated to bin 3.05. The gene models for most of these significant SNPs in this region were associated to enzymes such as glutathione transferases, S-formylglutathione hydrolase suggesting the significance of this region for cold stress tolerance in maize.
It has been widely understood that the cold stress adaptation in maize is a result of changes or modifications in cell structure, photosynthesis particularly PSII accompanied by changes in the development of maize and genes associated with these changes may contribute to cold stress tolerance. In the current study, the gene models for significant SNPs in the identified haplotypes for grain yield were associated with pentacotricopeptide repeat 60, ABC transporter protein, CID domain-containing protein, Zinc finger CCCH domain-containing protein 40. Several of these enzymes are associated with abiotic stress tolerance and/ or of cell wall constituents rendering them to be important in abiotic stress breeding. For instance, ABC transporter proteins are involved in maintaining cellular osmotic homeostasis and has been previously reported to contribute to tolerance to various abiotic stresses86. CID domain-containing protein and CCCH zinc-finger proteins regulate the adaptation of plants to abiotic stress. CCCH zinc-finger proteins improve the cold tolerance of plants by directly regulating the expression of downstream cold-related genes as transcription factors87. This further marks the importance of these regions in enhancing cold stress tolerance in maize.
The study also identified few hot spots where in significant SNPs for multiple traits for cold stress colocalized. The gene models associated with the significant SNPs within these regions have been previously associated with cold stress tolerance, such as serine/threonine kinaes, S-formyl glutathione hydrolase (SFGH), Calcium uniporter protein, Peptide transporter PTR2, kinesin-like protein KIN-4C, potassium channel proteins and transcription factors such as WRKY DNA-binding proteins. Several of these enzymes are associated with abiotic stress tolerance and/ or of cell wall constituents rendering them to be important in abiotic stress breeding. Families of Serine/Threonine protein kinases (SNRK-2) are known to enhance cold stress tolerance in Arabidopsis88. In addition, enhanced expression of few of SNRK-2 genes have also been reported under cold stress in maize89,90. Further, WRKY transcription factors constitute one of the largest transcription factor families in plants and play important role in plant growth and thermal stress response91. As an outcome of this study, SNP assays are being developed for the SNPs identified in these hotspots, and the gene models associated with the most significant SNPs for secondary traits to validate them in breeding populations for them to be used for selections in the breeding pipeline.
Conclusion
The study identified lines from the tropical maize breeding germplasm that are suitable for initiating cold stress tolerance breeding for developing and deploying hybrids directed at winter season cultivation of maize in South Asia. In addition, a suite of significant SNPs for grain yield and other secondary traits under cold stress in tropical maize germplasm were also identified in the study that could be of utility in these breeding pipelines. Several of these SNPs identified co-localized to QTLs previously reported for chilling tolerance, signifying their direct applicability in breeding programs post validation in independent breeding population. Further, genomic regions with cluster of SNPs for multiple cold related traits has also been identified particularly on chromosomes 1 (bin 1.01 &1.05), 4 (bin 4.05), 7 (bin 7.02) and chromosome8 (bin8.03) in this study, that would be of interest to further investigate. It was also observed that many of the SNPs identified explained minor effects suggesting the possibility of following a genomic selection approach, particularly in the initial stages of the breeding pipeline for enhanced and faster genetic gains.
Materials and methods
An association mapping panel was constituted involving 297 diverse maize inbred lines sourced from CIMMYT’s lowland tropical maize program. The entries were a subset of a large panel (424) of working germplasm92 that have been used in previous studies used for identifying genomic regions for various abiotic stresses. The elite lines active in the breeding program and constituting the working germplasm of CIMMYT-Asia program involving both tropical and subtropical lineage. This panel of 297 lines were test-crossed with a CIMMYT tester line with high general combining ability (CML451) and phenotyped under natural field screens for cold stress associated traits under seedling, vegetative and flowering stage over five locations between the years 2018 and 2020.
Experimental design
The test-cross progenies of association mapping panel involving 297 lines were planted using an alpha-lattice design with two replications across the four locations during the winter seasons of 2018 to 2020 across different target population of environments in India. The locations and planting dates within each locations were selected such that the lowest temperature period of the season coincided with the targeted crop growth stage viz., germination, vegetative and flowering stage (Table1). Each entry at the locations were planted in a 4-m long plot with a plant-to-plant spacing of 20 cm and a row-to-row spacing of 70 cm. Nitrogen (N) in the form of urea (60 kg ha−1) , Phosphorus ha−1 as single super phosphate (60 kg ha−1) and Potassium as muriate of potash (40 kg ha−1) along with 10 kg Zinc as zinc sulfate were applied as basal dose prior to planting. The second and third doses of N (each 30 kg ha−1) were side-dressed when plants were at knee-high and at tasselling stage, respectively. Pre-emergence application of pendimethalin and atrazine [both at 0.75 kg ha−1 active ingredient (a.i.), tank mixed] were used to keep the crop weed-free at early growth stages. Standard agronomic and plant protection measures were followed throughout the cropping period to prevent other biotic or abiotic stress affecting the trials.
Phenotyping
The complete set of replicated trials involving the 297 entries were planted during the winter season at 5 locations, with planting dates adjusted such that the minimum temperature (< 15 °C) coincided with the critical growth stages of the crop representing the off-season environmental conditions that are prevalent in these TPEs. The different planting dates at each of the location and the targeted crop stage in these locations are detailed in the Table 1. The weather parameters during the crop growth stage at these locations are presented in Supplementary material Fig. 1. Observations were recorded on a plot basis for major agronomic traits such as days to 50 percent male and female flowering, plant height and plot yield. The complete plots of the trials were harvested and shelled, and the total kernel weight along with the moisture percentage was recorded. The grain yields were then converted to per hectare adjusted at 12.5 percent moisture content.
Environmental variation
The trials were planted during the winter season at four locations (Table 1). Planting dates at each of these locations varied such that the crop could be exposed to low temperatures at varied growth stages (Table 1). The trial at location 1 (Ludhiana 2018) was planted during the second fortnight of December when the minimum daily temperatures reached below 4 °C and maximum daily temperature was < 20 °C (Supplimentary Fig. 1). The minimum temperatures (night) improved after a month but was still below < 10 °C. This trial location was used for studying the effect of low temperature on seedling germination and thus the plant stand. Two more trials were planted during October of the year 2018 at Locations 2 (L2: Samastipur 2018) & 3 (L3: Varanasi 2018) such that the pre flowering stage (V7 to VT) coincided with low temperatures. In both the locations, the temperatures during the first month of the planting and active vegetative growth stage were optimal (Tmax ~ 30 °C and Tmin ~ 15 °C). These trials experienced very low temperatures during the month of December and January (pre- flowering to flowering stage), when daily Tmin reached below 10 °C. The temperatures started to increase during the month of February. The trials in the second year of testing at locations 4 (L4: Ludhiana_2019) and 5 (L5: Varanasi_2019) were planted during November, such that the active vegetative stage of the trials coincided with low temperatures. As expected, the temperatures during the initial month of the planting were optimal (Tmax ~ 25 °C and Tmin ~ > 10 °C) and started to drop (Tmax < 20 °C and Tmin < 6 °C) from the second fortnight of December in both the locations. The sub-optimal minimum temperature (< 10 °C) regime continued for more than 3 months in location 4 (L4: Ludhiana_2019) extending the vegetative state of the crop.
Genotyping
The panel of lines studied were genotyped using the GBS platform at the Institute for Genomic Diversity, Cornell University, Ithaca, NY, USA93. A total of 955,690 SNPs were generated through GBS v2.7 platform that were further filtered for the analysis, based on the method described by Suwarno et al.94 with slight modifications. A call rate of > 0.85 and minor allelic frequency (MAF) > 0.05 criteria resulted in a set of 213,273 SNPs for association mapping analysis. Marker density for association mapping panel was one SNP per 4.27 kb. Structure of the association panel was determined using principal components, wherein the SNPs were further filtered based on a call rate criterion of 0.9 and MAF < 0.1. In addition, Linkage Disequilibrium (LD) pruning based on adjacent markers was also carried out resulting in a set of 63,151 SNPs for principal component analysis.
Statistical analysis
A linear mixed model was used for the analysis of the traits using the web handle META-R developed by CIMMYT95. The model for the analysis is represented as
Considering the strong relationship between grain yield and plant height, a covariate analysis was followed to obtain adjusted grain yield following the model
where, Yijkl are the trait of interest, µ is the mean effect, Envi is the effect of the ith environment Repi is the effect of the ith replicate, Blockj (EnviRepi) is the effect of the jth incomplete block within the ith replicate and environment, Genl is the effect of the lth genotype, Envi and Envi × Genl are the effects of the ith environment and the environment by genotype interaction, Cov is the effect of plant height and ε is the residual. All the effects were considered random except plant height which is considered fixed in the model.
The broad-sense heritability (h2) for individual and pooled dataset for the traits were estimated as \({\upomega }^{2}= {\upsigma }^{2}\mathrm{g }/({\upsigma }^{2}\mathrm{g }+{(\upsigma }^{2}\mathrm{e }/\mathrm{ R}))\) and \({\mathrm{h}}^{2}= {\upsigma }^{2}\mathrm{g }/({\upsigma }^{2}\mathrm{g }+{\upsigma }^{2}\mathrm{ge }/\mathrm{E }+ {\upsigma }^{2}\mathrm{e }/(\mathrm{E }\times \mathrm{ R}))\) respectively. Wherein, σ2g, σ2ge and σ2e represents variance components of genotype, genotype × environment and residuals, and number environments and replicates are represented as E and R.
Further to assess the stability of the genotypes across location, the grain yield from each for the entries tested were used to estimate coefficient of stability following Eberhart and Russell96. The genotypes with coefficient of regression close to 1 and low standard deviation were considered as adaptable and stable. The analysis was performed on CIMMYT data handle GEA-R97.
Association mapping
The analysis was carried out using multiple multi-locus models (Farm CPU, mrMLM, MLM, MLMM and BLINK) in GAPIT98,99,100,101,102. Association tests in between markers and association mapping panel phenotypes were corrected for population structure and kinship. The kinship matrix used as covariates were treated as random effects. The Quantile–quantile plots plotted using the observed − log 10P values and the expected − log 10P values for the primary trait of the study grain yield across the vegetative and cold stress environments were used to select the best model (BLINK)102 for association analysis based on observed minimal genomic inflation. A threshold of p value ≤ 1 × 10–5 was considered as a threshold for declaring a significant association. The associated SNPs were considered co-localized if found in proximity (within 1 Mb) to previously reported SNP for the trait or to be in the same chromosomal bin (www.maizegdb.org). In addition, the gene models with the putative SNP associations were identified from maize GDB genome browser at http://www.maizegdb.org (B73 RefGen_V2) and information on their prospective protein codes were obtained from Uniprot genome browser at http://uniprot.org.
Haplotype trait regression approach was also followed for the trait grain yield across environments. SNPs with a P value ≤ 0.0001for association in GWAS for grain yield in all the test locations were selected for haplotype detection and trait regression. Expectation maximization (EM) algorithm with 50 EM iterations, EM convergence tolerance of 0.0001, and a frequency threshold of 0.01 was used to estimate the haplotype frequency and block detection in SVS version 8.6.0. A step wise regression with forward elimination process was followed between the identified haplotype blocks and the trait grain yield across the four test locations.
Data availability
The phenotypic and the genotypic datasets used in the trials are deposited and available on CIMMYT Dataverse web handle (https://hdl.handle.net/11529/10548826). Additional information is also presented in the Supplementary Tables S1–S2.
References
FAO Stat. Food and Agricultural Organization of the United Nation. Economic and Social Department. Statistic Division (ESS). Retrieved from FAO STAT website: http://www.fao.org/about/whowe-are/departments/statistics-division/en (2018).
Prasanna, B. M. et al. Genetic variability and genotype× environment interactions for kernel iron and zinc concentrations in maize (Zea mays L.) genotypes. Indian J. Agric. Sci. 81, 704 (2011).
Lee, M. et al. Expanding the genetic map of maize with the intermated B73× Mo17 (IBM) population. Plant Mol. Biol. 48, 453–461 (2002).
Pertea, M., Kim, D., Pertea, G. M., Leek, J. T. & Salzberg, S. L. Transcript-level expression analysis of RNA-seq experiments with HISAT StringTie and Ballgown. Nat. Protocol 11, 1650–1667. https://doi.org/10.1038/nprot.2016.095 (2016).
Reeves, P. & Coupland, G. Analysis of flowering time control in Arabidopsis by comparison of double and triple mutants. Plant Physiol. 126, 1085–1091. https://doi.org/10.1104/pp.126.3.1085 (2001).
Aidun, V. L., Migus, W. N. & Hamilton, R. I. Use of inbred seedling cold tolerance to predict hybrid cold tolerance in maize (Zea mays L.). Can. J. Plant Sci. 71, 663–667 (1991).
Janda, T., Szalai, G., Tari, I. & Paldi, E. Hydroponic treatment with salicylic acid decreases the effects of chilling injury in maize (Zea mays L.) plants. Planta 208, 175–180 (1999).
Gillani, S. F. et al. Assessment of cold stress tolerance in maize through quantitative trait locus, genome-wide association study and transcriptome analysis. Notulae Botanicae Horti Agrobotanici Cluj-Napoca 49, 12525. https://doi.org/10.15835/nbha49412525 (2021).
Rodríguez, V. M. et al. Genetic regulation of cold-induced albinism in the maize inbred line A661. J. Exp. Bot. 64, 3657–3667 (2013).
Greaves, J. A. Improving suboptimal temperature tolerance in maize—the search for variation. J. Exp. Bot. 47, 307–323 (1996).
Leipner, J., Fracheboud, Y. & Stamp, P. Effect of growing season on the photosynthetic apparatus and leaf antioxidative defenses in two maize genotypes of different chilling tolerance. Environ. Exp. Bot. 42, 129–139 (1999).
Fracheboud, Y., Haldimann, P., Leipner, J. & Stamp, P. Chlorophyll fluorescence as a selection tool for cold tolerance of photosynthesis in maize (Zea mays L.). J. Exp. Bot. 50, 1533–1540 (1999).
Goddard, M. E., Wray, N. R., Verbyla, K. & Visscher, P. M. Estimating effects and making predictions from genome-wide marker data. Stat. Sci. 24, 517–529 (2009).
Waqas, M. A. et al. Thermal stresses in maize: Effects and management strategies. Plants 10, 293 (2021).
Li, P. et al. Transcriptomic profiling of the maize (Zea mays L.) leaf response to abiotic stresses at the seedling stage. Front. Plant Sci. 8, 290. https://doi.org/10.3389/fpls.2017.00290 (2017).
Burnett, A. C. & Kromdijk, J. Can we improve the chilling tolerance of maize photosynthesis through breeding?. J. Exp. Bot. 73, 3138–3156. https://doi.org/10.1093/jxb/erac045 (2022).
Yi, Q., Alvarez-Iglesias, L., Malvar, R. A., Romay, M. C. & Revilla, P. A worldwide maize panel revealed new genetic variation for cold tolerance. Theor. Appl. Genet. 134, 1083–1094 (2021).
Revilla, P., Malvar, R. A., Cartea, M. E., Butrón, A. & Ordás, A. Inheritance of cold tolerance at emergence and during early season growth in maize. Crop Sci. 40, 1579–1585 (2000).
Zaidi, P. H. et al. Morpho-physiological traits associated with cold stress tolerance in tropical maize (Zea mays L.). Maydica 55, 201–208 (2010).
Revilla, P. et al. Cold tolerance in two large maize inbred panels adapted to European climates. Crop Sci. 54, 1981–1991 (2014).
Strigens, A. et al. Association mapping for chilling tolerance in elite flint and dent maize inbred lines evaluated in growth chamber and field experiments. Plant Cell Environ. 36, 1871–1887 (2013).
Giauffret, C., Lothrop, J., Dorvillez, D., Gouesnard, B. & Derieux, M. Genotype× environment interactions in maize hybrids from temperate or highland tropical origin. Crop Sci. 40, 1004–1012 (2000).
Frascaroli, E. & Landi, P. Divergent selection in a maize population for germination at low temperature in controlled environment: Study of the direct response, of the trait inheritance and of correlated responses in the field. Theor. Appl. Genet. 126, 733–746 (2013).
Revilla, P., Butrón, A., Cartea, M., Malvar, R. & Ordás, A. Breeding for cold tolerance. In Abiotic Stresses. Plant Resistance Through Breeding and Molecular Approaches (eds Ashraf, M. & Harris, P. J. C.) 301–398 (CRC Press, Boca Raton, FL, 2005).
Sobkowiak, A. et al. Genome-wide transcriptomic analysis of response to low temperature reveals candidate genes determining divergent cold-sensitivity of maize inbred lines. Plant Mol. Bio. 85, 317–331 (2014).
Sobkowiak, A. et al. Molecular foundations of chilling-tolerance of modern maize. BMC Genom. 17, 1–22 (2016).
Leipner, J., Jompuk, C., Camp, K. H., Stamp, P. & Fracheboud, Y. QTL studies reveal little relevance of chilling-related seedling traits for yield in maize. Theor. Appl. Genet. 116, 555–562 (2008).
Fracheboud, Y., Ribaut, J. M., Vargas, M., Messmer, R. & Stamp, P. Identification of quantitative trait loci for cold-tolerance of photosynthesis in maize (Zea mays L). J. Exp. Bot. 53, 1967–1977 (2002).
Fracheboud, Y., Jompuk, C., Ribaut, J. M., Stamp, P. & Leipner, J. Genetic analysis of cold-tolerance of photosynthesis in maize. Plant Mol. Bio. 56, 241–253 (2004).
Jompuk, C., Fracheboud, Y., Stamp, P. & Leipner, J. Mapping of quantitative trait loci associated with chilling tolerance in maize (Zea mays L.) seedlings grown under field conditions. J. Exp. Bot. 56, 1153–1163 (2005).
Hund, A., Fracheboud, Y., Soldati, A., Frascaroli, E., Salvi, S. & Stamp, P. QTL controlling root and shoot traits of maize seedlings under cold stress. Theor. Appl. Genet. 109, 618–629 (2004).
Guerra-Peraza, O. et al. Temperature at night affects the genetic control of acclimation to cold in maize seedlings. Maydica 56, 367–377 (2012).
Presterl, T. et al. Quantitative trait loci for early plant vigour of maize grown in chilly environments. Theor. Appl. Genet. 114, 1059–1070 (2007).
Rodríguez, V. M., Butrón, A., Rady, M. O., Soengas, P. & Revilla, P. Identification of quantitative trait loci involved in the response to cold stress in maize (Zea mays L.). Mol. Breed. 33, 363–371 (2014).
Shi, Y. et al. Genetic dissection of seed vigour traits in maize (Zea mays L.) under low-temperature conditions. J. Genet. 95, 1017–1022 (2016).
Allam, M., Revilla, P., Djemel, A., Tracy, W. F. & Ordás, B. Identification of QTLs involved in cold tolerance in sweet× field corn. Euphytica 208, 353–365 (2016).
Zhou, X., Muhammad, I., Lan, H. & Xia, C. Recent advances in the analysis of cold tolerance in maize. Front. Plant Sci. 13, 866034. https://doi.org/10.3389/fpls.2022.866034 (2022).
Jończyk, M., Sobkowiak, A., Trzcinska-Danielewicz, J. & Sowiński, P. Chromatin-level differences elucidate potential determinants of contrasting levels of cold sensitivity in maize lines. Plant Mol. Biol. Rep. 39, 335–350 (2021).
Jończyk, M. et al. Global analysis of gene expression in maize leaves treated with low temperature. II. Combined effect of severe cold (8 C) and circadian rhythm. Plant Mol. Bio. 95, 279–302 (2017).
Revilla, P. et al. Association mapping for cold tolerance in two large maize inbred panels. BMC Plant Biol. 16, 1–10 (2016).
Huang, J. et al. Genome-w ide association analysis of ten chilling tolerance indices at the germination and seedling stages in maize. J. Integr. Plant Biol. 55, 735–744 (2013).
Hu, G. et al. Genome-wide association study identified multiple genetic loci on chilling resistance during germination in maize. Sci. Rep. 7, 1–11 (2017).
Zhang, H. et al. Identification of candidate tolerance genes to low-temperature during maize germination by GWAS and RNA-seq approaches. BMC Plant Biol. 20, 1–17 (2020).
Yi, Q., Malvar, R. A., Alvarez-Iglesias, L., Ordas, B. & Revilla, P. Dissecting the genetics of cold tolerance in a multiparental maize population. Theor. Appl. Genet. 133, 503–516. https://doi.org/10.1007/s00122-019-03482-2 (2020).
Zhao, X. et al. Understanding and comprehensive evaluation of cold resistance in the seedlings of multiple maize genotypes. Plants (Basel) 11, 1881. https://doi.org/10.3390/plants11141881 (2022).
Sowiński, P. et al. Maize response to low temperatures at the gene expression level: A critical survey of transcriptomic studies. Front. Plant Sci. 11, 576941. https://doi.org/10.3389/fpls.2020.576941 (2020).
Farooq, M., Aziz, T., Wahid, A., Lee, D. J. & Siddique, K. H. Chilling tolerance in maize: Agronomic and physiological approaches. Crop Pasture Sci. 60, 501–516 (2009).
Goering, R. N. Uncovering Candidate Cold Tolerance Genes in Maize (Zea mays). Departmental Honors Projects. 54. https://digitalcommons.hamline.edu/dhp/54 (2017).
Ben-Haj-Salah, H. & Tardieu, F. Temperature affects expansion rate of maize leaves without change in spatial distribution of cell length (analysis of the coordination between cell division and cell expansion). Plant Physiol. 109, 861–870 (1995).
Grzybowski, M. et al. Increased photosensitivity at early growth as a possible mechanism of maize adaptation to cold springs. J. Exp. Bot. 70, 2887–2904 (2019).
Fryer, M. J., Oxborough, K., Martin, B., Ort, D. R. & Baker, N. R. Factors associated with depression of photosynthetic quantum efficiency in maize at low growth temperature. Plant Physiol. 108, 761–767 (1995).
Janowiak, F. & Markowski, A. Changes in leaf water relations and injuries in maize seedlings induced by different chilling conditions. J. Agron. Crop Sci. 172, 19–28 (1994).
Wijewardana, C., Hock, M., Henry, B. & Reddy, K. R. Screening corn hybrids for cold tolerance using morphological traits for early-season seeding. Crop Sci. 55, 851–867 (2015).
Zeng, R. et al. Natural variation in a type-A response regulator confers maize chilling tolerance. Nat. Commun. 12, 4713. https://doi.org/10.1038/s41467-021-25001-y (2021).
Thakur, P., Kumar, S., Malik, J. A., Berger, J. D. & Nayyar, H. Cold stress effects on reproductive development in grain crops: An overview. Environ. Exp. Bot. 67, 429–443 (2010).
Bechoux, N., Bernier, G. & Lejeune, P. Environmental effects on the early stages of tassel morphogenesis in maize (Zea mays L.). Plant Cell Environ. 23, 91–98 (2000).
Rymen, B. et al. Cold nights impair leaf growth and cell cycle progression in maize through transcriptional changes of cell cycle genes. Plant Physiol. 143, 1429–1438 (2007).
Westgate, M. E. & Boyer, J. S. Osmotic adjustment and the inhibition of leaf, root, stem and silk growth at low water potentials in maize. Planta 164, 540–549 (1985).
Enders, T. A. et al. Classifying cold-stress responses of inbred maize seedlings using RGB imaging. Plant Direct 3, e00104. https://doi.org/10.1002/pld3.104 (2019).
Liu, F. et al. Molecular evolution and genetic variation of G2-like transcription factor genes in maize. PLoS ONE 11, e016763. https://doi.org/10.1371/journal.pone.0161763 (2016).
Leipner, J., Stamp, P. & Fracheboud, Y. Artificially increased ascorbate content affects zeaxanthin formation but not thermal energy dissipation or degradation of antioxidants during cold-induced photooxidative stress in maize leaves. Planta 210, 964–969 (2000).
Rodríguez, V. M., Butrón, A., Sandoya, G., Ordás, A. & Revilla, P. Combining maize base germplasm for cold tolerance breeding. Crop Sci. 47, 1467–1474 (2007).
Vinayan, M. T. et al. Genotype-by-environment interaction effects under heat stress in tropical maize. Agronomy https://doi.org/10.3390/agronomy10121998 (2020).
Hindu, V. et al. Identification and validation of genomic regions influencing kernel zinc and iron in maize. Theor. Appl. Genet. 131, 1443–1457. https://doi.org/10.1007/s00122-018-3089-3 (2018).
Zaidi, P. H. et al. Genomic regions associated with root traits under drought stress in tropical maize (Zea mays L.). PLoS ONE 11, e0164340. https://doi.org/10.1371/journal.pone.0164340 (2016).
Qi, W., Lu, L., Huang, S. & Song, R. Maize Dek44 encodes mitochondrial ribosomal protein L9 and is required for seed development. Plant Physiol. 180, 2106–2119. https://doi.org/10.1104/pp.19.00546 (2019).
Zhang, Y. et al. Genome-wide association study uncovers new genetic loci and candidate genes underlying seed chilling-germination in maize Genome-wide association study uncovers new genetic loci and candidate genes underlying seed chilling-germination in maize. Peer J. 9, e11707 (2021).
Kulik, A., Wawer, I., Krzywińska, E., Bucholc, M. & Dobrowolska, G. SnRK2 protein kinases—key regulators of plant response to abiotic stresses. OMICS 15, 859–872 (2011).
Huai, J. et al. Cloning and characterization of the SnRK2 gene family from Zea mays. Plant Cell Rep. 27, 1861–1868 (2008).
Zou, H. et al. Cloning and characterization of maize ZmSPK1, a homologue to nonfermenting1-related protein kinase2. Afr. J. Biotech. 5, 490–496 (2006).
Buckler, E. S. et al. The genetic architecture of maize flowering time. Science 325, 714–718 (2009).
Neuffer, M., Coe, E. & Wessler, S. Mutants of Maize (Cold Spring Harbor Laboratory Press, Cold Spring Harbor, 1997).
Colasanti, J., Yuan, Z. & Sundaresan, V. The indeterminate gene encodes a zinc finger protein and regulates a leaf-generated signal required for the transition to flowering in maize. Cell 93, 593–603 (1998).
Muszynski, M. G. et al. Delayed flowering1 encodes a basic leucine zipper protein that mediates floral inductive signals at the shoot apex in maize. Plant Physiol. 142, 1523–1536 (2006).
Ducrocq, S. et al. Fine mapping and haplotype structure analysis of a major flowering time quantitative trait locus on maize chromosome 10. Genetics 183, 1555–1563 (2009).
Salvi, S. et al. Conserved noncoding genomic sequences associated with a flowering-time quantitative trait locus in maize. Proc. Natl. Acad. Sci. U. S. A. 104, 11376–11381 (2007).
Shen, G. et al. Plant GDSL esterases/lipases: Evolutionary, physiological and molecular functions in plant development. Plants 11, 468 (2022).
An, X. et al. ZmMs30 encoding a novel GDSL lipase is essential for male fertility and valuable for hybrid breeding in maize. Mol. Plant. 12, 343–359 (2019).
Huo, Y. et al. IRREGULAR POLLEN EXINE2 encodes a GDSL lipase essential for male fertility in maize. Plant Physiol. 184, 1438–1454 (2020).
Tsugama, D., Fujino, K., Liu, S. & Takano, T. A GDSL-type esterase/lipase gene, GELP77, is necessary for pollen dissociation and fertility in Arabidopsis. Biochem. Biophys. Res. Commun. 526, 1036–1041 (2020).
Xu, M. Y. et al. Stress-induced early flowering is mediated by miR169 in Arabidopsis thaliana. J. Exp. Bot. 65, 89–101 (2014).
Bomblies, K. & Doebley, J. F. Pleiotropic effects of the duplicate maize FLORICAULA/LEAFY genes zfl1 and zfl2 on traits under selection during maize domestication. Genetics 172, 519–531 (2006).
Yang, J. et al. Molecular basis for TPR domain-mediated regulation of protein phosphatase 5. EMBO J 24, 1–10 (2005).
Sharma, M. & Pandey, G. K. Expansion and function of repeat domain proteins during stress and development in plants. Front Plant Sci. 6, 1218. https://doi.org/10.3389/fpls.2015.01218 (2016).
Chardon, F. et al. Genetic architecture of flowering time in maize as inferred from quantitative trait loci meta-analysis and synteny conservation with the rice genome. Genetic 168, 2169–2185. https://doi.org/10.1534/genetics.104.032375 (2004).
Cassani, E. et al. Characterization of the first dominant dwarf maize mutant carrying a single amino acid insertion in the VHYNP domain of the dwarf8 gene. Mol. Breed. 24, 375–385 (2009).
Galbiati, M. et al. Identification and analysis of maize mutants defining six new genes affecting plant stature. Maydica 47, 169–180 (2002).
Peiffer, J. A. et al. The genetic architecture of maize height. Genetics 196, 1337–1356. https://doi.org/10.1534/genetics.113.159152 (2014).
Guo, Z. et al. Genome-wide analysis of the ATP-binding cassette (ABC) transporter family in Zea mays L. and its response to heavy metal stresses. Int. J. Mol. Sci. 23, 2109 (2022).
Han, G., Qiao, Z., Li, Y., Wang, C. & Wang, B. The roles of CCCH zinc-finger proteins in plant abiotic stress tolerance. Int. J. Mol. Sci. 22, 8327. https://doi.org/10.3390/ijms22158327 (2021).
Wang, C.-T. et al. Maize WRKY transcription factor ZmWRKY106 confers drought and heat tolerance in transgenic plants. Int. J. Mol. Sci. 19, 3046. https://doi.org/10.3390/ijms19103046 (2018).
Vinayan, M. T. et al. Genome wide association study and genomic prediction for stover quality traits in tropical maize (Zea mays L.). Sci. Rep. 11, 1–14 (2021).
Elshire, R. J. et al. A robust, simple genotyping-by-sequencing (GBS) approach for high diversity species. PLoS ONE 6, e19379. https://doi.org/10.1371/journal.pone.0019379 (2011).
Suwarno, W. B., Pixley, K. V., Palacios-Rojas, N., Kaeppler, S. M. & Babu, R. Formation of heterotic groups and understanding genetic effects in a provitamin A biofortified maize breeding program. Crop Sci. 54, 14–24 (2014).
Alvarado, G., et al. META-R (Multi Environment Trial Analysis with R for Windows.) Version 6.0 http://hdl.handle.net/11529/10201 International Maize and Wheat Improvement Center (2016).
Eberhart, S. A. & Russell, W. A. Stability parameters for comparing varieties. Crop Sci. 6, 36–40 (1966).
Pacheco, Á., Vargas, M., Alvarado, G., Rodríguez, F., Crossa, J. & Burgueño, J. GEA-R (Genotype x Environment Analysis with R for Windows) Version 4.1. https://hdl.handle.net/11529/10203, CIMMYT Research Data & Software Repository Network, V16 (2015).
Yu, J. M. et al. A unified mixed-model method for association mapping that accounts for multiple levels of relatedness. Nat. Genet. 38, 203–208 (2006).
Segura, V. et al. An efficient multi-locus mixed-model approach for genome-wide association studies in structured populations. Nat. Genet. 44, 825–830 (2012).
Liu, X., Huang, M., Fan, B., Buckler, E. S. & Zhang, Z. Iterative Usage of fixed and random effect models for powerful and efficient genome-wide association studies. PLoS Genet. 12, e1005767 (2016).
Zhang, Y.-W. et al. mrMLM v4.0.2: An R platform for multi-locus genome-wide association studies. Genom. Proteom. Bioinform. 18, 481–487. https://doi.org/10.1016/j.gpb.2020.06.006 (2020).
Huang, M., Liu, X., Zhou, Y., Summers, R. M. & Zhang, Z. BLINK: A package for the next level of genome-wide association studies with both individuals and markers in the millions. Gigascience https://doi.org/10.1093/gigascience/giy154 (2019).
Acknowledgements
The authors greatly acknowledge the funding support from CGIAR Research Program on Maize (CRP-MAIZE) and One CGIAR Initative on Accelerated Breeding. The authors have all the necessary permissions to collect and/or evaluate the experimental materials used in this study.
Funding
This study was carried out under CGIAR Research Program on MAIZE (CRP-MAIZE) and One CGIAR Initiative on Accelerated Breeding.
Author information
Authors and Affiliations
Contributions
M.T.V. and P.H.Z.: conceptualization, investigation, and supervision. M.T.V. and K.N.S.: formal analysis. S.K., P.Z., M.T.V., J.P.S., K.S., R.S., G.T., A.S., S.S. and B.S.: methodology. M.T.V., P.H.Z., JPS, and KNS: visualization and writing – review and editing. M.T.V.: writing – original draft. All authors contributed to the article and approved the submitted version.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Shikha, K., Madhumal Thayil, V., Shahi, J.P. et al. Genomic-regions associated with cold stress tolerance in Asia-adapted tropical maize germplasm. Sci Rep 13, 6297 (2023). https://doi.org/10.1038/s41598-023-33250-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-023-33250-8
- Springer Nature Limited