Abstract
To date there are thirteen species validly assigned to the genus Anaerococcus. Most of the species in this genus are anaerobic and of human origin. Anaerococcus urinimassiliensis sp. nov., strain Marseille-P2143T is member of family Peptoniphilaceae, which was isolated from the urine of a 17-year-old boy affected by autoimmune hepatitis and membranoproliferative glomerulonephritis using the culturomic approach. In the current study, a taxono-genomics method was employed to describe this new species. The strain Marseille-P2143T was gram positive cocci with translucent colonies on blood agar. Its genome was 2,189,509 bp long with a 33.5 mol% G + C content and exhibited 98.48% 16S rRNA similarity with Anaerococcus provencensis strain 9,402,080. When Anaerococcus urinomassiliensis strain Marseill-P2143T is compared with closely related species, the values ranged from 71.23% with A. hydrogenalis strain DSM 7454T (NZ_ABXA01000052.1) to 90.64% with A. provencensis strain 9402080T (NZ_HG003688.1). This strain has implemented the repertoire of known bacteria of the human urinary tract.
Similar content being viewed by others
Introduction
The genus Anaerococcus belonging to the phylum Firmicutes, was first described in 20011. Members of this bacterial genus are mainly anaerobic gram-positive cocci2. They are mostly encountered in human vagina, but can also be detected in nostrils or skin3. Anaerococcus spp. were involved in human infections and were isolated from different sites of human body such as peritoneal, ovarian and cervical abscesses, an arthritic knee, bacteremia, foot ulcers, a sternal wound and vaginoses4,5,6. Actually, the genus Anaerococcus contains 13 species validly described with standing in nomenclature7. The culturomic concept has recently been developed in our laboratory as an alternative method to expand the human gut repertoire through the multiplication of culture conditions with a rapid identification method by matrix-assisted laser desorption/ionization time-of-flight mass spectrometry (MALDI-TOF MS)8,9,10,11. Isolation and identification of microorganisms by culturomic can be used for further studies12 as well as for their diagnosis and/or therapeutic potential. The systematic description of new bacterial species recovered from patients may contribute to the description of emerging infections, but can also led to other discoveries. For example, the strain Eubacterium limosum isolated by culture in the gut sample has been used to test biotransformation’s of specific pollutants, methoxychlor and dichlorodiphenyltrichloroethane (DDT) insecticides13. The studies conducted on probiotic Escherichia coli strain Nissle 1917, show that a visceral analgesic can be produced by bacterial strain, which could be the basis of the development of new visceral pain therapies14. Currently clinical trials using bacterial cocktails are being used to restore dybiosis by fecal transplantation.
We report here, through a taxono-genomics strategy15, the description of Anaerococcus urinimassiliensis strain Marseille-P2143 (= CSUR P2143 = DSM 103,473), a new bacterium isolated from the urine of a young boy affected by autoimmune hepatitis associated with membranoproliferative glomerulonephritis, and classified into Peptoniphilaceae family. This new bacterial species was shortly described in a new species announcement in 201616.
Material and methods
Sample collection and ethical considerations
In 2015, we isolated in the urine of a 17-year-old boy with autoimmune hepatitis and membranoproliferative glomerulonephritis, a bacterial strain belonging to the genus Anaerococcus that could not be identified at the species level. A signed and informed consent was collected from the patient and parents, and the study obtained approval from ethics committee of the Institut Fédératif de Recherche IFR48 under number 09-022. All the methods were carried out in accordance with relevant guidelines and regulations conformed to the Declaration of Helsinki.
Strain isolation and identification by MALDI-TOF MS
The initial growth was obtained after 10 days of incubation in an anaerobic blood culture vial (Becton Dickinson, Le Pont-de-Claix, France) supplemented with 5 mL of 0.2 μm filtered rumen fluid. A pure culture of strain Marseille-P2143 was then obtained after 48 h of incubation at 37 °C on 5% sheep blood–Columbia agar medium (bioMérieux, Marcy l'Etoile, France) in anaerobic atmosphere generated using the GENbag Anaer system (bioMérieux) as previously described16. Strain Marseille-P2143T was not identified by Matrix Assisted Laser Desorption Ionization-Time of Flight Mass Spectrometry (MALDI-TOF MS), after several attempts essayed as described elsewhere17. The screening was performed on a Microflex LT spectrometer (Bruker, Daltonics, Bremen, Germany) as previously reported18. The reference spectrum obtained (Fig. 1) was imported and analyzed using the Biotyper software (version 3.0)19 against the Bruker database, which was continually incremented with local URMS database (https://www.mediterranee-infection.com/urms-data-base/).
Strain identification and phylogenetic tree
In order to classify this bacterium, the 16S rRNA gene was amplified using the primer pair fD1 and rP2 (Eurogentec, Angers, France) and sequenced using the Big Dye Terminator v1.1 Cycle Sequencing Kit and 3500xLGenetic Analyzer capillary sequencer (Thermofisher, Saint-Aubin, France) as previously described12. The 16S rRNA nucleotide sequence was assembled and corrected using CodonCode Aligner software (http://www.codoncode.com). For phylogenetic analysis, sequences of the phylogenetically closest species were obtained after performing a BLASTn search within the NCBI Blastn 16S rRNA Sequence Reference Base for closest related species to calculate sequence similarities of the 16S rRNA genes (refseq_rna) (https://blast.ncbi.nlm.nih.gov/Blast.cgi?PROGRAM=blastn&PAGE_TYPE=BlastSearch&LINK_LOC=blasthome). “The All‐Species Living Tree" Project of Silva20. The alignment was performed using MUSCLE21. The evolutionary history was inferred using the Maximum Likelihood method based on the Tamura-Nei model22. The tree with the highest log likelihood (-5398.79) is shown. The percentage of trees in which the associated taxa clustered together is shown next to the branches. Initial tree(s) for the heuristic search were automatically obtained by applying Neighbor-Join and BioNJ algorithms to a matrix of pairwise distances estimated using the Maximum Composite Likelihood (MCL) approach, and then selecting the topology with superior log likelihood value. The tree is drawn to scale, with branch lengths measured in the number of substitutions per site. The analysis involved 18 nucleotide sequences. The codon positions included were 1st + 2nd + 3rd + Noncoding. All positions containing gaps and missing data were eliminated. There were a total of 1184 positions in the final dataset. Evolutionary analyses were conducted in MEGA software (version X) (https://www.megasoftware.net/)23.
Phenotypic characteristics and biochemical features
The optimum growth condition of the strain was determined by culturing the strain under different temperatures, atmospheres, PH and salinity. The strains were cultured and incubated under aerobic, anaerobic (GENbag anaer, bioMérieux Limited, France) and microaerophilic (GENbag Microaer, bioMérieux Limited, France) conditions on Columbia agar enriched with 5% sheep blood (bioMérieux Limited, France) at the following temperatures: 25, 28, 37, 45, and 55 °C. The pH conditions used were 5.5, 6, 6.5, 7, 7.5, 8, 8.5, 9, 9.5 and the salinity conditions used were the following: 0%, 5%,10%, 25%, 50%, 100%. The phenotypic characteristics of the strain such as Gram staining, motility, oxidase and catalase activities were determined using standard microbiological methods as previously described23. These phenotypic and biochemical characteristics were tested for strain Marseille-P2143 incubated at 37 °C for 48 h. The use of carbon sources was assayed with API 50 CH strips. The API 50 CH strips were interpreted after incubation at 37 °C for 24 h. Antibiotic susceptibility was determined using disc diffusion plate method according to the instructions of the CA-SFM / EUCAST 2020 (https://www.eucast.org/clinical_breakpoints_and_dosing/eucast_setting_breakpoints/), in reference to the EUCAST disk diffusion method for susceptibility testing of the Bacteroides fragilis group isolates24. Antibiotic discs used were the following: erythromycin (15 μg/ml), penicillin G (10 UI), doxycycline (30 μg/ml), rifampicin (30 μg/ml), vancomycin (30 μg/ml), clindamycin (15 μg/ml), fosfomycin (50 μg/ml), amoxicillin (25 μg/ml), colistin (15 μg/ml), gentamycin (500 μg/ml), amoxicillin-clavulanic acid (30 μg/ml), ceftriaxone (30 μg/ml), colistin (50 μg/ml), trimethoprim-sulfamethoxazole (25 μg/ml), oxacillin (5 μg/ml), imipenem (10 μg/ml), tobramycin (10 μg/ml), and metronidazole (4 μg/ml). For fatty acids analysis, the bacterial strains cultivated on cos medium after 48 h in aerobic condition, were collected in triplicates in three tubes with approximately the same amount of biomass and were then weighed. We obtained an average of 130 mg of biomass per tube and cellular fatty acid methyl ester (FAME) analysis was performed by Gas Chromatography/Mass Spectrometry (GC/MS) as described by Sasser et al.25. GC/MS analyses were carried out as previously described26. Spectral database search was performed using MS Search 2.0 operated with the Standard Reference Database 1A (NIST, Gaithersburg, USA) and the FAMEs mass spectral database (Wiley, Chichester, UK).
Genome sequencing and assembly
Genomic DNA was extracted using the EZ1 biorobot with the EZ1 DNA tissue kit (Qiagen, Hilden, Germany) and then sequenced on a MiSeq sequencer (Illumina Inc, San Diego, CA, USA) with the Nextera Mate Pair sample prep kit and Nextera XT Paired End (Illumina), as previously described27. The assembly was performed using a pipeline containing several softwares (Velvet28, SPAdes29 and SOAP Denovo30) and trimmed (MiSeq and Trimmomatic31 softwares) or untrimmed data (only MiSeq software). GapCloser software32 was used to reduce assembly gaps. Scaffolds < 800 base pairs (bp) and scaffolds with a depth value lower than 25% of the mean depth were removed. The best assembly was selected using different criteria (number of scaffolds, N50, number of N). The degree of genomic similarity of strain Marseille-P2143T (NZ_FQRX00000000.1) with closely related species Anaerococcus hydrogenalis DSM 7454T(NZ_ABXA01000052.1), Anaerococcus jeddahensis strain SB3T(NZ_CWHU01000001.1), Anaerococcus tetradius ATCC 35,098T (NZ_GG666394.1), Anaerococcus octavius strain NCTC9810T(UFTA01000001.1), Anaerococcus pacaensis 9403502T(NZ_HG326663.1), Anaerococcus prevotii strain NCTC11806T (UFSY01000001.1), Anaerococcus provencensis strain 9402080T(NZ_HG003688.1), Anaerococcus rubiinfantis MT16T(FAVH01000001.1), Anaerococcus senegalensis JC48T (NZ_HE578907.1), was estimated using the OrthoANI software33. These genomes were then aligned with scapper (https://github.com/tseemann/scapper). These aligned genomes were eventually used to build the phylogenetic tree using the Maximum likelihood method from MEGA X software34.
Genome annotation and analysis
The prediction was performed using prodigal in the open reading frame (ORF)35 with default parameters. Planned ORFs covering a sequencing gap region (containing N) were excluded. The bacterial proteome was predicted with BLASTP (E-value of 1e03, coverage of 0.7 and identity percentage of 30) against the database of orthologic group clusters (COG). If no matches were found, we searched the nr database36 using BLASTP with an E-value of 1e03, a coverage of 0.7 and an identity percentage of 30. An E-value of 1e05 was used only if the sequence length was less than 80 amino acids (aa). The domains maintained by the PFAM (PFAM-A and PFAM-B domains) were searched on each protein using the hmmscan analysis tool. RNAmmer37 and the tRNAScanSE tools38 were used to find rRNA and tRNA genes. When BLASTP E-value was lower than 1e-03 for alignment length greater than 80 amino acids, ORFans are identified. However, when alignment lengths smaller than 80 amino acids were obtained, an E-value of 1e-05 was used. Artemis39 was used for data management and visualization of genomic characteristics. The in-house MAGI software was used to analyze the average level of similarity of nucleotide sequences at the genome level. It calculated the average genomic identity of gene sequences (AGIOS) among the genomes compared15. This software combines Proteinortho software40 to detect orthologic proteins in pairwise genomic comparisons. Then, the corresponding genes were recovered and the average percentage identity of nucleotide sequences among the orthological ORFs was determined using the Needleman—Wunsch global alignment algorithm. We also used the Genome-to-Genome Distance Calculator Web service to calculate DNA: Digital DNA hybridization estimates (dDDH) with confidence intervals according to recommended parameters (Formula 2, BLAST)41.
The degree of genomic similarity of Marseille-P2143 with closely related species was estimated using the OrthoANI software42.
Ethical approval
All the methods were carried out in accordance with relevant guidelines and regulations conformed to Declaration of Helsinki.
Results and discussion
Identification of Strain Marseille-P2143
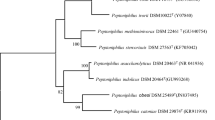
The mass spectrum of strain Marseille-P2143 was not present in the MALDI-TOF MS Bruker database. Thus, we were not able to identify this strain using this instrument. However, the 16S rRNA gene sequencing analysis indicated that it exhibited 98.48% sequence similarity with Anaerococcus provencensis strain 9402080T (Genbank accession number NR_133036.1), the phylogenetically closest species after blast in the NCBI database (Fig. 2). We consequently proposed to classify strain Marseille-P2143 Type strain as a new species within the genus Anaerococcus43.
Phylogenetic tree highlighting the position of Anaerococcus urinimassiliensis Marseille-P2143T sp. nov., with regard to other closely related species. Genbank accession numbers of 16S rRNA are indicated in parentheses. Sequences were aligned using MUSCLE with default parameters, phylogenetic inference were obtained using the neighbor joining (A) and Maximum likelihood (B) method and the MEGA X software. Bootstrap values obtained by repeating the analysis 50 times to generate a majority consensus tree are indicated at the nodes. The scale bar indicates a 2% and 5% nucleotide sequence divergence.
Phenotypic characteristics and biochemical features
The optimal growth of strain Marseille-P2143 was obtained after 5 day of culture at 37 °C under anaerobic conditions (anaeroGEN, Oxoid Ltd, Dardilly, France). Agar-grown colonies were small with a mean diameter of 50 μm and were translucent white. Bacterial cells were Gram-positive cocci ranging in diameter from 0.5 to 0.7 µm (Fig. 3). Catalase and oxidase activities were not observed. A brief description and characteristics of strain Marseille-P2143 are summarized in Table 1. Using API ZYM strip, positive reactions were observed for alkaline phosphatase, esterase, esterase lipase, leucine arylamidase, cystine arylamidase, trypsin, naphthol-AS-BI-phosphohydrolase, α-galactosidase and β-galactosidase. But negative reactions were noted with lipase, valine arylamidase, α-chymotrypsin, acid phosphatase, and β-glucuronidase. Using API 50 CH strip (bioMérieux), strain Marseille-P2143 was able to metabolize d-galactose, d-glucose, d-fructose, N-acetyl-glucosamine, salicin, d-maltose, d-lactose, and d-saccharose. Negative reactions were obtained for glycerol, d-mannose, methyl α-d-glucopyranoside, d-turanose, d-tagatose, erythritol, d-adonitol, d-mannose, mannitol, d-sorbitol, amygdalin, arbutin, esculin, d-melibiose, d-trehalose, inulin, d-melezitose, d-raffinose, starch, glycogen, d-arabitol, l-arabitol and Potassium 5-ketogluconate. When strain Marseille-P2143 was compared to Anaerococcus provencensis strain 9402080T the closest species with a validly published name, it differed in catalase activity (Table 2). The Antimicrobial susceptibility test according to the EUCAST showed that strain Marseille-P2143 was susceptible to Penicillin, Oxacillin, Amoxicillin, Amoxicillin-Clavulanic acid, Imipenem, Rifampin, Vancomycin, Clindamycin, Fosfomycin, and Tobramycin but resistant to Ceftriaxone, Erythromycin, Doxycycline, Gentamicin, Colistin and Trimethoprim-Sulfamethoxazole. Hexadecanoic acid was the most abundant fatty acid (57%). Unsaturated fatty acids 9-Octadecenoic acid and 9,12-Octadecadienoic were also detected with significant amounts. Tetradecanoic acid and Octadecanoic acid were minor fatty acids were quantified. By comparing strain Marseille-P2143 with related close species (Table 3), we obtain mainly a similar profile between species with C16:0 as the abundant fatty acid (> 50%).
Genome properties
The genome of Marseille-P2143 was 2,189,509 bp long with 33.5 mol% G + C content (Fig. 4). It was composed of 10 scaffolds (11 contigs). Of the 2077 predicted genes, 1952 were protein-coding genes and 61 were RNAs (4 genes are 5S rRNA, 3 genes are 16S rRNA, 5 genes are 23S rRNA, 46 genes are tRNA genes) 64 were pseudo genes.
Graphical circular map of the chromosome of Anaerococcus urinimassiliensis sp.nov. From outside to the center: genes on the forward strand colored by COG categories (only genes assigned to COG), genes on the reverse strand colored by COG categories (only gene assigned to COG), RNA genes (tRNAs green, rRNAs red), GC content and GC skew.
Genome comparison
The draft genome sequence of Anaerococcus urinimassiliensis (2,189,509 bp) was larger than that of A. prevotii strain NCTC11806T (UFSY01000001.1) (2,004,977 bp), A. pacaensis strain 9403502T(NZ_HG326663.1) (845,487 bp) and A. provencensis strain 9402080T(NZ_HG003688.1) (29,418). The G + C content of A. urinimassiliensis strain Marseille-P2143T (33.5 mol%) was smaller than those of A. provencensis strain 9402080T(NZ_HG003688.1) (29,418) (33.7%), A.pacaensis strain 9403502T(NZ_HG326663.1) (34.1%) and A. prevotii strain NCTC11806T (UFSY01000001.1) (35.6%). The gene content of A. urinimassiliensis strain Marseille-P2143 (2077genes) was smaller than those of A. provencensis strain 9402080T(NZ_HG003688.1) (2146) and A.pacaensis strain 9403502T(NZ_HG326663.1) (2223), but larger than those of A. prevotii strain NCTC11806T (UFSY01000001.1) (1894) respectively (Table 4). OrthoANI values among closely related species (Figs. 5) ranged from 69.52% between A. pacaensis strain 9403502T(NZ_HG326663.1) and A. rubiinfantis strain MT16T (FAVH01000001.1) to 92.77% between A. jeddahensis strain SB3T (NZ_CWHU01000001.1) and A. rubiinfantis strain MT16T (FAVH01000001.1). When Anaerococcus urinomassiliensis strain Marseill-P2143is compared with closely species, the values ranged from 71.23% with A. hydrogenalis strain DSM 7454T (NZ_ABXA01000052.1) to 90.64% with A. provencensis strain 9402080T (NZ_HG003688.1). The phylogenetic tree highlighting the position of the genome of strain Marseille-P2143 in relation to other closely related species with a validly published name is presented in (Fig. 6). Genes with putative function (by COGs) were 1796 for Anaerococcus urinimassiliensis strain Marseille-P2143T (81%), ,903 for A.provencensis strain 9402080T(NZ_HG003688.1) (82%), 1539 for A.prevotii strain NCTC11806T (UFSY01000001.1) (86) and 1867 for Anaerococcus pacaensis strain 9403502T(NZ_HG326663.1) (78%). Finally, 414, 426, 253 and 538 genes (19%,18%, 14% and 22%) were annotated as hypothetical proteins for Anaerococcus urinimassiliensis strain Marseille-P2143, A.provencensis strain 9402080T(NZ_HG003688.1), A.prevotii strain NCTC11806T (UFSY01000001.1) and A.pacaensis strain 9403502T(NZ_HG326663.1), respectively Table 5. Analysis of the Clusters of Orthologous Groups (COGs) categories shows that the mobile elements of the Marseille-P2143 genome appear to be more numerous than those of the genomes of A.provencensis strain 9402080T(NZ_HG003688.1), A.prevotii strain NCTC11806T (UFSY01000001.1), A.pacaensis strain 9403502T(NZ_HG326663.1) (82, 45, 16 and 48 in category [X], respectively).
Phylogenetic tree highlighting the position of Anaerococcus urinimassiliensis Marseille-P2143T sp. nov., with regard to other closely related species considering the genome. Genbank accession numbers of genome are indicated in parentheses. Sequences were aligned using scraper with default parameters, phylogenetic inference were obtained using the Maximum likelihood method and the MEGA X software. Bootstrap values obtained by repeating the analysis 50 times to generate a majority consensus tree are indicated at the nodes. The scale bar indicates a 1% nucleotide sequence divergence.
The 16S rRNA gene sequence obtained by sanger sequencing was identical to that obtained within the sequenced genome (Locus Taq : CJIAFGEE_00528, contic : FQRX01000011.1 , Per. Ident : 99.51% using Blast).
Conclusion
On the basis of unique phenotypic features, including the MALDI-TOF spectrum, a 16S rRNA sequence similarity lower than < 98.65% and, an OrthoANI and a DDH values lower than 95% and 70% respectively with the phylogenetically closest species with standing in nomenclature, we formally propose strain Marseille-P2143T as the type strain of Anaerococcus urinimassiliensis sp. nov., a new species within the genus Anaerococcus. Anaerococcus urinimassiliensis (u.ri.ni.mas.si.li.en’sis. N.L.adj.masc. urinimmassiliensis, composed of urini, from the latin urina, urine and massiliensis, from Massilia, the roman name of Marseille, France, where the strain Marseille-P2143 was first isolated. The colonies are thin, translucent and 50 µm in diameter. Cells are Gram-positive and anaerobic cocci. Cells have a diameter ranging from 0.5 to 0.7 µm. They do not produce catalase and oxidase, but exhibit alkaline phosphatase, leucine arylamidase, esterase lipase, α-galactosidase, naphthol-AS-BI-phosphohydrolase and β-galactosidase. d-glucose, d-galactose, d-maltose, d-lactose, d-fructose, N-acetyl-glucosamine, salicin, and sucrose are metabolized. The genome of strain Marseille-P2143 is 2,189,509 bp long with a 33.5 mol% G + C content. Its 16S rRNA gene sequence and whole-genome sequence are deposited in GenBank under accession numbers LN898272.1 and NZ_FQRX00000000.1, respectively. The type strain Marseille-P2143T (= CSUR P2143 = DSM 103,473) was isolated from the urine of a 17-year-old boy suffering from autoimmune hepatitis and membranoproliferative glomerulo-nephritis. This new species implements the repertoire of human urinary tract known bacteria44.
Description of Anaerococcus urinimassiliensis sp. nov
Anaerococcus urinimassiliensis (u.ri.ni.mas.si.li.en’sis. L. gen. masc. urini, of Urine and massiliensis referring to the Latin name of Marseille where strain Marseille-P2143 was cultivated). Cells are Gram-positive and non-motile, but they are negative for catalase and oxidase activities. They had a mean diameter of 0.6 µm. On blood agar after 48 h of incubation at 37 °C, colonies of strain Marseille-P2143 appear transluscent white. The optimum growth is observed at pH 7.5. Major cellular fatty acid was Hexadecanoic acid (57%), while unsaturated fatty acids such as 9-Octadecenoic acid and 9,12-Octadecadienoic were also detected with strain Marseille-P2143. The enzymes activities of alkaline phosphatase, esterase, esterase lipase, leucine arylamidase, cystine arylamidase, trypsin, naphthol-AS-BI-phosphohydrolase, α-galactosidase and β-galactosidase were positive using the API ZYM. Also, fructose, galactose, glucose, N-acetyl-glucosamine, salicin, maltose, lactose, and saccharose are fermented but not for mannose, methyl α-d-glucopyranoside, melibiose, turanose, erythritol, tagatose, adonitol, mannose, mannitol, arbutin, esculin, trehalose, sorbitol, inulin, melezitose and amygdalin. 16S rRNA and genome sequences of this new species are deposited in GenBank under accession numbers LN898272 and FQRX00000000, respectively. The genome size is 2.19 Mb with a G + C content at 33.47%. The type strain Marseille-P2143T (= CSUR P2143 = DSM 103,473). The Marseille-P2143 strain was isolated in the urine of a young boy suffering from autoimmune hepatitis membranoproliferative glomerulonephritis.
Deposit in culture collections
Strain Marseille-P2143T was deposited in the French culture collection center, Collection des Souches de l’Unité des Rickettsies (CSUR), under the number CSUR P2143. And in the DSM collection under the number 103473.
References
Ludwig, W., Schleifer, K. H. & Whitman, W. B. Revised road map to the phylum Firmicutes. In Bergey’s Manual of Systematic Bacteriology Vol. 2 (eds De Vos, P., Garrity, G. M., Jones, D. et al.) 1–13 (Springer, New York, 2009).
Ezaki, T. et al. Proposal of the genera Anaerococcus gen. nov., Peptoniphilus gen. nov. and Gallicola gen. nov. for members of the genus Peptostreptococcus. Int. J. Syst. Evol. Microbiol. 51, 1521–1528 (2001).
Song, Y., Liu, C. & Finegold, S. M. Peptoniphilus gorbachii sp. nov., Peptoniphilus olsenii sp. nov., and Anaerococcus murdochii sp. nov. isolated from clinical specimens of human origin. J. Clin. Microbiol. 45, 1746–1752 (2007).
Jain, S., Bui, V., Spencer, C. & Yee, L. Septic arthritis in a native joint due to Anaerococcus prevotii. J. Clin. Pathol. 61, 775–776 (2008).
La Scola, B., Fournier, P. E. & Raoult, D. Burden of emerging anaerobes in the MALDI-TOF and 16S rRNA gene sequencing era. Anaerobe 17, 106–112 (2011).
Pépin, J. et al. The complex vaginal flora of West African women with bacterial vaginosis. PLoS ONE 6, e25082 (2011).
Parte, A. C. LPSN-list of prokaryotic names with standing in nomenclature. Nucleic Acids Res. 42, D613–D616 (2014).
Lagier, J. C. et al. Microbial culturomics: paradigm shift in the human gut microbiome study. Clin. Microbiol. Infect. 18, 1185–1193 (2012).
Lagier, J. C. et al. The rebirth of culture in microbiology through the example of culturomics to study human gut microbiota. Clin. Microbiol. Rev. 28, 237–264 (2015).
Lagier, J. C. et al. Culture of previously uncultured members of the human gut microbiota by culturomics. Nat. Microbiol. 1, 16203 (2016).
Lagier, J. C. et al. Non-contiguous-finished genome sequence and description of Anaerococcus senegalensis sp. nov. Stand Genomic Sci. 6, 116–125 (2012).
Morel, A. S. et al. Complementarity between targeted real-time specific PCR and conventional broad-range 16S rDNA PCR in the syndrome-driven diagnosis of infectious diseases. Eur. J. Clin. Microbiol. Infect. Dis. 34, 561–570 (2015).
Yim, Y. et al. Reductive dechlorination of methoxychlor and DDT by human intestinal bacterium Eubacterium limosum under anaerobic conditions. Arch. Environ. Contam. Toxicol. 54, 406–411 (2008).
Pérez-Berezo, T. et al. Identification of an analgesic lipopeptide produced by the probiotic Escherichia coli strain Nissle 1917. Nat. Commun. 8(1), 1314 (2017).
Ramasamy, D. et al. A polyphasic strategy incorporating genomic data for the taxonomic description of novel bacterial species. Int. J. Syst. Evol. Microbiol. 64, 384–391 (2014).
Morand, A. et al. Anaerococcus urinomassiliensis sp. nov., isolated from a urine sample of a 17-year-old boy affected by autoimmune hepatitis and membranoproliferative glomerulonephritis. New Microbes New Infect. 6(13), 56–58 (2016).
Lo, C. I. et al. MALDI-TOF mass spectrometry: a powerful tool for clinical microbiology at Hôpital Principal de Dakar, Senegal (West Africa). PLoS ONE 10, e0145889 (2015).
Fall, B. et al. The ongoing revolution of MALDI-TOF mass spectrometry for microbiology reaches tropical Africa. Am. J. Trop. Med. Hyg. 92, 641–647 (2015).
Nagy, E., Becker, S., Kostrzewa, M., Barta, N. & Urbán, E. The value of MALDI-TOF MS for the identification of clinically relevant anaerobic bacteria in routine laboratories. J. Med. Microbiol. 61, 1393–1400 (2012).
Yilmaz, P. et al. The SILVA and “All-species Living Tree Project (LTP)” taxonomic frameworks. Nucleic Acids Res. 42, D643–D648 (2014).
Edgar, R. C. MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 32(5), 1792–1797 (2004).
Tamura, K. & Nei, M. Estimation of the number of nucleotide substitutions in the control region of mitochondrial DNA in humans and chimpanzees. Mol. Biol. Evol. 10(3), 512–526 (1993).
Jousimies-Somer, H. et al. Wadsworth-KTL Anaerobic Bacteriology Manual 6th edn. (Star Publishing, Belmont, CA, 2002).
Nagy, E. et al. Development of EUCAST disk diffusion method for susceptibility testing of the Bacteroides fragilis group isolates. Anaerobe 31, 65–71 (2015).
Sasser, M. Bacterial identification by gas chromatographic analysis of fatty acids methyl esters (GC-FAME) (MIDI, Technical Note, Newark, NY, 2006).
Dione, N. et al. Genome sequence and description of Anaerosalibacter massiliensis sp. nov. New Microbes New Infect. 10, 66–76 (2016).
Lo, C. I. et al. High-quality draft genome sequence and description of Haemophilus massiliensis sp. nov. Stand Genomic Sci. 11, 31 (2016).
Zerbino, D. R. & Birney, E. Velvet: algorithms for de novo short read assembly using de Bruijn graphs. Genome Res. 18, 821–829 (2008).
Bankevich, A. et al. SPAdes: a new genome assembly algorithm and its applications to single-cell sequencing. J. Comput. Biol. 19, 455–477 (2012).
Luo, R. et al. SOAPdenovo2: an empirically improved memory-efficient short-read de novo assembler. Gigascience. 1, 18 (2012).
Bolger, A. M., Lohse, M. & Usadel, B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30, 2114–2120 (2014).
Xu, G. C. et al. LR_Gapcloser: a tiling path-based gap closer that uses long reads to complete genome assembly. Gigascience. https://doi.org/10.1093/gigascience/giy157 (2019).
Lee, I., Ouk Kim, Y., Park, S. C. & Chun, J. OrthoANI: an improved algorithm and software for calculating average nucleotide identity. Int. J. Syst. Evol. Microbiol. 66, 1100–1103 (2016).
Kumar, S., Stecher, G., Li, M., Knyaz, C. & Tamura, K. MEGA X: molecular evolutionary genetics analysis across computing platforms. Mol. Biol. Evol. 35(6), 1547–1549 (2018).
Hyatt, D. et al. Prodigal: prokaryotic gene recognition and translation initiation site identification. BMC Bioinform. 11, 119 (2010).
Clark, K., Karsch-Mizrachi, I., Lipman, D. J., Ostell, J. & Sayers, E. W. GenBank. Nucleic Acids Res. 44, D67-72 (2016).
Lagesen, K. et al. RNAmmer: consistent and rapid annotation of ribosomal RNA genes. Nucleic Acids Res. 35, 3100–3108 (2007).
Lowe, T. M. & Chan, P. P. tRNAscan-SE on-line: integrating search and context for analysis of transfer RNA genes. Nucleic Acids Res. 44, W54-57 (2016).
Carver, T., Harris, S. R., Berriman, M., Parkhill, J. & McQuillan, J. A. Artemis: an integrated platform for visualization and analysis of high-throughput sequence-based experimental data. Bioinformatics 28, 464–469 (2012).
Lechner, M. et al. Proteinortho: detection of (co-)orthologs in large-scale analysis. BMC Bioinform. 12, 124 (2011).
Meier-Kolthoff, J. P., Göker, M., Spröer, C. & Klenk, H. P. When should a DDH experiment be mandatory in microbial taxonomy?. Arch. Microbiol. 195, 413–418 (2013).
Jain, C. et al. High throughput ANI analysis of 90K prokaryotic genomes reveals clear species boundaries. Nat. Commun. 9, 5114 (2018).
Kim, M. et al. Towards a taxonomic coherence between average nucleotide identity and 16S rRNA gene sequence similarity for species demarcation of prokaryotes. Int. J. Syst. Evol. Microbiol. 64(2), 346–351 (2014).
Morand, A. et al. Human bacterial repertoire of the urinary tract: a potential paradigm shift. J. Clin. Microbiol. 57, e00675-e718 (2019).
Acknowledgements
This work has benefited from support of the French State, managed by the “Agence Nationale pour la Recherche” including the “Programme d’Investissement d’Avenir” under the reference Méditerranée Infection 10-IAHU-03. We acknowledge Professor Florence Fenollar, Doctor Matthieu Million, Doctor Grégory Dubourd and Professor Chabrol for their significant help.
Author information
Authors and Affiliations
Contributions
D.R., M.T., J.C.L. and P.E.F. conceived and designed the experiments, M.T. and F.C. actively participated in the specimen collection, A.M., L.T., E.K.Y., I.I.N., C.I.L., F.C. and A.L. performed the experiments, A.M., L.T., A.L. and P.E.F. analysed the data, A.M., L.T., C.I.L. and P.E.F. wrote the manuscript. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Morand, A., Tall, M.L., Kuete Yimagou, E. et al. Anaerococcus urinimassiliensis sp. nov., a new bacterium isolated from human urine. Sci Rep 11, 2684 (2021). https://doi.org/10.1038/s41598-021-82420-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-021-82420-z
- Springer Nature Limited
This article is cited by
-
Human nasal microbiota shifts in healthy and chronic respiratory disease conditions
BMC Microbiology (2024)
-
Bioelectricity Production from Microbial Fuel Cell (MFC) Using Lysinibacillus xylanilyticus Strain nbpp1 as a Biocatalyst
Current Microbiology (2023)