Abstract
Environmental stresses greatly limit crop yield. With the increase in extreme weather events due to climate change and the constant pressure of diseases and pests, there is an urgent need to develop crop varieties that can tolerate multiple stresses. However, our knowledge of how plants broadly respond to stress is limited. Here, we explore the rice core stress response via meta-analysis of publicly available rice transcriptome data. Our results confirm that rice universally down-regulates photosynthesis in response to both abiotic and biotic stress. Rice also generally up-regulates hormone-responsive genes during stress response, most notably genes in the abscisic acid, jasmonic acid and salicylic acid pathways. We identified several promoter motifs that are likely involved in stress-responsive regulatory mechanisms in rice. With this work, we provide a list of candidate genes to study for improving rice stress tolerance in light of environmental stresses. This work also serves as a proof of concept to show that meta-analysis of diverse transcriptome data is a valid approach to develop robust hypotheses for how plants respond to stress.
Similar content being viewed by others
Introduction
Because plants are immobile, they must respond to and endure a wide variety of environmental and biotic stresses in the field. Both abiotic and biotic stresses cause major yield losses to crops1,2,3,4. It is therefore not surprising that many crop improvement programs focus on developing stress tolerant plant varieties5,6,7. Breeding tolerance for a single stress (e.g. drought, salinity, pathogen, etc.) or a single stress type (e.g. abiotic or biotic) may be risky because plants respond uniquely to different or simultaneous stresses, and increasing tolerance to one stress may be at the expense of tolerance to another4,8. With climate change, more extreme weather events are occurring, increasing the likelihood that plants experience multiple stresses in the field, including additional pressure from plant diseases9. There is, therefore, a need to understand the similarities and differences among stress response pathways to best optimize targeted crop improvement.
Plants respond to stress in a variety of ways. Common plant responses to avoid or tolerate abiotic stresses include stomatal closure, reduced photosynthesis, increased reactive oxygen scavenging activity, reduced leaf growth and increased root length10. Biotic stresses such as pathogens also cause plants to close stomata and reduce photosynthesis11,12. Other plant responses to pathogens include production of toxic compounds, including phytoalexins and reactive oxygen species, and induction of localized cell death13. Many of these responses are coordinated by phytohormones14,15. The hormones abscisic acid (ABA) and jasmonic acid (JA) are critical regulators of tolerance to abiotic stresses. For immunity to pathogens, plants primarily rely on salicylic acid (SA), JA and ethylene signaling. The abiotic stress response is regulated by many transcription factor (TF) families, both ABA-dependent and ABA-independent. The former includes ABA-induced basic leucine zipper (bZIP) TFs16,17. These TFs induce stomatal closure, expression of dehydration tolerance genes, and other adaptive physiological responses18,19,20,21,22. However, ABA often increases plant susceptibility in biotic interactions23,24,25,26,27 and frequently acts antagonistically with SA28,29,30.
With this study, we explore the rice transcriptome for a more thorough understanding of how rice regulates responses to multiple abiotic and biotic stresses. Previous studies have explored broad plant stress response by analyzing microarray data31,32. We expand on these studies with robust meta-analysis of publicly available rice RNA-Seq data sets. Our results reveal universally stress-regulated pathways, which we call the rice core stress response. The network of core stress-responsive genes presented here can be further explored for rice improvement in light of the need for tolerance to multiple environmental stresses. In addition to the valuable predictive transcriptome analysis for an important crop system, our approach can be easily expanded to other plant and crop systems.
Results
Meta-analysis of publicly available RNA-Seq data reveals the rice core stress response
To investigate the rice response to stress, we downloaded and analyzed publicly available RNA-Seq data sets representing rice transcriptome response to diverse abiotic and biotic stresses. These stresses include drought33, salt34, high and low temperature35,36, and infection with Xanthomonas oryzae pathovars oryzicola (Xoc) and oryzae (Xoo)37,38, Magnaporthe oryzae39, and Rice Stripe (RSV) and Dwarf (RDV) viruses40,41 (Table 1). All selected studies used stress-sensitive rice varieties. Four technical considerations were applied to choose RNA-Seq data sets: (1) there must be at least two replicates per treatment, (2) there must be untreated controls for each treatment, (3) tissue type was primarily above-ground, and (4) varieties were non-transgenic and non-mutant.
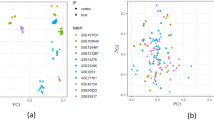
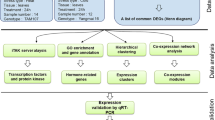
A standard pipeline for consistently processing all raw sequencing data files was used (Fig. 1a). Included in this pipeline were steps for removing low quality reads, aligning to the reference genome, and counting reads. The proportions of reads mapped to the reference genome were generally high, with a mean of 77.4% total reads mapped to loci across all samples (Table S1). We conducted differential gene expression analysis separately on each experiment (Fig. 1a). The number of differentially expressed genes (DEGs) varied widely depending on stress treatment, and ranged from 1,220 to 11,644 DEGs (Fig. 1b,c, Table S2).
Analyses reveal rice core stress responses to abiotic and biotic stresses. (a) Analysis pipeline used to conduct differential gene expression analysis and meta-analysis on publicly available data sets. Number of DEGs identified in all (b) abiotic and (c) biotic stress experiments. (d) MetaDEGs identified from meta-analyses. (e) Number of metaDEGs unique and common in abiotic and biotic meta-analyses up- (up arrow) and down-regulated (down arrow).
To explore the rice core response to abiotic stress, we used a meta-analysis to combine the results from all abiotic stress experiments (Fig. 1a). We found 5,863 meta-analysis-identified DEGs (metaDEGs) that were generally responsive to all abiotic stresses (Fig. 1d, Supplementary Data S1). We repeated this process to explore the core response to biotic stress, and found 2,154 metaDEGs generally responsive to all biotic stresses (Fig. 1d, Supplementary Data S1). Of the DEGs identified in the individual analyses, 10 to 43% were retained as metaDEGs (Table S2). The expression trends of the metaDEGs within individual studies followed the trends identified in the meta-analysis; that is, up- and down-regulated metaDEGs were mostly up- and down-regulated, respectively, within individual studies (Fig. S1). Therefore, this approach was valid for investigating rice core responses to abiotic and biotic stress.
To identify the rice response to all stresses, we investigated the overlap in expression patterns of the two sets of metaDEGs (Fig. 1e). We found all possible patterns of gene expression between abiotic and biotic stresses, including metaDEGs that were uniquely regulated by one stress type (abiotic or biotic), similarly regulated by both stress types, and oppositely regulated by both stress types (Fig. 1e, Table S3). Most metaDEGs were uniquely regulated by either abiotic or biotic stress. Interestingly, there were many more metaDEGs regulated similarly by both stress types (913 metaDEGs) than oppositely (88 metaDEGs). The annotations and median log2 fold changes of all 1,001 common metaDEGs are in Supplementary Data S2. Taken together, these results indicate there are: (1) genes responsive to a single stress type (abiotic or biotic), and (2) genes responsive to all stresses.
Stress altered regulation of photosynthesis-related genes in rice
To investigate the rice biological processes (BP) altered during stress, we evaluated the enrichment patterns of the 45 BP gene ontology (GO) terms in abiotic and biotic up- and down-regulated metaDEGs (Table S4). The GO terms ‘catabolic process’, ‘cell communication’, ‘embryo development’, ‘reproduction’, and ‘response to extracellular stimulus’ were all enriched within metaDEGs (relative to background genes) up-regulated by both abiotic and biotic stress (Fisher’s exact test FDR-corrected p ≤ 0.01, Table 2). The GO terms ‘photosynthesis’, ‘protein modification process’, and ‘response to external stimulus’ were all enriched within metaDEGs down-regulated by both stresses. Several GO terms were enriched exclusively in abiotic or biotic metaDEGs, but no GO terms were enriched in genes oppositely regulated by stress type.
There were 85 metaDEGs annotated with the GO term ‘photosynthesis’. These metaDEGs were generally down-regulated in individual transcriptome studies (Fig. S2). Rice down-regulated photosynthesis-annotated metaDEGs in response to drought, heat, cold, Xoc, M. oryzae, and RDV. Conversely, salt and RSV did not regulate these metaDEGs, and, in the study used, Xoo up-regulated them. These results indicate that altered regulation of photosynthetic pathways is a common rice response to stress.
Stress up-regulated rice phytohormone-induced genes
Because phytohormones are regulators of plant responses, we investigated how stress responses influenced phytohormone-induced genes. Abiotic metaDEGs responsive to ABA, auxin, JA and SA were more up-regulated than expected by random chance as determined by the χ2 goodness of fit test (p ≤ 0.05, Fig. 2, Table S5). Biotic metaDEGs in all hormone-responsive pathways were more up-regulated than expected. Response to ABA was the most significantly up-regulated hormone pathway in both abiotic and biotic metaDEGs, indicating that ABA signaling is likely important to the core stress response.
Rice hormone-responsive genes were generally up-regulated by stress. Observed number of up-regulated hormone-responsive metaDEGs is shown vs. the number expected to be up-regulated by random chance. Asterisks denote numbers observed differed significantly from numbers expected as determined by the χ2 goodness of fit test (***p < 10−14, **p < 10−6, *p < 0.005, see Table S5 for all p-values).
There were 408 and 228 genes responsive to JA and/or SA in abiotic and biotic metaDEGs, respectively (Fig. 3, Supplementary Data S3). The expression of these genes indicate that during either stress type, JA and SA signaling are increased (Fig. 3). Only three small clusters identified within abiotic metaDEGs (Fig. 3a, labeled C1–C3) and two small clusters within biotic metaDEGs (Fig. 3b, labeled C4–C5) did not follow this trend; genes in these clusters were regulated oppositely by stress and hormones (JA and/or SA). Interestingly, in the Xoo study used for this validation, the expression of JA- and SA-responsive genes was generally opposite of all other biotic stress responses. JA and SA response were a larger component of the biotic stress response (10.6% of biotic metaDEGs) than of the abiotic stress response (7.0% of abiotic metaDEGs). Many of the JA- and/or SA-responsive genes were also responsive to ABA (Fig. 3). Genes responsive to JA and/or SA, but not responsive to ABA, were still up-regulated more than expected by random chance (Table S5). However, this was a much smaller proportion of metaDEGs (1.4 and 3.5% of abiotic and biotic metaDEGs, respectively). Taken together, these results indicate that in responses to any stress, rice orchestrates responses via phytohormones.
Signaling downstream of JA and SA is increased during stress. Gene expression (log2 fold changes) of JA- and SA-responsive metaDEGs for (a) abiotic stress and (b) biotic stresses relative to controls (columns) are shown on the right in yellow (down-regulated), black (not regulated) and cyan (up-regulated). Hormone regulatory patterns of JA- and SA-responsive metaDEGs are shown on the left in magenta (down-regulated), black (not regulated; n.r.) and white (up-regulated). Clusters of genes regulated oppositely of hormone pathways are indicated by the orange squares (C1 through C5).
Discovery of promoter motifs important to the stress response
We performed de novo promoter motif enrichment analysis to identify potential stress-responsive regulatory elements. There were 22 and 17 motifs discovered in the abiotic and biotic metaDEGs, respectively (Table S6). GO term analysis revealed six motifs that are likely to be involved in stress-responsive pathways (Fig. 4). Many of these motifs contained a sequence similar to the ACGT core sequence of the ABA responsive element (ABRE), an upstream bZIP TF binding sequence42, indicating a possible role for bZIP TFs in the core stress response. Of the 21 bZIP TFs we identified as metaDEGs, 17 were up-regulated in response to abiotic stress (Supplementary Data S1), including bZIP23 (MSU: LOC_Os02g52780) and bZIP46 (MSU: LOC_Os06g10880), which are key players in ABA response43,44. Biotic stress only up-regulated three bZIP TFs. The enrichment of ABRE-like motifs in the promoters of biotic stress-induced metaDEGs suggests that even though there are fewer bZIP TFs responsive to biotic stresses than abiotic stresses, bZIP TFs may still act as critical regulators of response to biotic stress. One bZIP TF (MSU: LOC_Os08g38020) was a potential node of antagonistic cross-talk, up-regulated by abiotic stress and down-regulated by biotic stress. Taken together, these results indicate that rice utilizes ACGT-bZIP TF to regulate response to both abiotic and biotic stress, and identify bZIP elements as key nodes for further studies.
Pre-processed publicly available gene expression data validates meta-analysis results
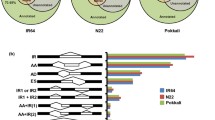
To validate the results of the meta-analysis, nine publicly available pre-processed gene expression studies37,45,46,47,48,49,50,51,52 were examined for the trends expected from our previous analysis (Table 3). With one exception, all studies fit the expected trends; i.e., up- and down-regulated metaDEGs were more up- and down-regulated than expected by random chance, respectively, as determined by the χ2 goodness of fit test (p ≤ 0.05, Fig. 5a,b, Table S7). The study that did not fit the expected trend (GSE57950 drought) had two time-points, with the earlier time-point (1 d after stress) not fitting the expected trend in down-regulated metaDEGs. As in the meta-analysis, photosynthesis genes were mostly down-regulated (Fig. 5c, Table S8). Three studies did not significantly alter photosynthesis gene expression (GSE42096 heat, GSE74465 drought 1 h, GSE107425 drought), and one study up-regulated this pathway (GSE57950 drought 1 d). In the later time-point of the latter study (GSE57950 drought 3d), plants down-regulated photosynthesis-annotated genes, suggesting there may be some temporal effects of drought on altered regulation of photosynthesis, particularly as leaves dehydrate after continued drought. In study GSE108504, rice strongly down-regulated photosynthesis-annotated genes in response to Xoo (Fig. 5c), opposite to the set used in the training data, where these genes were up-regulated by Xoo (Fig. S2). These results validate our meta-analysis approach to finding the rice core stress response.
Publicly available gene expression studies validated meta-analysis results. (a) Up- and (b) Down-regulated metaDEGs and (c) photosynthesis-annotated genes generally followed expected trends in pre-processed publicly available gene expression datasets. n.s. indicates the counts observed did not differ significantly from counts expected as determined by the χ2 goodness of fit test (p > 0.05, see Tables S7 and S8 for all p-values).
Discussion
A variety of environmental stresses affect plants in the field and can limit crop yield. To endure these stresses, plants respond with coordinated changes to their transcriptome. While these changes are dependent on the specific stress experienced, our results indicate that there is a rice core response to all stresses. With our meta-analysis of publicly available RNA-Seq data of rice experiencing various abiotic and biotic stresses, we identified 5,863 and 2,154 genes that are differentially regulated by abiotic stress and biotic stress, respectively (Fig. 1, Supplementary Data S1). Of these, 913 genes were similarly regulated by both abiotic and biotic stress, while 88 were regulated oppositely (Table S3, Supplementary Data S2). A different study utilized differential expression analysis of rice microarray data to identify genes commonly regulated by abiotic and biotic stresses, and found 40 rice genes that were responsive to both abiotic and biotic stresses32. Our meta-analysis of RNA-Seq data identified more of the rice core stress response than this previous comparative microarray analysis. We also validated our meta-analysis approach using additional publicly available studies not used in the training sets; through this validation, we identified sets of stress-responsive genes similar to those found in the meta-analysis (Fig. 5, Tables S7 and S8).
Although the reference genome is annotated with only 45 BP GO terms, we identified several BPs that were altered by stress, including ‘catabolic process’, ‘cell communication’, ‘embryo development’, ‘reproduction’, and ‘response to extracellular stimulus’, which were all up-regulated by stress, and ‘photosynthesis’, ‘protein modification process’, and ‘response to external stimulus’, which were all down-regulated by stress. Photosynthesis is known to be down-regulated by abiotic stresses such as drought, cold, and heat stress53,54,55,56. This is likely a protective mechanism against plant photooxidative damage during stress56,57. In stress tolerant varieties of rice, photosynthetic efficiency is restored, and up-regulation of photosynthesis is physiologically important for yield stability58,59. Consistent with these findings, overexpression of a master regulator of photosynthesis enhanced rice tolerance to drought60. A range of biotic stresses, including bacterial, viral, and fungal pathogens, also inhibit photosynthesis in plants61,62,63,64,65. It is hypothesized that the photosynthesis pathway is a hub of cross-talk in growth and defense trade-offs during plant-pathogen interactions66. Studying the roles of the photosynthesis-regulated metaDEGs identified in this study may facilitate the development of stress tolerant varieties of rice.
Various stresses positively induced phytohormone pathways (Fig. 2). Abiotic stress up-regulated genes responsive to ABA, auxin, JA, and SA, while biotic stress up-regulated genes responsive to the same hormones plus cytokinin and ethylene. The ABA, JA, and SA pathways were the most significantly up-regulated hormone pathways in both abiotic and biotic stress. ABA, JA and SA signaling regulate response to abiotic stresses31,54,55,67. While JA and SA are positive regulators, ABA tends to be a negative regulator of resistance to pathogens25,68,69,70. ABA is also important to inter-kingdom signaling among pathogens and plants. For example, synthesis of ABA by the fungal pathogen M. oryzae during interactions with rice is necessary for pathogen virulence71. Plant-synthesized ABA promotes rice susceptibility to the bacterial pathogen X. oryzae pv. oryzae (Xoo) and even induces swimming in the bacteria26,72. While our results show that both ABA and SA are induced during response to biotic stress, ABA-induced susceptibility to Xoo is due to ABA suppressing SA-mediated defense26. We previously hypothesized that ABA is a node of cross-talk in the rice response to simultaneous high temperature stress and X. oryzae infection35. The results from our current study show that cross-talk among ABA, JA and SA response pathways makes the contribution of each hormone to the rice transcriptome unclear (Fig. 3). Notably, ABA-regulated genes appear to dominate the hormone response during stress. That is, of the metaDEGs responsive to JA and SA, most were also responsive to ABA (Fig. 3, Table S5). These intertwined pathways are critical to plant stress responses, which frequently occur simultaneously, emphasizing that additional study of hormonal cross-talk is needed to provide insights into how to improve plant health.
Our results open the path to future avenues of research, including both in silico and in planta studies. We immediately provide candidate genes for studying multiple stress responses in rice. For example, the prevalence of enriched ABRE-like promoter motifs suggest that the bZIP TFs identified here are good candidate regulators of stress responses (Fig. 4, Supplementary Data S1). Our analysis only used studies with rice plants that were sensitive (susceptible) to the different stresses. Future researchers can expand on this work by analyzing the regulation of metaDEGs in studies with stress-tolerant rice varieties. We only found 88 oppositely regulated metaDEGs between abiotic and biotic stresses, but it is likely that stress tolerance and sensitivity oppositely regulate many more genes. The resources and approach provided with this work will allow for a deeper understanding of rice strategies for overcoming stresses.
We present this work as a proof of concept: meta-analysis of diverse transcriptomic data sets is a valid and robust approach to develop hypotheses for how plants respond to stress in general. It is also possible to expand our approach into other systems. For example, with the wealth of publicly available Arabidopsis transcriptome data, researchers can repeat this analysis to identify candidate regulators of Arabidopsis stress response. In systems with few or no publicly available transcriptome studies, the analysis we describe enables researchers to design transcriptome studies from the ground up to study stress response in their systems. Even while limited by the available rice stress-responsive transcriptome data, with multiple tissue types, host cultivars, and few replicates per treatment (Table 2), real trends were identified, indicating it is possible to design experiments in less well-studied plant systems to use with our approach.
To summarize, publicly available rice transcriptome data were used to identify genes and pathways regulated by abiotic stress, biotic stress, and both stress types. We confirmed that photosynthesis is a generally down-regulated pathway in response to all stress types. We also identified stress-induced plant hormone-responsive genes, particularly genes downstream of ABA, JA and SA. With this work, we provide a list of candidate genes to study for improving rice stress tolerance, and thus yield, in light of environmental stresses. This study provides a valid approach to ask additional questions with respect to how plants respond to stress, including but not limited to (1) how tolerant rice varieties respond to stress and (2) how other plants respond to stress.
Methods
RNA-Seq Data Acquisition and Processing
Raw sequence data for all accessions were downloaded from NCBI Sequence Read Archive using the SRA Toolkit (https://github.com/ncbi/sra-tools). Adapter sequences and low quality reads were removed with Trimmomatic v0.3673. Reads were mapped to the MSU RGAP 7.0 rice reference genome74 with STAR v2.575 and counted using HTSeq v0.9.176.
Differential Gene Expression and Meta-Analyses
Differential gene expression analyses were conducted using the Bioconductor package edgeR77,78. For single analyses, genes were considered differentially expressed if the FDR-adjusted p-values were ≤0.01. For meta-analyses, Fisher’s sum of logs method (as discussed by Rau et al.79 and implemented in the R package metap v0.8; https://cran.r-project.org/web/packages/metap/index.html) was used to combine unadjusted p-values. The p.adjust function in R80 was used to adjust the combined p-values for multiple testing with the ‘fdr’ method. Genes were considered differentially expressed in meta-analyses if the adjusted p-values were ≤0.01 and the absolute value of the median log2 fold change for all studies within the analysis was ≥1. GO terms were considered enriched within a metaDEG set if the odds ratio estimates relative to background genes was >1 and the FDR-corrected p-values from Fisher’s exact test were ≤0.01.
Phytohormone-responsive Gene Analysis
Known hormone-responsive genes were from Garg et al.81. The chisq.test function in R was used for χ2 goodness of fit test, with a p-value threshold of 0.05, to determine if number of hormone-responsive up- and down-regulated genes were as expected due to random chance. For the χ2 tests, the expected number of up- and down-regulated genes was proportional to the total number of up- and down-regulated genes in the background set.
De novo Promoter Motif Discovery
Promoter motifs and associated GO terms were discovered with DREME82 and GOMo83 respectively, using 500 bp regions upstream of putative transcription start sites. Fisher’s exact test with a p-value threshold of 0.01 was used to determine whether motifs were enriched in metaDEG sets.
Validation with Pre-processed Gene Expression Studies
Pre-processed gene expression studies were acquired from NCBI Gene Expression Omnibus. Because many of these studies lacked replicates, regulatory patterns of genes were estimated by finding the ratio of normalized expression value of treatment to control, disregarding log2 fold changes with absolute value < 1. Studies GSE67588 and GSE108504 were normalized by calculating number of gene reads per millions of total reads. The χ2 goodness of fit test with a p-value threshold of 0.05 was used to determine whether the counts of up- and down-regulated were as expected by random chance. For the χ2 tests, the expected number of up- and down-regulated genes was proportional to the total number of up- and down-regulated genes in the background set.
Data Availability
The datasets generated during and/or analyzed during the current study are available from the corresponding author on reasonable request.
References
Mafakheri, A., Siosemardeh, A., Bahramnejad, B., Struik, P. & Sohrabi, Y. Effect of drought stress on yield, proline and chlorophyll contents in three chickpea cultivars. Aust. J. Crop Sci. 4, 580 (2010).
Lobell, D. B., Bänziger, M., Magorokosho, C. & Vivek, B. Nonlinear heat effects on African maize as evidenced by historical yield trials. Nat. Clim. Change 1, 42 (2011).
Semenov, M. A. & Shewry, P. R. Modelling predicts that heat stress, not drought, will increase vulnerability of wheat in Europe. Sci. Rep. 1, 66 (2011).
Atkinson, N. J. & Urwin, P. E. The interaction of plant biotic and abiotic stresses: from genes to the field. J. Exp. Bot. 63, 3523–3543 (2012).
Jongdee, B., Pantuwan, G., Fukai, S. & Fischer, K. Improving drought tolerance in rainfed lowland rice: an example from Thailand. Agr. Water Manage. 80, 225–240 (2006).
Ashraf, M. & Akram, N. A. Improving salinity tolerance of plants through conventional breeding and genetic engineering: an analytical comparison. Biotechnol. Adv. 27, 744–752 (2009).
Fukuoka, S. et al. Gene pyramiding enhances durable blast disease resistance in rice. Sci. Rep. 5, 7773 (2015).
Mittler, R. Abiotic stress, the field environment and stress combination. Trends Plant Sci. 11, 15–19 (2006).
Garrett, K. A., Dendy, S. P., Frank, E. E., Rouse, M. N. & Travers, S. E. Climate change effects on plant disease: genomes to ecosystems. Annu. Rev. Phytopathol. 44, 489–509 (2006).
Maiti, R. & Satya, P. Research advances in major cereal crops for adaptation to abiotic stresses. GM Crops Food 5, 259–279 (2014).
Bilgin, D. D. et al. Biotic stress globally downregulates photosynthesis genes. Plant Cell Environ. 33, 1597–1613 (2010).
Melotto, M., Underwood, W., Koczan, J., Nomura, K. & He, S. Y. Plant stomata function in innate immunity against bacterial invasion. Cell 126, 969–980 (2006).
Wojtaszek, P. Oxidative burst: an early plant response to pathogen infection. Biochem. J. 322, 681–692 (1997).
Nguyen, D., Rieu, I., Mariani, C. & van Dam, N. M. How plants handle multiple stresses: hormonal interactions underlying responses to abiotic stress and insect herbivory. Plant Mol. Biol. 91, 727–740 (2016).
Shigenaga, A. M. & Argueso, C. T. In Semin. Cell Dev. Biol. 174–189 (Elsevier).
Banerjee, A. & Roychoudhury, A. Abscisic-acid-dependent basic leucine zipper (bZIP) transcription factors in plant abiotic stress. Protoplasma 254, 3–16 (2017).
Yoshida, T. et al. Four Arabidopsis AREB/ABF transcription factors function predominantly in gene expression downstream of SnRK2 kinases in abscisic acid signalling in response to osmotic stress. Plant Cell Env. 38, 35–49 (2015).
Kim, T.-H., Böhmer, M., Hu, H., Nishimura, N. & Schroeder, J. I. Guard cell signal transduction network: advances in understanding abscisic acid, CO2, and Ca2+ signaling. Annu. Rev. Plant Biol. 61, 561–591 (2010).
Silva, V. A. et al. Reciprocal grafting between clones with contrasting drought tolerance suggests a key role of abscisic acid in coffee acclimation to drought stress. Plant Growth Regul. 85, 221–229 (2018).
Huang, X. et al. The antagonistic action of abscisic acid and cytokinin signaling mediates drought stress response in Arabidopsis. Mol. Plant 11, 970–982 (2018).
Wang, L., Zhu, J., Li, X., Wang, S. & Wu, J. Salt and drought stress and ABA responses related to bZIP genes from V. radiata and V. angularis. Gene 651, 152–160 (2018).
Ghosh, T. K. et al. Abscisic acid‐induced gene expression in the liverwort Marchantia polymorpha is mediated by evolutionarily conserved promoter elements. Physiol. Plantarum 156, 407–420 (2016).
Xiong, L. & Yang, Y. Disease resistance and abiotic stress tolerance in rice are inversely modulated by an abscisic acid–inducible mitogen-activated protein kinase. Plant Cell 15, 745–759 (2003).
Yasuda, M. et al. Antagonistic interaction between systemic acquired resistance and the abscisic acid–mediated abiotic stress response in Arabidopsis. Plant Cell 20, 1678–1692 (2008).
Lievens, L., Pollier, J., Goossens, A., Beyaert, R. & Staal, J. Abscisic acid as pathogen effector and immune regulator. Front. Plant Sci. 8, 587 (2017).
Xu, J., Audenaert, K., Hofte, M. & De Vleesschauwer, D. Abscisic acid promotes susceptibility to the rice leaf blight pathogen Xanthomonas oryzae pv. oryzae by suppressing salicylic acid-mediated defenses. PLOS One 8, e67413 (2013).
Peskan‐Berghöfer, T. et al. Sustained exposure to abscisic acid enhances the colonization potential of the mutualist fungus Piriformospora indica on Arabidopsis thaliana roots. New Phytol. 208, 873–886 (2015).
de Torres Zabala, M., Bennett, M. H., Truman, W. H. & Grant, M. R. Antagonism between salicylic and abscisic acid reflects early host–pathogen conflict and moulds plant defence responses. Plant J. 59, 375–386 (2009).
Jiang, C.-J. et al. Abscisic acid interacts antagonistically with salicylic acid signaling pathway in rice–Magnaporthe grisea. interaction. Mol. Plant Microbe In. 23, 791–798 (2010).
Ding, Y., Dommel, M. & Mou, Z. Abscisic acid promotes proteasome‐mediated degradation of the transcription coactivator NPR 1 in Arabidopsis thaliana. Plant J. 86, 20–34 (2016).
Hahn, A. et al. Plant core environmental stress response genes are systemically coordinated during abiotic stresses. Int. J. Mol. Sci. 14, 7617–7641 (2013).
Narsai, R. et al. Antagonistic, overlapping and distinct responses to biotic stress in rice (Oryza sativa) and interactions with abiotic stress. BMC Genomics 14, 93 (2013).
Galbiati, F. et al. Hd3a, RFT1 and Ehd1 integrate photoperiodic and drought stress signals to delay the floral transition in rice. Plant Cell Env. 39, 1982–1993 (2016).
Wang, S. et al. Integrated RNA sequencing and QTL mapping to identify candidate genes from Oryza rufipogon associated with salt tolerance at the seedling stage. Front. Plant Sci. 8, 1427 (2017).
Cohen, S. P. et al. RNA-Seq analysis reveals insight into enhanced rice Xa7-mediated bacterial blight resistance at high temperature. PLOS One 12, e0187625 (2017).
Shen, Y. et al. Transcriptome dynamics through alternative polyadenylation in developmental and environmental responses in plants revealed by deep sequencing. Genome Res. 21, 1478–1486 (2011).
Wilkins, K. E., Booher, N. J., Wang, L. & Bogdanove, A. J. TAL effectors and activation of predicted host targets distinguish Asian from African strains of the rice pathogen Xanthomonas oryzae pv. oryzicola while strict conservation suggests universal importance of five TAL effectors. Front. Plant Sci. 6, 536 (2015).
Zhang, F. et al. Comparative transcriptome profiling of a rice line carrying Xa39 and its parents triggered by Xanthomonas oryzae pv. oryzae provides novel insights into the broad-spectrum hypersensitive response. BMC Genomics 16, 111 (2015).
Huang, H. et al. Increase of fungal pathogenicity and role of plant glutamine in nitrogen-induced susceptibility (NIS) to rice blast. Front. Plant Sci. 8, (265 (2017).
Yang, J., Zhang, F., Li, J., Chen, J.-P. & Zhang, H.-M. Integrative analysis of the microRNAome and transcriptome illuminates the response of susceptible rice plants to rice stripe virus. PLOS One 11, e0146946 (2016).
Zhao, S. et al. A viral protein promotes host SAMS1 activity and ethylene production for the benefit of virus infection. eLife 6, e27529 (2017).
Gomez-Porras, J. L., Riano-Pachon, D. M., Dreyer, I., Mayer, J. E. & Mueller-Roeber, B. Genome-wide analysis of ABA-responsive elements ABRE and CE3 reveals divergent patterns in Arabidopsis and rice. BMC Genomics 8, 260, https://doi.org/10.1186/1471-2164-8-260 (2007).
Xiang, Y., Tang, N., Du, H., Ye, H. & Xiong, L. Characterization of OsbZIP23 as a key player of the basic leucine zipper transcription factor family for conferring abscisic acid sensitivity and salinity and drought tolerance in rice. Plant Physiol. 148, 1938–1952, https://doi.org/10.1104/pp.108.128199 (2008).
Lu, G., Gao, C., Zheng, X. & Han, B. Identification of OsbZIP72 as a positive regulator of ABA response and drought tolerance in rice. Planta 229, 605–615, https://doi.org/10.1007/s00425-008-0857-3 (2009).
Wang, D. et al. Nucleolar DEAD-box RNA helicase TOGR1 regulates thermotolerant growth as a pre-rRNA chaperone in rice. PLOS Genetics 12, e1005844 (2016).
Huang, L. et al. Comparative transcriptome sequencing of tolerant rice introgression line and its parents in response to drought stress. BMC Genomics 15, 1026 (2014).
Garg, R., Chevala, V. N., Shankar, R. & Jain, M. Divergent DNA methylation patterns associated with gene expression in rice cultivars with contrasting drought and salinity stress response. Sci. Rep. 5, 14922 (2015).
Jung, I. et al. Clustering and evolutionary analysis of small RNAs identify regulatory siRNA clusters induced under drought stress in rice. BMC Syst. Biol. 10, 115 (2016).
Zong, W. et al. Feedback regulation of ABA signaling and biosynthesis by a bZIP transcription factor targets drought resistance related genes. Plant Physiol. 171, 2810–2825 (2016).
Fu, J. et al. OsJAZ1 attenuates drought resistance by regulating JA and ABA signaling in rice. Front. Plant Sci. 8, 2108 (2017).
Bidzinski, P. et al. Transcriptional basis of drought-induced susceptibility to the rice blast fungus Magnaporthe oryzae. Front. Plant Sci. 7, 1558 (2016).
Tran, T. T. et al. Functional analysis of African Xanthomonas oryzae pv. oryzae TALomes reveals a new susceptibility gene in bacterial leaf blight of rice. Plos Pathog. 14, e1007092 (2018).
Pandey, N. et al. CAMTA 1 regulates drought responses in Arabidopsis thaliana. BMC Genomics 14, 216 (2013).
Maruyama, K. et al. Integrated analysis of the effects of cold and dehydration on rice metabolites, phytohormones, and gene transcripts. Plant Physiol. 164, 1759–1771 (2014).
Todaka, D. et al. Temporal and spatial changes in gene expression, metabolite accumulation and phytohormone content in rice seedlings grown under drought stress conditions. Plant J. 90, 61–78 (2017).
Brestic, M., Zivcak, M., Kunderlikova, K. & Allakhverdiev, S. I. High temperature specifically affects the photoprotective responses of chlorophyll b-deficient wheat mutant lines. Photosynth. Res. 130, 251–266 (2016).
Yan, K. et al. Dissection of photosynthetic electron transport process in sweet sorghum under heat stress. Plos One 8, e62100 (2013).
Li, J. et al. Influence of drought stress on photosynthetic characteristics and protective enzymes of potato at seedling stage. J. Saudi Soc. Agr. Sci. 16, 82–88 (2017).
Zhang, Z.-F., Li, Y.-Y. & Xiao, B.-Z. Comparative transcriptome analysis highlights the crucial roles of photosynthetic system in drought stress adaptation in upland rice. Sci. Rep. 6, 19349 (2016).
Ambavaram, M. M. et al. Coordinated regulation of photosynthesis in rice increases yield and tolerance to environmental stress. Nat. Commun. 5, 5302 (2014).
Akimoto‐Tomiyama, C., Tanabe, S., Kajiwara, H., Minami, E. & Ochiai, H. Loss of chloroplast‐localized protein phosphatase 2Cs in Arabidopsis thaliana leads to enhancement of plant immunity and resistance to Xanthomonas campestris pv. campestris infection. Mol. Plant Pathol. 19, 1184–1195 (2018).
Girija, A. M. et al. Rice leaf transcriptional profiling suggests a functional interplay between Xanthomonas oryzae pv. oryzae lipopolysaccharide and extracellular polysaccharide in modulation of defense responses during infection. Mol. Plant Microbe In. 30, 16–27 (2017).
Cheng, D.-D. et al. Photoinhibition and photoinhibition-like damage to the photosynthetic apparatus in tobacco leaves induced by Pseudomonas syringae pv. tabaci under light and dark conditions. BMC Plant Biol. 16, 29 (2016).
Pérez‐Clemente, R., Montoliu, A., Vives, V., López‐Climent, M. & Gómez‐Cadenas, A. Photosynthetic and antioxidant responses of Mexican lime (Citrus aurantifolia) plants to Citrus tristeza virus infection. Plant Pathol. 64, 16–24 (2015).
Ghosh, S., Kanwar, P. & Jha, G. Alterations in rice chloroplast integrity, photosynthesis and metabolome associated with pathogenesis of Rhizoctonia solani. Sci. Rep. 7, 41610 (2017).
Kangasjärvi, S., Tikkanen, M., Durian, G. & Aro, E.-M. Photosynthetic light reactions–An adjustable hub in basic production and plant immunity signaling. Plant Physiol. Bioch. 81, 128–134 (2014).
Sah, S. K., Reddy, K. R. & Li, J. Abscisic acid and abiotic stress tolerance in crop plants. Front. Plant Sci. 7, 571 (2016).
Creelman, R. A. & Mullet, J. E. Jasmonic acid distribution and action in plants: regulation during development and response to biotic and abiotic stress. Proc. Natl. Acad, Sci. USA 92, 4114–4119 (1995).
Klessig, D. F. & Malamy, J. The salicylic acid signal in plants. Plant Mol. Biol. 26, 1439–1458 (1994).
Klessig, D. F., Choi, H. W. & Dempsey, D. M. A. Systemic acquired resistance and salicylic acid: past, present and future. Mol. Plant Microbe In. 31, 871–888 (2018).
Spence, C. A., Lakshmanan, V., Donofrio, N. & Bais, H. P. Crucial roles of abscisic acid biogenesis in virulence of rice blast fungus Magnaporthe oryzae. Front. Plant Sci. 6, 1082 (2015).
Xu, J. et al. Phytohormone-mediated interkingdom signaling shapes the outcome of rice-Xanthomonas oryzae pv. oryzae interactions. BMC Plant Biol. 15, 10 (2015).
Bolger, A. M., Lohse, M. & Usadel, B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30, 2114–2120, https://doi.org/10.1093/bioinformatics/btu170 (2014).
Kawahara, Y. et al. Improvement of the Oryza sativa Nipponbare reference genome using next generation sequence and optical map data. Rice 6, 4, https://doi.org/10.1186/1939-8433-6-4 (2013).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21, https://doi.org/10.1093/bioinformatics/bts635 (2013).
Anders, S., Pyl, P. T. & Huber, W. HTSeq–a Python framework to work with high-throughput sequencing data. Bioinformatics 31, 166–169, https://doi.org/10.1093/bioinformatics/btu638 (2015).
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140, https://doi.org/10.1093/bioinformatics/btp616 (2010).
McCarthy, D. J., Chen, Y. & Smyth, G. K. Differential expression analysis of multifactor RNA-Seq experiments with respect to biological variation. Nucleic Acids Res. 40, 4288–4297, https://doi.org/10.1093/nar/gks042 (2012).
Rau, A., Marot, G. & Jaffrezic, F. Differential meta-analysis of RNA-seq data from multiple studies. BMC Bioinformatics 15, 91, https://doi.org/10.1186/1471-2105-15-91 (2014).
R Core Team. R: A Language and Environment for Statistical Computing, <Available from, https://www.R-project.org/> (2016).
Garg, R., Tyagi, A. K. & Jain, M. Microarray analysis reveals overlapping and specific transcriptional responses to different plant hormones in rice. Plant Signal. Behav. 7, 951–956, https://doi.org/10.4161/psb.20910 (2012).
Bailey, T. L. DREME: motif discovery in transcription factor ChIP-seq data. Bioinformatics 27, 1653–1659, https://doi.org/10.1093/bioinformatics/btr261 (2011).
Buske, F. A., Boden, M., Bauer, D. C. & Bailey, T. L. Assigning roles to DNA regulatory motifs using comparative genomics. Bioinformatics 26, 860–866, https://doi.org/10.1093/bioinformatics/btq049 (2010).
Acknowledgements
The authors acknowledge U.S. National Science Foundation (IOS-1444511, J.E.L.) for support of S.P.C. The authors thank Vamsi J. Nalam (Colorado State University) for important discussion and Alvaro L. Pérez-Quintero (Colorado State University) for critical reading of the manuscript. The authors also thank all generators of the publicly available data used within this study, without which this study would not be possible.
Author information
Authors and Affiliations
Contributions
Under the supervision of J.E.L., S.P.C. developed the workflow described, conducted the data analysis, prepared the figures, and wrote the main manuscript text. Both authors critically edited the manuscript and approved the final version.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Cohen, S.P., Leach, J.E. Abiotic and biotic stresses induce a core transcriptome response in rice. Sci Rep 9, 6273 (2019). https://doi.org/10.1038/s41598-019-42731-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-019-42731-8
- Springer Nature Limited
This article is cited by
-
Functional annotation and meta-analysis of maize transcriptomes reveal genes involved in biotic and abiotic stress
BMC Genomics (2024)
-
Comparisons of two receptor-MAPK pathways in a single cell-type reveal mechanisms of signalling specificity
Nature Plants (2024)
-
Identification of responsive genes to multiple abiotic stresses in rice (Oryza sativa): a meta-analysis of transcriptomics data
Scientific Reports (2024)
-
Current status of molecular rice breeding for durable and broad-spectrum resistance to major diseases and insect pests
Theoretical and Applied Genetics (2024)
-
Transcriptional Regulation of Biotic and Abiotic Stress Responses: Challenges and Potential Mechanism for Stress Tolerance and Chickpea Improvement
Tropical Plant Biology (2024)